Abstract
Background:
5-fluorouracil (5-FU) is a widely used drug for cancer treatment, but its effect and underlying mechanisms on osteosarcoma (OS) cells remain unclear.
Methods:
U2OS and MG63 cells were treated with 0, 50, 100, and 500 μM 5-FU. MTS and flow cytometry were used to examine the effect of 5-FU on cell viability and apoptosis, respectively. Circular RNA (circRNA) expression was detected using RNA sequencing and quantitative real-time PCR (qPCR). Differentially expressed circRNAs were further subjected to the Kyoto Encyclopedia of Genes and Genomes (KEGG) and gene ontology (GO) analysis to predict their functions. A circRNA–miRNA–mRNA interaction network was generated to analyze the regulatory networks of 5-FU–induced differentially expressed circRNAs. Western blotting (WB) was used to verify the protein in the downstream of circRNAs.
Results:
5-FU inhibited the cell viability of the MG63 cells in a concentration-dependent manner. The most significant effect was observed in the cells treated with 500 μM 5-FU. Apoptosis was also increased in the MG63 cells after 500 μM 5-FU treatment for 3 days. RNA sequencing results showed that 183 differentially expressed circRNAs (172 upregulated and 11 downregulated) in 5-FU–treated cells. KEGG and GO analysis showed that the differentially expressed circRNAs were primarily enriched in proliferation-, apoptosis-, and metabolism-related functions. qPCR was used to verify the most upregulated and downregulated circRNAs. The circRNA–miRNA–mRNA interaction network showed that these 8 circRNAs had a sizable regulatory network that links a series of genes involved in tumor suppression.
Conclusion:
5-FU treatment resulted in the differentially expressed circRNAs that were proliferation- and apoptosis-associated and were involved in the 5-FU–induced inhibition of tumor proliferation in OS cells.
Introduction
Osteosarcoma (OS) is the most frequently occurring primary malignant tumor, and it can cause cancer mortality in children and adolescents. OS is characterized by uncomplicated metastasis. Approximately 10%–20% of the patients with OS show metastasis, especially to the respiratory system. Treatment for OS includes surgical removal and systemic chemotherapy to control the micrometastatic disease. Five-year event-free survival for local patients with OS is approximately 70%. 1 However, the 5-year survival rate in patients with recurrent OS and lung metastases is significantly reduced to 30%. 2 Research concerning treatment for and mechanisms of OS is still warranted.
5-fluorouracil (5-FU) is a chemotherapy drug that is widely used for the treatment of serial tumors, including colorectal cancer, gastrointestinal cancer, breast cancer, and head/neck cance. 3 -6 Studies have shown that 5-FU inhibits the proliferation of the OS U2OS cell line. 7 However, the mechanism of 5-FU’s inhibitory effect on OS has not been entirely characterized.
Circular RNA (circRNA), which is a newly identified class of endogenous RNA, forms a covalent closed-loop structure by head-to-tail contact, which is a cyclical product of mRNA precursor mRNA, and act as miRNA sponges. In contrast to the traditional linear RNA, the circRNA is a highly stable circular form that does not have a typical 5’ cap and 3’ polyA tail. 8,9 CircRNAs are widely expressed in many types of tumors, including OS, and play a significant role in regulating tumor genesis, progression, chemotherapy, and resistance. 10 -12 It has also been suggested that the circRNA is involved in regulating the OS proliferation, apoptosis, migration, and invasion. 13 -15 However, it is yet to be reported whether the circRNA is involved in the therapeutic effect of 5-FU in chemotherapy.
The present study, in which a high-throughput sequencing was used, investigated the changes in the expression profile of the circRNAs in OS cells following 5-FU treatment. Hence, this study provided deep insights concerning the effects of 5-FU on OS cell behavior and its underlying mechanisms.
Materials and Methods
Cell Culture
The 2 OS cell lines, U2OS and MG63, were purchased from Cellcook Inc. (Guangzhou, China). The cells were cultured at 37°C in Dulbecco’s Modified Eagle Medium (DMEM; Invitrogen, CA, USA) containing 10% fetal bovine serum (FBS; Invitrogen, CA, USA) in a humidified incubator containing 5% carbon dioxide (CO2).
Cell Viability (MTS Assay)
The cells were cultured in a 96-well plate at a density of 0.5 × 104 cells/well for 16 h. Then, the cells were treated with 0, 50, 100, or 500 μM 5-FU (Aladdin, Shanghai, China). After 1, 2, and 3 d, the cells were collected and analyzed using the 3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium (MTS) assay, according to the manufacturer’s instructions (Promega, Beijing, China).
