Abstract
Keywords
Introduction
Head and neck squamous cell carcinoma (HNSCC) is the 6th most common cancer worldwide, with nearly 1 320 000 new cases and 440 000 new deaths each year.1,2 HNSCC occurs in the head and neck, which has a complex anatomical structure and involves many organs. Excessive proliferation of HNSCC can have a great impact on the patient's appearance and basic physiological functions such as swallowing, language, and breathing, significantly affecting the quality of life. Many lymph nodes exist in the head and neck, therefore, cancer cells are prone to lymph node metastasis. Most patients with advanced HNSCC have local metastasis at initial diagnosis. 3 The prognosis of patients with advanced HNSCC is poor even when treated with surgery combined with radiotherapy and chemotherapy; the 5-year survival rate is less than 50%. 4 Therefore, exploration of the underlying mechanisms affecting the proliferation and migration of HNSCC is essential for the diagnosis and treatment of HNSCC.
Heat shock protein 90 (HSP90) is a class of ATP-dependent, highly conserved molecular chaperone proteins. When cells are stressed by external stimuli, HSP90 utilizes the energy generated by ATP hydrolysis to participate in the correct folding and stabilization of important regulatory proteins in various physiological and biochemical processes. In this way, HSP90 indirectly participates in cell cycle regulation, proliferation, migration, and apoptosis.5–8 Previous studies have revealed that HSP90 is highly expressed in a variety of cancers, and is associated with poor prognosis.9–11 HSP90 may play a tumor-promoting role by stabilizing various abnormal and highly expressed oncogenic proteins in cancer. Therefore, HSP90 is a potential molecular target for cancer treatment.12,13 HSP90 inhibitors can prevent the correct folding and stabilization of HSP90 client proteins, leading to their ubiquitination and degradation through the proteasome pathway. Therefore, they have become a promising synergistic anti-cancer drug.14–16 In mammalian cells, HSP90 has 4 subtypes: heat shock protein 90 alpha family class A member 1 (HSP90AA1), heat shock protein 90 alpha family class B member 1 (HSP90AB1), glucose-regulated protein 94 (GRP94), and TNF receptor-associated protein 1 (TRAP1). 17 Previous studies mainly focused on HSP90 as a whole, and studies on its subtypes are few. In fact, there has been no systematic study on the role of one HSP90 subtype in the progression of HNSCC so far. Recent studies have demonstrated that HSP90AB1 is highly expressed in many cancers and is significantly correlated with prognosis. Wang et al 12 found that HSP90AB1 was overexpressed in gastric cancers and correlated with the poor prognosis, proliferation, and invasion via the interaction of low-density lipoprotein receptor-related protein 5 (LRP5) and inhibited ubiquitin-mediated degradation of LRP5. Meng et al 18 found that HSP90AB1 induced endothelial cell-dependent tumor angiogenesis by activating vascular endothelial growth factor receptors (VEGFRs) transcription in hepatocellular carcinoma. Rong et al 19 found that the upregulation of HSP90AB1 was associated with poor postsurgical survival time and lymphatic metastasis in lung cancer patients. However, the role of HSP90AB1 in HNSCC is still poorly understood. Therefore, exploration of the significance of HSP90AB1 in the occurrence and development of HNSCC is crucial.
The Warburg effect has been widely studied as a common feature of malignant tumors, whereby even with a sufficient oxygen supply, cancer cells tend to obtain energy through glycolysis rather than oxidative phosphorylation. 20 Glycolysis in HNSCC is significantly higher than in normal tissues,21,22 increasing glucose uptake and lactic acid production, thus affecting cancer cell growth, proliferation, angiogenesis, and invasion.23–25 Currently, inhibition of glycolysis in malignant tumors is a feasible tumor treatment strategy.26,27 However, very few studies have analyzed the direct relationship between HSP90 and glycolysis in cancer. Considering HSP90 has a wide range of client proteins, we speculated it may also play a regulatory role in aerobic glycolysis in malignant tumors.
Here, we hypothesized that HSP90AB1 also functions as an oncogene in HSNCC. We began by investigating the HSP90AB1 expression in HNSCC and analyzing its relationship with clinicopathological parameters. Subsequently, we constructed 3 stable HSP90AB1 knockdown HNSCC cell lines and studied the effects of HSP90AB1 knockdown on their proliferation and migration. Finally, we detected the changes in glycolysis levels in 3 HNSCC cell lines and preliminarily explored its possible mechanism.
Materials and Methods
Cell Culture
The human HNSCC cell lines CAL27, HSC3, and HN4 were purchased from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). The JHU011 cell line was obtained from Johns Hopkins University (MD, USA) and the SCC4, SCC9, and FaDu cell lines were obtained from the American Type Culture Collection (ATCC; VA, USA). The human immortalized keratinocyte line HaCaT was also purchased from ATCC. The FaDu cells were cultured in minimal essential medium (MEM; Gibco, NY, USA) and the rest of the cell lines were cultured in Dulbecco's modified Eagle's medium (DMEM; Gibco, NY, USA) supplemented with 10% fetal bovine serum (FBS; Biological Industries, Israel), 100 U mL−1 penicillin and 100 mg mL−1 streptomycin (KeyGen, Jiangsu, China) at 37 °C in a humidified atmosphere containing 5% CO2 and 20% O2.
