Abstract
Background:
PTPRG antisense RNA 1 has been well-documented to exert an oncogenic role in diverse neoplasms. However, the precise role of PTPRG antisense RNA 1 in regulating radiosensitivity of nonsmall cell lung cancer cells remains largely elusive.
Methods:
Expression levels of PTPRG antisense RNA 1 and miR-200c-3p in nonsmall cell lung cancer tissues and cells were detected by quantitative real-time polymerase chain reaction, while transcription factor 4 expression was examined by immunohistochemistry and Western blot. After nonsmall cell lung cancer cells were exposed to X-ray with different doses in vitro, Cell Counting Kit-8 assay and colony formation assay were conducted to determine the influence of PTPRG antisense RNA 1 on cell viability. Interaction between miR-200c-3p and PTPRG antisense RNA 1 as well as transcription factor 4 was investigated by dual luciferase reporter assay.
Result:
In nonsmall cell lung cancer tissues, the expressions of PTPRG antisense RNA 1 and transcription factor 4 were significantly upregulated, whereas the expression of miR-200c-3p was downregulated. It was also proved that PTPRG antisense RNA 1 and 3′-untranslated region of transcription factor 4 can bind to miR-200c-3p. Under X-ray irradiation, overexpressed PTPRG antisense RNA 1 could promote the viability and enhance the radioresistance of nonsmall cell lung cancer cells, and this effect was partially weakened by miR-200c-3p mimics. Transcription factor 4 was identified as a target gene of miR-200c-3p, which could be positively regulated by PTPRG antisense RNA 1.
Conclusion:
PTPRG antisense RNA 1 reduces the radiosensitivity of nonsmall cell lung cancer cells via modulating miR-200c-3p/TCF4 axis.
Introduction
Lung cancer is the leading cause of cancer-related mortality in both male and female worldwide. 1,2 Nonsmall cell lung cancer (NSCLC) is the main histological type of lung cancer, accounting for nearly 80% of the total number of lung cancer. 3,4 Unfortunately, a majority of patients are diagnosed at an advanced stage, resulting in a high mortality rate. 5 The radiation technology offers symptomatic relief and improvement in survival for patients with NSCLC. 6 -9 However, radiation resistance hinders the further advancement of radiotherapy efficacy and leads to treatment failure in patients. 10 Hence, an in-depth investigation into the underlying mechanism of radioresistance in NSCLC cells is highly desirable to improve survival rate and help to determine optimized treatment for NSCLC.
Long-chain noncoding RNA (lncRNA) is a class of noncoding RNA that is more than 200 nucleotides without protein-coding ability. 11 -15 Mounting studies indicate that lncRNA wields enormous influence in biological processes, such as the proliferation, apoptosis, and tumorigenesis. 16 -19 The altered lncRNA expression can be used as an indicator to evaluate the tumorigenesis, progression and relapse of multiple cancers, including NSCLC. 20 For instance, the expression of lncRNA TP73-AS1 was significantly elevated in NSCLC that predicted shorter survival time in patients 21 ; lncRNA AWPPH can facilitate the proliferation of NSCLC cells and suppress the apoptosis. 22 Moreover, lncRNA is also involved in the regulation of radiosensitivity of tumor cells. For instance, lncRNA PVT1 induces radioresistance in NSCLC cells 23 ; high expression of lncRNA GACAT3 can impede metastasis of cancer cells and improve the effect of radiotherapy. 24 LncRNA PTPRG-AS1 has been shown to be significantly overexpressed in breast cancer and nasopharyngeal carcinoma and induce radiation resistance in nasopharyngeal carcinoma cells. 25,26 Nevertheless, there is at present no defined role and detailed mechanism of PTPRG-AS1 in regulating radiosensitivity in NSCLC cells.
MicroRNA (miRNA) is a class of small noncoding RNA, containing 18 to 25 nucleotides that specifically binds to the target gene 3′-untranslated region (3′-UTR) to cause translation inhibition or messenger RNA (mRNA) degradation, thus regulating in gene expression. 27,28 The abnormal expression of miRNA is closely related to cell differentiation, stress response, proliferation, apoptosis, and other biological behaviors, thus wielding its impact on the tumorigenesis and progression of diverse cancers. 29 -31 Accumulating evidences have verified that miR-200 family members, including miR-200a, miR-200b, and miR-200c, exert an antitumor role in NSCLC. 32 Specifically, miR-200c restrains epithelial–mesenchymal transition and metastasis of lung cancer through targeted regulation of HMGB1 expression. 33 In addition, miR-200c can enhance the radiosensitivity of breast cancer and NSCLC cells. 34,35 MiR-200c-3p has been proven to be involved in the biological behaviors as a tumor inhibitor in a variety of tumors, like renal clear cell carcinoma and glioma. 36,37 However, the role of miR-200c-3p in modulating the radiosensitivity of NSCLC cells and its regulatory mechanism remain poorly clarified.
A line of evidences have validated the abnormal activation of transcription factor 4 (TCF4) plays a pivotal role in the tumorigenesis, cancer progression, and radiotherapy resistance of multiple malignant tumors. 38 -41 It is reported that silencing TCF4 can induce the radiosensitivity of cancer cells to X-ray. 42 Interestingly, bioinformatics analysis (based on StarBase database and TargetScan database) suggests that there are potential binding sites between miR-200c-3p and PTPRG-AS1 as well as the 3′-UTR of TCF4. The purpose of this study was to investigate the role of PTPRG-AS1 in radiosensitivity of NSCLC cells via regulating miR-200c-3p/TCF4.
Materials and Methods
Patients’ Recruitment
This study was conducted in accordance with the ethical standards of the Declaration of Helsinki. All patients involved gave informed consent to the study and signed a written consent form. Our study was approved by the Ethics Review Committee of Shanxi Cancer Hospital (Approval No. 2017-00325). In all, 35 cases of NSCLC tissues (16 cases of squamous cell carcinoma and 19 cases of adenocarcinoma) and matched nontumor tissues were procured from Shanxi Cancer Hospital (Taiyuan, China). All cancer tissues were pathologically confirmed as NSCLC, and no other malignant tumors were found.
