Abstract
Introduction
MicroRNAs (miRs) may be important regulators of risk for type 2 diabetes (T2D). Circulating miRs may provide information about which individuals are at risk for T2D. The purpose of this study was to assess longitudinal associations between circulating miR expression and variability in fasting blood glucose (FBG) and to identify miR-targeted genes and biological pathways.
Methods
Variability in FBG was estimated using standard deviation from participants (n = 20) in a previously completed yoga trial. Expression of 402 miRs was measured using hydrogel particle lithography. MirTarBase was used to identify mRNAs, and miRPathDB was used to identify pathways targeted by differentially expressed miRs.
Results
Six circulating miRs (miR-192, miR-197, miR-206, miR-424, miR-486, and miR-93) were associated with variability in FBG and targeted 143 genes and 23 Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways. Six mRNAs (AKT1, CCND1, ESR1, FASN, SMAD7, and VEGFA) were targeted by at least two miRs and four of those were located in miR-targeted KEGG pathways.
Conclusions
Circulating miRs are associated with variability in FBG in individuals at risk for T2D. Further studies are needed to determine whether miRs may be prodromal biomarkers that can identify which individuals are at greatest risk to progress to T2D and which biological pathways underlie this risk.
Introduction
Circulating miRs found in serum and plasma are easily measured in blood and have the potential to be clinically useful biomarkers for risk for development of type 2 diabetes (T2D), exhibiting changes in expression levels prior to the onset of prediabetes and T2D. 1,2 The goal of this study was to characterize the associations between expression levels of circulating miRs and changes in fasting blood glucose (FBG) over time in individuals at risk for T2D. To the best of our knowledge, this is the first study to integrate repeated measures of FBG over time with expression levels of miRs in order to better understand which individuals are likely to progress toward T2D over time and to identify the biological pathways that may be targeted in these individuals.
Methods
Study sample
The study sample included participants from the previously completed Practicing Restorative Yoga versus Stretching for the Metabolic Syndrome (PRYSMS) clinical trial (clinicaltrials.gov identifier: NCT01024816). A randomly selected subset of participants with high (n = 10) versus low (n = 10) variability in FBG over the 12-month intervention, matched for age, sex, and study intervention arm, were included. Blood was collected by venipuncture into EDTA-containing vacutainers, centrifuged at 4°C to separate plasma from cellular components, and stored at −80°C. The Firefly Bioworks Multiplex Circulating MicroRNA Assay (Abcam, MA) was used for direct quantification of miRs from plasma in two separate groups of 10 participants each (Supplementary Materials). Quantitative polymerase chain reaction (qPCR) was performed to validate expression of a subset of miR targets using standard protocols (Supplementary Materials). Descriptive statistics and Student’s t-test were used to evaluate demographic and clinical characteristics of participants (Stata version 13, College Station, TX). Variability in FBG over time was estimated by calculating the standard deviation across 5 time-points over 12-months with imputation of missing values using the mean within participant (Stata version 15, College Station, TX). Differential expression of individual miRs was determined using Student’s t-test (Stata version 16). False discovery rate (FDR) threshold of 0.2 was applied. The miRTarBase database (Release 7.0) was used to identify messenger RNA (mRNA) targets of significantly differentially expressed miRs. 3 The miRPathDB database (Release 1.0) was used to identify KEGG pathways targeted by the miRs found to be significantly differentially expressed. 4
Results
Participant characteristics
There were no significant differences in baseline demographic or clinical characteristics between the individuals who showed variable compared to stable FBG over 12 months (Table S1). The standard deviation of FBG in the variable group ranged from 12 to 25 mg/dL and that of the stable group ranged from 3 to 5 mg/dL. A total of 30 miRs were differentially expressed in the first group of participants, and 28 miRs were differentially expressed in the second group of participants (FDR < 0.2 for both groups). Six miRs (i.e., miR-192, miR-197, miR-206, miR-424, miR-486, and miR-93) showed overlap between both groups. Validation by qPCR for a subset of these overlapping miRs (i.e. miR-192, miR-486, and miR-93) confirmed our initial findings, showing the same direction of differential expression for all three miR targets (data not shown).