Clinical Specimens
Eight cases of fresh-frozen tumor samples together with matched adjacent normal samples and 68 cases of formalin-fixed, paraffin-embedded (FFPE) tumor samples from HNSCC patients who underwent radical surgery at Nanjing Stomatological Hospital were acquired from the Biobank of Nanjing Stomatological Hospital. The fresh-frozen samples were used for western blot analysis, while the FFPE samples were used for immunohistochemical staining. The collection of patient samples was approved by the Ethics Committee of Nanjing Stomatological Hospital, Nanjing University. All patients provided written informed consent before sample collection.
Bioinformatics Analysis
The RNA-seq transcriptome data normalized by the Expectation-Maximization (RSEM) approach of HNSCC samples and normal control samples were downloaded from the TCGA data portal (https://portal.gdc.cancer.gov/). In addition, the corresponding clinical data including clinical stage, T stage, lymph node metastasis, and survival information of HNSCC patients were also acquired. The co-expression relationship between HSP90AB1 and various key proteins in the glycolysis pathway was obtained through Gene Expression Profiling Interactive Analysis (GEPIA). 28 Gene Set Enrichment Analysis (GSEA) was used to investigate the biological pathway difference between high and low HSP90AB1 patients. The reference gene set was “Hallmark.v7.4.gmt.”
Western Blotting
Western blotting was performed as described previously 29 with the following modifications. The following primary antibodies were used: rabbit anti-HSP90AB1 (1:4000, 11405-1-AP, Proteintech, Hubei, China), rabbit anti-E-cadherin (1:10 000, 20874-1-AP, Proteintech, Hubei, China), mouse anti-Vimentin (1:10 000, 60330-1-Ig, Proteintech, Hubei, China), rabbit anti-PCNA (1:4000, 10205-2-AP, Proteintech, Hubei, China), mouse anti-β-actin (1:10 000, 66009-1-Ig, Proteintech, Hubei, China), rabbit anti-Akt (pan) (1:1000, 4691S, Cell Signaling Technology, MA, USA), and rabbit anti-Phospho-Akt (Ser473) (1:1000, 4060S, Cell Signaling Technology, MA, USA). The secondary antibodies used in this study included goat anti-mouse IgG (H + L) HRP (1:5000, SA00001-1, Proteintech, Hubei, China) and goat anti-rabbit IgG (H + L) HRP (1:5000, SA00001-2, Proteintech, Hubei, China).
RNA Extraction and qRT-PCR
Total cellular RNA was extracted using TRIZOL reagent (Invitrogen, CA, USA) following a standard protocol. The PrimeScript RT reagent kit with gDNA Eraser (Takara, Liaoning, China) was used for RNA reverse transcription. Quantitative real-time PCR (qRT-PCR) was performed on an Applied Biosystems ViiA™7 instrument using a ChamQ Universal SYBR qPCR Master Mix (Vazyme, Jiangsu, China) according to the manufacturer’s protocol. Gene expression was normalized to β-actin expression. The quantification of mRNA was performed using the 2−△△Ct method. The sequences of primers used are listed in Supplemental Table S1.
Immunohistochemical Analysis
Immunohistochemistry (IHC) was performed as described previously 29 with the following modifications. The sections were incubated overnight with rabbit anti-HSP90AB1 (1:500, 11405-1-AP, Proteintech, Hubei, China). The IHC staining scores were calculated based on the following criteria: (a) the positive area was divided into 4 degrees (1, ≤ 25%; 2, 25%-50%; 3, 50%-75%; and 4, > 75%-100%); (b) the staining intensity was divided into 4 degrees (1, negative; 2, week; 3, moderate; and 4, strong). The scores were multiplied to produce the total score. Those with a total score > 9 were classified into the high HSP90AB1 expression group and the rest into the low HSP90AB1 expression group.
Lentiviruses and Transfection
Knockdown of HSP90AB1 in HSNCC cell lines was achieved using a lentiviral vector (Genechem, Shanghai, China) carrying a short-hairpin RNA (shRNA) specific for HSP90AB1, according to the manufacturer’s instructions. The sequence of the HSP90AB1 shRNA was: 5′-AGTAAACTAAGGGTGTCAACTCGAGTTGACACCCTTAGTTTACTGC-3′, while the sequence of the negative control shRNA was: 5′-TTCTCCGAACGTGTCACGT-3′.
