Abstract
Growing evidence has suggested that CD155 participates in the regulation of many biological processes ranging cell growth, invasion, and migration from regulation of immune responses in most malignances. However, the impact of prognostic value and CD115-related immune response on the survival in multiple cancers remains incompletely clear. In our study, we assessed the prognostic significance and immune-associated mechanism of CD155 based on data from multiple databases and methods, including UCSC Xena, Oncomine, PrognoScan. We identified that CD155 was commonly upregulated in most human cancers, and High expression of CD155 was closely correlated with unfavorable clinical outcomes in 10/33 of human cancers, while CD155 at low level was responsible for better survival in KICH and PAAD. More intriguingly, CD155 expression had a significant interaction with immune function in several tumors by analyzing Tumor mutational burden and microsatellite in stability, immune score and stromal score. The correlation between immune infiltration and CD155 expression also indicated that CD155 expression positively correlated with CD4+ T cells in Head and Neck squamous cell carcinoma, Lung adenocarcinoma and Colon adenocarcinoma, while had inversely interaction with CD8+ T in Kidney renal clear cell carcinoma and Head and Neck squamous cell carcinoma as well as Tregs in Skin Cutaneous Melanoma, Head and Neck squamous cell carcinoma and Bladder Urothelial Carcinoma. These findings indicate CD155 correlates with cancer immunotherapy function. In conclusions, our observations revealed CD155 might function as immune-associated system in the development of human cancers, and acted as a promising prognostic and therapeutic target against human cancers.
Introduction
Cancer is one of main causes of morbidity and mortality globally among people in both high-income countries and less developed countries. 1 -4 Based on agency for Research on Cancer (IARC), there may be an increase in 22 million cancer cases and 13 million cancer-related deaths in 2030. 5 Recently, immunotherapy has reported to be now a remarkably effective therapeutic approach for anti-tumor. 6 -11 However, the impact of immune response on the survival of multiple cancers remains incompletely clear; therefore, it is crucial to identify immune-related prognostic biomarkers.
Poliovirus receptor (PVR, CD155), a member of the immunoglobulin (Ig) superfamily, participates in the regulation of many biological processes ranging cell growth, invasion, and migration from cell adhesion, and modulation of immune responses in the development of malignances. 12 -15 PVR is significantly upregulated in most human cancers, and its high activity tightly correlates with poor prognosis. 16 -18 Recently, PVR, which has been identified by receptors expressed on T and NK cells, is also recognized as a promising immunotherapy target. 19 -22 It has been proposed that in the tumor microenvironment the balance between CD155/DNAM-1and CD155/TIGIT/CD96 contrasting signals contributes to modulate NK cell effector functions. 23 -27 These results display that CD155 has functionally indispensable roles in T and NK cells. However, the deeper mechanisms understanding of CD155 in tumor progression deserves further explored.
In this present study, we investigated the expression profiling and prognostic value of CD155 in multiple types of cancer based on several databases and identified the interaction of CD155 with Tumor mutational burden (TMB), microsatellite in stability (MSI), immune score, stromal score and tumor-infiltrating immune cells in the different tumor microenvironments.
Material and Methods
Patients and Specimens
The RNA-Seq FPKM data, somatic mutation data, and corresponding clinical data with multiple human cancers of The Cancer Genome Atlas (TCGA) were obtained from UCSC Xena (https://xena.ucsc.edu/). 28 We collected expression profile data of gene in 33 solid tumors, more than 10000 samples from TCGA project.
Oncomine Database Analysis
We used the Oncomine database, which included 715 gene expression datasets from 86,733 caners and normal samples, to examine different expression of CD155 in distinct types of malignances (http://www.oncomine.org). 29 The results obtained from the database are set up with threshold of p-value < 0.05, fold change >1.5, and gene ranking of Top 10%.
