Abstract
Background:
Osteosarcoma is the most leading primary malignancy of the bone in adolescents all over the world. Long non-coding RNA (lncRNA) 91 H has been reported to participated in multiple cancers. Meanwhile, lncRNA 91 H has been proved to be upregulated in osteosarcoma. However, the function of 91 H in osteosarcoma remains unclear.
Methods:
Gene and protein expressions in osteosarcoma cells were detected by qRT-PCR and western blot, respectively. Cell viability was tested by CCK-8 assay. Ki67 staining was used to measure cell proliferation. Cell apoptosis and cycle were assessed by flow cytometry. In addition, transwell assay was used to detect cell migration and invasion. Furthermore, Methylation-specific PCR (MSP) was performed to test the methylation of CDK4 promoter. Finally, xenograft mice model was established to explore the role of 91 H in osteosarcoma in vivo.
Results:
Knockdown of 91 H significantly inhibited the growth of osteosarcoma cells via inducing the cell apoptosis. In addition, 91 H siRNA notably suppressed the migration and invasion of osteosarcoma cells. Meanwhile, knockdown of 91 H inhibited the progression of osteosarcoma via inducing methylation of CDK4 promoter. Furthermore, 91 H knockdown obviously induced G1 arrest in osteosarcoma cells via inhibition of PCNA and Cyclin D1. Finally, knockdown of 91 H notably inhibited the tumor growth of osteosarcoma in vivo.
Conclusion:
knockdown of 91 H suppressed the tumorigenesis of osteosarcoma via inducing methylation of CDK4 promoter in vitro and in vivo. Thus, 91 H may serve as a new target for the treatment of osteosarcoma.
Keywords
Introduction
Osteosarcoma is the most leading primary malignant bone tumor of mesenchymal origin which character with osteoblastic differentiation and malignant osteoid formation. 1 In addition, osteosarco ma is usually accompanied by local swelling and pain. 2,3 Nowadays, osteosarcoma tends to occur in adolescent who ages from 10 to 25 years. 4 However, the pathogenesis of osteosarcoma remains unclear. The current treatments contain intricate limb-sparing surgeries and chemotherapy. 5 Despite the great advances in osteosarcoma treatment, patients still diagnosed with metastasis and relapse accompanied by low survival rate. 6 Hence, it is necessary to explore effective ways for the treatment of osteosarcoma.
Long non-coding RNAs (lncRNAs) are a class of non-coding RNA transcripts with the length of 200 nucleotides. 7 LncRNAs often play key roles during the progression of multiple diseases. 8 -10 In addition, recent studies have shown a close correlation between lncRNAs and osteosarcoma. 4,11 For example, silencing of lncRNA XIST could promote the migration and invasion of osteosarcoma cells via mediation of miR-153/SNAI1 axis. 12 Meanwhile, lncRNA 91 H is known to be upregulated in osteosarcoma. 13 Moreover, it has been reported that lncRNA 91 H could promote the tumorigenesis of colorectal cancer, 14,15 and acted as a promoter in breast cancer. 16 However, the mechanism by which 91 H regulates the tumorigenesis of osteosarcoma remains unclear.
In this research, we aimed to investigate the function of 91 H in osteosarcoma with the purpose of exploring new molecular target for the treatment of osteosarcoma.
Material and Methods
Cell Culture
Osteosarcoma cell lines (MG63 and U2OS) were obtained from American Type Culture Collection (ATCC, Manassas, VA, USA). All cell lines were cultured in RPMI-1640 medium (Gibco, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (Gibco) and penicillin (100 U/mL). In addition, cells were cultured in the condition of 37°C and 5% CO2.
Tissue Collection
Five pairs of osteosarcoma tissues and adjacent normal tissues were collected from General Hospital of Ningxia Medical University between August 2020 and December 2020. Clinical and pathological data and tissue collection of these patients were collected with their written informed consent. This research was approved by the Institutional Ethical Committee of General Hospital of Ningxia Medical University.