The miRTarBase tool 3 identified a total of 143 mRNAs with strong experimental evidence of being targets of these six miRs (Table S2). MiR-192 targets 41 mRNAs, miR-197 targets 1 mRNA, miR-206 targets 33 mRNAs, miR-424 targets 31 mRNAs, miR-486 targets 6 mRNAs, and miR-93 targets 31 mRNAs. There were six mRNAs that were targeted by two miRs: AKT serine/threonine kinase 1 (AKT1; miR-192 and miR-206), cyclin D1 (CCND1; miR-206 and miR-424), estrogen receptor 1 (ESR1; miR-192 and miR-206), fatty acid synthase (FASN; miR-424 and miR-486), SMAD family member 7 (SMAD7; miR-93 and miR-424), and vascular endothelial growth factor A (VEGFA; miR-93 and miR-206).
Overlapping mRNA and KEGG pathway targets.
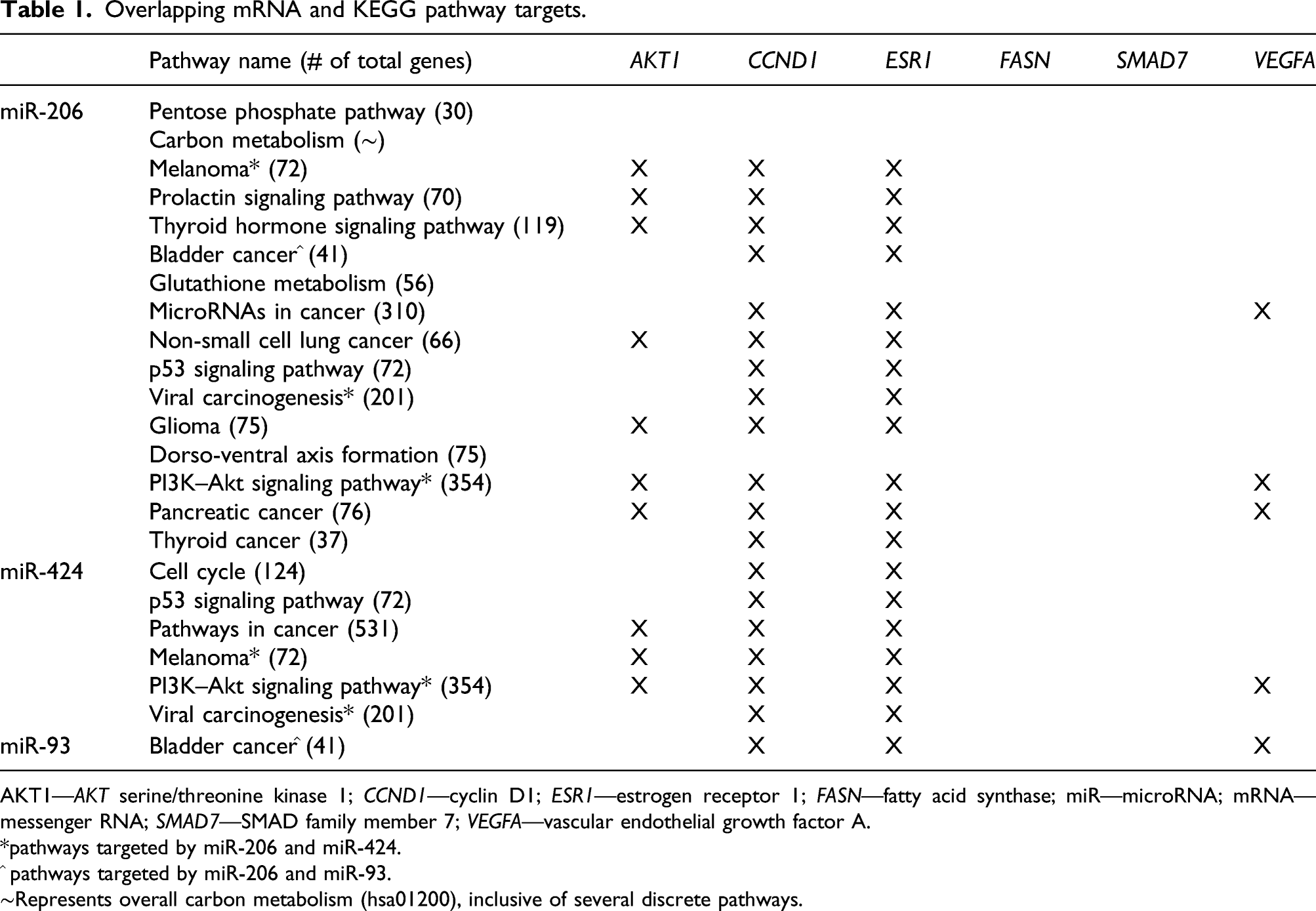
AKT1—AKT serine/threonine kinase 1; CCND1—cyclin D1; ESR1—estrogen receptor 1; FASN—fatty acid synthase; miR—microRNA; mRNA—messenger RNA; SMAD7—SMAD family member 7; VEGFA—vascular endothelial growth factor A.