The HNSCC cells were seeded in 6-well plates at a density of 5 × 104 cells per well and grown to approximately 20% to 30% confluence. Then, 10 μL lentivirus solution (7 × 108 TU/mL) was added to the cells for 12 h and then replaced with a fresh culture medium. The stably transfected HNSCC cells were screened with a medium containing Puromycin 2HCI (5 μg/mL, Selleck, Shanghai, China) for 2 weeks. The efficiency of transfection was confirmed by fluorescence microscope, western blotting, and qRT-PCR.
Cell Proliferation Assay
Cell counting kit-8 (CCK-8) assay was performed using a CCK-8 kit (Bimake, TX, USA) according to the manufacturer’s instructions. In brief, cells were seeded at a density of 5 × 103 cells/well in 96-well plates in 100 μL complete culture medium and cultured successively for 24, 48, 72, and 96 h. Next, CCK-8 (10 μL) was added to each well and after 30 min, the optical density (OD) values were detected at 450 nm with a microplate reader (SpectraMax M3; Molecular Device, CA, USA).
5-Ethynyl-2′-deoxyuridine (EdU) incorporation assay was performed using a Cell-Light ™ EdU Apollo567 In Vitro Kit (RiboBio, Guangdong, China) according to the manufacturer’s instructions, as described previously. 29
For colony formation assays, cells were plated into 6-well plates at a density of 1 × 103 cells/well. After 10 days, the supernatant was discarded and the cells were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet (Beyotime, Shanghai, China). Images were captured using an EPSON V330 scanner (EPSON, Nagano, Japan).
Cell Migration Assay
Migration assays were carried out using a transwell system with 6.5 mm insert chambers (Corning, NY, USA). Cells were seeded with 0.1 mL serum-free culture medium on the top chamber at a density of 1 × 104 cells per well. A complete culture medium (20% FBS, 500 μL/well) was added to the bottom chamber. After incubation for 24 h at 37 °C, the cells remaining on the upper surface were completely removed and the migrated cells were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet (Beyotime, Shanghai, China), photographed, and quantified.
The wound healing assay was performed as described below. Cells were seeded in six-well plates and grown to confluence. Then, 200-μL pipette tips were used to make artificial wounds. The gaps of the wound edges were captured under the microscope after 36 h and quantified to assess cell migration using ImageJ software.
Xenograft Tumor Model
The BALB/c male nude mice (n = 5, 5-6 weeks old) were purchased from Nanjing Biomedical Research Institute of Nanjing University (Jiangsu, China) and maintained in a specific pathogen-free environment. After knockdown of HSP90AB1, CAL27 cells were injected into the dorsal flanks of each mouse subcutaneously (1 × 107/mL, 0.1 mL/site, shHSP90AB1 in the left flank and shNC in the right flank). After 28 days, the mice were sacrificed, and the tumors were dissected and weighed.
Glycolysis Analysis
To measure the glycolytic activity in cells, corresponding assay kits were purchased from the Nanjing Jiancheng Bioengineering Institute (Jiangsu, China) to detect the content of ATP (A095-2-1), pyruvate (A081-1-1), and lactic acid (A019-2-1) and the activity of hexokinase (A077-3-1), pyruvate kinase (A076-1-1), and lactate dehydrogenase (A020-1-2).
Statistical Analysis
All statistical analyses and statistical graphs were done using GraphPad Prism 9.0 (GraphPad Software Inc., CA, USA). All data are expressed as the mean ± standard deviation (SD). Overall survival distributions were estimated according to the Kaplan–Meier method and were compared with the log-rank test. The correlation between the HSP90AB1 expression and clinicopathological parameters in HNSCC tissues was evaluated by Fisher's exact test. Other statistical differences were measured using Student’s two-tailed t-test or one-way ANOVA for multiple comparisons. For all statistical analyses, P < .05 was considered statistically significant.
Ethics Approval
Institutional review board approval was obtained from the Nanjing Stomatological Hospital Ethics Committee (approval number: D2103022). All animal experiments and experimental protocols were in accordance with the rules of the Animal Ethical and Welfare Committee of Nanjing University (approval number: SYXK2019-0056).
Results
HSP90AB1 was up-Regulated in HNSCC
We first compared the quantitative data of relative HSP90AB1 expression levels between 502 HNSCC and 44 normal head and neck tissues downloaded from the TCGA database. Results showed that the HSP90AB1 expression level in HNSCC was significantly higher than in normal head and neck tissues (Figure 1A). Subsequently, we extracted proteins from a human immortalized keratinocyte line (HaCaT) and 7 head and neck squamous cell carcinoma cell lines (CAL27, HSC3, SCC4, SCC9, FaDu, HN4, and JHU011), and compared the HSP90AB1 expression levels. We found the 7 HNSCC cell lines had increased HSP90AB1 protein expression compared to the HaCaT cell line, to varying degrees (Figure 1B). The results of qRT-PCR also showed that HSP90AB1 mRNA expression was higher in HNSCC cell lines than that in the HaCaT cell line (Figure 1C). Moreover, we compared HSP90AB1 protein expression in 8 patient sample pairs of HNSCC tissues and adjacent normal tissues. Results showed that the HSP90AB1 protein expression in HNSCC tissues was higher than in adjacent normal tissues (Figure 1D). Taken together, these findings suggested that HSP90AB1 was up-regulated in HNSCC.