Quantitative Real Time Polymerase Chain Reaction (qRT-PCR)
Total RNA was extracted from osteosarcoma cell lines using TRIzol reagent (TaKaRa, Tokyo, Japan) according to the manufacturer’s protocol. cDNA was synthesized using the reverse transcription kit (TaKaRa, Ver.3.0) according to the manufacturer’s protocol. qRT-PCR was performed in an ABI7500 system using SYBR Green methods. Moreover, qRT-PCR was performed in triplicate with the following protocol: 2 min at 94°C, followed by 35 cycles (30 s at 94°C and 45 s at 55°C). The primers for all the genes were obtained from GenePharma (Shanghai, China). 2-△△CT method was used to quantify the data. The internal reference gene (β-actin) was used for normalization. The primers for 91 H were: forward: 5’-GACCGTTGTTAGCACGCCTT-3’ and reverse: 5’-CACGTATCGGTCCGTGTTGG-3’; the primers for CDK4 were: forward: 5’-AGGGCGGAGAGAACGGTG-3’ and reverse: 5’-AACTGGGTCCTGTGGTGCGTA-3’ and the primers for β-actin were: forward: 5’-CCTGCGAAACACCTTGATCG-3’ and reverse: 5’-TCGTCATGTTCCCCACTTCG-3’.
CCK-8 Assay
Cell counting kit-8 assay (CCK-8, Beyotime, Shanghai, China) was used for testing the cell viability. MG63 or U2OS cells were plated into 96-well plates with a density of 5 × 103 cells per well and treated with negative control (NC) or 91 H siRNA2 for 0, 24, 48 and 72 h, respectively. Then, osteosarcoma cells were treated with 10 µl CCK-8 reagents for another 2 h at 37°C. Finally, the absorbance of cells was measured at 450 nm using a microplate reader (Thermo Fisher Scientific, Waltham, MA, USA).
Cell Transfection
siRNAs targeted against 91 H (91 H siRNA1 and 91 H siRNA2; 10 nM) and a negative control siRNA (NC) were purchased from Guangzhou RiboBio Co., Ltd. and transfected into osteosarcoma cells (5 × 103) using Lipofectamine® 2000 (Thermo Fisher Scientific, Inc.), according to the manufacturer’s instructions. Cells were incubated at 37°C for 6 h before subsequent experiments were performed. Transfection efficiency was determined using reverse transcription-quantitative PCR (qRT-PCR). The sequences of the siRNAs were as follows: Negative control siRNA, 5’-UUCUCCGAACGUGUCACGUTT-3’; 91 H siRNA1, 5’-CCGGUUGGACAAUGAGCCC-3’; and 91 H siRNA2, 5’-CCCTTUGCGGGGGCGAGT-3’.
CDK4 overexpression plasmid was obtained from GenePharma (Shanghai, China). Osteosarcoma cells were transfected with CDK4 overexpression plasmid (pcDNA3.1-CDK4, Genepharma, Shanghai, China) or empty vector (pcDNA3.1, Genepharma, Shanghai, China) by using lipofectamine 2000 (Invitrogen) in accordance with the manufacturer’s instructions. The transfection efficiency was determined by western blot.
Western Blotting
Osteosarcoma cells were isolated from cell lysates by using RIPA buffer and quantified by BCA protein assay kit (Beyotime, Shanghai, China). Proteins were separated with 10% SDS-PAGE and transferred onto PVDF (Bio-Rad) membranes. After blocking with 3% skim milk for 1 h, the membranes were incubated with primary antibodies at 4°C overnight. The primary antibodies were as follows: anti-Bax (1:1000, Abcam, CA, USA), anti-XIAP (1:1000, Abcam), anti-cleaved caspase 3 (Abcam, 1:1000), anti-CDK4 (Abcam, 1:1000), anti-PCNA (Abcam, 1:1000), anti-cyclin D1 (Abcam, 1:1000) and β-actin (Abcam, 1:1000). Then, the membranes were incubated with horseradish peroxidase (HRP)-conjugated goat anti-rabbit IgG polyclonal secondary antibody (Abcam, 1:5000) at room temperature for 1 h. Membranes were scanned by using an Odyssey Imaging System and analyzed with Odyssey v2.0 software (LICOR Biosciences, Lincoln, NE, USA). β-actin was used as an internal control.
