Abstract
The CellSearch system is the only FDA approved and successful used detection technology for circulating tumor cells(CTCs). However, the process for identification of CTCs by CellSearch appear to damage the cells, which may adversely affects subsequent molecular biology assays. We aimed to explore and establish a membrane-preserving method for immunofluorescence identification of CTCs that keeping the isolated cells intact. 98 patients with lung cancer were enrolled, and the efficacy of clinical detection of CTCs was examined. Based on the CellSearch principle, we optimized an anti-EpCAM antibody and improved cell membrane rupture. A 5 ml peripheral blood sample was used to enrich CTCs with EpCAM immunomagnetic beads. Fluorescence signals were amplified with secondary antibodies against anti-EpCAM antibody attached on immunomagnetic beads. After identifying CTCs, single CTCs were isolated by micromanipulation. To confirm CTCs, genomic DNA was extracted and amplified at the single cell level to sequence 72 target genes of lung cancer and analyze the mutation copy number variations (CNVs) and gene mutations. A goat anti-mouse polyclonal antibody conjugated with Dylight 488 was selected to stain tumor cells. We identified CTCs based on EpCAM+ and CD45+ cells to exclude white blood cells. In the 98 lung cancer patients, the detection rate of CTCs (≥1 CTC) per 5 ml blood was 87.76%, the number of detections was 1–36, and the median was 2. By sequencing 72 lung cancer-associated genes, we found a high level of CNVs and gene mutations characteristic of tumor cells. We established a new CTCs staining scheme that significantly improves the detection rate and allows further analysis of CTCs characteristics at the genetic level.
Keywords
Introduction
According to the “Global Cancer Statistics Data for 2018” report published online by the American Journal of Cancer Society’s Journal of Clinician Cancer, lung cancer is the most malignant tumor with the highest morbidity and mortality worldwide. 1 The final diagnosis of lung cancer depends on histopathology. Lung cancer tissue is mainly obtained by lung biopsy, lung resection surgery and cytological samples. There are certain risks during the operations, and it is difficult to obtain lung cancer tissue. Liquid biopsy analysis may provide continuous molecular information on tumor lesions. This is particularly helpful for early detection of small lesions and personalized management of cancer patients, beneficial to stratify patients into targeted therapies and monitor treatment effects in real time, and determine treatment goals and drug resistance mechanisms.
Circulating tumor cells (CTCs) are tumor cells shed from the primary tumor, which pass through blood vessels or lymphatic vessels and reach distant tissues or organs through blood circulation. 2 Ashworth 3 first described the presence of circulating tumor cells in the peripheral blood of cancer patients in 1869. Subsequently, Paget 4 proposed the “seed and soil” hypothesis in 1889, while research interest in its clinical application has only expanded in recent years. Moreover, a new technology platform has been developed to isolate CTCs for follow-up analysis. CTCs in blood are extremely rare, only 1–10 per ml of peripheral blood, 5 which clearly limits the detection of CTCs. In the past 20 years with the development of cell isolation and identification technologies, detection of CTCs has become a research hotspot. Although circulating tumor DNA (ctDNA) is approved for epidermal growth factor receptor (EGFR) genetic testing for lung cancer, its information is limited. Compared with ctDNA, circulating tumor RNA, and exosomes, CTCs provide more complete information on the cell morphology, transcriptome, and proteome. CTCs have been detected in many types of cancer, including breast, prostate, lung, colorectal, and ovarian cancers. 6 -9 The detection, sorting, and downstream analysis of circulating tumor cells facilitate cancer diagnosis, prognosis judgment, and treatment strategy selection, 7 which represent the cutting-edge technology and key direction of cancer research.
