Abstract
Keywords
Introduction
COVID-19, acute respiratory syndrome caused by the SARS-CoV-2 virus, 1 has resulted in a global health emergency. The high propensity of the virus to mutate and form vaccine-resistant strains calls for action at an early juncture. Delta and Omicron are two such strains with over 30 mutations that have come under the World Health Organization’s (WHO) scanner owing to their transmissibility and vaccine resistance. 2 This has been followed by high death rates across the globe. 3 Against this threat, over 400 therapeutic agents are being investigated, including corticosteroids, dexamethasone, and Remdesivir. 4 Corticosteroids and immunosuppressants were the first to be contemplated for severe illness based on their effectiveness in reversing abnormal systemic inflammatory responses. Notably, Remdesivir shortened the recovery time from 15 to 11 days in severe cases and received emergency use authorization from the US Food and Drug Administration. 5 Yet, clinical trials more often have inconclusive results, sometimes without any effect compared to standard therapy or placebo. 5 Emerging studies point out the efficacy of Remdesivir against Delta and Omicron mutations. 6
Literature review
Ongoing developments in digital technology, especially AI and ML, are benefiting medicine in a big way. ML is helping in three substantial areas: clinical trials for non-COVID medicines, repurposing non-COVID-19 medicines for COVID-19, and creating new therapeutic agents. 7 For instance, Lam et al. (2021) used decision trees to examine data from 2364 hospitalized patients in ten US hospitals and demonstrated that both Remdesivir and corticosteroids were beneficial to survival time. 4 Likewise, Gao et al. (2022) employed ML for the prediction of corticosteroid therapy outcomes in 666 patients from two hospitals in China with good accuracy through the application of models like Gradient Boosting Decision Tree (GBDT), Neural Network (NN), and Logistic Regression (LR). 8
Aghajani et al. (2022) compared eight therapies for hospitalized COVID-19 patients in Iran involving 861 patients. Their research illustrated the efficacy of regimens containing Remdesivir and Favipiravir in minimizing death risks, particularly in critically ill patients who needed intensive care. 9 Kuno et al. (2021) employed ML algorithms in predicting mortality rates of 1571 US hospitalized patients and determined six significant variables affecting the risk of mortality. 10 Blasiak et al. (2020) also investigated a combination of Remdesivir and ritonavir and lopinavir through AI algorithms and identified a triple therapy that had 6.5-fold better performance than Remdesivir monotherapy. 5 The research strives to create a precise analytical description of the impact of different treatments on the duration of hospitalization in Ahvaz, Iran, hospitalized patients. In contrast to previous research,11–19 it includes a broad range of treatment regimens and medications, with enhanced methodology. With hospitalization duration instead of mortality rates, the research demonstrates the pandemic’s complex nuances and tries to fill gaps in the evaluation of treatment effectiveness.
Review of studies related to the efficacy of drugs in the treatment of COVID-19.

The drugs remdesivir, dexamethasone, and corticosteroids were investigated in most of the reviewed studies. These drugs demonstrated positive outcomes in treating patients with COVID-19 and in reducing mortality. The literature review reveals that the patient populations were predominantly male, ranging from 53 to 75 percent, and within the middle-aged to elderly demographic, generally over 50 years old. This pattern indicates the high-risk populations that were predominant in these COVID-19 treatment studies. The results from the reviewed studies suggest that the efficacy of the drugs is dependent on various clinical factors. These factors include disease severity, drug dosage, and specific patient characteristics.
The present study aims to identify key clinical and pharmacological factors that influence the duration of hospitalization among patients with COVID-19, utilizing ML techniques. The analysis focuses on determining which patient characteristics and treatment variables are most strongly associated with prolonged or shortened hospital LOS. It is hypothesized that early administration of certain medications, along with baseline clinical indicators such as comorbidity burden and vital signs, significantly affects the LOS. Furthermore, this study evaluates the potential utility of ML models to support clinical decision-making and improve hospital resource management.
Materials and method
Dataset
Overview of patient parameters and clinical data.