Flow Cytometry
The 6-well plates seeded with cells at a density of 0.3 × 106 cells/well were cultured for 16 h. Here, the cells were incubated with 500 μM 5-FU and with no 5-FU (negative control or NC). After 3 d, the cells were collected and centrifuged. Approximately 1 × 106 cells were resuspended in phosphate-buffered saline (PBS) and incubated with annexin V5 and propidium iodide (PI) for flow cytometry and imaging for 15 min at room temperature in the dark. Apoptosis was then analyzed using the BD LSR II flow cytometry (BD Biosciences, CA, USA).
Total RNA Isolation and Sequencing
Total RNA was isolated from the cells, which were treated with 500 μM 5-FU for 3 d and without 5-FU treatment, and then was treated with the TRIzol reagent (Invitrogen, CA, USA) according to the standard protocol. Total RNA quality was assessed using an Agilent 2100 Bioanalyzer (Agilent, CA, USA). Before RNA-sequencing (RNA-seq), DNase I (Epicentre, WI, USA) was added to the total RNA mixture to remove the contaminant DNAs. Next, a Ribo-zero™ rRNA Removal Kit (Epicentre, WI, USA) was used to remove the ribosomal RNAs (rRNA). RNase R digestion (Epicentre, WI, USA) was subsequently performed to digest the linear RNAs. CircRNA libraries were finally generated using the NEBNext® Ultra™ Directional RNA Library Prep Kit for Illumina® (NEB, MA, USA) following the manufacturer’s instructions. Next, the RNA-seq library was sequenced using the HiSeq 2500 platform (Illumina, CA, USA).
Quantitative Real-Time PCR (qPCR)
qPCR was performed to identify the dysregulation in the circRNA and mRNA expression. For this aim, total RNAs isolated previously were converted into cDNAs by reverse transcription using the PrimeScript RT Master Mix (Takara, Dalian, China) and random primers. Divergent primers were designed to specifically target the back splicing site of each circRNA (Table 1). cDNAs were then amplified using the SYBR Premix Ex Taq II (Takara, Dalian, China) according to the manufacture’s recommendations. GAPDH was used for circRNA and mRNA quantification. qPCR was accomplished by incubating the reaction mixture at an initial melting at 95°C for 3 min, followed by melting at 40 cycles at 95°C for 15 s, and a final elongation at 60°C for 30 s. The relative expression of the circRNAs and mRNAs were calculated using the 2-ΔΔCt method. The experiments were repeated in 3 times.
Primers Used in the Study.

Bioinformatics
After RNA-seq was completed, the reads generated by the Illumina sequencer were filtered, and the resulting clean data were aligned to the human GRCh37/hg19 reference genome with the Burrows-Wheeler Alignment Tool (version 0.7.13). The circRNAs were predicted and annotated using the CIRI software. For the quantification of the circRNA expression, the circRNA reads were calculated in the reads per kilobase million (RPKM) unit. Reads for the NC and 5-FU groups were then normalized to the total number of reads. Differentially expressed circRNAs were identified as those with a fold change of log2 (5-FU/NC) >1 and an adjusted P-value of < 0.05, which was adjusted using the Benjamini–Hochberg approach to control the false discovery rate (FDR).
Besides, heatmaps were drawn using the Cluster 3.0 software (USA) to demonstrate the circRNAs that were differentially expressed between the NC and 5-FU cells. The GO and KEGG enrichment analysis were performed by a web based software toolkit, GOEAST, and the online database (http://www.genome.jp/kegg/), respectively. Default parameters were used for the GO analysis. An adjusted P-value of < 0.05 and a gene count of ≥ 2 were set as the criteria for the KEGG analysis.
Additionally, an R software package was used to predict the miRNA targets of the 4 most upregulated and 4 most downregulated circRNAs and their candidate downstream target mRNAs to construct circRNAs–miRNAs–mRNA interaction networks. Then, the networks were drawn using the Cytoscape software (version 2.8.3, http://www.cytoscape.org/).
Western Blotting
Total proteins were extracted using a protein extraction buffer (#MDL91201; MDL, Beijing, China) following the manufacturer’s instructions. The protein concentration was determined using the bicinchoninic acid assay (BCA). Total proteins from each group were separated on 10%–15% SDS-PAGE gels and transferred onto 0.22 µm polyvinylidene fluoride (PVDF) membranes (Millipore, MA, USA). The gels were incubated at room temperature with diluted primary antibodies and stored overnight at 4°C. Then, they were washed 3 times with PBS for 10 min and incubated at room temperature with diluted secondary antibodies for 2 h. The final development of the immune complexes was performed using the ChemiDoc MP chemiluminescence imaging system (#170-8280; Bio-Rad, CA, USA) using the manufacturer’s instructions. Actin was used as an internal standard. Band intensities from at least 3 different biological replicates were used for the statistical analysis using the Image-Pro Plus software (version 6.0). In the WB analysis, anti-CCND2 (#67048-1-Ig; Proteintech, IL, USA; 1:2000), anti-BCL2 (#3498; CST, MA, USA; 1:1000), and anti-BAX (#50599-2-Ig; Proteintech; 1:1000) were employed as the antibodies.