CTCs are considered to be the cause of the transfer, and therefore many researchers are exploring their potential as prognostic indicators. The first observation indicating the prognostic value of CTCs was in 2004 using the CellSearch system to count CTCs in patients with metastatic breast cancer. The results showed that CTCs were related to progression-free survival and overall survival (OS). 10 Many studies have shown that quantification of CTCs can be used to predict the patient response to drugs and overall survival rate, 11 which is closely associated with OS rates of patients with metastatic lung cancer, 12 prostate cancer, 13 and breast cancer. 14
Enriched CTCs can be genomic amplified by fluorescence in situ hybridization (FISH), immunocytochemistry (ICC) for protein labeling, and reverse transcription polymerase chain reaction (RT-PCR)/quantitative RT-PCR (qRT-PCR) to quantify specific RNA and DNA sequences. To detect CTCs, ICC is used in most enrichment techniques, and there are various methods to characterize CTCs. 15 Among all enrichment methods, the most successful and widely used CTC technology is the CellSearch system approved by the Food and Drug Administration (FDA, USA) for prognostic evaluation of breast cancer in 2012. Its working principle is to enrich CTCs by epithelial marker epithelial cell adhesion molecule (EpCAM) immunomagnetic beads and then recognize CTCs with a fluorescent anti-cytokeratin (CK) antibody and exclude leukocytes with a fluorescent anti-CD45 antibody.
However, identification of CTCs by CK immunofluorescence and single cell whole exon sequencing have revealed that only 25%–80% of the exon region can be captured, 16 and the cells enriched using CellSearch appear to be more apoptotic and unlikely to generate a FISH signal. 17 Because CK is located in the cytoplasm, the membrane-breaking procedure to stain CK may cause degradation of genomic DNA, which will adversely affect subsequent molecular biological assays. Therefore, in this study, we omitted staining for the intracellular epithelial marker CK and identified CTCs using EpCAM magnetic beads to enrich CTCs and improve staining the cell membrane protein EpCAM to confirm its epithelial source properties. Exploring the use of membrane-preserving methods for immunofluorescence staining and identification of CTCs and incorporating EpCAM CTC separation technology will likely promote further in-depth genetic and molecular biology research.
Methods
Blood Specimens and Cell Preparation
114 patients were admitted to the Beijing Chest Hospital. Peripheral venous blood was collected from 3 healthy volunteers. Ethylene diamine tetraacetic acid (EDTA) anticoagulated venous blood (5 ml) was stored at room temperature and processed within 2 hours. This study was approved by the ethics committee of Beijing Chest Hospital. The lung cancer cell line NCI-H2009 was purchased from the Cell Resource Center of the Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences. NCI-H2009 cells were cultured in 10% FBS/RPMI-1640 medium.
Anti-CD326 (EpCAM) Antibody Staining
NCI-H2009 cells were collected, washed with phosphate buffer saline (PBS), and then incubated with or without 100 µl CD326 magnetic beads (Miltenyi Biotec Inc., Bergisch Gladbach, Germany) at 4°C for 30 minutes. After washing twice, the cells were incubated with anti-CD326 monoclonal antibodies CD326-FITC (3H) (prepared in-house), CD326-AF488 (9C4), CD326-Biotin (9C4) (Biolegend, San Diego, CA, USA), and CD326-Viobright 515 (Miltenyi Biotec Inc.), streptavidin-BB515 (BD Biosciences, San Jose, CA, USA) and anti-mouse Dylight 488 polyclonal antibody (pAb) (Ab98757, Abcam, Cambridge, MA, USA) at 1 µg each for 30 minutes at 4°C. Cells were washed twice, centrifuged at 300 × g for 5 minutes, and observed under a fluorescence microscope.
Anti-Mouse pAb and Leukocyte Binding Assay
To determine whether the pAb bind to white blood cells (WBCs). A 1 ml sample of anti-coagulated peripheral blood was collected from healthy individuals, red blood cells were lysed, and 100 µl CD326 magnetic beads was added, followed by mixing and incubation at 4°C for 30 minutes. After washing, the WBCs were resuspended with PBS. The WBCs were then incubated with 1 µg anti-mouse pAb-Dylight 488 at 4°C for 30 minutes. After washing twice in PBS, centrifuged at 300 × g for 5 minutes, the cells were resuspended and observed under a fluorescence microscope.