Cell Apoptosis Analysis
MG63 or U2OS cells were trypsinized, washed with phosphate buffered saline and resuspended in Annexin V Binding Buffer. After that, cells were stained with 5 µl Annexin V and propidium (PI) in the dark for 15 min. Cells were analyzed using flow cytometer (BD, Franklin Lake, NJ, USA) to test the cell apoptosis rate.
Immunofluorescence
Osteosarcoma cells were seeded in 24-well plates overnight. Next, cells were treated with NC or 91 H siRNA2 for 72 h. Cells were blocked with 10% goat serum for 30 min at room temperature and then incubated with anti-Ki67 antibody (Abcam; 1:1000) at 4°C overnight. After that, cells were incubated with goat anti-rabbit IgG (Abcam; 1:5000) at 37°C for 1 h. The nuclei were stained with DAPI (Beyotime, Shanghai, China) for 5 min. Finally, cells were observed under a fluorescence microscope (Olympus CX23, Tokyo, Japan).
Transwell Assay
Briefly, 24-well Transwell plates (Corning, New York, NY, USA) were used for cell invasion and migration detection. For the cell migration assay, 2 × 105 osteosarcoma cells were seeded into the upper chambers of the 24-well plates in 200 µL of serum-free RPMI 1640 medium supplemented with 0.2% bovine serum albumin. The lower chambers contained RPMI 1640 medium supplemented with 1% FBS. After 24 h of incubation at 37°C, the non-migrating cells were gently removed from the upper side of each chamber with a cotton swab, while the cells that had migrated were fixed with 95% alcohol for 10 min and stained with 0.1% crystal violet (Sigma, Grand Island, NY, USA) for 5 min. Finally, images were captured, and the cells were counted under an inverted light microscope (Olympus) at 400 x magnification.
For the invasion assay, the upper chambers of the 24-well plates were pretreated with 50 µL of Matrigel (12.5 mg/L), and the wells were pretreated with Matrigel (BD Biosciences, Franklin Lake, NJ, USA). Then, osteosarcoma cells (1 × 106 cells/ml) in FBS-free medium were seeded into the upper chambers. The lower chambers contained RPMI 1640 medium supplemented with 1% FBS. The cells were incubated at 37°C for 24 h, and cells that attached to the underside of the membrane were fixed and stained with a 0.1% crystal violet solution. Finally, images were captured, and the number of invading cells was counted under a microscope at 400 x magnification.
Methylation-Specific PCR (MSP)
For MSP detection, a pair of primers for amplifying methylated CpG targets was obtained based on the CpG island sequence of the CDK4 promoter-proximal elements. The sequences were as follows: 5’-TAGTTAGTTACGTGTTCGTCGCG-3’ (forward) and 5’-AACTACCAAAAAAACGCGCG-3’ (reverse). Other sequences of primers (performed to amplify unmethylated CpG targets) were as follows: 5’-TGGTAGTTAGTTATGTGTTTGTTGTG-3’ (forward) and 5’-ACCAACTACCAAAAAAACACACA-3’ (reverse). MSP was performed via a standard PCR machine. PCR products were resolved by agarose gel electrophoresis and stained with EB.
Cell Cycle Distribution
Osteosarcoma cells were treated with NC or 91 H siRNA2 for 72 h. Then, the cell cycle distribution was detected by flow cytometry (BD) as previously described. 17 Briefly, cell cycle detection was performed using Cycletest Plus DNA Reagent Kit (BD, NJ, USA). Osteosarcoma cells were harvested and counted with a hemocytometer. Cells (5 × 105) were fixed, permeabilized, and stained in accordance with the manufacturers’ instructions, and the sample was analyzed by flow cytometry using a FACSCalibur measuring FL2 area versus total counts. The data were analyzed using ModFit (http://mycyte.org/) and FlowJo (http://mycyte.org/) softwares to generate the percentages of cells in G1, S, and G2 to M phases of the cell cycle.
In Vivo Study
BALB/c nude mice (6-8 weeks old) were purchased from Vital River (Beijing, China). The mice were housed within a dedicated SPF facility (4 mice per group). Osteosarcoma cells were subcutaneously transplanted into mouse according to the previous reference. 18 When the tumor volume reached about 150 mm3, control or 91 H siRNA2 (50 nM) was injected intra-tumor twice weekly. The tumor volume was measured weekly according to the formula: Length×Width×Width/2. At the end of the experiments, mice were sacrificed and the tumors were collected and weighted. All in vivo experiments were performed in accordance with National Institutes of Health guide for the care and use of laboratory animals, following a protocol approved by the Ethics Committees of General Hospital of Ningxia Medical University.