*pathways targeted by miR-206 and miR-424.
^ pathways targeted by miR-206 and miR-93.
∼Represents overall carbon metabolism (hsa01200), inclusive of several discrete pathways.
Discussion
The potential clinical applications of this study include identification of novel biomarkers that detect risk for T2D earlier than changes in FBG, and discovery of the biological mechanisms that underlie risk for T2D and may be future therapeutic targets. For many miRs, the biological functions are currently poorly defined. There exists evidence for roles of the miRs identified in this study in the pathophysiology of T2D. A prior genome-wide association study found an association between insulin-like growth factor binding protein 5 (IGFBP5), which is an mRNA target of miR-197, and adipose tissue in women. 5 MiR-424 has been associated with insulin resistance 6 and pancreatic β-cell function, 7 as well as islet autoimmunity.8,9 MiR-206 may be related to regulation of glucose metabolism in mice. 10 MiR-93 also has some evidence for a role in processes related to T2D, including insulin resistance and subsequent polycystic ovarian syndrome. 11 These prior studies suggest a plausible role for involvement of these specific miRs in T2D that merits further investigation.
Among the mRNAs targeted by multiple miRs that were identified by this study, we identified AKT1 as an mRNA target of miR-192 and miR-206. This gene is also located in 8 of the 19 identified pathways targeted by differentially expressed miRs, including pathways targeted by miR-206. This gene is highly expressed in pancreatic tissue as well as other tissues related to the pathophysiology of T2D, including liver. Numerous studies provided experimental evidence for a role of AKT1 in the pathophysiology of T2D. 12 An mRNA targeted by multiple miRs was VEGFA, which is located in the PI3K–Akt signaling pathway that is targeted by miR-206 and miR-424. Numerous studies reported evidence that this pathway is relevant to the pathophysiology of T2D with activity in the pancreas, skeletal muscle, adipose tissue, and liver. 12 A primary function of VEGFA is to regulate vascular endothelial function, including in diabetic nephropathy and retinopathy, and this gene is a validated target of miR-126. FASN was an identified target of miR-424 and miR-486 that is expressed almost exclusively in adipose tissue. There are several studies that identified experimental evidence to support a role for FASN in some aspect of risk for or complications from T2D. 13 The gene SYK, which is targeted by miR-486, is also located in pathways where FASN is present (i.e., fatty acid biosynthesis, fatty acid metabolism, and insulin signaling pathways). SMAD7 is targeted by miR-93 and miR-424, but is not located in any of the 19 identified targeted pathways. However, this gene is highly expressed in thyroid and urinary/bladder tissues, and the thyroid hormone signaling (miR-206), thyroid cancer (miR-206), and bladder cancer (miR-206, miR-93) pathways were identified targets of the differentially expressed miRs in this study.