The inclusion criteria consisted of patients aged 18 and above, hospitalized for over 24 h, and those receiving both Remdesivir and corticosteroids, while classified as moderate or acute based on the national early warning chart.
In this study, patients were divided into two groups, which we refer to as Experiment 1 and Experiment 2. This categorization is based on the National Early Warning (NEWS) guideline, which classifies patients into acute patients (Experiment 1) and moderate patients (Experiment 2) according to respiratory rate, consciousness indicators, temperature, and PO2.20,21 This study included 215 patients in Experiment 1, consisting of 52.5% women and 47.5% men, and 1578 patients in Experiment 2, comprising 51.8% women and 48.2% men.
All experiments and simulations were conducted separately for these two groups to evaluate treatment efficacy and outcomes systematically. This approach ensures a detailed analysis of drug effects on different patient conditions. The initial pool of 106 factors was identified through a systematic review of existing literature, from which a preliminary checklist of key variables was compiled.
To ensure clinical validity, these factors were further evaluated by three COVID-19 specialists. Treatment parameters were determined based on standard protocols widely used in Iran. Subsequently, feature selection algorithms were applied to extract the most significant predictive factors. Patients who received corticosteroids or immunomodulatory agents (e.g., tocilizumab) within 14 days before admission were excluded. Those with advanced comorbidities (end-stage renal disease on dialysis, decompensated liver failure, or active malignancy under treatment) were also excluded. Chronic conditions not meeting these severity criteria (e.g., stable CKD, controlled HIV without immunomodulators) were permitted.
Methodology
The proposed method, outlined in Figure 1, begins with correlation analysis to identify the relationship between patient features and hospitalization duration, aiming to select effective features while eliminating less relevant ones. Patients were then categorized into training and evaluation samples for classifier development and assessment. Effective drugs and features were input into these classifiers for training, followed by extracting hospitalization durations using classification methods. The proposed method for estimating the duration of hospitalization using ML approaches.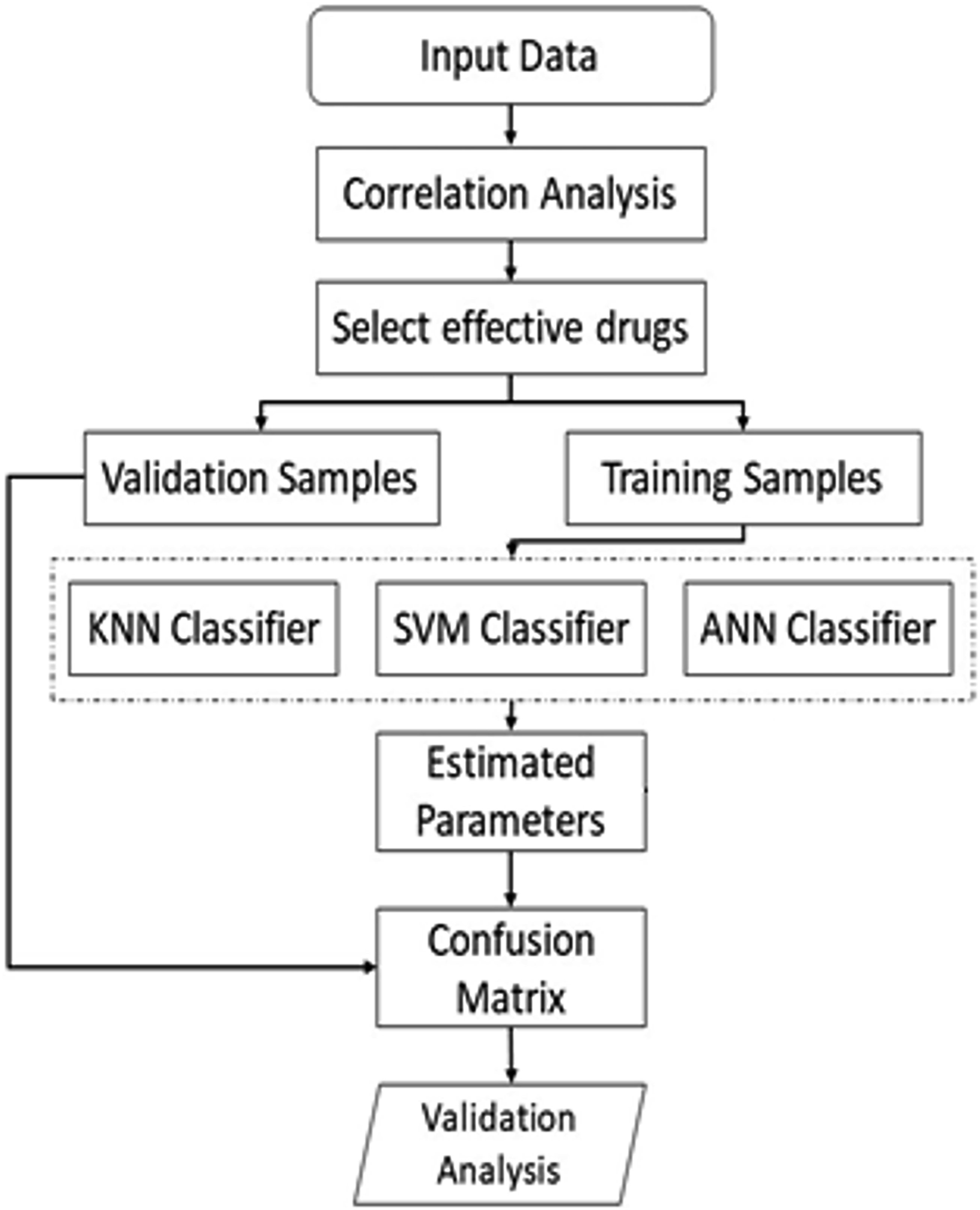
A confusion matrix was generated based on classifier results and evaluation samples. Finally, patient group features were analyzed using mean and standard deviation. Pearson’s correlation coefficient, a key statistical tool ranging from −1 to 1, was used to measure feature relationships, providing insight into both intensity and type (direct or inverse). Each step is detailed further in subsequent sections.
The Pearson’s correlation coefficient is determined using the equation as follows.22,23
In the above relationship, the variables
Training and testing samples selection