HSP90AB1 was overexpressed in HNSCC cells and tissues. (A) Quantitative data of relative HSP90AB1 expression levels between 502 HNSCC and 44 normal head and neck tissues downloaded from the TCGA database were analyzed and compared. (B, C) Protein expression levels of HSP90AB1 in a HaCaT and 7 HNSCC cell lines (CAL27, HSC3, SCC4, SCC9, FaDu, HN4, and JHU011) were detected by western blotting (B) and qRT-PCR (C). (D) Protein expression levels of HSP90AB1 in 8 paired fresh-frozen HNSCC and matched adjacent normal samples were detected by western blotting. N normal tissue, T tumor tissue. Error bars represent the mean ± SD of 3 replicates. *P < .05; **P < .01; ***P < .001.
HSP90AB1 Expression was Associated With T Stage, Lymph Node Metastasis, and Prognosis in HNSCC
To explore the correlation between HSP90AB1 expression and clinicopathological parameters in HNSCC, we first analyzed the quantitative data downloaded from the TCGA database. We compared the HSP90AB1 expression levels of HNSCC in patients with different clinicopathologic parameters and found that HSP90AB1 expression in HNSCC was significantly correlated with T grade and lymph node metastasis, but not with histological grade. HSP90AB1 expression in HNSCC with T3/T4 grade and lymph node metastasis was higher than in those with T1/T2 grade and no lymph node metastasis (Figure 2A). In addition, Kaplan–Meier survival analysis of 499 patients with follow-up data showed that higher HSP90AB1 expression was significantly correlated with decreased overall survival in HNSCC patients (Figure 2B). To further verify this finding, we performed IHC on HNSCC tissues from 68 patients with complete pathological and follow-up data. Patients were divided into 2 groups, a high HSP90AB1 expression group, and a low HSP90AB1 expression group, according to the scores calculated from the product of positive area score and staining intensity score (Figure 2C). Surprisingly, the analysis yielded similar results to the bioinformatics analysis above. HNSCC patients with T3/T4 grade and lymph node metastasis showed significantly higher HSP90AB1 expression levels than those with T1/T2 and no lymph node metastasis, and no obvious correlation between histological grade and HSP90AB1 expression was observed (Figure 2D). Kaplan–Meier survival analysis also suggested that low HSP90AB1 expression was correlated with poor prognosis in HNSCC (Figure 2E). Overall, these findings strongly indicated that HSP90AB1 expression was correlated with T grade and lymph node metastasis and could be used as a prognostic factor in HNSCC.

The expression level of HSP90AB1 was associated with T stage, lymph node metastasis, and prognosis in HNSCC. (A, B) The relationship between HSP90AB1 expression levels and histological grade, T grade, and lymph node metastasis in HNSCC was analyzed. Kaplan–Meier survival analysis of HNSCC patients with low and high expression levels of HSP90AB1 was performed. The data was from the TCGA database. LNM lymph node metastasis. (C) Representative figures of the scores for the different positive area (P) and staining intensity (I) after HSP90AB1 IHC in HNSCC tissues were shown. (D) Graphical representation of the statistical distribution of HSP90AB1 with regard to histological grade, T grade, and lymph node metastasis in HNSCC tissues. (E) Kaplan–Meier survival analysis of HNSCC patients with low and high expression levels of HSP90AB1 was performed. #P > .05; *P < .05; **P < .01.
Knockdown of HSP90AB1 Inhibited the Proliferation and Migration of HNSCC Cells
In view of the above results showing that HSP90AB1 expression was related to HNSCC T stage and lymph node metastasis, we hypothesized that HSP90AB1 regulates the proliferation and migration of HNSCC. We, therefore, selected a tongue squamous cell carcinoma cell line (CAL27), pharyngeal squamous cell carcinoma cell line (FaDu), and laryngeal carcinoma cell line (JHU011) for phenotypic experiments. We constructed stable HSP90AB1 knockdown cell lines via lentivirus and verified the knockdown efficiency by western blotting. Results showed that HSP90AB1 protein expression in the shHSP90AB1 groups was knocked down by more than 50% compared with the shNC groups, in all 3 cell lines (Figure 3A). HSP90AB1 mRNA expression was detected by qRT-PCR and showed a significant decrease in the shHSP90AB1 groups compared with the shNC (Figure 3B). These results verified that the knockdown efficiency of HSP90AB1 was satisfactory. In order to check whether the HSP90AB1 knockdown had an off-target effect and caused any alteration in other HSP90 subtypes, we also detected the mRNA expressions of HSP90AA1, GRP94, and TRAP1. The results showed that although the mRNA levels of GRP94 and TRAP1 in CAL27 and JHU011 shHSP90AB1 groups were slightly higher than those in shNC groups, and the mRNA level of TRAP1 in FaDu shHSP90AB1 group was slightly higher than that in shNC group, the differences were within an acceptable range (Figure 3C). Based on these results, we believed that the HSP90AB1 knockdown was successful and had no off-target effect, which allowed us to conduct subsequent phenotypic experiments.