Immunohistochemistry
Paraffin-coated NSCLC tissues and adjacent tissues were sliced, dewaxed, and hydrated by xylene. The slices were incubated with 0.3% hydrogen peroxide solution at room temperature for 30 minutes. Afterward, the slices were then incubated with the primary antibody (Anti-TCF4, Abcam, ab185736, 1:100) at 4 °C overnight and then with the secondary antibody (goat-antirabbit immunoglobulin G [IgG]) at 37 °C for 1 hour. Subsequently, the slices were rinsed thoroughly with phosphate buffered saline. Then the color development was performed using 3,3’-diaminobenzidine (Hubei Biossci Biotechnology Co, Ltd.). The scoring standard of immunohistochemistry was completed by pathologists in the Department of Pathology, Shanxi Cancer Hospital.
Cell Lines and Cell Culture
Human lung cancer cell lines (H157, A549, H838, H460, and H1299) and normal bronchial epithelial cell lines (16HBE cells) were purchased from the American Type Culture Collection. All cells were cultured in Dulbecco Modified Eagle’s medium (Hyclone) supplemented with 10% heat-inactivated fetal bovine serum (Invitrogen), 100 U/mL penicillin, and 100 μg/mL streptomycin in 5% CO2 at 37 °C. The culture medium was replaced at an interval of 3 to 4 days.
Cell Transfection
Small interfering RNA (siRNA) control, siRNAs against PTPRG-AS1 (si-PTPRG-AS1), pcDNA empty vector (NC), pcDNA-PTPRG-AS1, miRNA control (miR-con), miR-200c-3p mimic, and miR-200c-3p inhibitor were purchased from GenePharma Co, Ltd. H157 and H1299 cells were inoculated into 6-well plate at a density of 3 × 105/mL and incubated at 37 °C for 24 hours with 5% CO2. H157 and H1299 cells were transfected using Lipofectamine 3000 (Invitrogen; Thermo Fisher Scientific, Inc.) according to the supplier’s instructions. Following that, the cells were cultured at 37 °C and 5% CO2 for 24 hours for further analyses. After that, the transfection efficiency was detected by quantitative real-time polymerase chain reaction (qRT-PCR).
Ionizing Radiation
The NSCLC cells in logarithmic growth phase were collected. Transfected NSCLC cells were irradiated with linear accelerator (Varian Medical Systems) at room temperature. The cells were exposed to X-ray with different doses (0, 2, 4, 6, and 8 Gy) with a 6-MV photon beam at a dose rate of 3.5 Gy/min and 160 kv X-ray energy. After that, cells were harvested for subsequent experiments. As for the time course experiment, the cells were exposed to X-ray at the dose of 4 Gy, and then the cells irradiated by X-ray were collected for qRT-PCR analysis every 3 hours within 24 hours.
Real-Time Polymerase Chain Reaction
Total RNA from tissue and cultured cells was isolated by TRIzol Reagent (Invitrogen); 1 µg total RNA was reversely transcribed into complementary DNA (cDNA) using SuperScript First Strand cDNA System (Invitrogen). Then real-time polymerase chain reaction (PCR) was carried out. Quantitative RT-PCR cycling was initially performed at 95 °C for 2 minutes, then 40 cycles at the conditions of 95 °C for 15 seconds, 60 °C for 15 seconds, and 68 °C for 20 seconds. The relative expressions of PTPRG-AS1 and miR-200c-3p were calculated using 2−ΔΔCT method. The primer sequences are shown as follows: PTPRG-AS1: 5′-AAGCCAAGCAGTCAGAAGCA-3′ (forward) and 5′-GAGCCCTGACAGCCTAATGA-3′ (reverse); miR-200c-3p: 5′-UAAUACUGCCGGGUAAUGAUGCA-3′ (forward) and 5′-CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTCCATCAT-3′ (reverse); U6: CTCGCTTCGGCAGCACA (forward) and AACGCTTCACGAATTTGCGT (reverse).
Western Blot
The cells were lysed with radioimmunoprecipitation assay buffer (Beyotime Biotechnology). After high-speed centrifugation (10 minutes, 12 000 rpm, 4 °C), the supernatant was collected and heated in a water bath at 100 °C for 10 minutes to denature the protein. The content of the protein was determined by bicinchoninic acid kit (Beyotime Biotech). After that, sodium dodecyl sulfate-polyacrylamide gel electrophoresis was performed, and the protein was electrophoretically transferred to polyvinylidene difluoride (PVDF) membrane (Millipore). After the PVDF membrane was washed with Tris buffered saline Tween (TBST) solution, TCF4 antibody (Abcam, ab185736, 1:1000) and β-actin antibody (Abcam, ab20272, 1:100) were added to incubate the membrane for 12 hours at 4 °C. Then, the secondary antibody goat antirabbit IgG H&L (Abcam, ab150077, 1:1000) was added to incubate the PVDF membrane at room temperature for 1 hour. Then, TBST was used to wash the membrane 3 times. Ultimately, color rendering was performed using hypersensitive electrochemiluminescence (Biossci Biotechnology Co, Ltd).
Cell Counting Kit-8 Assay
H157 and H1299 cells in logarithmic phase were harvested. Afterward, cells were cultured in 96-well plates for different times (24, 48, 72, and 96 hours). At each time point, 10 µL enhanced Cell Counting Kit 8 (CCK8; Biossci Biotechnology Co, Ltd) was added and the culture was continued. After 1 hour, the absorbance value (OD value) of each well was measured at 450 nm on a microplate reader.
Colony Formation Assay
The transfected NSCLC cells were inoculated into a 6-well plate and irradiated with a specified single dose of X-ray (0, 2, 4, 6, or 8 Gy). After incubation at 37 °C for 2 weeks, the cells were fixed with 10% methanol and stained with 0.1% crystal violet for 15 minutes. After the plate was washed and dried, the number of colonies formed was observed and recorded under a microscope.