Training and test data were randomly selected from the entire patient dataset, dividing them into two categories: training (70%) and test (30%).25,26 The training data was further classified into three groups based on hospitalization duration: Class 1 (less than 5 days), Class 2 (6–10 days), and Class 3 (more than 11 days). This categorization facilitated a structured approach to model development and evaluation. The samples in Experiments 1 and 2 were randomly assigned. In the runs of Experiment 1, 150 samples were selected for training and 65 for the test using the k-fold cross-validation method. The distribution of the training samples across classes 1 to 3 was 40, 51, and 59, respectively. The distribution of the test samples for classes 1 to 3 was 16, 20, and 29, respectively. In Experiment 2, 1105 and 473 samples were randomly allocated for training and testing, respectively. The training set distribution for classes 1 to 3 was 467, 372, and 266. The test set distribution for these classes was 204, 155, and 114.
Classification
After identifying the most effective features influencing hospitalization duration, a prediction model was developed using three common ML methods: Support Vector Machine (SVM), k-Nearest Neighbors (kNN), and Artificial Neural Networks (ANN). In our investigation, we selected and fine-tuned these popular class of ML algorithms based on their track record of success in clinical prediction tasks, availability of medical interpretability, and quality of fit to structured tabular data. We selected SVM because previous COVID-19 studies (Lam et al.) have found that it performs well in predicting treatment outcomes. In addition, it performs well in a high-dimensional space, which is essential with a dataset with 106 features, and it tends to perform well while avoiding overfitting because of SVM’s margin maximization principle. Furthermore, SVM performs well with non-linear relationships using kernel tricks and demonstrates high flexibility in modeling complex clinical data. The method of SVM is a supervised learning algorithm that separates data points with a hyperplane that has a maximum margin.27–29
It uses nonlinear optimization techniques to determine the optimal hyperplane. Different kernel functions, such as exponential, polynomial, and sigmoid, can be employed to solve the optimization equation. Key parameters for optimizing SVM performance include kernel type, kernel size, and initial margin values. Bayesian inference was utilized in this study to fine-tune these parameters, minimizing error through Gaussian process modeling and Bayesian probability updates. 27
kNN offers explainability in its distance-based predictions so that clinicians can intuitively check “neighbor” cases and develop trust in the model’s decisions. It does not require a training phase, and therefore it is very agile to new data, especially precious in the midst of the fast-moving information during a pandemic. kNN has also been found to be helpful in drug efficacy studies and other clinical applications. The kNN algorithm is a supervised learning technique used for classification and regression.30–32 It identifies the nearest neighbors of a sample and predicts outcomes by averaging their values. The hyperparameter k for the k-NN classifier was set to 1 after several tests from k = 1 to 4. This choice aligns with common practice for benchmarking feature extraction methods, 33 providing a strong baseline that tests the inherent similarity structure of the learned feature space. Furthermore, a preliminary cross-validation check on a subset of the data confirmed that performance plateaued near k = 1, with larger values of k offering no significant improvement in accuracy. An exhaustive search algorithm was used, ensuring all distances were calculated to find the nearest neighbor.