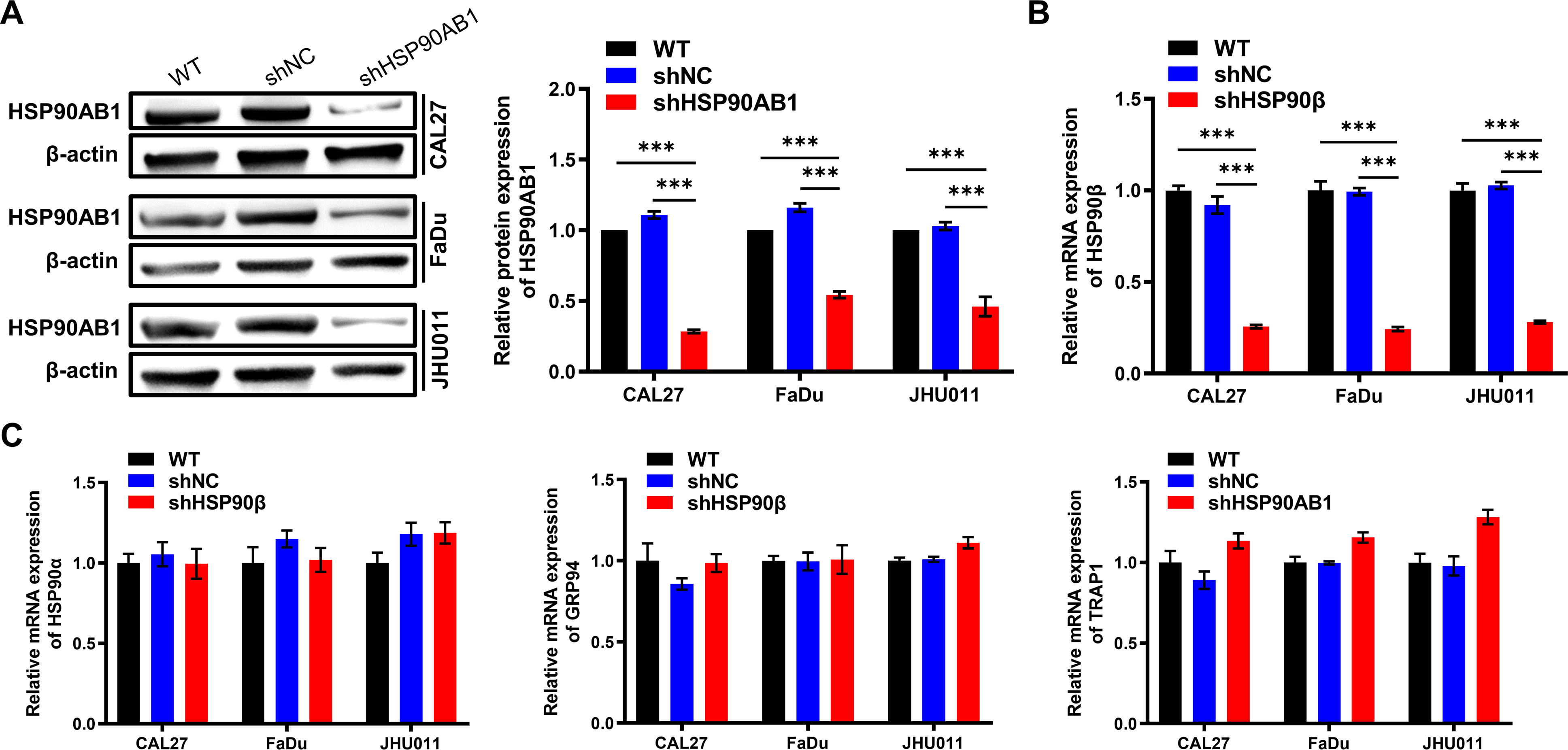
Verification of HSP90AB1 knockdown efficiency in HNSCC cells. (A) Changes in HSP90AB1 protein expression in 3 HNSCC cell lines after being transfected with lentivirus were detected by western blotting. (B) Changes in HSP90AB1 mRNA expression in 3 HNSCC cell lines after being transfected with lentivirus were detected by qRT-PCR. (C) Changes in mRNA expression of HSP90AA1, GRP94, and TRAP1 in 3 HNSCC cell lines after being transfected with lentivirus were detected by qRT-PCR. Error bars represent the mean ± SD of 3 replicates. ***P < .001.
