Abstract
Background and aim:
Many reported studies have been conducted to investigate the association of angiotensin II type 1 receptor (AT1R) A1166C gene polymorphism with myocardial infarction (MI) susceptibility. However, the results from those reports are still conflicting. This meta-analysis was performed to study the relationship between AT1R A1166C gene polymorphism and MI risk.
Method:
The databases of PubMed, Embase, and Cochrane Library were searched as of 1 March 2012, and eligible investigations were recruited into this meta-analysis.
Results:
Eighteen investigations were identified for the analysis of association between AT1R A1166C gene polymorphism and MI risk, 11 in Caucasians, three in Asians, two in Africans, one in the population of Brazil and one in the population of Durban, South Africa . There was a marked association between AT1R C allele and MI susceptibility for overall populations (odds ratio (OR)=1.12, 95% confidence interval (CI): 1.01–1.25, p=0.03), and AT1R AA genotype was associated with a lower risk of MI in overall populations (OR=0.87, 95% CI: 0.78–0.98, p=0.02). However, AT1R A1166C gene polymorphism was not associated with MI risk in the sub-groups of Caucasians, Asians, Africans, Brazil and Durban populations.
Conclusions:
C allele is a risk factor for the MI susceptibility in overall populations, and AA genotype might be a protective factor against the MI risk in overall populations. However, more case-control association investigations on larger, stratified populations are required in the future.
Introduction
Myocardial infarction (MI) is a public health problem, and it brings various health problems and threatens the life of patients. MI is a multifactorial disease with a complex pathogenesis where lifestyle, individual genetic background and environmental risk factors are involved. 1 However, the pathogenesis of MI remains difficult to elucidate at the moment. Early diagnosis for MI risk is difficult as there is no well-documented indicator which can predict the onset of MI. In past decades, lots of studies found that gene polymorphism might be associated with the risk of some diseases and might become a potential marker for early diagnosis of the onset of some diseases.2–5
The angiotensin II receptor includes two subtypes: type 1 receptor (angiotensin II type 1 receptor, AT1R) and type 2 receptor. 6 The AT1R, which is the primary pathogenic effector for angiotensin II, is a member of the G-protein-coupled receptor superfamily expressed in most tissues, where receptor activation leads to vasoconstriction, water retention and vascular smooth muscle cell proliferation and hypertrophy. 7 The AT1R is present predominantly in vascular smooth muscle cells and because of the physiological role of the AT1R in the cardiovascular system, the gene that encodes this molecule is an excellent candidate in the etiology of cardiovascular diseases 8 AT1R A1166C gene polymorphism, an important mutation of AT1R, might be implicated in the etiology of MI susceptibility and had been investigated in numerous epidemiologic studies. Zhang et al. 9 performed the first meta-analysis to study the association of the AT1R gene A1166C polymorphisms with MI risk, and found an association of the C allele of AT1R gene A1166C polymorphisms with MI risk. This meta-analysis was performed to update the conclusion of the relationship between AT1R A1166C gene polymorphism and MI susceptibility, with the intention to provide a much more reliable finding of the significance of this association. Furthermore, this meta-analysis first explored the association of AT1R A1166C gene polymorphism with MI susceptibility for different ethnicities.
Materials and methods
Search strategy
Search strategy for the association of AT1R A1166C gene polymorphism with MI risk
Relevant studies were sought from the electronic databases of PubMed, Embase, and Cochrane Library on 1 March 2012. The retrieval strategy was that (angiotensin II type-1 receptor) AND polymorphism AND (myocardial infarction) were entered into these databases. The search in PubMed or Embase was limited to English language. Additional articles were identified through references cited in the retrieved articles.
Inclusion and exclusion criteria
Inclusion and exclusion criteria for MI
Inclusion criteria were (a) a case–control study; (b) the outcome had to be MI; (c) there had to be at least two comparison groups (MI group vs control group); (d) investigation on the association of AT1R A1166C gene polymorphism with MI risk.
Exclusion criteria were (a) review articles and editorials; (b) case reports; (c) investigation did not provide detailed data of AT1R A1166C genotype distributions; (d) preliminary results not about AT1R A1166C gene polymorphism (investigation on the association of other gene polymorphism, such as angiotensin-converting enzyme (ACE) insertion/deletion gene polymorphism, with MI risk) or MI risk (investigation on the association of AT1R A1166C gene polymorphism with the risk of other diseases, such as renal disease); (e) investigating the role of AT1R inhibitors in diseases.