Statistical Analysis
Data were expressed as mean ± standard deviation (SD) and analyzed with SPSS (version 19.0) (SPSS Inc., IL, USA). The student’s t-test was used to compare the differences between the 2 groups. Comparisons among more than 3 groups were analyzed with one-way analysis of variance (ANOVA) with Bonferroni post-hoc test. A p-value of < 0.05 was considered statistically significant.
Results
5-FU Inhibited the OS Cell Activity and Promoted the Cell Apoptosis
The effects of different concentrations of 5-FU (0, 50, 100, and 500 μM) on the U2OS and MG63 cell proliferation were examined by performing the MTS assay. The viability of the MG63 cells treated with 5-FU for 3 d was significantly lower than that of the NC. The viability was significantly lower in the cells treated with 500 µM 5-FU compared to those treated with 100 μM 5-FU, and in the cells treated with 100 µM 5-FU compared to those treated with 50 μM 5-FU (Figure 1A). Nevertheless, the viability of the U2OS cells treated with or without 5-FU was not significantly different; therefore, the MG63 cells were selected for further evaluations. Cytometry analysis was done to analyze the apoptosis in the MG63 cells after 3 d of treatment with 500 μM 5-FU. The results indicated that apoptosis levels were higher in the MG63 cells with 5-FU treatment than that of no 5-FU treatment (Figure 1B and C).
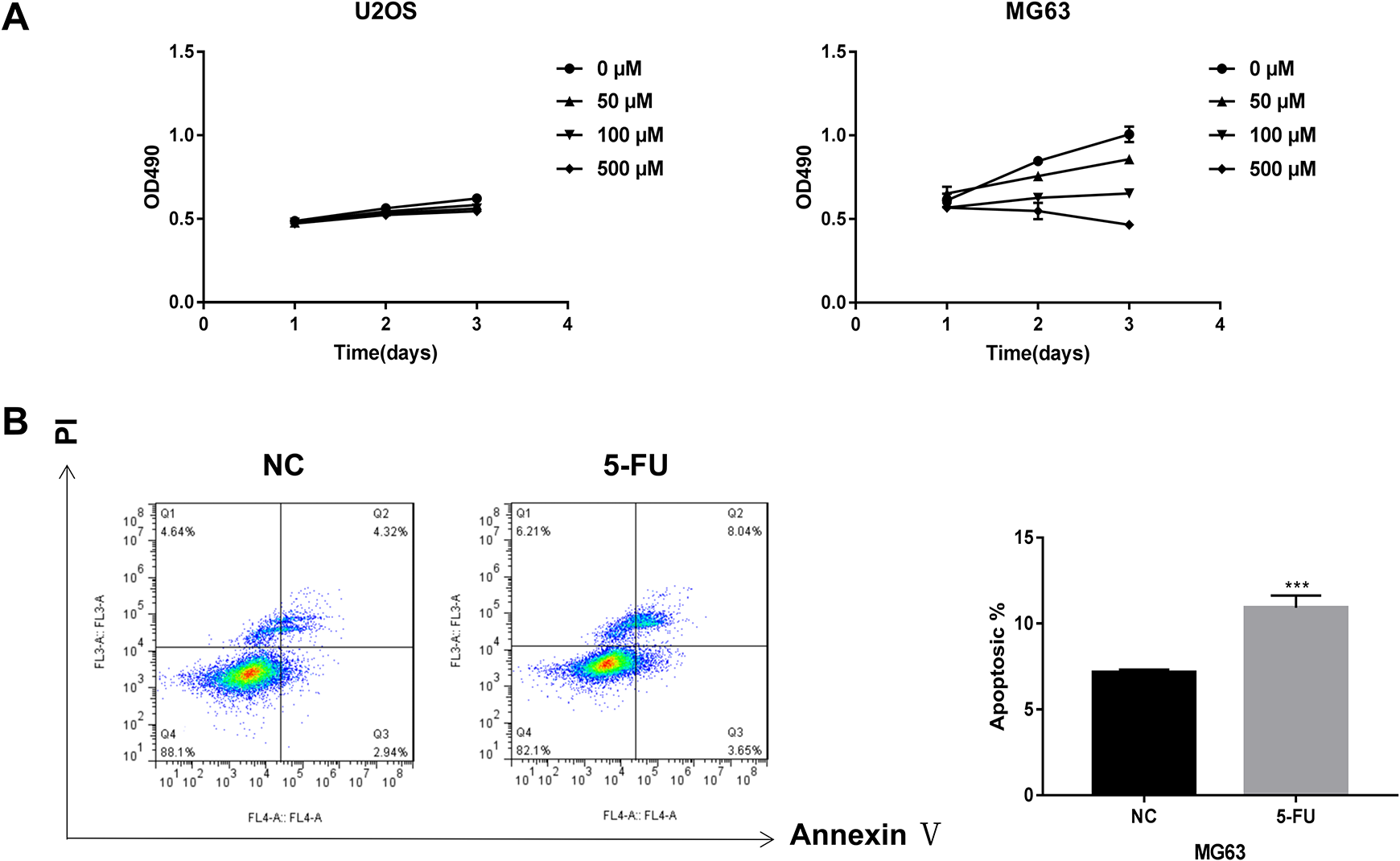
The effects of 5-fluorouracil (5-FU) on cell activity and apoptosis in the osteosarcoma (OS) cells. (A) Cell activity was determined by the MTS assay in U2OS and MG63 cells treated with 0, 50, 100, and 500 μM of 5-FU for 1, 2, and 3 d. (B) The result of flow cytometry showed a more substantial proportion of apoptotic MG63 cells treated with 5-FU for 3 d.
Overview of circRNA Profile Affected by 5-FU Treatment in the MG63 Cells
RNA-seq was performed to detect the changes in the expression profile of the circRNAs of the MG63 cells after 500 μM 5-FU treatment for 3 d. The results showed that a total of 183 circRNAs were differentially expressed in the treated cells. Of these, 172 were upregulated, and 11 were downregulated (Figure 2A). The length of the differentially expressed circRNAs was mostly restricted in the 0–1500 nt range. The length of the circRNAs was concentrated in the 300-nt range (Figure 2B). The upregulated and downregulated circRNAs were then sorted by chromosomal location. The results indicated that the upregulated circRNAs were present mostly on the chromosomes, whereas the downregulated circRNAs were almost distributed, except chromosomes 7, 19, X, and Y (Figure 2C).