ANN was also included because it can manage complex interactions, e.g., drug interactions and comorbidities, by deep hierarchies of features. Its scalability also makes it suitable for use when dataset size is large—here, 1578 patients. ANNs have also proved to be clinically important in other research, e.g., Gao et al., who successfully applied them for the prediction of corticosteroid outcomes. The ANN method employs a multilayer perceptron (MLP) architecture for data prediction and classification.34–36 The network consists of an input layer, output layer, and hidden layers, with neurons interconnected across layers. A fully connected neural network was implemented, featuring 10 layers, 106 input features, and three outputs. The ReLU activation function (ReLU(x) = max (0,x)) was used for inner layers, while the softmax function was applied to the output layer for classification purposes. Parameters were initialized using the Glorot method, and weights were updated using the quasi-Newton LBFGS algorithm.
36
The softmax function, defined by the equation where xi represents the ith input and k is the number of classes, was used in the output layer for classification. These methods collectively aimed to predict hospitalization duration effectively and provide insights into treatment efficacy. The output layer is the classification layer where the softmax function is used. The equation for the softmax function is as follows, where xi represents the ith input and k is the number of classes:
These algorithmic selections were driven by their complementary strengths: SVM for generalization and detection of non-linear relationships; kNN as a clear, easy-to-interpret baseline; and ANN to detect subtle interactions between the data. Decision trees, for instance, were excluded due to their limitations compared to the dimensionality and interpretability of the data. Additional information and parameterization of these algorithms are available in Supplementary Materials.24–26
Evaluation
Finally, the performance of the developed models was rigorously evaluated using standard classification metrics: sensitivity (recall), specificity, accuracy, and precision. To ensure reliable and unbiased estimation of model performance, we employed 10-fold cross-validation.37,38 This method partitions the dataset into 10 equal subsets (folds), iteratively using nine folds for training and one for testing, repeating until each fold serves as the testing set once. The final metrics were averaged across all folds to provide robust performance estimates.
Results
Correlation Coefficients for All Parameters with LOS in the first group of patents (*Statistically Significant = p < 0.05).

Correlation Coefficients for All Parameters with LOS in the second group of patents (*Statistically Significant = p < 0.05).