Statistical Analysis
All experiments were performed in at least 3 independent experiments, and all data are presented as the means ± standard deviation (SD). The comparison between 2 groups was analyzed by unpaired Student’s t-test. The comparisons among multiple groups were made with one-way analysis of variance (ANOVA) followed by Tukey test (GraphPad Prism7). P < 0.05 was considered to indicate a statistically significant result.
Results
Knockdown of 91 H Significantly Suppressed the Proliferation of Osteosarcoma Cells
To investigate the role of 91 H in osteosarcoma, qRT-PCR was performed. As indicated in Figure 1A, the expression of 91 H was significantly upregulated in osteosarcoma tissues, compared with adjacent normal tissues. Meanwhile, the expression of 91 H in osteosarcoma cells was significantly downregulated by 91 H siRNAs (Figure 1B). Since 91 H siRNA2 exhibited better transfection efficiency, this siRNA was used in following experiments. In addition, silencing of 91 H notably decreased the viability of osteosarcoma cells (Figure 1C). Furthermore, the result of Ki67 staining revealed that 91 H siRNA2 greatly inhibited the proliferation of osteosarcoma cells (Figure 1D and 1E). All these data showed that knockdown of 91 H significantly suppressed the proliferation of osteosarcoma cells.

Silencing of 91 H Notably Inhibited the Migration and Invasion of Osteosarcoma Cells
In order to detect the effect of 91 H on cell migration and invasion, transwell was used. The data indicated the migration of osteosarcoma cells was notably inhibited by 91 H siRNA2 (Figure 2A). Consistently, knockdown of 91 H significantly suppressed the invasion of osteosarcoma cells (Figure 2B). Taken together, silencing of 91 H notably inhibited the migration and invasion of osteosarcoma cells.

91 H siRNA2 Notably Induced the Apoptosis of Osteosarcoma Cells