Survival Analysis and Clinical Correlation Analysis
The Survival package of R was utilized to identify the prognosis value of CD155 involved with different types of malignances. 30 This test was based on the Kaplan-Meier method and Univariate Cox regression analysis, respectively. Hazard ratio (HRs) and 95% confidence intervals (CIs) were appied with describe the relative risk. P < 0.05 was considered as a statistical difference. Clinical stage was used to analyze the correlation between gene expression and clinical characteristics.
PrognoScan Database Analysis
The interaction between survival time of patients in human cancers and elevated CD155 expression was determined by the PrognoScan database (http://www.abren.net/PrognoScan/). 31 Hazard ratio (H.R.s) and 95% confidence intervals (CIs) were used to describe the relative risk. Corrected P-value < 0.05 was regarded as a statistically significant difference.
Tumor Mutational Burden (TMB) and Microsatellite in Stability (MSI) Analysis
TMB and MSI have also been recognized as biomarkers in the response of anti-PD1/PD-L1 monotherapy outcomes. 32 TMB of each patient was calculated by somatic mutation data of 33 cancers. MSI data download from the research of Bonneville etc. 33
CD155 Modulate the Tumor Microenvironment in Human Cancers
The tumor microenvironment of 33 kinds of cancer were evaluated by the “ESTIMATE” R package. 34 CIBERSORT (https://cibersort.stanford.edu/)was utilized by performing integrative analysis between tumor-infiltrating immune cells and CD155 expression in human cancers. 35
Gene Set Enrichment Analysis
We unitized gene set enrichment analysis (GSEA) 36,37 to investigate the possible mechanisms and functions of CD155, including Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment analysis. The number of random sample permutations was set at 1000, and the significance threshold was set at p < 0.05.
Statistical Analyses
Statistical analyses were performed with R software (https://www.r-project.org/; Version 3.6.1). The Wilcox test was used for differential analysis of gene expression. The most cancers of overall survival (OS), Disease−specific survival (DSS), Disease−free interval (DFI) and Progression−free interval (PFI) of the CD155 was analyzed using Kaplan-Meier (KM) and univariate Cox regression analysis, respectively. Additionally, Hazard ratio (H.R.s) and 95% confidence intervals (CIs) were used to describe the relative risk. P-value < 0.05 was regarded as a statistically significant difference. The correlation between gene expression and TMB, MSI, TME were tested by Pearson Correlation coefficient analysis.
Results
CD155 mRNA Expression in Multiple Human Cancers
We observed that CD155 was differently expressed in multiple human tumor-tissues, and that CD155 was an increase in 12 cancers (BLCA, CHOL, COAD, ESCA, HNSC, KICH, KIRP, LIHC, PRAD, READ, STAD, and UCEC), while the down-regulation in LUSC and THCA (Figure 1A). Next, we utilized the Oncomine database, which was made up to 715 datasets and 86,733 samples, to examine if the CD155 expression is activated in human cancers than paired normal samples. As was seen from Figure 1, CD155 mRNA expression in most human tumors was consistent with above results (Figure 1B). This result implied that overexpressed CD155 were identified in multiple tumors.

Pan-cancer expression profiling analysis of CD155(PVR). A, The boxplot shows the pan-cancer expression profiling of CD155 in human cancers. The below row refers to the standard abbreviations of tumors in TCGA. P-value Significant Codes: ***≤0.001 ≤**≤0.01 _ *≤0.05. B, CD155 was highly expressed in 14 cancer data sets and low expressed in 9 cancer data sets in the Oncomine database.