Dual Luciferase Reporter Gene Assay
Dual luciferase reporter gene assay was applied to validate the targeting relationships between miR-200c-3p and TCF4, as well as miR-200c-3p and PTPRG-AS1. Wild type (WT) or mutant type (MUT) PTPRG-AS1/TCF4 sequence was subcloned into pGL3 Basic vector (Promega). MiR-200c-3p or miR-con was then cotransfected with WT or MUT vector into H157 and H1299 cells, respectively. The transfected cells were then inoculated on a 24-well plate with 5000 cells per well. Luciferase activity was measured 48 hours later with dual luciferase reporter assay system (Promega).
Statistical Analysis
SPSS 21.0 (IBM Corp) was adopted for statistical analysis. Data were presented as mean ± standard deviation. Student t test was carried out to analyze the difference of measurement data. Chi-square test was performed to analyze the difference of counting data. A P value <.05 indicated statistical significance.
Results
Significant Correlations Among the Expression Levels of PTPRG-AS1, miR-200c-3p, and TCF4 in NSCLC
To get insight into the correlations among PTPRG-AS1, miR-200c-3p, and TCF4, RT-PCR was performed to detect the expression levels of PTPRG-AS1, miR-200c-3p, and TCF4 mRNA in NSCLC tissues and matched normal tissues. We found that NSCLC tissues exhibited high expression levels of PTPRG-AS1 and TCF4 mRNA and low expression level of miR-200c-3p (Figure 1A-C). The subsequent correlation analysis showed that there was a negative correlation between PTPRG-AS1 and miR-200c-3p (Figure 1D, R = −0.719, P < 0.001) as well as miR-200c-3p and TCF4 (Figure 1E, R = −0.6486, P < 0.001), and a positive correlation was found between PTPRG-AS1 and TCF4 (Figure 1F, R = 0.5065, P < .01). The immunohistochemistry indicated that the expression level of TCF4 protein in cancer tissues was significantly higher than that in adjacent normal tissues (Figure 1G). Through chi-square test, we discovered that high TCF4 protein expression was associated with higher PTPRG-AS1 expression (Table 1, chi-square value = 4.8802, P = 0.0271), while high TCF4 protein expression was associated with lower expression level of miR-200c-3p (Table 2, chi-square value = 4.8043, P = 0.0283). Our data supported that there were possible regulatory relationships among PTPRG-AS1, miR-200c-3p, and TCF4.
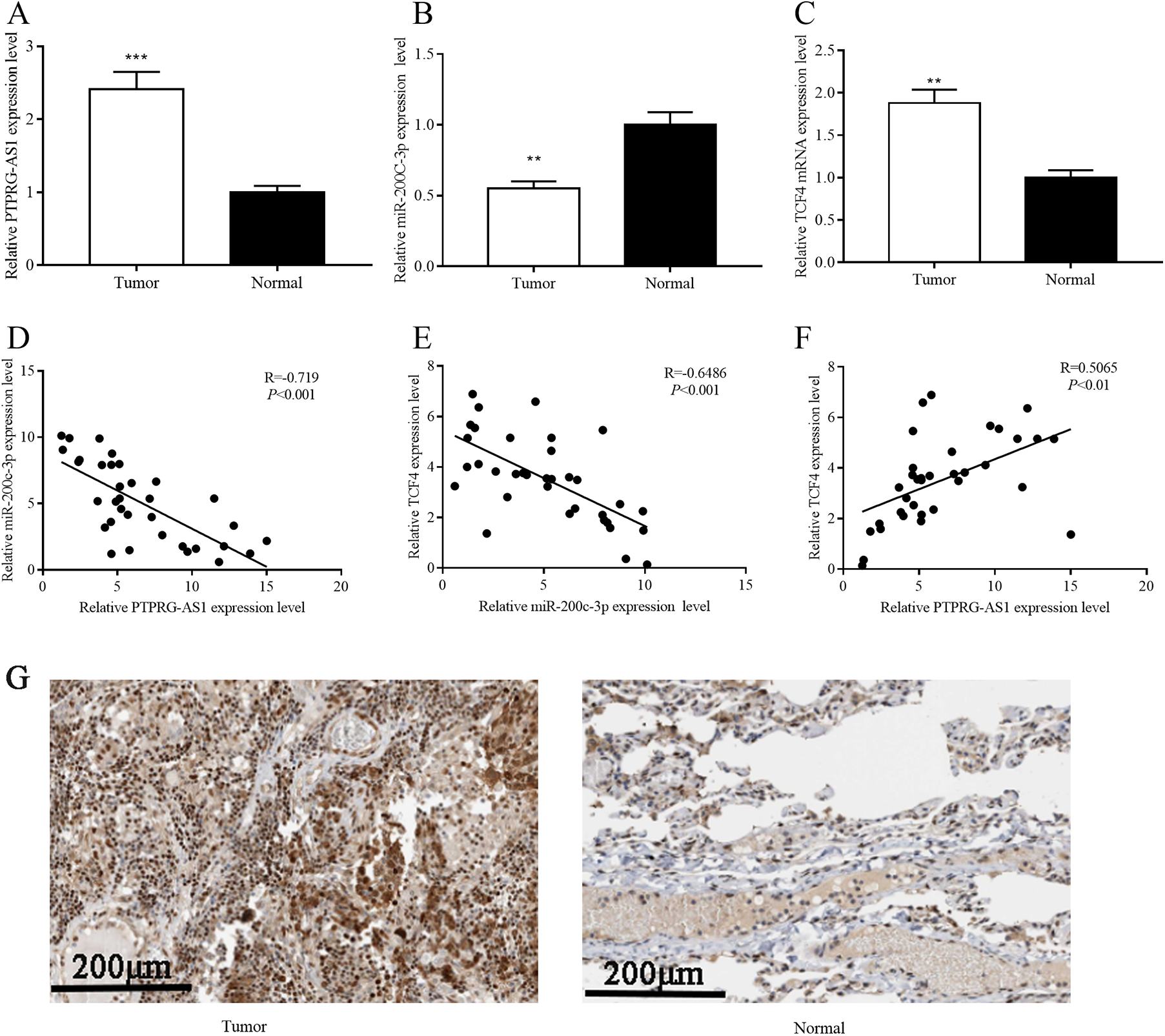
Correlations among PTPRG-AS1 expression, miR-200c-3p expression, and TCF4 expression in NSCLC. A-C, The expressions of PTPRG-AS1, miR-200c-3p, and TCF4 mRNA in NSCLC tissues were detected by qRT-PCR. The expression level of PTPRG-AS1 was negatively correlated with the expression of miR-200c-3p in NSCLC samples. The expression level of miR-200c-3p was negatively correlated with that of TCF4 mRNA in NSCLC samples. The expression level of TCF4 mRNA was positively correlated with the expression level of PTPRG-AS1 in NSCLC samples. The expressions of TCF4 protein in NSCLC tissues and adjacent normal tissues were detected by immunohistochemistry. ** and *** represent P < .01 and P < .001, respectively. mRNA indicates messenger RNA; NSCLC, nonsmall cell lung cancer; qRT-PCR, quantitative real-time polymerase chain reaction; TCF4, transcription factor 4.