The CCK-8 assay showed that downregulation of HSP90AB1 significantly inhibited the growth of HSNCC cells compared with cells transfected with control vectors (Figure 4A). Then, we examined cell proliferation through EdU incorporation assay and found that stable HSP90AB1 knockdown cells showed lower EdU fluorescence positive rates, which revealed that HSP90AB1 knockdown markedly reduced the proliferation of HNSCC cells compared to controls (Figure 4B). Similarly, colony formation of HSP90AB1-depleted cells was remarkably suppressed compared with the shNC groups (Figure 4C). Next, we established a xenograft model of HNSCC by subcutaneously injecting stably transfected HSP90AB1-knockdown and control CAL27 cells into the left and right sides of the same nude mice. After 28 days, we sacrificed the mice, dissected and weighed the tumors, and found that tumors developing from the HSP90AB1-knockdown cells were significantly smaller than those developing from the control cells (Figure 4D). Moreover, as illustrated by the transwell migration assay and wound healing assay (Figure 5A and B), migrated cell numbers and migration distances of shHSP90AB1 groups were fewer than those of shNC groups, indicating that HSP90AB1 knockdown also decreased the migration ability of HNSCC cells. Finally, we examined the expression of proliferation and migration marker proteins in HNSCC cells by western blotting. We found that after the knockdown of HSP90AB1, proliferating cell nuclear antigen (PCNA) expression decreased observably in CAL27 and JHU011 cell lines, and E-cadherin expression increased, Vimentin expression decreased observably in all 3 HNSCC cell lines (Figure 5C). These results strongly indicated that HSP90AB1 was vital for the proliferation and migration of HNSCC cells, and knockdown of HSP90AB1 inhibited the proliferation and migration of HNSCC cells.

Knockdown of HSP90AB1 inhibited the proliferation of HNSCC cells. (A-C) Changes in cell proliferation ability of 3 HNSCC cell lines after HSP90AB1 knockdown were measured by CCK-8 assays (A), EdU incorporation assay (B), and colony formation assay (C). Error bars represent the mean ± SD of 3 replicates. (D) Images of subcutaneous tumors from mice injected with HSP90AB1-knockdown and control stably transfected CAL27 cells. Tumors were weighed and compared to analyze the effect of HSP90AB1 knockdown on the proliferation of CAL27 cells. *P < .05; **P < .01; ***P < .001.

Knockdown of HSP90AB1 inhibited the migration of HNSCC cells. (A, B) Changes in cell migration ability of 3 HNSCC cell lines after HSP90AB1 knockdown were measured by transwell migration assays (A) and wound healing assays (B). Error bars represent the mean ± SD of 3 replicates. (C) The proliferation and migration marker proteins (E-cadherin, Vimentin, and PCNA) in 3 HNSCC cell lines were detected by western blotting. #P > .05; *P < .05; **P < .01; ***P < .001.
Knockdown of HSP90AB1 Inhibited Glycolysis in HNSCC Cells
We performed GSEA analysis between high and low HSP90AB1 patients, and found that glycolysis might be correlated with the expression of HSP90AB1 in HNSCC (Figure 6A). It is known that the enhancement of aerobic glycolysis can promote the proliferation and migration of cancer cells,23,24 while inhibiting glycolysis can suppress malignant phenotypes of cancers.26,27 As a result, we conjectured that HSP90AB1 knockdown could cause inhibition of glycolysis in HNSCC cells. We analyzed the co-expression relationship between HSP90AB1 and various key proteins in the glycolysis pathway through GEPIA and found that the expression of HSP90AB1 was positively associated with the expression of phosphoglycerate kinase 1 (PGK1), enolase 1 (ENO1), pyruvate kinase M1/2 (PKM), and lactate dehydrogenase A (LDHA) (Figure 6B). Then, we detected the mRNA expression of 8 key glycolytic enzymes: hexokinase 2 (HK2), phosphofructokinase, liver type (PFKL), aldolase (ALDOA), PGK1, ENO1, enolase 2 (ENO2), pyruvate kinase M2 (PKM2), and LDHA in 3 cell lines via qRT-PCR. The results showed that except for ENO1 in the CAL27 cell line and PFKL in the JHU011 cell line, the mRNA expression levels of 8 key glycolytic enzymes were significantly decreased after HSP90AB1 knockdown in 3 HNSCC cell lines (Figure 6C). Moreover, we tested the content of several products and the activity of key glycolytic enzymes using assay kits. The results showed that in 3 HNSCC cell lines, the concentration of ATP, pyruvate (PA), and lactic acid (LD), and the activity of hexokinase (HK), pyruvate kinase (PK), and lactate dehydrogenase (LDH) were significantly reduced after HSP90AB1 knockdown (Figure 6D). These data suggested that the knockdown of HSP90AB1 suppressed the overall glycolysis level of HNSCC cells. The PI3K-Akt-mTOR pathway is an important upstream pathway of glycolysis. 30 Therefore, we detected the protein expression levels of pan-Akt and phospho-Akt (Ser473) in 3 HNSCC cell lines and found that there was no visible change in the expression of pan-Akt after HSP90AB1 knockdown, while the expression of phospho-Akt (Ser473) was significantly downregulated (Figure 6E). This result suggested that the decrease of glycolysis level after HSP90AB1 knockdown may be due to the inhibition of the PI3K-Akt-mTOR pathway caused by the degradation of phospho-Akt.