RNA-seq data showed the differentially expressed circRNAs in the 5-fluorouracil (5-FU) treated MG63 cells. (A) Hierarchical clustering analysis showed the circRNAs that were differentially expressed between 5-FU treated and untreated MG63 cells. The blue and the red color represented downregulation and upregulation, respectively. The length (B) and chromosome distribution (C) of the differentially expressed circRNAs.
The KEGG and GO analyses were used to predict the functions of the differentially expressed circRNAs’ target mRNAs. The KEGG analysis showed that differentially expressed circRNAs were primarily enriched in proliferation-, apoptosis-, and nucleic acid metabolism-related pathways, such as cell cycle, endocytosis, and MAPK signaling (Figure 3A). Similarly, the GO analysis showed that differentially expressed circRNAs were primarily enriched in terms of proliferation, apoptosis, and nucleic acid metabolism (Figure 3B).

The functional analysis of differentially expressed circRNAs’ target mRNAs. The KEGG (A) and GO (B) analyses were used to predict the function of the differentially expressed circRNAs’ target mRNAs.
Validation of the Differentially Expressed circRNAs
Table 2 showed the 4 most upregulated and downregulated circRNAs as ranked by a fold change. Among these differentially expressed circRNAs, the most upregulated was the hsa_circ_0001013, followed by hsa_circ_0034616, hsa_circ_00116639, and hsa_circ_FAM172A. On the other hand, the 4 most downregulated circRNAs were the hsa_circ_0005694, hsa_circ_0007386, hsa_circ_0008362, and hsa_circ_0073817 (Table 2). qPCR results showed that expression of hsa_circ_0001013, hsa_circ_0034616, hsa_circ_00116639, and hsa_circ_FAM172A in the MG63 cells treated with 500 μM 5-FU for 3 d was significantly higher compared to the NC. The expressions of hsa_circ_0005694, hsa_circ_0007386, hsa_circ_0008362, and hsa_circ_0073817 were downregulated in the cells treated with 5-FU compared to those found in the NC (Figure 4).
Differentially Expressed circRNAs in the 5-FU–treated Cells.