Prognostic Potential of CD155 in Several Solid Tumors
First, the KM plotter split patients by CD155 media expression were used to examine the prognosis of CD155 in cancers within the RNA sequencing data from TCGA. High expression of CD155, afforded poor OS in BLCA, BRCA, CESC, HNBC, KIRP, LGG, LUAD, MESO, SARC and THCA (P = 0.007, 0.036, 0.009, 0.011, 0.024, 0.002, 0.004, 0.005, 0.014, 0.027, respectively), while high expression of CD155 was significantly associated with a favorable prognosis in KICH and PAAD patients (P = 0.027, 0.005; Figure 2). The univariate Cox regression analysis shown that overexpressing CD155 possessed a high risk in ACC, BLCA, CESC, HNSC, KIRP, LGG, LUAD, MESO, PRAD, SARC, THCA, (HR = 1.9, 95%CI:1.164-3.104, P = 0.010; HR = 1.31, 95%CI:1.102-1.575, P = 0.003; HR = 1.51, 95%CI:1.160-1.957, P = 0.002; HR = 1.39, 95%CI:1.140-1.695, P = 0.001; HR = 2.42, 95%CI:1.660-3.529, P < 0.001; HR = 2.183, 95%CI:1.539-3.097, P < 0.001; HR = 1.45, 95%CI:1.187-1.773, P = <0.001, HR = 1.525, 95%CI:1.149-2.025, P = 0.004; HR = 5.07, 95%CI:1.379-18.696, P = 0.015; HR = 1.31, 95%CI:1.07-1.693, P = 0.037; HR = 4.39, 95%CI:1.266-15.222, P = 0.020, respectively; Table 1).
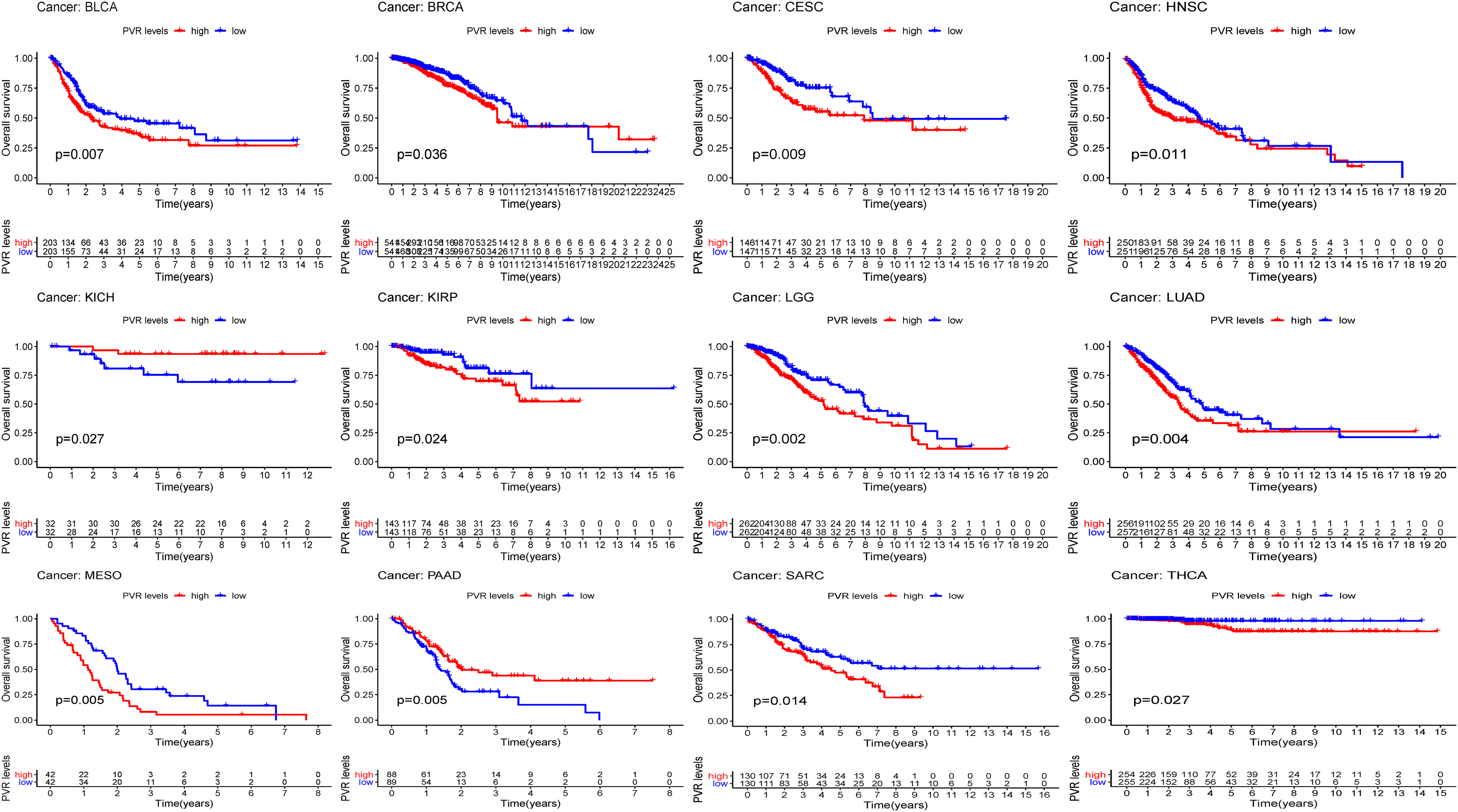