Correlation Between IHC Staining Score of PTPRG-AS1 and TCF4 in NSCLC Samples.

Abbreviations: IHC, immunohistochemical; NSCLC, nonsmall cell lung cancer; TCF4, transcription factor 4.
Correlation Between IHC Staining Score of miR-200c-3p and TCF4 in NSCLC Samples.
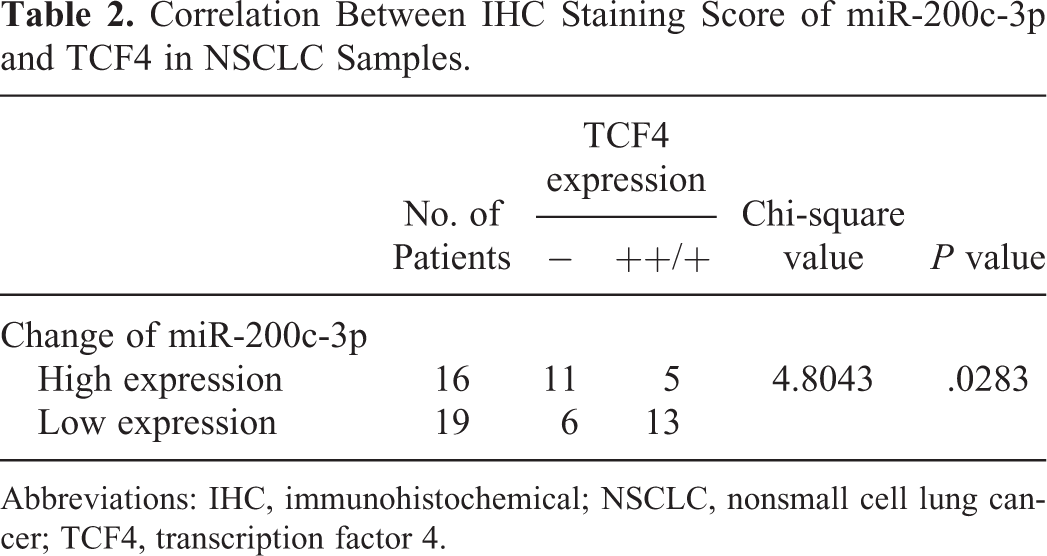
Abbreviations: IHC, immunohistochemical; NSCLC, nonsmall cell lung cancer; TCF4, transcription factor 4.
The Expression Level of PTPRG-AS1 Was Significantly Increased While miR-200c-3p Was Notably Decreased in NSCLC Cells Exposed to X-Ray
Next, the expression levels of PTPRG-AS1 and miR-200c-3p in 5 kinds of NSCLC cells (H157, A549, H838, H460 and H1299 cells) were detected by RT-PCR. Compared with normal bronchial epithelial cell line (16HBE cell line), the expressions of PTPRG-AS1 was significantly higher in the above 5 kinds of NSCLC cells, while the expression of miR-200c-3p was decreased (Figure 2A and B). To determine whether radiation can cause changes in the expressions of PTPRG-AS1 and miR-200c-3p in H157 and H1299 cells, H157 cells and H1299 cells were exposed to 4 Gy radiation. Subsequently, the expression level of PTPRG-AS1 was detected by RT-PCR every 3 hours. The RT-PCR showed that compared with the control group (0 Gy), the expression level of PTPRG-AS1 in H157 cells and H1299 cells was significantly increased after irradiation (Figure 2C), while the expression of miR-200c-3p was suppressed (Figure 2D). The above results showed that PTPRG-AS1 and miR-200c-3p displayed opposite changes under irradiation, which suggested that the radiosensitivity of NSCLC cells was impeded by PTPRG-AS1 but facilitated by miR-200c-3p.