Knockdown of HSP90AB1 inhibited glycolysis in HNSCC cells. (A) GSEA analysis between high and low HSP90AB1 patients revealed that glycolysis might be correlated with the expression of HSP90AB1 in HNSCC. (B) Co-expression relationship between HSP90AB1 and several key glycolytic proteins (PGK1, ENO1, PKM, LDHA) in HNSCC was obtained through GEPIA. (C) Changes in mRNA expression levels of key glycolytic enzymes (HK2, PFKL, ALDOA, PGK1, ENO1, ENO2, PKM2, LDHA) in 3 HNSCC cell lines after HSP90AB1 knockdown were detected by qRT-PCR. (D) Changes in the content of important products (ATP, PA, LD) and activity of key glycolytic enzymes (HK, PK, LDH) in 3 HNSCC cell lines after HSP90AB1 knockdown were detected by corresponding assay kits. (E) The protein expression levels of pan-Akt and phospho-Akt (Ser473) in 3 HNSCC cell lines were detected by western blotting. Error bars represent the mean ± SD of 3 replicates. #P > .05; *P < .05; **P < .01; ***P < .001.
Discussion
In this study, we found that HSP90AB1 was up-regulated in HNSCC, and the expression of HSP90AB1 was related to T grade, lymph node metastasis, and prognosis of HNSCC. Knockdown of overexpressed HSP90AB1 inhibited the proliferation, migration, and glycolysis of HNSCC. HSP90AB1 is a molecular chaperone protein that can fold and stabilize client proteins, thus contributing to their normal activity. Additionally, most of its client proteins are important regulatory proteins involved in a variety of physiological and biochemical processes. 31 Increasing evidence shows that HSP90AB1 is highly expressed in cancers, may participate in stabilizing the reprogramming of oncogenic proteins, and plays a very important role in the occurrence and development of cancer.12,18 Taking lung cancer as an example, the study showed that the HSP90AB1 expression level in lung cancer was higher than that in normal tissues, and was related to the pathological grade, lymphatic infiltration, and prognosis of lung cancer. 19 The expression of HSP90AB1 in bronchoalveolar lavage fluid of patients with lung cancer was also significantly higher than that of patients with benign lung diseases and was closely related to the pathological grade and lymphatic infiltration of lung cancer. 32 In addition, the content of HSP90AB1 in the malignant pleural effusion of lung cancer patients was also related to the malignant biological behavior of the tumors. 33 However, the functions and mechanisms of HSP90AB1 in the progression of HNSCC are still unclear. We first confirmed the up-regulation of HSP90AB1 in HNSCC via bioinformatics, cell, and tissue experiments. Next, we verified that HSP90AB1 expression level is related to T grade, lymph node metastasis, and prognosis of HNSCC through bioinformatics and immunohistochemical analysis. We then constructed 3 stable HSP90AB1 knockdown HNSCC cell lines by lentiviral transfection rather than HSP90 inhibitors, thus excluding the effects of other subtypes of HSP90. Through the loss of function experiments, we confirmed that knockdown of HSP90AB1 inhibited the proliferation and migration of HNSCC cells at the molecular, cellular, and whole animal levels. Combined with previous studies, our data support the idea of a cancer-promoting role of HSP90AB1 in a variety of cancers.