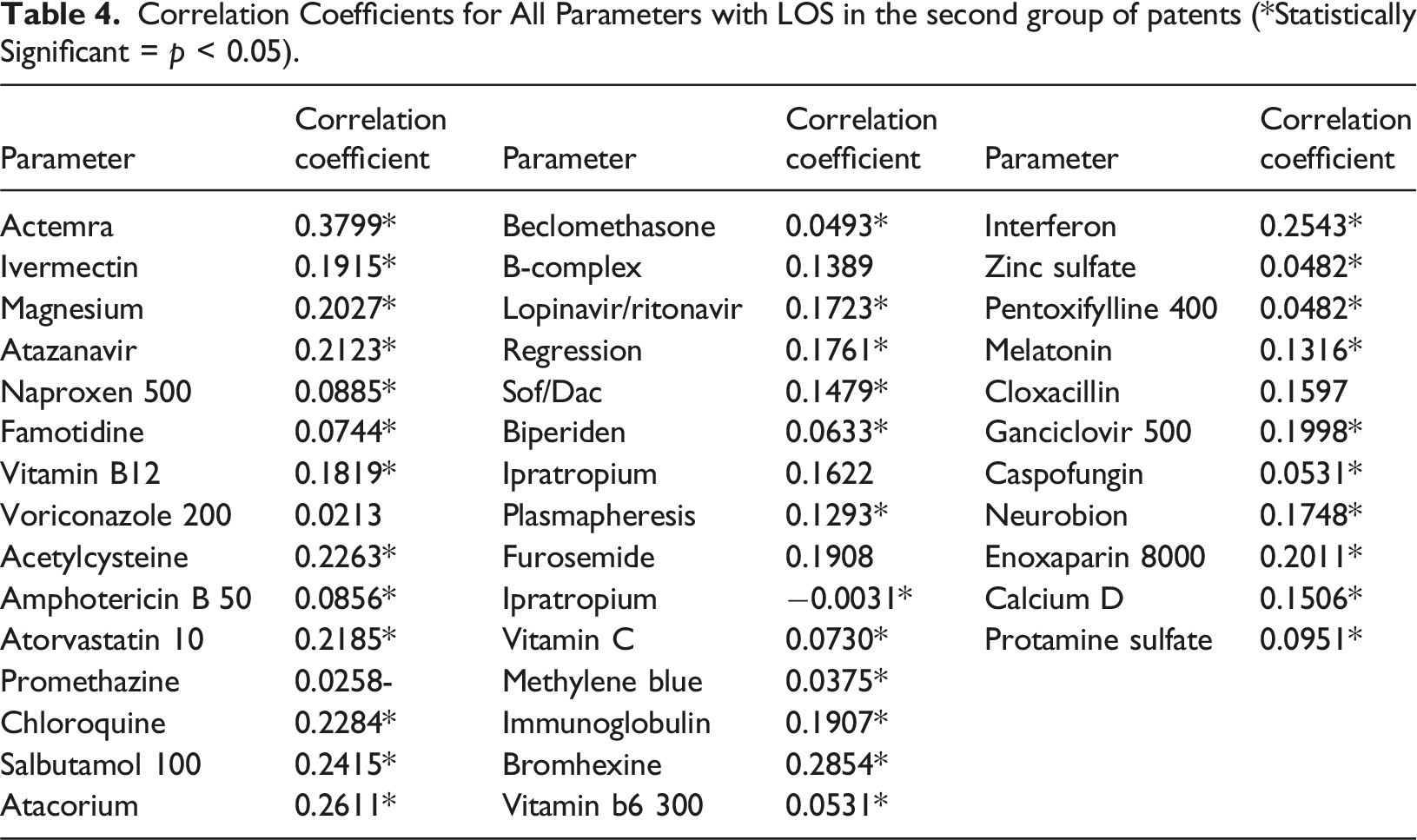
The observed correlation patterns are clinically insightful and align with established pharmacological knowledge. The strong positive correlation of Actemra (Tocilizumab) with increased hospitalization duration is highly plausible, as it is a specialized immunosuppressant primarily administered to severely ill COVID-19 patients experiencing cytokine storms. Its presence is thus a proxy for disease severity rather than a cause of longer stay. Similarly, the association of Amphotericin B, a potent antifungal, likely reflects complications from secondary fungal infections (e.g., mucormycosis), which necessitate extended treatment protocols. Conversely, the lack of significance for common medications like B-Complex or Furosemide is expected, as they are routinely administered for general supportive care and are not specific markers of disease progression. This analysis confirms that our feature selection method successfully identified biomarkers reflective of critical patient states and complex clinical pathways, thereby enhancing the clinical relevance of our predictive model.
Model performance using the testing dataset in experiment (1).

Model performance using the testing dataset in experiment (2).

K-fold cross-validation further confirmed the robustness of the classifiers. The lowest average k-fold error was observed for SVM in both experiments, as presented in Tables S4 and S5.