To investigate the effect of 91 H on cell apoptosis, flow cytometry was performed. As revealed in Figure 3A and 3B, knockdown of 91 H obviously induced the apoptosis of osteosarcoma cells. Moreover, U2OS cells were more sensitive to 91 H siRNA2. Therefore, U2OS cells were selected of use in the subsequent experiments. Meanwhile, the expressions of Bax and cleaved caspase 3 in U2OS cells were significantly upregulated by 91 H siRNAs (Figure 3C-3E). In contrast, 91 H knockdown significantly decreased the expression of anti-apoptotic protein (XIAP) in U2OS cells (Figure 3C and 3F). To sum up, 91 H siRNA2 notably induced the apoptosis of osteosarcoma cells.

91 H siRNA2 Suppressed the Tumorigenesis of Osteosarcoma via Inducing the Methylation of CDK4 Promoter
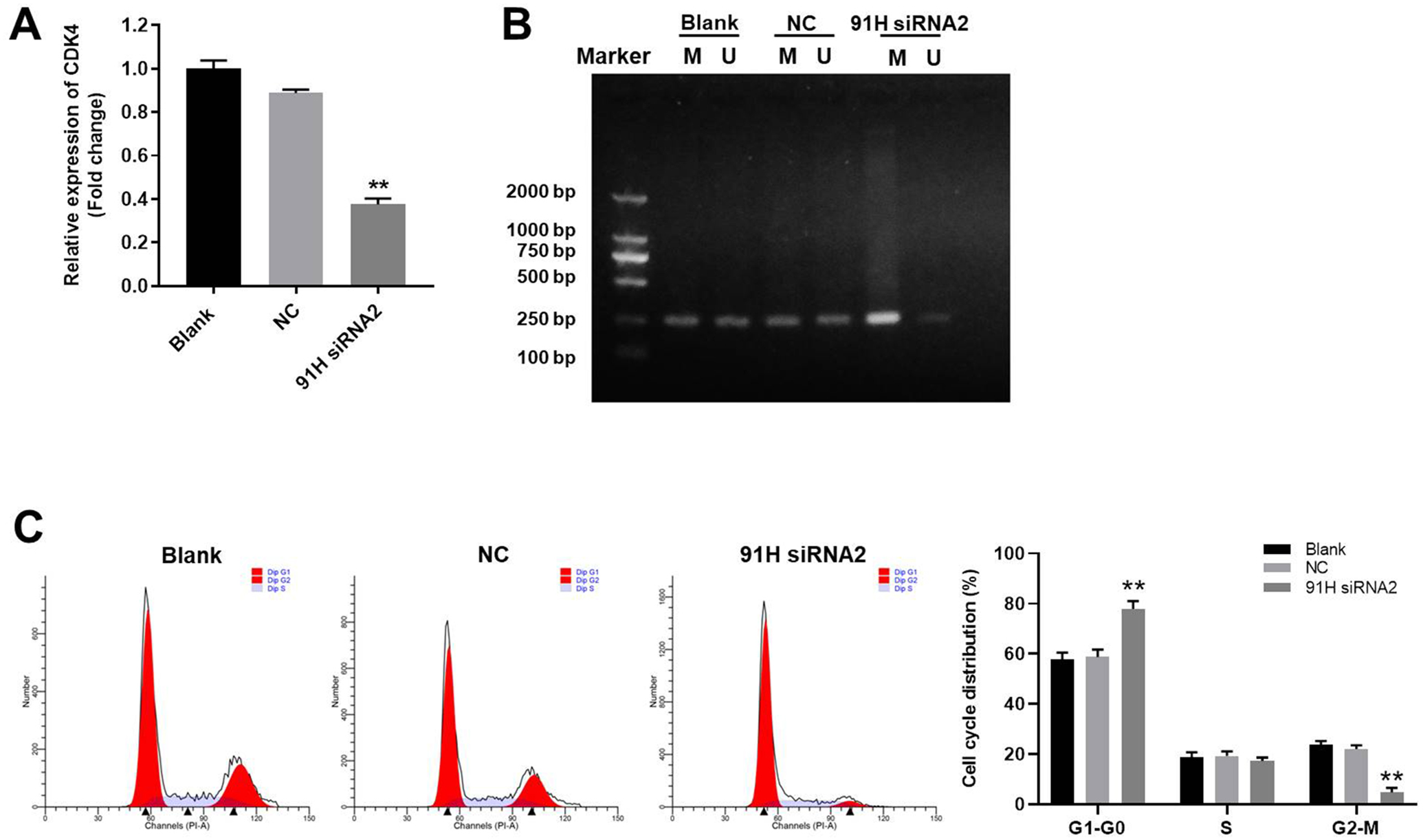
Next, qRT-PCR was used to detect the gene expression. The result showed that expression of CDK4 in U2OS cells was notably decreased by 91 H siRNA2 (Figure 4A). In addition, the data of MSP indicated that the methylation of CDK4 promoter in U2OS cells was greatly induced by 91 H siRNA2 (Figure 4B). Since CDK4 is known to be a key mediator in cell cycle, 19 flow cytometry was then used to detect the cell cycle distribution. As we expected, knockdown of 91 H notably induced G1 arrest in U2OS cells (Figure 4C). All these data suggested that 91 H siRNA2 inhibited the growth of osteosarcoma cells via inducing the methylation of CDK4 promoter.
91 H siRNA2 Induced G1 Arrest in Osteosarcoma Cells Via Downregulation of Cyclin D1, PCNA and CDK4
In order to explore the mechanism by which 91 H siRNA2 mediated the cell cycle distribution of osteosarcoma, western blot was performed. As revealed in Figure 5A-5D, the expressions of CDK4, Cyclin D1 and PCNA in U2OS cells were significantly downregulated in the presence of 91 H siRNA2. To sum up, 91 H siRNA2 induced G1 arrest in osteosarcoma cells via downregulation of Cyclin D1, PCNA and CDK4.

Knockdown of 91 H Significantly Inhibited the Tumor Growth of Osteosarcoma In Vivo
Next, xenograft mice model was established to investigate the role of 91 H during the tumorigenesis of osteosarcoma in vivo. As revealed in Figure 6A and 6B, tumor sizes of mice were significantly decreased by 91 H knockdown. Moreover, 91 H knockdown notably reduced the tumor weight in mice (Figure 6C). Taken together, knockdown of 91 H significantly inhibited the tumor growth of osteosarcoma in vivo.