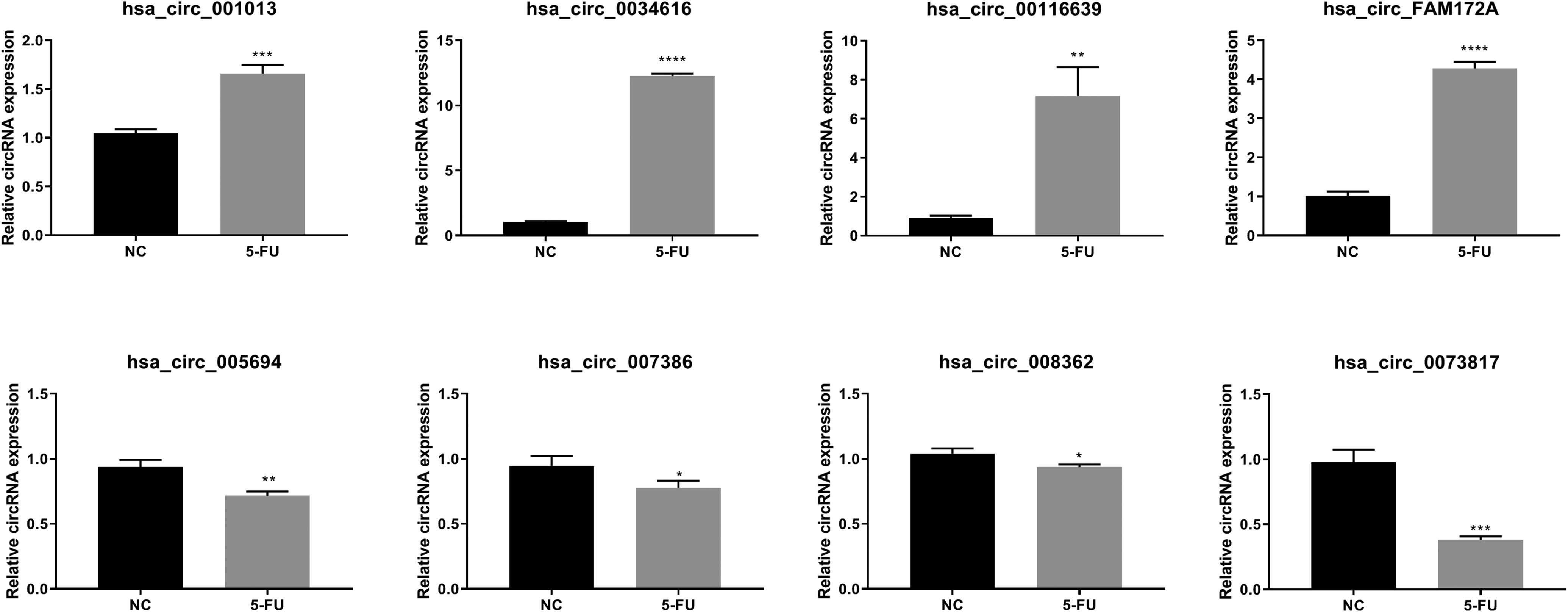
qPCR analysis verified the 4 upregulated and 4 downregulated circRNAs in 5-fluorouracil (5-FU) treated or untreated MG63 cells. NC = untreated MG63 cells, 5-FU = 5-FU treated MG63 cells. Asterisks indicated significance at **** p ≤ 0.0001; *** p ≤ 0.001; ** p ≤ 0.01; * p ≤ 0.05.
Networks of Differentially Expressed circRNAs in OS Cell Proliferation and Apoptosis
The target miRNAs of the 8 circRNAs that were most upregulated and downregulated were predicted. The miRNAs were correlated with the target mRNAs associated with cell proliferation and apoptosis to identify the circRNA-miRNA-mRNA networks. It was found that the cell proliferation network of the differentially expressed circRNAs contained a total of 122 miRNAs and 145 mRNAs (Figure 5 and Table S1), and the apoptosis network contained 79 miRNAs and 51 mRNAs (Figure 6 and Table S2).

The circRNA-miRNA-mRNA interaction network showing the potential regulatory network linking the 8 most upregulated (hsa_circ_0001013, hsa_circ_0034616, hsa_circ_00116639, and hsa_circ_FAM172A) and downregulated (hsa_circ_0005694, hsa_circ_0007386, hsa_circ_0008362, and hsa_circ_0073817) circRNAs and proliferation-associated genes.