Data extraction and synthesis
Two investigators extracted the following information from each eligible study independently: first author’s surname, year of publication and the number of cases and controls for AT1R A1166C genotypes. The frequency of C allele was calculated for the case group or the control group, from the corresponding genotype distribution. The results were compared and disagreements were resolved by discussion.
Statistical analysis
Cochrane Review Manager Version 5 (Cochrane Library, UK) was used to calculate the available data from each investigation. The pooled statistic was counted using the fixed effects model, but a random effects model was performed when the p value of heterogeneity test was less than 0.1.10,11 Results were expressed with odds ratio (OR) for dichotomous data, and 95% confidence interval (CI) were also calculated.12,13 For the pooled OR to be statistically significant, p<0.05 was required. I2 was used to test the heterogeneity among the included studies.14,15 In order to avoid excessive comparisons, the OR was calculated by using three methods: method 1, allele comparison (C allele vs A allele); method 2, comparing CC homozygous with the other two combinations (CC vs CA+AA); method 3, comparing AA genotype with the other two combinations (AA vs CC+CA). A chi-square (χ2) test using a web-based program was applied to determine if genotype distributions of the control population reported conformed to Hardy-Weinberg equilibrium (HWE; p<0.05 was considered significant), and if the genotype distributions in the controls significantly deviated from HWE then the study was excluded from our sensitive analysis. The Begg adjusted rank correlation test 16 and the Egger regression asymmetry test 17 were used for exploring publication bias (p<0.1 was considered significant) when the sample number was more than 10. All descriptive data were expressed as mean±standard deviation (SD).
Results
Study characteristics for MI
A total of 115 articles were retrieved in PubMed (70), Embase (43), and Cochrane Library (2). Ninety-seven articles were excluded for the following reasons: review articles, editorials, case reports, preliminary result not about AT1R A1166C gene polymorphism or MI, investigating the role of AT1R inhibitors in diseases. Eighteen articles were recruited into our investigation for the relationship between AT1R A1166C gene polymorphism and MI susceptibility (Figure 1). The data of interest were extracted: first author’s surname, year of publication and the number of cases and controls for AT1R A1166C genotypes (Table 1). Those 18 investigations18–35 contained 6794 cases and 6204 controls, including 11 studies of Caucasians,18–28 three studies of Asians,29,30,35 two studies of Africans,31,34 one study in the population of Durban in South Africa, 32 and one study in the Brazilian population. 33 The average frequency of C allele distribution in patients with MI for overall populations was 24.74% and the average frequency in controls was 22.58%. The average frequency of C allele distribution in Caucasians was 29.43% in cases and 27.83% for controls. Furthermore, the average frequency of C allele distribution in Asians with MI was 15.11%, and the average frequency in controls was 11.27%. In African population, the average frequency of C allele distribution in MI patients was 22.05%, and the average frequency in controls was 17.92%. In Durban population, the frequency of C allele distribution in MI patients was 4.38%, and the frequency in controls was 5.18%. In Brazil population, the frequency of C allele distribution in MI patients was 27.73%, and the frequency in controls was 25.48%. The average frequency of C allele in patients with MI for overall populations was similar to that in controls (MI/controls=1.10). The ratio of MI/control for the average frequency of C allele distribution in Asians or Africans was notably higher than that in Caucasians, the Durban population, or the Brazil population (Asians: MI/control=1.34; Africans: MI/control=1.23; Caucasians: MI/control=1.06; Durban population: MI/control=0.85; Brazil population: MI/control=1.09).

Flow chart for our meta-analysis.
Characteristics of the studies evaluating the effects of angiotensin II type 1 receptor (AT1R) A1166C gene polymorphism on myocardial infarction (MI) risk.

HWE: Hardy–Weinberg equilibrium.
Association of AT1R A1166C gene polymorphism with MI risk
In this meta-analysis, a significant association between C allele and MI risk was observed in overall populations (Figure 2(A); Table 2). However, the CC genotype was not associated with the MI susceptibility (Figure 2(B); Table 2). Interestingly, AA genotype might be a protective factor against the risk of MI in overall populations (Figure 2(C); Table 2).