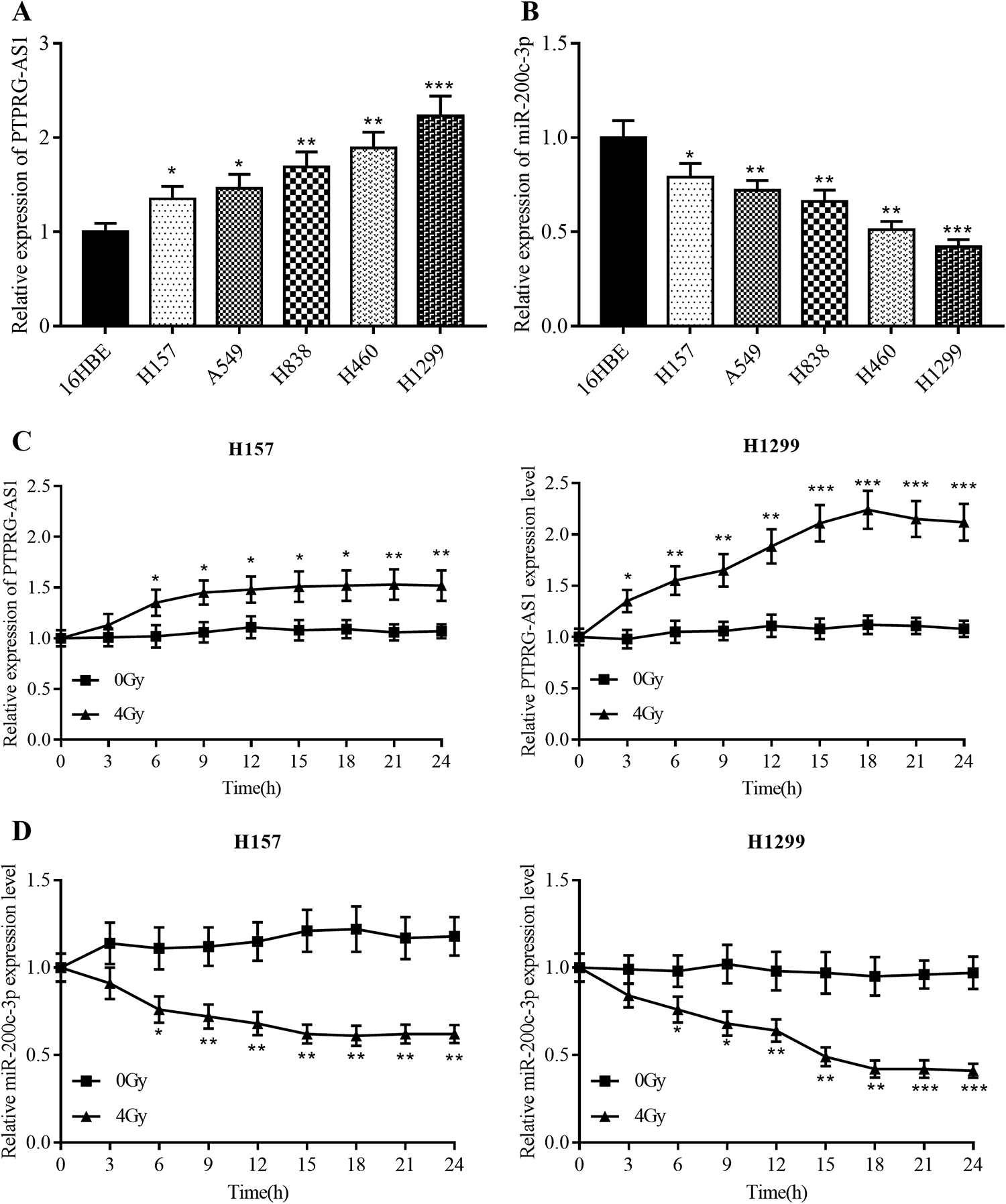
Under irradiation, the expression of PTPRG-AS1 increased significantly, while the expression of miR-200c-3p decreased notably. A and B, The expressions of PTPRG-AS1 and miR-200c-3p in 5 NSCLC cell lines and normal bronchial epithelial cell line 16HBE were detected by RT-PCR. C and D, After treated with X-ray (4 Gy), the expressions of PTPRG-AS1 and miR-200c-3p in H157 cells and H1299 cells were detected every 3 hours. *, **, and *** represent P < .05, P < .01, and P < .001, respectively. NSCLC indicates nonsmall cell lung cancer; RT-PCR, real-time polymerase chain reaction.
PTPRG-AS1 Plays a Prominent Role in Reducing the Radiosensitivity of NSCLC Cells
From the above results, we observed that the expression of PTPRG-AS1 in NSCLC cells was significantly upregulated after radiation, so we made a hypothesis that PTPRG-AS1 could affect the radiosensitivity of NSCLC cells. Among the 5 NSCLC cells, the expression level of PTPRG-AS1 was the lowest in H157 cells and the highest in H1299 cells (Figure 2A), so we transfected PTPRG-AS1 plasmids into H157 cells and PTPRG-AS1 siRNA into H1299 cells. Real-time polymerase chain reaction showed that the cell model with PTPRG-AS1 overexpressed and the cell model with PTPRG-AS1 knocked down were successfully constructed (Figure 3A). On this basis, we detected the viability of H157 and H1299 cells by CCK8 assay. The results showed that the overexpression of PTPRG-AS1 significantly increased the viability of H157 cells, while knocking down PTPRG-AS1 reduced the viability of H1299. This informed us that PTPRG-AS1 could accelerate the proliferation of NSCLC (Figure 3B). Subsequently, the colony number formed by H157 cells and H1299 cells irradiated by different doses of rays was detected. The results demonstrated that overexpressed PTPRG-AS1 would increase the number of colony formation and knockdown PTPRG-AS1 had the opposite effect (Figure 3C, Supplementary Figure 1). Additionally, we transfected the PTPRG-AS1 overexpression plasmid into 16HBE cells. The colony formation experiment confirmed that the overexpression of PTPRG-AS1 significantly promoted the radiation resistance of 16HBE cells (Supplementary Figure 2). These data verified that PTPRG-AS1 could promote the viability of NSCLC cells and reduce the sensitivity of NSCLC cells to radiation.