Table 5 presents the performance of the SVM, k-NN, and ANN models on the testing dataset for one sample run selected from multiple runs in Experiment 1. While all models demonstrated high and comparable overall accuracy, ranging from 82.7% to 92.2%, a detailed examination of per-class precision and recall reveals significant distinctions in their predictive characteristics. The ANN model emerged as the most robust overall, achieving perfect precision (100%) for Class 1 and the highest recall for Class 3 (96.30%), indicating its superior reliability in correctly identifying these classes without false positives and missing the fewest true instances, respectively. Conversely, the KNN model excelled at recalling Class 2 samples (94.74%) but suffered from lower precision on that class (81.82%), suggesting a tendency to over-predict it. The SVM model showed strong precision for Class 1 (92.86%) but exhibited the lowest recall for Class 2 (65.38%), highlighting a potential weakness in capturing all members of that specific class. This analysis underscores that model selection should be guided not merely by overall accuracy but by the specific precision-recall trade-offs most critical to the application domain.
Table 6 details the comparative performance of the SVM, KNN, and ANN models. The SVM and ANN models demonstrate exceptional and nearly equivalent overall accuracy, at 98.73% and 98.09% respectively, significantly outperforming the KNN model (87.31%). This performance gap is further elucidated by the per-class metrics. Both SVM and ANN achieve remarkably high precision and recall across all three classes, with the SVM exhibiting perfect recall for Class 2 (100%) and perfect precision for Class 3 (100%), and the ANN showing near-perfect recall for Class 2 (99.42%). In stark contrast, the KNN model shows considerably lower performance on these granular metrics, particularly in precision for Class 2 (78.98%) and recall for Class 1 (86.12%), indicating a higher propensity for misclassification. These results clearly establish SVM and ANN as the superior and highly reliable models for this specific task, while the KNN algorithm is markedly less effective in this comparative analysis.
Discussion
This study employed ML algorithms to predict the length of hospital stay for COVID-19 patients using clinical and treatment characteristics. Among the evaluated models (SVM, kNN, and ANN), the SVM algorithm demonstrated the highest predictive accuracy, highlighting its potential utility in clinical decision-support systems for optimizing patient management during the pandemic. 39
The current study and the work by Kuno et al. 40 both utilize ML to address key challenges in managing COVID-19 patients, though with different objectives and methodological approaches. While our research focused on predicting hospital length of stay (LOS) among Iranian patients based on treatment protocols, Kuno et al. developed a model to estimate in-hospital mortality among patients treated with steroids and remdesivir in the United States. Despite these differing aims, several important parallels and insights emerge, illustrating the adaptability and potential of ML in this field. Both studies employed advanced ML techniques capable of handling high-dimensional clinical data. In our work, we achieved prediction accuracies of up to 98% using tuned SVM, kNN, and ANN. Kuno et al., on the other hand, used LightGBM with LASSO for feature selection and SHAP for interpretability, achieving AUC scores between 0.88 and 0.91. A critical aspect of both studies was feature selection: our analysis identified Actemra and Bromhexine as strongly associated with LOS, while Kuno et al. pinpointed age, ICU admission, and intubation as key predictors of mortality. These differences highlight that the most relevant variables for COVID-19 outcomes are highly context-specific. The success of ML in these studies highlights its potential for personalized medicine. For instance, our LOS model could be adapted to optimize discharge planning, while Kuno et al.’s tool could guide intensive care transitions. Together, these studies illustrate the transformative role of AI in pandemic response.