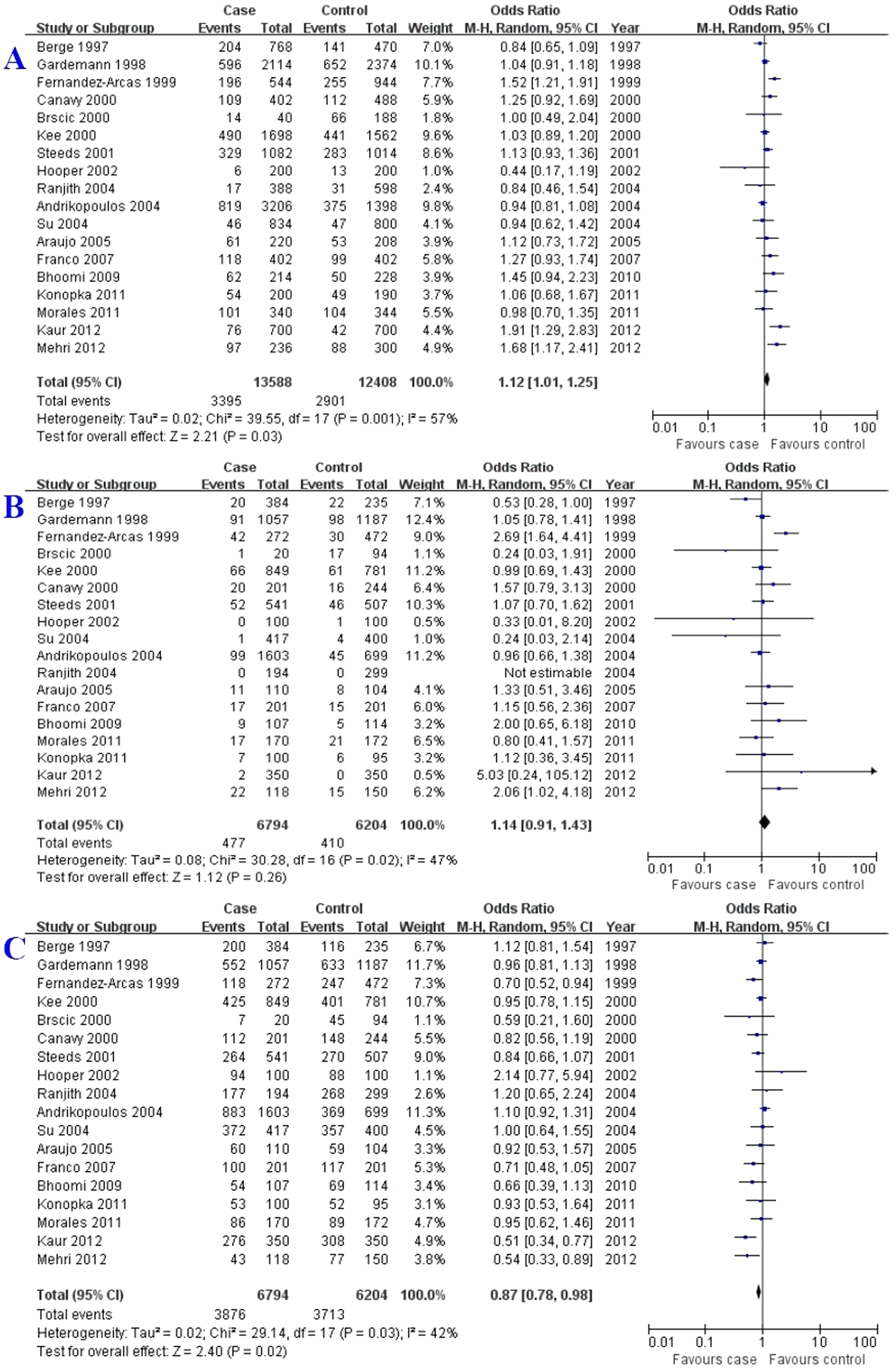
Association of AT1R A1166C gene polymorphism and MI susceptibility in overall populations: (A) C vs A; (B) CC vs CA+AA; (C) AA vs AC+CC.
Meta-analysis of the association of angiotensin II type 1 receptor (AT1R) A1166C gene polymorphism with risk of myocardial infarction (MI).