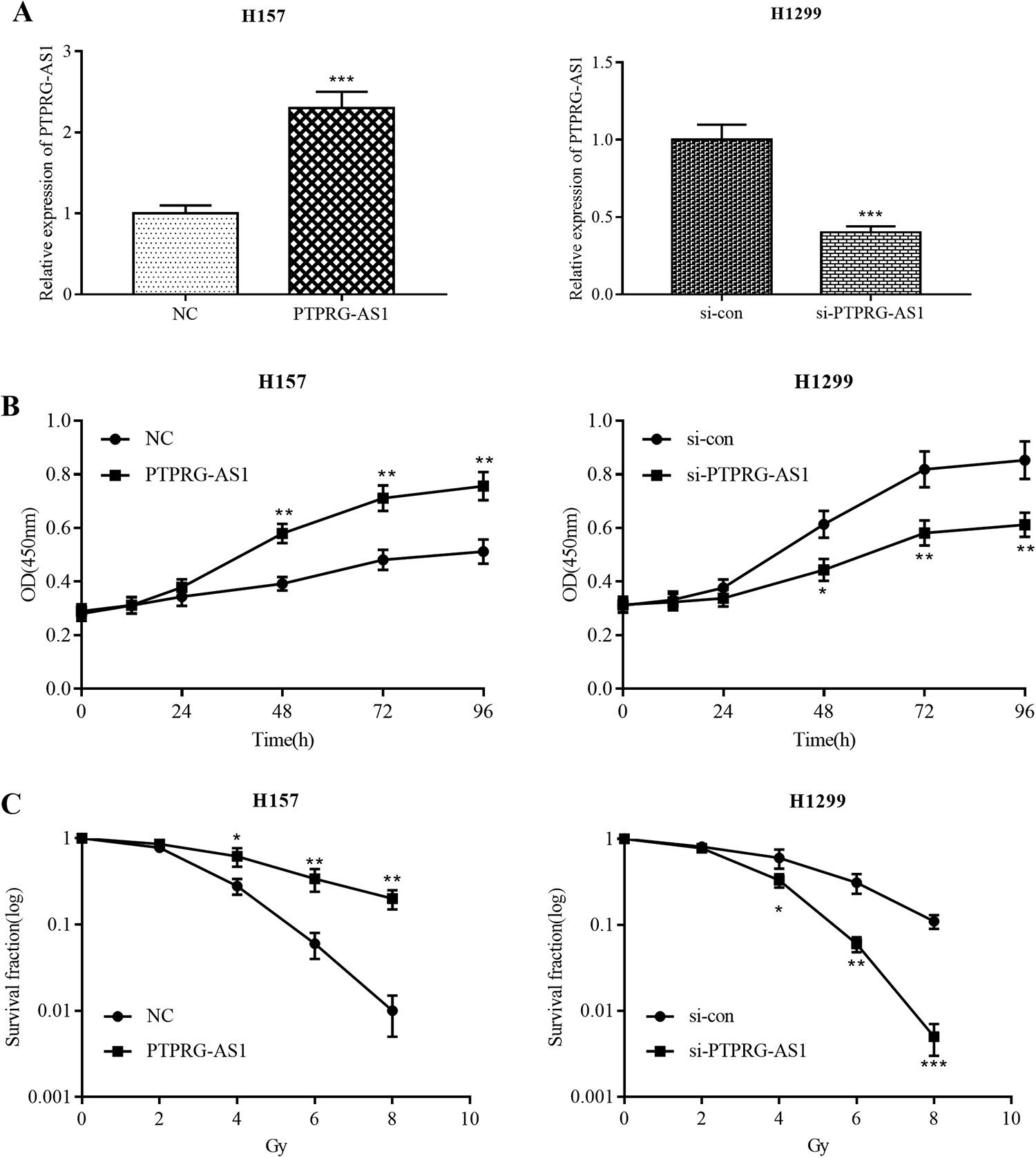
PTPRG-AS1 regulates the viability and radiotherapy resistance of NSCLC cells. A, The NSCLC cells were transfected with pcDNA-NC/pcDNA-PTPRG-AS1, control siRNA/PTPRG-AS1 siRNA, and then the transfection efficiency was detected by RT-PCR. B, The viability of H157 cells and H1299 cells was detected by CCK8 assay. C, After treated with X-ray (0, 2, 4, 6, 8 Gy), the radiosensitivity of NSCLC cells was detected by colony formation assay. *, **, and *** represent P < .05, P < .01, and P < .001, respectively. CCK8 indicates Cell Counting Kit 8; NSCLC, nonsmall cell lung cancer; siRNA, small interfering RNA.
MiR-200c-3p Can Interact Directly With PTPRG-AS1, and Its Expression Level is Negatively Regulated by PTPRG-AS1
Bioinformatics predicts that miR-200c-3p was one of the targeted miRNAs of PTPRG-AS1, and its binding site was shown (Figure 4A). To figure out whether miR-200c-3p can bind directly to PTPRG-AS1, luciferase report vector containing WT or MUT PTPRG-AS1 sequences was constructed. The results showed that miR-200c-3p markedly decreased the luciferase activity of pGL3-PTPRG-AS1-WT but had no significant effect on pGL3-PTPRG-AS1-MUT (Figure 4B). In addition, H157 cells and H1299 cells were transfected with PTPRG-AS1 overexpression plasmid and control plasmid, respectively. The results suggested that PTPRG-AS1 overexpression significantly repressed the expression of miR-200c-3p; conversely, H157 and H1299 transfected with PTPRG-AS1 siRNA displayed an increased miR-200c-3p expression (Figure 4C). Taken together, miR-200c-3p can interact directly with PTPRG-AS1, and its expression level is negatively regulated by PTPRG-AS1.

The interaction between PTPRG-AS1 and miR-200c-3p. A, The binding site between PTPRG-AS1 and miR-200c-3p. B, The predicted binding site between PTPRG-AS1 and miR-200c-3p was validated by dual luciferase reporter gene assay. C, H157 cells and H1299 cells were transfected with pcDNA-NC, pcDNA-PTPRG-AS1, control siRNA, and sPTPRG-AS1 siRNA, respectively, and the expression of miR-200c-3p was detected by RT-PCR. *** represents P < .001. RT-PCR indicates real-time polymerase chain reaction; siRNA, small interfering RNA.
MiR-200c-3p Could Reverse the Function of PTPRG-AS1
To further explore the potential mechanism of radiosensitivity regulated by PTPRG-AS1 in NSCLC cells, H157 cells were transfected with pcDNA-NC, pcDNA-PTPRG-AS1, and pcDNA-PTPRG-AS1 + miR-200c-3p mimics respectively; H1299 cells were transfected with siRNA control, PTPRG-AS1 siRNA, and PTPRG-AS1 siRNA + miR-200c-3p inhibitors, respectively. The results of CCK-8 assay indicated that the inhibitory effect of PTPRG-AS1 knockdown on the viability of NSCLC cells was weakened by miR-200c-3p inhibitors; consistently, miR-200c-3p mimics reversed the effect of PTPRG-AS1 on increasing cell activity (Figure 5A). Moreover, we treated the above groups of cells with different doses of X-ray (0, 2, 4, 6, and 8 Gy). It is worth noting that miR-200c-3p mimics partially weakens the radioresistance of NSCLC cells induced by PTPRG-AS1; miR-200c-3p inhibitors reversed the radiosensitivity induced by PTPRG-AS1 knockdown (Figure 5B, Supplementary Figure 3). Our data supported that PTPRG-AS1 reduced the radiosensitivity of NSCLC cells by regulating the expression of miR-200c-3p.