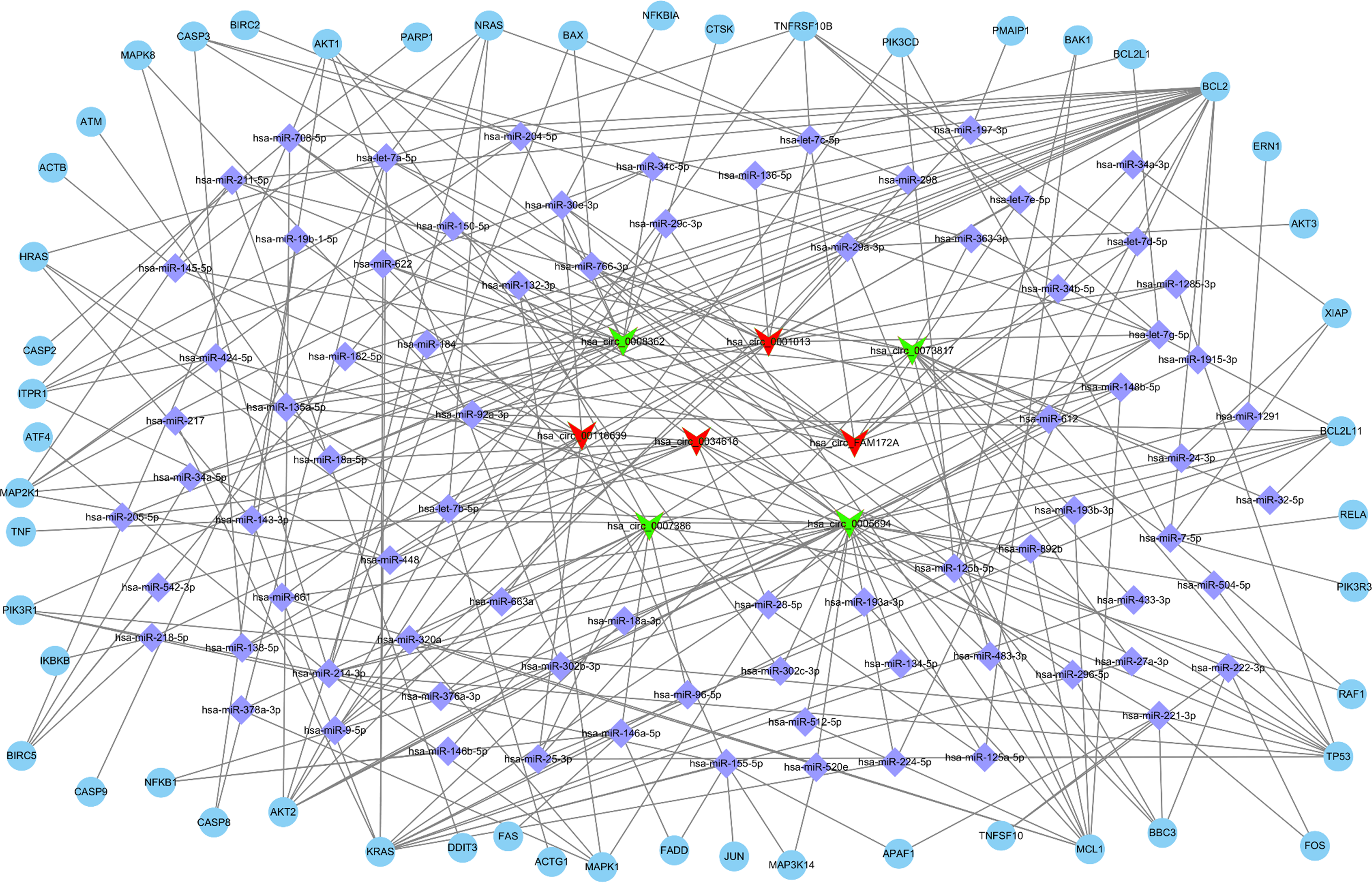
The circRNA-miRNA-mRNA interaction network showing the potential regulatory network linking the 8 most different regulated (hsa_circ_0001013, hsa_circ_0034616, hsa_circ_00116639, and hsa_circ_FAM172A) and downregulated (hsa_circ_0005694, hsa_circ_0007386, hsa_circ_0008362, and hsa_circ_0073817) circRNAs and apoptosis-associated genes.
Validation of the mRNA From Downstream of the 3 Selected circRNAs by qPCR and WB
Here, the downstream of hsa_circ_0005694, hsa_circ_0073817 and hsa_circ_0001013 was selected for further study due to the genes that participated in cell proliferation and apoptosis. The status of the base pairing of circRNA, miRNA, and mRNA was shown in Figure 7. The qPCR and WB of the mRNAs in the downstream of the circRNAs also showed that the changes of the mRNA expressions were consistent with the circRNAs, which were upstream of the miRNAs (Figure 8). These results could also verify the relationship between the circRNA, miRNA, and mRNA, as the hypothesis shown in Figure 7.