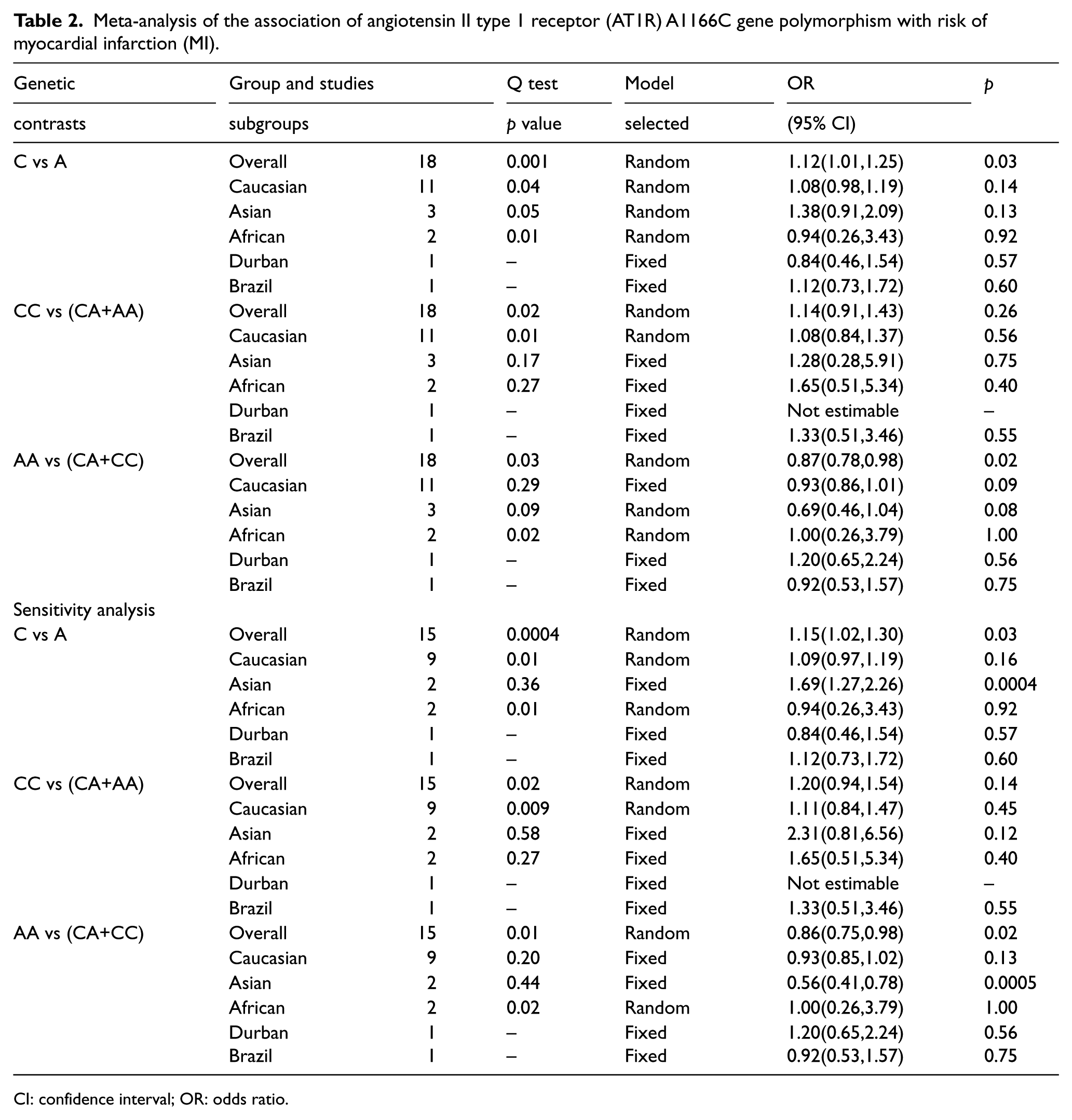
CI: confidence interval; OR: odds ratio.
Difference in race might have affected the results of our analysis for the association of AT1R A1166C gene polymorphism with MI susceptibility. In order to evaluate this effect, we divided the population by ethnicity. There was no association between AT1R A1166C gene polymorphism and MI susceptibility in Caucasians, Asians, Africans, populations of Durban and Brazil (Table 2).