Prognostic value of CD155 of pan-cancer by Kaplan-Meier method in overall survival.
Univariate Cox Regression Analysis.

Except for OS, the trend of DSS, DFI and PFI was consistent with above results (Figures 3 and 4). Additionally, Overview of these results was seen from above figures, there is an apparent heterogeneity among multiple human cancers. This analysis indicated that CD155 might be a worse prognostic factor in multiple cancers. Moreover, the prognostic value of CD155 in cancer patients has also been confirmed in the PrognoScan database (Table 2).

Prognostic value of CD155 in Disease−specific survival. A, Kaplan-Meier plots show the result of pan-cancer Disease−specific survival analysis. B, Forest plot shows the result of pan-cancer Disease−specific survival analysis of CD155.
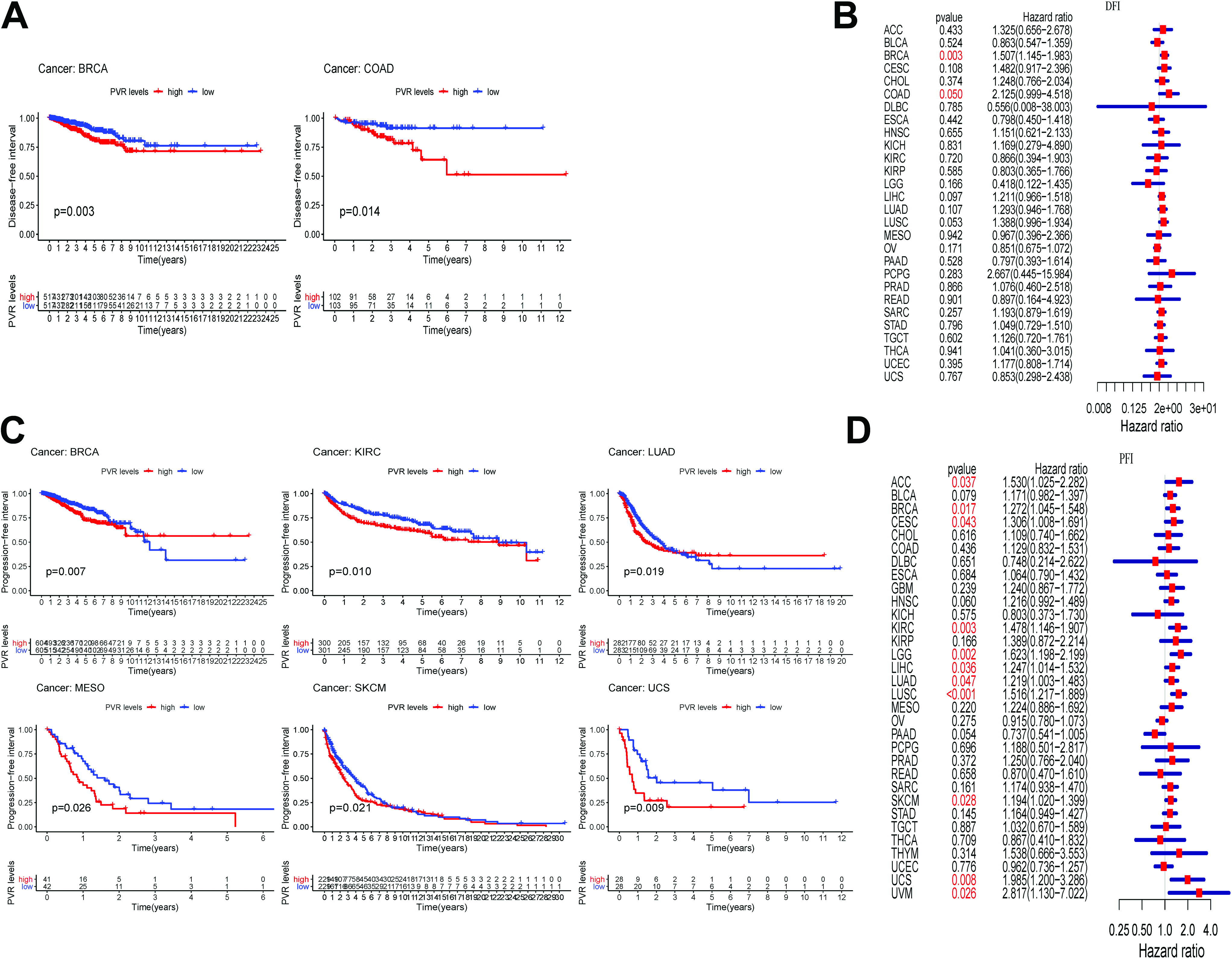
Prognostic value of CD155 in Disease−free interval and Progression−free interval analysis. A and C, Survival curves show the result of pan-cancer Disease−free interval and Progression−free interval analysis of CD155. B and D, Forest plots show the result of pan-cancer Disease−free interval and Progression−free interval analysis of CD155.
The Prognostic Value of CD155 in Human Cancer in the PrognoScan Database.