The base pairing of 3 pairs selected from the circRNA-miRNA-mRNA interaction network. The 3 selected circRNAs were included hsa_circ_0005694, hsa_circ_0073817 and hsa_circ_0001013, which could interacted with mRNA through base pairing with miRNA.

The results of qPCR and WB analyses of the mRNA from downstream of 3 pair selected circRNAs, including hsa_circ_0005694, hsa_circ_0073817 and hsa_circ_0001013. NC = untreated MG63 cells, 5-FU = 5-FU treated MG63 cells. Asterisks indicated significance at **** p ≤ 0.0001; ** p ≤ 0.01.
Discussion
The present study demonstrated that 500 μM 5-FU inhibited MG63 cell proliferation and promoted apoptosis. A comparison of the circRNA sequencing results showed the differential expression levels in 500 μM 5-FU treated cells and the untreated cells. The potential function of differentially expressed circRNAs predicted by the KEGG and GO analyses indicated the associations with cell proliferation-, apoptosis-, and metabolism-related functions. The 4 most upregulated and 4 most downregulated circRNAs were verified by qPCR. The circRNA-miRNA-mRNA interaction network and the cell proliferation- and apoptosis-related mRNAs analysis revealed that the significant differentially expressed circRNAs could be associated with cell proliferation and apoptosis in OS.
5-FU treatment has been widely used in colorectal cancer and shown significant therapeutic effects. 16 Moreover, 5-FU is often used as a positive control to study the antitumor effect of other drugs in OS. 7,17 However, the studies have mainly focused on U2OS cells, and few studies have demonstrated the antitumor effects of 5-FU in MG63 cells.
The present study provided evidence that 500 μM 5-FU treatment for 3 d inhibited MG63 cell proliferation and promoted apoptosis. RNA-seq data showed that the circRNAs were differentially expressed after 500 μM 5-FU treatment for 3 d. The circRNA targets were primarily enriched in the proliferation functions, such as cell cycle and the MAPK signaling pathway, and the apoptotic pathways, such as endocytosis and metabolic pathways. These results were consistent with the previous antitumor studies, similarly using 5-FU, for example, 5-FU induced cellular DNA synthesis, 2/M phase arrest, and apoptosis. 18 The MAPK pathway was also found as a vital tumor suppressor pathway in OS, 19 and the MAPK activation induced the apoptosis in OS cells. 20 In other tumors, 5-FU also inhibited the MAPK-related pathways and tumor activity, 21 and increased the chemosensitivity to 5-FU. 21 In another study, the mechanism of the anti-tumor effect of 5-FU was revealed via the inhibition of thymidylate synthase and incorporating its metabolites into RNA and DNA, thereby disrupting processing and function of normal RNA and DNA. 3 Moreover, some changes in the endocytic signals were associated with cancer behavior. 22,23 Therefore, it was hypothesized that the anti-tumor effects of 5-FU observed in the present study, and the changes in circRNA’s expression profile might be related to associated proliferation and apoptosis pathways. A previous study has shown that some dysregulated circRNAs gave rise to sustained proliferative signaling and evasion of proliferation suppressors in multiple cancers. 24
Accumulating evidence has also indicated a crucial role of circRNA in regulating OS cell proliferation. For example, Yanbin and Jing demonstrated that the circ_101356/circ_0004846, termed as circSAMD4, was preferentially expressed in OS tissues and could facilitate OS cell proliferation. 25 Another work presented that the circ0001564 was significantly overexpressed in OS tissue and cell lines, and knockdown of the circ_001564 could inhibit OS cell proliferation and apoptosis by sponging the miR-29c-3p. 13 Moreover, it was found that the circNT5C2 accelerated OS cell proliferation and metastasis by targeting the miR-448. 10 These previous studies found that the interaction between the circRNAs and the miRNAs was significant.
The present study also verified the expressions of the 4 most upregulated (hsa_circ_0005694, hsa_circ_0007386, hsa_circ_0008362, and hsa_circ_0073817) and 4 most downregulated (hsa_circ_0001013, hsa_circ_0034616, hsa_circ_00116639, and hsa_circ_FAM172A) circRNAs. The expressions and functions of these circRNAs have not yet been reported in the literature. However, a study by Wang et al. identified that lncRNA MIAT competed with critical RNAs to target miR-150-5p, thus regulating OS cell function. 26 Wu et al. confirmed the involvement of hsa_circ_0000263 in modulating downstream target genes of miR-150-5p. 27 In our study, miR-150-5p was also predicted to be the target of hsa_circ_0005694, which was downregulated in MG63 cells after 500 μM 5-FU treatment for 3 d. Thus, has_circ_0005694 and hsa_circ_0000263 might compete with lncRNA MIAT to target miR-150-5p, thereby regulating the function of OS cells undergoing 5-FU treatment.