Detection of Lung Cancer Cells and CTCs in Peripheral Blood
In a spiking experiment, NCI-H2009 cells (100 cells) were added to 5 ml anti-coagulated peripheral blood from healthy individuals, erythrocytes were lysed, and the remaining cells were washed, centrifuged at 300 × g for 5 minutes, and then incubated with CD326 magnetic beads. Then, an auto MACS pro platform (Miltenyi Biotec Inc.) was used to enrich CTCs. Samples were incubated at 4°C for 30 minutes with 1 µg anti-mouse pAb-Dylight 488 and 1 µg Hoechst 33342 (Sigma, Saint Louis, Missouri, USA). After centrifugal washing (at 300 × g for 5 minutes), the samples were incubated for 30 minutes at 4°C with 2 µl anti-human CD45 antibody (Abcam) and 1 µg Hoechst 33342. After centrifugal washing (at 300 × g for 5 minutes), cells were transferred to a glass slide and a CellEctor Plus (Molecular Machines & Industries, Flughofstrasse, Glattbrugg, Switzerland) fluorescence microscope was used to identify lung cancer cells.
All patients with suspected or confirmed lung cancer were enrolled in this study for detection CTCs. CTCs were isolated from 5 ml ant-icoagulated peripheral blood of patients as described above. Then the CellEctor Plus (Molecular Machines & Industries) was used to identify immunofluorescence and isolate individual CTCs. Under the microscope, tumor cells from patients were identified by complete morphology, round or ellipse, and met the following fluorescent staining criteria: EpCAM+/Hoechst+/CD45− cells. The isolated single CTCs were stored at −80°C until further analysis.
Extraction, Amplification, and Quality Control of Whole Genomic DNA from CTCs
Whole genomic DNA of CTCs samples was extracted and amplified with a REPLI-g Single Cell Kit (cat. no. 150343; Qiagen GmbH, Hilden, Germany). The obtained whole genome DNA was specifically amplified by primers for 8 housekeeping genes: CYB5A forward, 5′-GGC AACGCT TAG ACT CTG TGT G-3′ and reverse, 5′-CTG CCC TTGGCC TAA CTA ACC T-3′; PRPH forward, 5′-GTT CCT CAAGAA GCT GCA CGA G-3′ and reverse, 5′-CGT TAG ACT CTGGAT CTG GCG T-3′; GABARAPL2 forward, 5′-CCA GCCAAT TCA TGA GTC GGT G-3′ and reverse, 5′-CCT GAC AACTCG CAA GTA GCA C-3′; ACTG1 forward, 5′-GCT CAATGG GGT ACT TCA GGG T-3′ and reverse, 5′-GTG GAC GTTACG TAA AAG GCC C-3′; NDUFA7 forward, 5′-TGC TCTGGA TGT GAA GAT GCC A-3′ and reverse, 5′-TTC CAG GTAAAT CCA GCC CAG G-3′; UQCRC1 forward, 5′-CAG CCAGTC AGC ATC ATC CAA C-3′ and reverse, 5′-GAA AGCCGG ATT GCG GTA ACA T-3′; MYC forward, 5′-GGA TAGCTC TGC AAG GGG AGA G-3′ and reverse, 5′-TCG TCGCAG TAG AAA TAC GGC T-3′; MIF forward, 5′-AGAAGT CAG GCA CGT AGC TCA G-3′ and reverse, 5′-GGCACG TTG GTG TTT ACG ATG A-3′. Agarose gel electrophoresis was used to observe the amplified bands. If ≥5 bands, the next step of NGS sequencing was performed.
Library Construction, Next-Generation Sequencing and Data Analysis
Total 20 CTC samples that matched WBCs with quality DNAs were obtained from 6 patients for 72 gene-targeted sequencing. The library was constructed using QIAseq Targeted DNA Human Lung Cancer Panel Kit (cat. DHS-005Z, Qiagen GmbH, Hilden, Germany) in accordance with the manufacturer’s protocol. Q-PCR method was used to accurately quantify the effective concentration of the library. The qualified libraries were carried out on the Illumina NovaSeq 6000 platform. The sequencing data quality control and variant calling steps were performed with QIAGEN GeneGlobe Data Analysis Center. Sufficient sequencing quality of each sample were ensured by checking them if it contained at least 5 million reads.