The function of PTPRG-AS1 was dependent on miR-200c-3p. A, H157 cells were transfected with pcDNA-NC, pcDNA-PTPRG-AS1, and pcDNA-PTPRG-AS1 + miR-200c-3pmimcs, respectively; and H1299 cells were transfected with control siRNA, PTPRG-AS1 siRNA, and PTPRG-AS1 siRNA + miR-200c-3p inhibitors, respectively. Then the cell viability was detected by CCK8. B, After treated with X-ray (0, 2, 4, 6, 8 Gy), the radiosensitivity of NSCLC cells in each group was detected by colony formation assay. *, **, and *** represent P < .05, P < .01, and P < .001, respectively. CCK8 indicates Cell Counting Kit 8; NSCLC, nonsmall cell lung cancer; siRNA, small interfering RNA.
MiR-200c-3p Targeted 3′-UTR of TCF4 Directly
TargetScan database indicated that TCF4 was one of the candidate targets of miR-200c-3p, and their binding site was shown (Figure 6A). Real-time polymerase chain reaction and Western blot showed that TCF4 mRNA and protein decreased notably after miR-con and miR-200c-3p mimics were transfected into H157 and H1299 cells, respectively (Figure 6B and C). The results of dual luciferase reporter assay informed us that miR-200c-3p could bind specifically to 3′-UTR of TCF4 (Figure 6D). Western blot demonstrated that TCF4 expression was markedly increased in H157 and H1299 transfected with PTPRG-AS1 (Figure 6E). Subsequently, we cotransferred the PTPRG-AS1 overexpression plasmid and TCF4 siRNA into H157 cells. Colony formation assay showed that knocking down TCF4 significantly inhibited the viability of H157 cells overexpressing PTPRG-AS1 under radiation (4 Gy; Supplementary Figure 4A). In addition, miR-200c-3p inhibitors and TCF4 siRNA were cotransferred into H157 cells. We found that knocking down TCF4 significantly inhibited the viability of H157 cells with miR-200c-3p inhibited under radiation (4 Gy; Supplementary Figure 4B). Collectively, we come to a conclusion that TCF4 is the downstream gene of miR-200c-3p, and its expression level is negatively regulated by miR-200c-3p in a direct manner while positively modulated by PTPRG-AS1 indirectly. These data indicate that PTRPG-AS1 increases the radioresistance of NSCLC cell via regulating miR-200c-3p and TCF4 (Figure 7).

Transcription factor 4 was regulated by PTPRG-AS1 and miR-200c-3p. A, The binding sites between 3′-UTR of TCF4 and miR-200c-3p. B, After H157 and H1299 cells were transfected with miR-con or miR-200c-3p, the expression of TCF4 mRNA was detected by qRT-PCR .C, The expression of TCF4 was detected by Western blot after H157 and H1299 cells were transfected with miR-con or miR-200c-3p. D, The predicted binding sites between 3′-UTR of TCF4 and miR-200c-3p were validated by dual luciferase reporter assay. E, After PTPRG-AS1 was overexpressed in H157 cells and H1299 cells, the expression of TCF4 was detected by Western blot. *** represents P < .001. mRNA indicates messenger RNA; qRT-PCR, quantitative real-time polymerase chain reaction; TCF4, transcription factor 4; 3′-UTR, 3′-untranslated region.

Graphic Abstract: PTPRG-AS1 promotes the radioresistance of NSCLC cells via regulating miR-200c-3p and TCF4. NSCLC indicates nonsmall cell lung cancer; TCF4, transcription factor 4.
Discussion
Radiotherapy is currently considered to be one of the most effective methods for the treatment of patients with advanced NSCLC. 42,43 Unfortunately, radiation resistance of NSCLC cells caused by mixed factors is a major obstacle. 44 In this study, we found that the high expression of PTPRG-AS1 in NSCLC tissues and cells was negatively correlated with the expression of miR-200c-3p but positively correlated with the expression of TCF4. The aberrant expressions of PTPRG-AS1, miR-200c-3p, and TCF4 are closely linked to the radiosensitivity of NSCLC cells. Specifically, knocking down PTPRG-AS1 increases the radiosensitivity of NSCLC cells via miR-200c-3p/TCF4 axis, while upregulating PTPRG-AS1 has the opposite effect.
Long-chain noncoding RNA is a key regulator involved in the regulation of proliferation, apoptosis, metastasis, and radiation resistance of a variety of tumor cells, including NSCLC cells. 12 -15,45 -48 The studies on lncRNA can provide a basis for early diagnosis, evaluation of prognosis, monitoring progression, and new treatment strategies for NSCLC. 49,50 For example, LINC00152 promotes the proliferation, migration, invasion, and reduces the radiosensitivity of NSCLC cells. 51 Long-chain noncoding RNA GAS5 can inhibit the tumorigenesis of NSCLC and enhance the radiosensitivity of cancer cells. 52 In this study, we observed that NSCLC tissues and cell lines displayed high expression of PTPRG-AS1. What’s more, CCK8 assay showed that, after PTPRG-AS1 was knocked down, the viability of NSCLC cells was significantly decreased, whereas the overexpression of PTPRG-AS1 promoted the proliferation of NSCLC cells. Further, under a range of doses of X-ray irradiation, we found that the colony formation of NSCLC cells was increased significantly after the overexpression of PTPRG-AS1, while the knockdown of PTPRG-AS1 exhibited the opposite effect. These above results suggested that PTPRG-AS1 enhanced the viability of NSCLC cells and induced radioresistance.