Sensitivity analysis
Three studies (two in Caucasians22,25 and one in Asians 29 ) for which the genotype distributions in the controls were significantly deviated from HWE were excluded.
In the sensitivity analysis for overall populations, we found that the results were similar with those in the non-sensitivity analysis. A significant association between C allele and MI risk was observed in overall populations, and AA genotype might be a protective factor against the risk of MI in overall populations (Table 2).
In Caucasians, the results in sensitivity analysis were similar to those in the non-sensitivity analysis (Table 2). Furthermore, the results from sensitivity analysis in Africans, the Durban population and the Brazil population were the same as those from the non-sensitivity analysis (Table 2). Interestingly, in Asians, there was a significant association between C allele and MI risk, and AA genotype might be a protective factor against the risk of MI (Table 2). These results from the Asian studies had a sensitivity analysis inconsistent with those in the non-sensitivity analysis.
Testing for publication bias for the analysis of association of AT1R A1166C gene polymorphism and MI susceptibility in overall populations, and in Caucasians
There was no significant publication bias in overall populations (Begg: p=0.967; Egger: p=0.893; Figure 3(A)) and in Caucasians (Begg: p=0.161; Egger: p=0.112; Figure 3(B)).

Publication bias for the analysis of association of angiotensin II type 1 receptor (AT1R) A1166C gene polymorphism and myocardial infarction (MI) susceptibility: (A) publication bias for overall populations; (B) publication bias for Caucasians.
Discussion
The AT1R is the primary pathogenic effector for angiotensin II, which takes part in the etiology of cardiovascular diseases, such as MI. In our study, we found that AT1R A1166C gene polymorphism was associated with MI susceptibility in overall populations. In our meta-analysis, there was a significant association between the C allele and MI risk in overall populations, and AA genotype was associated with a lower risk of MI in overall populations. However, CC genotype was not associated with MI risk for overall populations. We also performed a sensitivity analysis, and found that the results for overall populations were similar to those in the non-sensitivity analysis. There was no publication bias in overall populations. The results from the overall populations might be robust to some extent.
According to the duration time of the MI, MI can be divided into acute MI and ‘old’ MI, etc. Diagnosis of definite AMI was based on World Health Organization (WHO) criteria, i.e. patients should have any two of the following three conditions: resting anginal chest pain lasting more than 30 min, ST segment elevation >1 mm above baseline in at least two leads of a standard 12-lead electrocardiogram, and creatinine phosphokinase (CPK) and CPK-MB elevation to at least twice the upper limit of the normal range. Diagnosis of definite old MI was based on WHO criteria, and the duration of the MI was more than three months. In this study, we also performed a meta-analysis to investigate the association of AT1R A1166C gene polymorphism with acute MI/old MI susceptibility. Ten reports19,23,24,26,29,30,32–35 were included for acute MI, and two reports25,27 were included for old MI. Whether the MI in the other studies was acute MI or old MI was not clear. In our study, we found that the results for acute MI were similar to those from MI (acute MI: C allele: OR=1.17, 95% CI: 1.01–1.34, p=0.03; CC genotype: OR=1.10, 95% CI: 0.92–1.32, p=0.28; AA genotype: OR=0.84, 95% CI: 0.71–0.99, p=0.03). However, AT1R A1166C gene polymorphism was not associated with old MI susceptibility (old MI: C allele: OR=1.02, 95% CI: 0.89–1.17, p=0.77; CC genotype: OR=0.95, 95% CI: 0.69–1.30, p=0.74; AA genotype: OR=0.95, 95% CI: 0.80–1.13, p=0.58). The number of included studies for old MI was small, and the results might be less robust.
The frequency of the C allele in some studies29,31,32,35 was very low compared to that in other included studies, and those studies were excluded from our further meta-analysis. The frequency of C allele in the case group of the study from Mehri et al. 34 was much higher than that in the control group (41.1% vs 29.33%), and this study was also excluded from our further meta-analysis. We re-ran the meta-analysis and found that the pooled OR for the C allele was favorable to the case group, although the difference was not statistically significant (OR=1.09, 95% CI: 0.99–1.19, p=0.07). Furthermore, the pooled OR for the AA allele was favorable to the control group, although the difference was not statistically significant (OR=0.93, 95% CI: 0.86–1.00, p=0.05). However, CC genotype was not associated with the risk of MI (p=0.24). More well-designed studies should be conducted in the future.