Results
Anti-CD326 Antibody Optimization
Five different fluorescent anti-CD326 antibodies were used for immunofluorescence staining of human lung cancer cell line NCI-H2009. There were 2 types: direct-labeled and enhanced signal antibodies. Direct-labeled antibodies were CD326-FITC (3H), CD326-AF488 (9C4), and CD326-Viobright515 and enhanced signal antibodies were CD326-Biotin + streptavidin-BB515 and anti-mouse pAb-Dylight 488. The immunomagnetic bead group without CD326 was directly stained with a fluorescent antibody. The CD326 magnetic bead group was first incubated with CD326 magnetic beads at 4°C for 30 minutes, washed twice, and then stained with a fluorescent antibody. The 5 anti-CD326 antibodies emitted fluorescent in the 2 groups with or without CD326 magnetic beads, and the fluorescence intensity of the CD326 magnetic bead group was weaker than that of the corresponding group without CD326 magnetic beads. The signal-enhanced biotinylated antibody group and polyclonal antibody group had significantly stronger fluorescence intensities than the direct-labeled fluorescent antibody group. However, in the biotinylated antibody group, the fluorescence intensity was decreased after adding CD326 magnetic beads. In contrast, the polyclonal antibody group (anti-mouse pAb-Dylight 488) had the best staining with clear cell contours (Figure 1A).
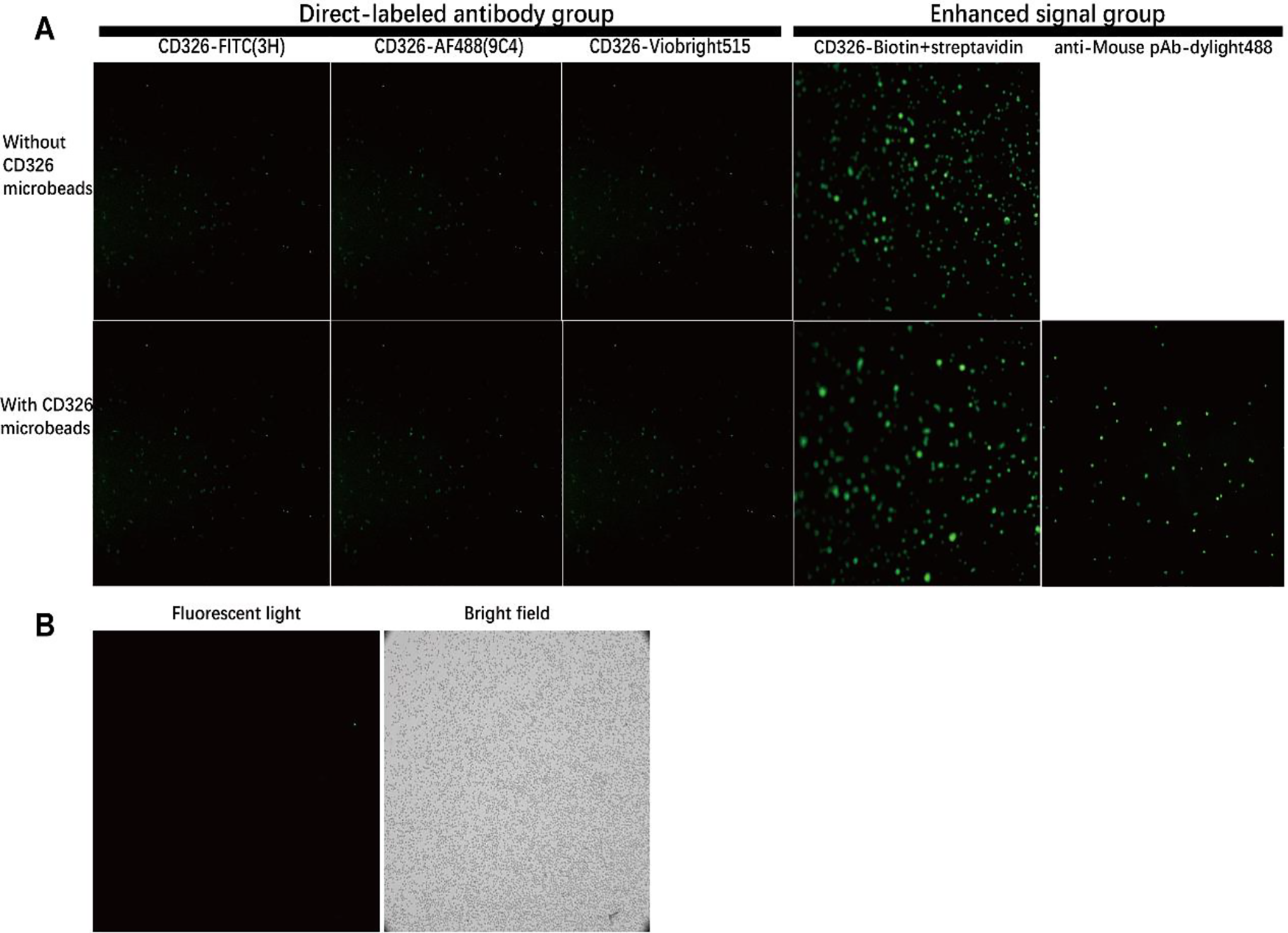
Immunofluorescence staining of human lung cancer cell line NCI-H2009 and peripheral blood leukocyte. A, The group without CD326 immunomagnetic beads was directly stained with fluorescent antibody. The group with CD326 immunomagnetic beads was incubated with CD326 immunomagnetic beads at 4°C for 30 min, washed twice by centrifugation and then stained with fluorescent antibody (10 × 10 times). B, Anti-mouse pAb-Dylight488 binding test with peripheral blood leukocyte. Erythrocytes were removed from human peripheral blood with erythrocyte lysate, and CD326 immunomagnetic beads were added for incubation at 4°C for 30 min. After washing for 2 times, anti-mouse pAb-Dylight488 was used for staining (10 × 10 times).