Significance of CD 155 in Different Stages of Distinct Cancers
To further confirm the prognostic value of CD155, we investigated differential expression of CD155 in different tumor stages. As shown in Figure 5, the expression level of CD155 was significantly different in the clinical stages of various tumors. Therefore, CD155 might affect the prognosis of distinct cancers.

The boxplot shows the correlation between the expression of PVR and the pathological grade in ACC, BLCA, BRCA, HNSC, KICH, LUSC, SKCM and STAD. The label at the top of the picture corresponds to the pathological stage of the patient.
Estimation of Tumor Mutational Burden (TMB) and Microsatellite in Stability (MSI)
We found that CD155 expression level had a significant correlation with the TMB in these tumors by Pearson’s correlation test (ACC r = 0.287, BRCA r = 0.149, COAD r = −0.140, ESCA r = 0.246, KIRP r = −0.168, LGG r = 0.237, LIHC r = 0.104, LUAD r = 0.247, OV r = 0.135, SARC r = 0.268, SKCM r = 0.094, THYM r = 0.341, UCEC r = 0.360, UCS r = 0.397; Figure 6A). On the other hand, it has a significant correlation with MSI in these cancers (BRCA r = 0.071, COAD r = −0.157, KICH r = 0.32, LUSC r = 0.113, PAAD r = 0.216, READ r = −0.19, SARC R = 0.130, UCEC r = 0.265 UVM r = 0.281; Figure 6B).