Zhou et al. also demonstrated that hsa-let-7 g was highly expressed in OS tissues, and high expression of hsa-let-7 g promoted OS occurrence by downregulating the HOXB1 and activating the NF-kB pathway. 28 In our study, hsa-let-7 g was predicted to be the target of hsa_circ_0001013, which was upregulated in the MG63 cells treated with 500 μM 5-FU for 3 d. Thus, hsa_circ_0001013 could also be a critical factor in the regulation of the HOXB1 and activation of the NF-kB pathway during the 5-FU treatment.
The regulatory networks of the differentially expressed circRNAs suggested a role in proliferation and apoptosis-related processes. Ye et al. reported that miR-18a-5p was recognized as a functional target of FER1L4, which plays a crucial role in modulation of cell apoptosis and epithelial-mesenchymal transition, 29 in OS. In this study, miR-18a-5p was also predicted as a target of hsa_circ_0034616 and hsa_circ_0008362, which were upregulated and downregulated, respectively, in the MG63 cells with 500 μM 5-FUtreatment for 3 d. In light of previous research, it could be hypothesized that the upregulated and downregulated circRNAs could work in unison to control the cell apoptosis process during the 5-FU treatment by interacting with the same miRNAs. The miR-125a-5p/VEGFA and the hsa-miR-132-3p/MAPK1 regulatory relationships in the cell proliferation network were verified in other studies to promote tumor growth. 30,31 Besides, interactions between hsa-182-5p/BCL2 and hsa-miR-193a-3p/MCL1 in the apoptotic network induced tumor cell apoptosis and had tumor-suppressive effects. 32,33 A previous study showed that let-7d-overexpression reduced cell proliferation by decreasing CCND2, encoding cyclin D2, which is strongly expressed in human granulosa cell tumors. 34,35 The expression of Bax and Bcl-2 could be regulated in contrary by β-caryophyllene in MG63 cell via reactive oxygen species (ROS)-induced apoptotic mitochondrial pathway. 36 Therefore, in our study, the network constructed with the obtained results verified the interaction of circRNAs, miRNAs, and mRNA. The relationship between the circRNAs, miRNAs, and mRNAs, and the expression changes of them were also verified by qPCR and WB in our present study. Enhanced relationships between circRNAs, miRNAs, and mRNA during 5-FU treatment induced cell proliferation and apoptosis further indicated the role of critical circRNA expression in the 5-FU induced inhibition of OS proliferation.
In summary, although this study did not explicitly analyze the function of specific circRNAs, current sequencing and bioinformatics analysis results showed that the circRNA was associated with 5-FU induced inhibition of OS proliferation. The circRNAs were differentially expressed in 5-FU treated cells compared to the untreated cells. Notably, the most differentially expressed circRNAs had a large proliferation- and apoptosis-related regulatory network, further indicating their important role in 5-FU induced inhibition of OS proliferation.
Supplemental Material
Supplemental Material, Table_S1 - The Analysis of Differentially Expressed circRNAs Under the Antiproliferative Effect From 5-Fluorouracil on Osteosarcoma Cells
Supplemental Material, Table_S1 for The Analysis of Differentially Expressed circRNAs Under the Antiproliferative Effect From 5-Fluorouracil on Osteosarcoma Cells by AiJun Huang, LiPing Chen, YiMing Wang, ShuQiang Ma, Song Jin, Hanzhou Cai, Xingzhong Huang, Hongbo Zhang, ZengRong Wang, Kun Lin and Fangsiyu Lin in Technology in Cancer Research & Treatment
Supplemental Material
Supplemental Material, Table_S2 - The Analysis of Differentially Expressed circRNAs Under the Antiproliferative Effect From 5-Fluorouracil on Osteosarcoma Cells
Supplemental Material, Table_S2 for The Analysis of Differentially Expressed circRNAs Under the Antiproliferative Effect From 5-Fluorouracil on Osteosarcoma Cells by AiJun Huang, LiPing Chen, YiMing Wang, ShuQiang Ma, Song Jin, Hanzhou Cai, Xingzhong Huang, Hongbo Zhang, ZengRong Wang, Kun Lin and Fangsiyu Lin in Technology in Cancer Research & Treatment
Footnotes
Abbreviations
Authors’ Note
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by LiPing Chen, YiMing Wang, ShuQiang Ma, Song Jin, Hanzhou Cai, Xingzhong Huang, Hongbo Zhang, ZengRong Wang, Kun Lin and Fangsiyu Lin. The first draft of the manuscript was written by AiJun Huang and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript. Our study did not require an ethical board approval because it did not contain human or animal trials.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