The selected fluorescent anti-CD326 antibody was used for immunofluorescence staining of peripheral blood CTCs, which may be affected by leukocytes in peripheral blood. To exclude binding of antibodies to peripheral blood leukocytes, we stained peripheral blood leukocytes with a fluorescent anti-CD326 antibody. Using a standard peripheral blood CTC-staining process, we found that peripheral blood leukocytes did not bind to the fluorescent anti-CD326 antibody, which ruled out the possibility of non-specific binding of anti-mouse pAb to leukocytes (Figure 1B). Finally, anti-mouse pAb-Dylight 488 was selected as the fluorescent antibody to detect CD326 protein on the surface of lung cancer cells.
CTCs Dyeing Scheme Optimized by Incorporation of NCI-H2009 Lung Cancer Cells
After optimization, EpCAM (goat anti-mouse pAb Dylight 488), CD45 [anti-CD45 antibody (MEM-28) APC], and nuclear dye Hoechst 33342 were used for staining. NCI-H2009 lung cancer cells were added to peripheral blood of healthy individuals. After erythrocyte lysis, CD326 magnetic beads enriched lung cancer cells, followed by staining with fluorescent antibodies (Figure 2). EpCAM staining was bright, the state was clear, the lung cancer cells all emitted fluorescence, and no non-specific binding to white blood cells was found. Figure 2 shows the results of EpCAM (CD326) staining, indicating that the use of mouse anti-human CD326 magnetic beads as the primary antibody and goat anti-mouse pAb (Dylight 488) as the secondary antibody avoided fluorescent anti-CD326 antibody and anti-CD326 magnetic bead epitope competition.

The lung cancer cell line NCI-H2009 spiked with peripheral blood was optimized for staining scheme. After enriched by CD326 immunomagnetic beads, stained with immunofluorescent antibody (goat anti-mouse pAb dylight 488), CD45 (anti-CD45 antibody [MEM-28] APC), nuclear dye Hoechst 33342 (20 × 10 times).