The radar plots show the correlation between the expression of PVR and the TMB, MSI. A, The expression of PVR associated with TMB in ACC, BRCA, COAD, ESCA, KIRP, LGG, LUAD, OV, SARC, SKCM, THYM, UCEC, UCS. B, The expression of PVR associated with MSI in BRCA, COAD, KICH, LUSC, PAAD, READ, SARC, UCEC, UVM. Significant Codes: ***≤ 0.001 ≤**≤0.01 _ *≤0.05.
CD155 Modulate the Immune Infiltration of Tumor Microenvironment in Human Cancers
After identifying the immunotherapy potential of CD155, we further observated that these genes expression in many cancers were significantly correlated with the immune score and stromal score (Figure 7 and Supplement Figure 1). More importantly, we investigated the correlation between immune infiltration and CD155 expression (Figure 8 Supplement Figure 2). As seen from above figures. CD155 expression level had a strongly positive correlation with CD4+ T cells in HNSC, LUAD and COAD as well as NK cells in HNSC, while inversely interaction with CD8+ T in KIRC and HNSC as well as Tregs in SCKM, HNSC and BLCA. Collectively, these findings indicate that CD155 correlates with cancer immunotherapy function.

The expression level of PVR was correlated with immune score and stromal score in human TME.

The expression level of PVR was correlated with immune cell infiltration in human cancer.
Functional Analysis
To identify the molecular mechanisms underlying the prognostic performance of CD155, we performed Gene Set Enrichment Analysis. As seen from Figure 9, GSEA analysis was performed to participate in the modulation of gene ontological (GO) enrichment and KEGG analysis. As shown in Table 3, the enrichment results were that the common GO functional categories (P < 0.05) were obviously enriched in humoral immune response, immune receptor activity, immunoglobulin-complex-circulating, and T cell and B cell related functions, while KEGG analysis exhibited that the fundamental pathway functional categories (P < 0.05) were significantly clustered in natural killer cell mediated cytotoxicity, pathways in cancer, regulation of Autophagy, Cytokine-cytokine receptor interaction, and cell cycle (Table 3). These results suggested that CD155 might affect multiple patients with poor prognosis by biological processes and pathways above.

GSEA analysis of CD155 in human cancer.
Gene Set Enrichment Analysis of CD155 in Human Cancer.

Discussion
Cancer immunotherapy has been one of the most effectively therapeutic approach for cancer therapy. 38 -41 CD155 (PVR/necl5), a member of the nectin-like family of adhesion molecules, is widely overexpressed across most malignances and has been linked to unfavorable clinical outcomes. 17,42 -45 Intriguingly, previous results reported on literature are almost similar to ours. Meanwhile, CD155 has been shown to play important tumor cell-extrinsic roles by modulating the function of tumor-infiltrating lymphocytes via interactions with both stimulatory and inhibitory receptors on T and NK cells. 19,46 Overall, our study explores insights in understanding the potential role of CD155 in tumor immunotherapy and its use as a cancer biomarker.
In our present study, for the first time, we identified that the CD155 is significantly elevated in 33 tumors based on the TCGA datasets, which is consistent with studies in hepatocellular carcinoma, 47 Pancreatic Cancer, 18,48 and colorectal carcinoma. 49 Next, KM plotter analysis and Univariate Cox regression analysis revealed that the higher expression of CD155 were remarkably correlated with its worse OS, DSS, DFI and PFI in many cancers, indicating a diagnostic and therapeutic role in the initiation of these cancers.