Similar to lncRNA, miRNA is involved in multiple cancers and functions as the regulator of radiosensitivity. 52,53 For instance, miR-99a enhances the radiosensitivity of NSCLC cells by targeting the expression of mammalian target of rapamycin. 54 Upregulated miR-328-3p makes NSCLC cells more sensitive to radiotherapy. 55 These studies suggest that miRNAs can be used as markers for evaluation of radiation resistance or potential targets to improve the effectiveness of radiotherapy in NSCLC treatment. In this study, it was demonstrated that miR-200c-3p was lowly expressed in NSCLC tissues and cells. Bioinformatics analysis predicted the existence of binding sites between PTPRG-AS1 and miR-200c-3p. Dual luciferase reporter assay confirmed that PTPRG-AS1 could sponge miR-200c-3p. In addition, in NSCLC cells, overexpressed PTPRG-AS1 could inhibit the expression of miR-200c-3p, while knockdown of PTPRG-AS1 would increase the expression level of miR-200c-3p. Subsequently, it was demonstrated that the viability of NSCLC cells with overexpressed PTPRG-AS1 was significantly inhibited after cotransfected with miR-200c-3p, while the viability of NSCLC cells with PTPRG-AS1 knocked down was significantly enhanced after cotransfection of miR-200c-3p inhibitor. Then, colony formation assay indicated that the inhibition of miR-200c-3p reversed the radiosensitivity induced by the knockdown of PTPRG-AS1. Therefore, we conclude that PTPRG-AS1 reduces the radiosensitivity of NSCLC cells by regulating the expression of miR-200c-3p.
Recent studies have reported the significance of TCF4 in regulating the biological behavior of NSCLC cells, including the sensitivity of cells to radiotherapy. 39,56,57 Overexpression of TCF4 contributes to the hyperactivation of Wnt/β-catenin signal pathway, which is a crucial event in the radioresistance of NSCLC cells. 39 In this study, TargetScan database predicted that there were 2 binding sites between miR-200c-3p and 3′-UTR of TCF4. Subsequently, we found that TCF4 in NSCLC cells transfected with miR-200c-3p mimics was significantly downregulated both at miRNA and protein levels. It is worth noting that miR-200c-3p could markedly reduce the luciferase activity of WT TCF4 reporter but exerts no impact on that of mutated TCF4 reporter, which validated that TCF4 was a target gene of miR-200c-3p. Furthermore, it was demonstrated that the expression of TCF4 was increased by PTPRG-AS1 overexpression. Collectively, we made a conclusion that TCF4 was negatively regulated by miR-200c-3p and positively modulated by PTPRG-AS1 in NSCLC cells, by which PTPRG-AS1/miR-200c-3p modulates the radioresistance of NSCLC cells.
In summary, our findings highlight a novel lncRNA-regulated mechanism in the radioresistance of NSCLC: PTPRG-AS1 enhances radiation resistance of NSCLC cells by regulating miR-200c-3p/TCF4 axis. This work helps clarify the mechanism of NSCLC progression and provides theoretical basis for improving the efficiency of radiotherapy in NSCLC.
Supplemental Material
Supplemental Material, Supplementary_figure_1 - Long Noncoding RNA PTPRG Antisense RNA 1 Reduces Radiosensitivity of Nonsmall Cell Lung Cancer Cells Via Regulating MiR-200c-3p/TCF4
Supplemental Material, Supplementary_figure_1 for Long Noncoding RNA PTPRG Antisense RNA 1 Reduces Radiosensitivity of Nonsmall Cell Lung Cancer Cells Via Regulating MiR-200c-3p/TCF4 by Qiang Ma, Rungui Niu, Wei Huang, Liangshan Da, Yanlei Tang, Daowen Jiang, Yanfeng Xi and Congjun Zhang in Technology in Cancer Research & Treatment
Supplemental Material
Supplemental Material, Supplementary_figure_2 - Long Noncoding RNA PTPRG Antisense RNA 1 Reduces Radiosensitivity of Nonsmall Cell Lung Cancer Cells Via Regulating MiR-200c-3p/TCF4
Supplemental Material, Supplementary_figure_2 for Long Noncoding RNA PTPRG Antisense RNA 1 Reduces Radiosensitivity of Nonsmall Cell Lung Cancer Cells Via Regulating MiR-200c-3p/TCF4 by Qiang Ma, Rungui Niu, Wei Huang, Liangshan Da, Yanlei Tang, Daowen Jiang, Yanfeng Xi and Congjun Zhang in Technology in Cancer Research & Treatment
Supplemental Material
Supplemental Material, Supplementary_figure_3 - Long Noncoding RNA PTPRG Antisense RNA 1 Reduces Radiosensitivity of Nonsmall Cell Lung Cancer Cells Via Regulating MiR-200c-3p/TCF4
Supplemental Material, Supplementary_figure_3 for Long Noncoding RNA PTPRG Antisense RNA 1 Reduces Radiosensitivity of Nonsmall Cell Lung Cancer Cells Via Regulating MiR-200c-3p/TCF4 by Qiang Ma, Rungui Niu, Wei Huang, Liangshan Da, Yanlei Tang, Daowen Jiang, Yanfeng Xi and Congjun Zhang in Technology in Cancer Research & Treatment
Supplemental Material
Supplemental Material, Supplementary_figure_4 - Long Noncoding RNA PTPRG Antisense RNA 1 Reduces Radiosensitivity of Nonsmall Cell Lung Cancer Cells Via Regulating MiR-200c-3p/TCF4
Supplemental Material, Supplementary_figure_4 for Long Noncoding RNA PTPRG Antisense RNA 1 Reduces Radiosensitivity of Nonsmall Cell Lung Cancer Cells Via Regulating MiR-200c-3p/TCF4 by Qiang Ma, Rungui Niu, Wei Huang, Liangshan Da, Yanlei Tang, Daowen Jiang, Yanfeng Xi and Congjun Zhang in Technology in Cancer Research & Treatment
Footnotes
Authors’ Note
RGN, CJZ, DWJ, and YFX conceived and designed the experiments. QM, RGN, YLT, and YFX performed the experiments. QM, WH, and LSD revised the manuscript after peer-review. YFX performed the statistical analysis. QM, RGN, and YFX wrote the paper. All authors read and approved the final manuscript. The data used to support the findings of this study are available from the corresponding author upon request. Daowen Jiang is also affiliated with Department of Chest Surgery, Zhongshan Hospital, Fudan University, Shanghai, China.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
Abbreviations
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