Fluorescence staining images of CTCs and white blood cell (WBC) in peripheral blood of lung cancer patients. The identification standard of CTCs is EpCAM+/Hoechst+/CD45−, and the identification standard of WBC is EpCAM−/Hoechst+/CD45+ (20 × 10 times).
Identification of CTCs in Patients With Lung Cancer
Peripheral blood of patients with lung cancer was enriched for CTCs by erythrocyte lysis and CD326 immunomagnetic beads, and then CTCs were detected using the optimized staining protocol described above. The criteria for CTCs were EpCAM+/Hoechst+/CD45−, and those for white blood cells were EpCAM-/Hoechst+/CD45+. Figure 3 shows the imaging of CTCs in patients with lung cancer under a fluorescence microscope.
Basic Information of Lung Cancer Patients
Peripheral blood CTCs were detected in 114 patients with suspected or confirmed lung cancer. Among them, 98 cases were diagnosed with lung cancer, 3 cases were diagnosed with other malignant tumors, 2 cases were diagnosed with infectious diseases, and 11 cases were undiagnosed. A follow-up analysis was performed on 98 patients with lung cancer. The basic information of patients was shown in Table 1.
Clinical Data of Patients With Lung Cancer.

CTC Clinical Test Results
Among 98 patients diagnosed with lung cancer, 85 patients had ≥1 CTCs in peripheral blood, and the overall detection rate was 87.76%. The number of CTCs detected was 1–36 with a median of 2, and 1–3 accounted for 70.6% (Figure 4A). Among the detected CTCs, the proportions of clinical stages were 13.0% of stage I patients, 6.0% of stage II patients, 21.0% of stage III patients, 60.0% of stage IV patients, 19.0% of stage I + II patients, and 81.0% of stage III + Ⅳ (Figure 4B). In each clinical stage, the detection rate of CTCs was 84.62% in stage I, 83.33% in stage II, and 85.71% in stage III, 89.66% in stage Ⅳ, 84.21% in stage Ⅰ + Ⅱ, and 88.61% in stage Ⅲ + Ⅳ (Figure 4C). Among the various pathological subtypes, the detection rates of CTCs were 90.48% for adenocarcinoma, 88.24% for squamous cell carcinoma, 80.00% for small cell carcinoma, and 75.00% for other types (Figure 4D).

Detection of peripheral blood CTCs in patients with lung cancer. A, The number of CTCs in the peripheral blood of patients with lung cancer and the number of patients; B, The distribution of clinical stages in patients with CTCs detected; C, The detection rate of CTCs in each clinical stage; D, The detection rate of CTCs in various pathological subtypes. AD, adenocarcinoma; SC, squamous cell carcinoma; SCC, small cell carcinoma.
CTCs Gene Variation Analysis
Single CTCs and paired WBCs were isolated for whole genomic DNA extraction and amplification. After quality control, we selected twenty CTCs samples and paired WBCs from 6 patients for targeted sequencing of 72 lung cancer-related genes and filtered out the same genetic variations in paired leukocytes. One WBC sequencing failure. Finally, effective data were obtained from fifteen CTC samples and paired WBC, and 5 CTC samples were controlled without WBCs. The number of gene mutations annotated in the Catalogue of Somatic Mutations in Cancer (COSMIC) database for a single CTC sample was between 1-50 (Table 2). Next, from the perspective of copy number variation (CNV) and gene mutation, CTCs were identified by gene-level variation.
Gene Mutations Detected in Single CTCs From Patients.