Previous studies confirmed that the glycolysis level in HNSCC is significantly higher than in normal tissues.21,22 Many studies have shown that enhanced glycolysis can promote the malignant phenotype of cancer,23–25 and drugs that inhibit glycolysis have been used in clinical cancer treatment.26,27 Considering the extensive regulatory function of HSP90, we hypothesized that it also plays a regulatory role in aerobic glycolysis in malignant tumors. Only a few studies on the relationship between HSP90 and glycolysis in cancer have been carried out. Cao et al 34 confirmed the synergistic anticancer effect of HSP90 and glycolysis inhibitors in pancreatic cancer. Crouch et al 35 found that in breast cancer, ectopic HSP90 expression was the highest in glycolytic tumors, and HSP90 inhibition was more effective in glycolytic tumors. Subramanian et al 36 found that a novel C-terminal HSP90 inhibitor (KU757) can effectively treat lenvatinib-resistant cells by targeting glycolysis. Liu et al 37 reported that a class of inhibitors of the calcineurin-NFAT pathway (YZ129 and its derivatives) suppressed glycolysis by competitive binding with HSP90 in glioblastoma. These studies suggest a regulatory relationship between HSP90 and glycolysis in cancer. For the first time, our study demonstrated that knockdown of HSP90AB1 alone reduced the glycolysis levels in HNSCC cells. Unfortunately, because a specific method to enhance the overall level of glycolysis in cells currently doesn't exist, rescue experiments could not be carried out to verify that HSP90AB1 affects the proliferation and migration of HNSCC partly by regulating glycolysis. Some studies showed that HSP90 promoted the glycolysis and proliferation of cancers by directly interacting with key glycolytic proteins. 13 However, considering our research showed that after HSP90AB1 knockdown, glycolysis was suppressed as a whole rather than some specific glycolytic enzymes, we speculated that inhibition of HSP90AB1 suppresses glycolysis mainly by reducing the stability of key pathway proteins upstream of glycolysis and promoting their degradation, rather than directly acting on specific glycolytic proteins. Signaling upstream of glycolysis mainly includes the PI3K-Akt-mTOR pathway, HIF-1α, c-MYC, and mutant p53,30,38 and it was reported that HSP90 can interact with these proteins, stabilizing their structure and enabling their function.39–42 Therefore, we hypothesize that HSP90AB1 may maintain a high aerobic glycolytic level in HNSCC by stabilizing the key pathway proteins upstream of glycolysis, thus indirectly regulating the proliferation and migration of HNSCC. Here, we studied the expression changes of pan-Akt and phospho-Akt after HSP90AB1 knockdown, and we surprisingly found that the expression of phospho-Akt was significantly downregulated, while pan-Akt was not. Studies indicated that HSP90 could bind to and maintain the phospho-Akt, thus keeping the activation of the PI3K-Akt-mTOR pathway, while inhibition of HSP90 would cause the degradation of phospho-Akt.43,44 Our findings support and enrich this theory. Furthermore, previous studies usually reduced the expression of all subtypes of HSP90 through HSP90 inhibitors, while our study revealed that just knocking down HSP90AB1 alone can inhibit the expression of phospho-Akt. In addition, the activation of the PI3K-Akt-mTOR pathway is also an important factor promoting the proliferation and migration of cancer cells.44,45 Further studies are needed to fully reveal the deep mechanism.
In summary, this study systematically explored the expression characteristics and role of HSP90AB1 in HNSCC for the first time. The results supported HSP90AB1's role as a cancer-promoting gene which had been confirmed by other relevant studies, suggesting that HSP90AB1 had the potential to become a tumor marker and be used to evaluate the prognosis of HNSCC. We found that HSP90AB1 could participate in the progress of HNSCC by promoting proliferation and migration, revealing the anti-cancer target value of HSP90AB1. This study also confirmed the regulatory relationship between HSP90AB1 and glycolysis in cancer for the first time, and preliminarily explored the mechanism, revealing that HSP90AB1 might promote the glycolysis of HNSCC by maintaining the Akt pathway.
Conclusion
The expression of HSP90AB1 was up-regulated in HNSCC and was associated with T stage, lymph node metastasis, and prognosis. HSP90AB1 could promote the proliferation and migration of HNSCC and might promote the glycolysis of HNSCC by maintaining the Akt pathway. HSP90AB1 plays a role in promoting cancer in the development of HNSCC and has the potential to become a therapeutic target.
Supplemental Material
sj-docx-1-tct-10.1177_15330338221118202 - Supplemental material for HSP90AB1 Promotes the Proliferation, Migration, and Glycolysis of Head and Neck Squamous Cell Carcinoma
Supplemental material, sj-docx-1-tct-10.1177_15330338221118202 for HSP90AB1 Promotes the Proliferation, Migration, and Glycolysis of Head and Neck Squamous Cell Carcinoma by Hongbo Zhang, Xiteng Yin, Xinyu Zhang, Meng Zhou, Wenguang Xu, Zheng Wei, Chuanhui Song, Shengwei Han and Wei Han in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Authors’ Note
Hongbo Zhang and Xiteng Yin conceived of the presented idea, and carried out the experiments. Xinyu Zhang performed the statistical analysis. Wenguang Xu performed the bioinformatics analysis. Meng Zhou contributed to the statistical analysis and funding acquisition. Chuanhui Song and Zheng Wei contributed technical support and experimental guidance. Shengwei Han and Wei Han supervised all the work. All authors wrote and revised the initial manuscript and approved the submitted version.
Data Availability
The data generated during the study are available from the corresponding author on reasonable request.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics Approval
Institutional review board approval was obtained from the Nanjing Stomatological Hospital Ethics Committee (approval number: D2103022). All animal experiments and experimental protocols were in accordance with the Animal Care and Use Committee of the Medical School of Nanjing University (approval number: SYXK2019-0056).
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Jiangsu Province (grant no. BK20210080).
Informed Consent
Written informed consent was obtained from all individual participants included in the study.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