Another study associated with immune function is that CD155 expression in malignances is closely correlated with TMB and MSI, especially for BRCA, COAD and UCEC, indicating that CD155 may serve as a biomarker predicting anti-PD1/PD-L1 monotherapy outcomes 50 -53 in these cancers. The immune score, stromal score has also significantly interaction with CD155 in BLCA, BRCA, CESC, HNSC, KIRC, LUAD, and SKCM. In reviewing reports, the stromal and immune cells has indispensable roles in driving TME of multiple cancers. 34,54 These consequences further confirm that CD155 plays a vitally significant part in the tumor progression by immune system. Lastly, results of immune infiltration demonstrate that CD155 expression level in diverse immune infiltration cell has a strongly positive correlation with CD4+ T cells in HNSC, LUAD and COAD as well as NK cells in HNSC, while inversely interaction with CD8+ T (KIRC and HNSC) and Tregs (SCKM, HNSC and BLCA). There is also evidence that CD155/CD226 interaction can attenuate the generation of CD8+ T cells by regulating NK T-cell differentiation. 55 In addition, the inhibition of TIGIT/CD155 signaling in HNSC significantly impeded the immunosuppression of Tregs by inactivating the output of suppressive cytokines. 56 However, this was not consistent with our predicting analysis in HNSC, suggesting that the underlying mechanism possibly needs to be further explored. Together, our studies provide possible mechanisms, which explain why CD155 expression correlates with cancer immune modulation.
In addition, by performing functional enrichment analysis, we can identify its underlying functions and previously predict unappreciated biological processes associated with human cancers. The functional analysis result of CD155-related genes indicated that it was relevant to humoral immune response, immune receptor activity, natural killer cell mediated cytotoxicity, pathways in cancer, regulation of Autophagy, Cytokine-cytokine receptor interaction, which also contribute to validate immune-related mechanisms by which CD155 can promote carcinogenesis and progression of tumor.
The above results show that the association between poor prognosis and cancer patients may be mainly because of lower immune repose and inhibited immune reactive in TME, which can promote overcome tumor immune suppression and enhance anti-tumor immunity. According to these changes, most tumors with malignance may benefit from immunotherapy and chemotherapy. However, some limitations are supposed to be highlighted in our investigation. First, the biological function of 7 identified genes, especially the association with immunofiltration, should be verified in the assays. Next, only limited data can be used for performance assessment, and it is necessary to collect more data of the robustness of CD115 in the future.
In summary, for the first time, we comprehensively explore the correlation between CD155 expression and prognostic potential among 33 tumors. Our findings also revealed that CD155 might function as immune-associated biomarker in the development of human cancers. Consequently, CD155 could be used as a promising prognostic biomarker and potential novel therapeutic target against human cancers.
Supplemental Material
Supplemental Material, sj-tif-1-tct-10.1177_1533033820980088 - CD155-Prognostic and Immunotherapeutic Implications Based on Multiple Analyses of Databases Across 33 Human Cancers
Supplemental Material, sj-tif-1-tct-10.1177_1533033820980088 for CD155-Prognostic and Immunotherapeutic Implications Based on Multiple Analyses of Databases Across 33 Human Cancers by Hongpan Zhang, Zhihao Yang, Guobo, Lu Cao and BangXian Tan in Technology in Cancer Research & Treatment
Supplemental Material
Supplemental Material, sj-tif-2-tct-10.1177_1533033820980088 - CD155-Prognostic and Immunotherapeutic Implications Based on Multiple Analyses of Databases Across 33 Human Cancers
Supplemental Material, sj-tif-2-tct-10.1177_1533033820980088 for CD155-Prognostic and Immunotherapeutic Implications Based on Multiple Analyses of Databases Across 33 Human Cancers by Hongpan Zhang, Zhihao Yang, Guobo, Lu Cao and BangXian Tan in Technology in Cancer Research & Treatment
Footnotes
Authors’ Note
Hongpan Zhang and Zhihao Yang conceived, designed and wrote the study. Zhang Hongpan and Yang Zhihao contributed equally. Du Guobo and Cao Lu contributed to the revision and statistical analysis of the background of the article. Tan Bangxian finally revised the article and provided financial support for the publication of the article. The data used in our study were obtained from public databases UCSC, TCGA, ONCOMINE, and PrognoScan, therefore, ethical approval was not required.
Acknowledgments
We thank the UCSC, TCGA, ONCOMINE, and
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
Abbreviation
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