Determination of CTCs by Gene CNV
The normal cell copy number (CN) is 2. When CN is greater than or less than 2, especially when the difference is obvious, it is considered to be CNV. Among the 72 genes in the paired WBCs of each CTC sample, the number of genes with CN ≠ 2 was 29–72 (Figure 5D). In CTC 100-6 as an example, there were 59 genes with CN ≠ 2. Among them, CN was 17 for discoidin domain receptor 2 (DDR2), and 9 for tumor necrosis factor alpha induced protein 3 (TNFAIP3), and CN was obviously >2, which was consistent with the characteristics of tumor cells (Figure 5A–C). The presence of a large number of CNVs in each CTC sample was a manifestation of chromosomal instability. The CNV level variation was more obvious at the level of single tumor cells. Thus, CNV variation is an important feature of tumor cells.

CNV and somatic mutations of lung cancer-associated genes in CTCs samples. A, B and C are all from CNV analysis of the 100-6 CTCs samples versus paired leukocytes. A, Copy number (CN) ≠ 2 of all genes in the 100-6 CTCs samples. B and C are the CN of genes DDR2 and TNFAIP3, respectively, which are significantly higher than 2. D, The number of genes with copies ≠ 2 in 15 CTCs samples. E, distribution of the number of somatic mutations in single CTCs sample.
Determination of CTCs by Gene Mutation Sites
Each CTC sample was analyzed for 72 gene mutations (associated with lung cancer), which were filtered out in paired leukocytes to obtain a high-quality somatic mutation gene distribution. The numbers of somatic mutations in a single CTC sample was 1–16 (Figure 5E and Table 2). All CTC samples contained more than one somatic mutations of lung cancer-related genes, which confirmed CTCs at the level of gene mutations.
Discussion
More than 90% of cancer deaths are caused by cancer cell metastases. 18 CTCs are essential for the prognosis of tumor metastasis. Krebs et al reported that CellSearch detected 5 CTCs in 7.5 ml of blood from advanced NSCLC patients as an unfavorable prognostic factor, and changes in the number of CTCs after a single standard chemotherapy cycle predicted survival outcomes. 19 Small cell lung cancer is a solid tumor with the largest number of CTCs reported so far, 0–44896 cells per 7.5 ml of blood, and the corresponding CellSearch CTC count cutoff value was set to 50 CTCs per 7.5 ml of blood. 12 CTCs facilitate the prognosis and management of the disease and patients may benefit from alternative therapies during treatment. 10 We isolated CTCs based on CTC detection technology. In this process, we also determined the number of CTCs to be potentially used for clinical prognosis evaluation.
We used the classic CellSearch system staining scheme to break the membrane and stain CK, and found that the extracted genomic DNA was extremely poor, which was consistent a previous reports. 20 This seriously affected the subsequent analysis. Cells expressing EpCAM on the cell surface were still enriched for CTCs in this optimization because it is hardly expressed on leukocytes and expressed on the surface of most cancer cells derived from epithelial cells, while blood does not contain cells derived from epithelial cells in health individuals. When optimizing the staining scheme, the membrane-breaking step for staining was simplified. As for approximately 10% of cells were lost in one centrifugation, the experimental steps were simplified to reduce cell loss during the centrifugation, which may increase the recovery rate of cells. The increase of EpCAM signals by the polyclonal antibody not only shortened the experimental time, but also ensured a high detection efficiency. In particular, the quality of the extracted genomic DNA was improved significantly, which facilitated subsequent molecular biology analysis. In the spiking experiment, the recovery rate of lung cancer cells by the original membrane-breaking fixed cell method was 15%, whereas that of the improved method was 31%.