91 H Knockdown-Induced Cell Growth Inhibition Was Partially Reversed by CDK4 Overexpression
In order to confirm the role of 91 H in osteosarcoma, rescue experiment was performed. As demonstrated in Figure 7A, the expression of CDK4 in osteosarcoma cells was notably upregulated by CDK4 overexpression. Moreover, viability of osteosarcoma cells was significantly inhibited by knockdown of 91 H, while this phenomenon was partially reversed in the presence of CDK4 overexpression (Figure 7B). In addition, 91 H knockdown-induced cell apoptosis was notably inhibited by CDK4 OE (Figure 7C). Consistently, the expression of CDK4 in osteosarcoma cells was significantly inhibited by 91 H knockdown, while this phenomenon was reversed by CDK4 overexpression (Figure 7D). Altogether, 91 H knockdown-induced cell growth inhibition was partially reversed by CDK4 overexpression.

Discussion
It has been reported that 91 H plays key roles in some cancers. 14,20,21 In addition, 91 H has been confirmed to be upregulated in osteosarcoma. 22 The present study found that knockdown of 91 H significantly inhibited the tumorigenesis of osteosarcoma via regulation of CDK4 promoter methylation. Thus, this research further explored the function of 91 H in osteosarcoma, suggesting that 91 H could act as a key mediator in osteosarcoma. According to a previous reference, 16 91 H could exert oncogenic properties in breast cancer. Our study was consistent to this research, indicating that 91 H might act as a promoter in malignancies.
Bax, XIAP and cleaved caspase 3 are known to be key mediators during cell apoptosis. 18,23,24 Meanwhile, Bax and cleaved caspase 3 are regarded as a promoter during apoptosis, 25,26 while XIAP plays an inhibitory role. 27 Zhong GB et al found that SNHG8 promoted the proliferation of osteosarcoma cells via mediation of Bax, XIAP and cleaved caspase 3. 28 Consistently, the present study suggested that 91 H siRNA2 induced the apoptosis of osteosarcoma cells via upregulating Bax and cleaved caspase 3 and downregulating XIAP. CDK4 is considered as an important mediator in cell growth. 29 In addition, CDK4 has been confirmed to be correlated with the tumorigenesis of osteosarcoma. 30,31 We also found 91 H knockdown could promote CDK4 methylation in osteosarcoma cells. Meanwhile, according to Zhang J et al, 32 piperine inhibited the tumorigenesis of osteosarcoma via mediation of CDK1. CDK1 is a key mediator in G2/M phase cell cycle. 33 However, 91 H siRNA2 induced G1 phase arrest in osteosarcoma cells. The different function between 91 H and piperine may due to their different roles in tumorgenesis.
Frankly speaking, there are some limitations in this research. First, the correlation between miRNAs and 91 H in tumorigenesis of osteosarcoma remains unclear. In addition, the interaction between miRNAs and CDK4 methylation is not clear yet. Thus, more investigations are needed in the future. In conclusion, knockdown of 91 H significantly inhibited the tumorigenesis of osteosarcoma via methylation of CDK4 promoter. Therefore, 91 H might serve as a key mediator during the tumorgenesis of osteosarcoma.
Footnotes
Author Contribution
Study design, literature research, experimental study was performed by Suoli Cheng, Jianping Zheng and Jiandang Shi; data acquisition, data analysis and statistical analysis were performed by Suoli Cheng and Jiandang Shi. All the authors were responsible for guarantor of integrity of entire study, manuscript preparation and manuscript editing. Jiandang Shi finished the submission.
Suoli Cheng and Jianping Zheng are authors contributed equally to this research.
Jiandang Shi have PhD degree.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Statement
In vivo study was approved by the Ethics Committees of General Hospital of Ningxia Medical University (approval no. GHNMU20200728). In addition, this research does not include human samples.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Ningxia Natural Science Foundation Project (No. 2019AAC03170, No. 2018A0854 and No. 2019AAC03193) and Ningxia Hui Autonomous Region Key R&D Plan (No. 2019BEG03036).