Ethnic variables might have affected the evaluation of the association of gene polymorphism with the susceptibility of disease.36,37 In this meta-analysis, we also tested the ethnic variable in the evaluation of association of AT1R A1166C gene polymorphism with MI susceptibility. Most of the recruited studies were from Caucasians. We found that the AT1R A1166C gene polymorphism was not associated with MI susceptibility in Caucasians. Furthermore, the results for Caucasians were similar to those in the non-sensitivity analysis. There was no publication bias in studies of Caucasians. The difference of the C allele distribution between the MI group and control group in Caucasians was not notable (MI/control=1.06). To sum up the findings mentioned above, AT1R A1166C gene polymorphism was not associated with MI susceptibility in Caucasians, and the result might be robust to some extent.
In Caucasians, Berget al. 18 reported that CC homozygosity appeared to be associated with MI in the Norwegian population. Fernandez-Arcas et al. 20 found that the CC genotype is an independent risk factor for MI. Canavy et al. 21 reported the prevalence of the C allele carriers being higher in patients with MI than in control individuals. However, some other studies reported that AT1R A1166C gene polymorphism was not associated with the MI risk19,22–25,27 AT1R A1166C gene polymorphism might be not associated with MI susceptibility in Caucasians. However, more studies in Caucasians should be performed in the future.
In Asians, AT1R A1166C gene polymorphism might be not associated with MI susceptibility. However, in the sensitivity analysis, there was a significant association between the C allele and MI risk, and AA genotype was associated with a lower risk of MI. These results from the sensitivity analysis were inconsistent with those in the non-sensitivity analysis. However, there were only three studies in our meta-analysis for Asians, and only two studies in the sensitivity analysis. The difference of C allele distribution between MI group and control group in Asians was notable (MI/control=1.34). To sum up those findings mentioned above, AT1R A1166C gene polymorphism might be associated with the MI susceptibility in Asians.
In Asians, Su et al. 29 reported that AT1R A1166C gene polymorphism might affect the MI susceptibility in Chinese. Kaur et al. 35 found that the AT1R 1166A/C gene polymorphism had an association with AMI among north Indian patients. However, Bhoomi et al. 30 reported that the AT1R A1166C CC genotype is not a risk factor for MI in patients in a South Indian population. The number of the included studies for Asians was small, and it would be difficult for us to draw a robust conclusion for Asians. More studies in Asians should be performed in the future.
In Africans, the Durban population and the Brazilian population, the conclusions from the sensitivity analysis were inconsistent with those in the non-sensitivity analysis. There was no association between AT1R A1166C gene polymorphism and MI susceptibility in Africans, the Durban population, and Brazilian population. The number of the included studies for Africans, the Durban population, or Brazilian population was much too small, and it would be difficult for us to draw a robust conclusion from them. Further studies in Africans, the Durban population, and the Brazilian population should be performed.
Interestingly, Zhang et al. 9 performed the first meta-analysis to study the association of the AT1R gene A1166C polymorphisms with MI risk, and found an association of the C allele of AT1R gene A1166C polymorphisms with MI risk in overall populations. It was similar with our result in this meta-analysis. In this meta-analysis, we found that the AT1R A1166C gene polymorphism was not associated with the MI susceptibility in Caucasians, and there was no significant publication bias. Interestingly, AT1R A1166C gene polymorphism might be associated with the MI susceptibility in Asians in the sensitivity analysis, and the frequency of C allele in the MI group was marked higher than that in the control group in Asians. However, more studies for Asians should be performed in the future.
Those findings mentioned above should be regarded cautiously because many other aspects, such as heterogeneity of enrolled cases, limited statistical power, variable study designs and different interventions, were likely to affect the results. Furthermore, whether the AT1R A1166C gene polymorphism is just linked with other discrete loci involved in the occurrence of MI is not clear at the moment.
In conclusion, C allele is a risk factor for the MI susceptibility in overall populations, and AA genotype might be a protective factor against the MI risk in overall populations. Furthermore, AT1R C allele/AA genotype might be associated with MI susceptibility in Asians. However, more case-control association investigations into larger, stratified populations are required in the future.
Footnotes
Conflict of interest
None declared.
Funding
This research received no specific grant from any funding agency in the public, commercial, or not-for-profit sectors.