Punnoose et al reported that the CellSearch platform detected peripheral blood CTCs in patients with non-small cell lung cancer at a detection rate of 76%, median of 2, and range of 0–77. 21 Our detection rate was 87.76%, which is higher than their results. The median number of CTCs detected was the same (both 2). The detection rate of the CellSearch system generally does not exceed 40%. 20 The detection rate of CTCs was 22% in stage I–IIIA NSCLC patients, 22 the median detection rate was 21% in stage III–IV patients, 23 and the detection rate was 76% in stage IV patients. 24 Our detection rates were 84.21% in stage Ⅰ + Ⅱ patients and 88.61% in stage Ⅲ + Ⅳ patients, which were much higher than their results especially in early stage patients. Although a low cut-off value is set in our study (≥1 CTC), this is also related to the optimization of our immunofluorescence staining method. The heterogeneity of EpCAM protein expression on CTCs membranes may lead to changes in the ability to detect these cells in patients. By increasing the magnetic load of cells with a low antigen density, the difference in selection efficiency due to changes in the density of EpCAM antigens in CTCs can be reduced. 25 The fluorescent secondary polyclonal antibody used in this study significantly enhanced the staining and increased the detection rate of CTCs with low expression of EpCAM. The detection rate of CTCs in early lung cancer was higher, although usually less than 2 CTCs. However, the number of early lung cancer patients enrolled was small and CTCs detection in early lung cancer patients is worthy of further study. 22
Phylogenetic analysis confirmed that the evolution of CTCs and metastases originate from a part of the same specific branch. 26 CTCs enriched in the peripheral blood of breast cancer, melanoma, non-small cell lung cancer, and small cell lung cancer patients form tumors in immunodeficient mice, confirming their tumorigenic potential. 27 -29 At present, surgical treatment of stage I lung cancer is very effective, but the 5-year recurrence rate is still as high as 20%. 30 Our study found that the detection rate of CTCs in stage I patients was 84.62%. Although the number was small, the results suggested that CTCs already exist in the peripheral blood of patients with early lung cancer, which may lead to distant metastasis. This phenomenon may explain why early lung cancer remains after surgery and there is a possibility of relapse.
Potential limitations of this separation technology are as follows: The separation technology greatly improved the detection rate, but only CTCs expressing EpCAM protein were detected and CTCs that did not express EpCAM were undetected, such as epithelial-mesenchymal transitioned CTCs, interstitial CTCs, and tumor stem cells. CTCs can also express CD45 protein. 31 and our detection method excluded such cells. Additionally, although the detection rate was improved, the number of detected CTCs was not increased significantly with 1–3 CTCs accounting for the majority, which is similar to previous data from the CellSearch system. Further research is still needed to increase the detection rate of more phenotypic CTCs. The separation technology was based on epithelial identification. The isolated CTCs may be mixed with epithelial cells contaminating blood during collection. Chemi et al sequenced 6 CTCs isolated by the CellSearch system and identified 3 as circulating epithelial cells. 26 Additionally, in patients with benign colon disease, both the CellSearch system and CK19-EPISPOT test detected circulating epithelial cells that met the “tumor cell” standard. 32 Therefore, the criteria for epithelial labeling to identify CTCs at the protein level are not completely reliable including non-driving gene mutation, so circulating epithelial cells mistaken for CTCs cannot be ignored. We detected gene mutations by next-generation sequencing and confirmed that CTCs were tumor cells in terms of gene mutation sites and CNV. The 11 tested CTC samples consisted of all tumor cells and no circulating epithelial cells were found. Although the detected 72 gene variants indicating lung cancer cell properties were filtered using white blood cell controls, the tumor cell properties of individual CTCs may be excluded, especially CTCs with few mutations.
In summary, using the principle of the CellSearch system after optimization, we established a new CTC separation technology based on EpCAM expression, which improves the detection rate of CTCs and extraction quality of whole genomic DNA from cells. It is also suitable for further molecular biology research and subsequent sequencing.
Footnotes
Abbreviations
Acknowledgments
We would like to acknowledge Xiaojue Wang, Zhuohong Yan, Ling Yi, Bin Yang, Panjian Wei, Chang Jiang for their contributions during the execution of this study.
Ethics Approval
This study was approved by the ethics committee of Beijing Chest Hospital (reference YJS-2019004).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants of “Study on the Application of Clinical Characteristics of Capital” to HTZ (no. Z151100004015104) and to SCZ (no. Z161100000516107) from Beijing Municipal Science and Technology Project, Beijing University’s high-precision discipline construction project, Capital Medical University and Peking Union Medical College clinical medicine discipline co-construction project (11920703), Research on Key Technologies of Lung Cancer Diagnosis and Biotherapy in Beijing Health Planning Commission (PXM2018_026271_000002).