The group characterized by low variability in FBG also had a less healthy metabolic profile overall. The individual miRs identified in this study may be capturing the co-occurring risk factors that characterize metabolic syndrome, as opposed to isolated changes in FBG. We did not find strong experimental evidence that the differentially expressed miRs in this study target the KEGG pathways that are annotated to T2D. However, a limitation of pathway annotation databases is that naming conventions may erroneously bias toward a specific disease. In addition, our focus on target mRNA genes requiring strong experimental evidence may have resulted in the exclusion of additional genes that may be enriched in miR-targeted and T2D pathways. Also, we assessed extracellular miRs from plasma, which may not reflect the underlying expression levels in discrete tissue types.
Prior studies focused on miRs associated with risk for T2D have primarily assessed miR expression at a single time point. The purpose of this study was to build on prior cross-sectional findings and begin to explore longitudinal relationships between circulating miRs and FBG over 1 year in individuals at risk for T2D. We sought to build on observational studies by providing in silico analysis of the functional mRNA and pathway targets of the miRs identified in this study. We validated one miR that was previously identified in cross-sectional studies focused on T2D, five miRs with associations with T2D and related mechanisms, and one miR with less well-characterized biological function. There is strong experimental evidence that these differentially expressed miRs target mRNA known to be involved in the pathophysiology of T2D as well as biological pathways that include mRNA related to the pathophysiology of T2D. This study provided further evidence that miRs may be biomarkers associated with prodromal risk for T2D, and may provide insights about the underlying pathophysiology of T2D, and inform potential therapeutic targets for treatments to reduce risk for T2D.
Supplemental Material
sj-pdf-1-dvr-10.1177_14791641211055837 – Supplemental Material for Circulating microRNAs are associated with variability in fasting blood glucose over 12-months and target pathways related to type 2 diabetes: A pilot study
Supplemental Material, sj-pdf-1-dvr-10.1177_14791641211055837 for Circulating microRNAs are associated with variability in fasting blood glucose over 12-months and target pathways related to type 2 diabetes: A pilot study by Elena Flowers, Isabel E Allen, Alka M Kanaya and Bradley E Aouizerat in Diabetes & Vascular Disease Research
Supplemental Material
sj-pdf-2-dvr-10.1177_14791641211055837 – Supplemental Material for Circulating microRNAs are associated with variability in fasting blood glucose over 12-months and target pathways related to type 2 diabetes: A pilot study
Supplemental Material, sj-pdf-2-dvr-10.1177_14791641211055837 for Circulating microRNAs are associated with variability in fasting blood glucose over 12-months and target pathways related to type 2 diabetes: A pilot study by Elena Flowers, Isabel E Allen, Alka M Kanaya and Bradley E Aouizerat in Diabetes & Vascular Disease Research
Supplemental Material
sj-pdf-3-dvr-10.1177_14791641211055837 – Supplemental Material for Circulating microRNAs are associated with variability in fasting blood glucose over 12-months and target pathways related to type 2 diabetes: A pilot study
Supplemental Material, sj-pdf-3-dvr-10.1177_14791641211055837 for Circulating microRNAs are associated with variability in fasting blood glucose over 12-months and target pathways related to type 2 diabetes: A pilot study by Elena Flowers, Isabel E Allen, Alka M Kanaya and Bradley E Aouizerat in Diabetes & Vascular Disease Research
Footnotes
Acknowledgements
Molecular data collection was performed by the Helen Diller Family Comprehensive Cancer Center Laboratory for Cell Analysis Shared Resource Facility (P30CA082103).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Dr Flowers who was supported by the National Center for Advancing Translational Sciences of the National Institutes of Health grant number KL2TR000143 and the Hellman Family Foundation. The PRYSMS study was supported by the National Center for Complementary and Alternative Medicine of the National Institutes of Health grant number R01AT004569. Molecular data collection from the PRYSMS study was supported by the National Institute of Diabetes, Digestive, and Kidney Disease of the National Institutes of Health grant number R21DK117346. Dr Kanaya is supported by National Heart Lung, and Blood Institute of the National Institutes of Health grant number 2K24HL112827.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