Blasiak et al. (2020) conducted a study in Singapore investigating the use of remdesivir in combination with ritonavir and lopinavir as a potentially effective therapeutic regimen against severe acute respiratory syndrome coronavirus (SARS-CoV-2) through the application of artificial intelligence. 5 The study aimed to identify a clinically optimal drug combination using AI-driven algorithms. For this purpose, the researchers employed the IDentif.AI platform, which is designed to rapidly generate optimized combination therapies for infectious diseases, including COVID-19. The findings indicated that the most effective therapeutic strategy for COVID-19 patients was a triple-drug regimen consisting of remdesivir, ritonavir, and lopinavir, which demonstrated a 6.5-fold improvement in efficacy compared to remdesivir monotherapy. Conversely, the results suggested that hydroxychloroquine and azithromycin were relatively ineffective in treating the disease. 5 It is noteworthy, however, that the technical details and underlying algorithms of the IDentif.AI platform were not disclosed, limiting the possibility of direct comparison with other computational approaches. Moreover, Aghajani et al. (2022) conducted a study in Iran comparing the effectiveness of eight different therapeutic regimens for hospitalized COVID-19 patients. The study was based on data from 861 patients. The treatment protocols investigated encompassed eight major therapeutic strategies used for COVID-19 management. Using statistical analyses, the study demonstrated that protocols including remdesivir or favipiravir were more effective than hydroxychloroquine in reducing the risk of mortality among hospitalized patients, particularly in critical cases, defined as patients admitted to the intensive care unit or requiring mechanical ventilation. It should be noted that the study by Aghajani et al. did not utilize ML techniques; rather, it relied exclusively on traditional statistical methods to evaluate treatment protocols. In terms of sample size, this study included a larger patient population compared to previous investigations by the same research group. 9
The application of ML in this context not only enhances predictive accuracy but also supports early identification of patients at higher risk of prolonged hospitalization. This has critical implications for hospital resource management, especially in overburdened health systems. Moreover, our model highlights modifiable treatment variables that can be integrated into real-time clinical decision-making, offering a pathway toward more personalized and data-driven care.
Our findings have several important implications for clinical practice and health policy. Notably, the analysis identified certain medications, such as corticosteroids and anticoagulants, as being associated with shorter hospitalization durations, consistent with existing global treatment guidelines for moderate to severe COVID-19 cases. 12 Conversely, treatments such as hydroxychloroquine, which were more common during the early phases of the pandemic, showed no consistent benefit and were later de-emphasized in international protocols. These findings not only validate the effectiveness of key components of current treatment strategies but also provide real-world evidence from a non-Western healthcare setting. As treatment protocols evolve, ML-based analyses like ours can serve as ongoing tools to evaluate their effectiveness across diverse populations.
The evidence strongly supports the inclusion of Actemra (tocilizumab) in Iran’s national COVID-19 treatment guideline for patients with moderate-to-severe disease, particularly those without substantial comorbidities. 41 This is supported by two key considerations that explain to its clinical and pragmatic benefits. First, Actemra was associated with a significant reduction in hospital length of stay (LOS). This reduction not only means improved patient outcomes but also translates into valuable cost savings for healthcare systems. Among all treatments that were compared, Actemra boasted the largest value for correlation with improved outcomes (r = 0.38, p < 0.05), demonstrating its clinical utility and effectiveness in the treatment of severe COVID-19 cases. Second, its single-dose regimen is advantageous in resource-limited settings like Iran because it is less difficult to deliver and less demanding in logistics vis-a-vis multi-dosing regimens.
A couple of significant steps should be initiated to facilitate successful integration of Actemra into mainstream clinical practice. These are formal training sessions for clinicians regarding proper indications, dosing regimens, and monitoring steps so that the drug is appropriately and effectively administered. Moreover, distinct budget allocations by Iran’s Health Insurance Organization are needed to guarantee even access in the whole healthcare system. Moreover, integrating outcome monitoring systems into hospital electronic health records (EHRs) will be essential to track efficacy, adverse events, and compliance and hence guarantee an evidence-based, continuous assessment of its impact.
This approach aligns with global guidance in COVID-19 management, emphasizing evidence-based interventions tailored to available local capacities. Introduction of Actemra into national guidelines with adequate supplementary measures could well revolutionize the therapeutic outcomes for severe cases of COVID-19 in Iran.
While our ML models demonstrated high predictive accuracy in identifying factors associated with COVID-19 hospitalization duration, their clinical utility hinges not only on performance but also on interpretability. Transparent models are essential to foster clinician trust and enable actionable decision-making. 40 For instance, while Actemra and Bromhexine emerged as strongly correlated with reduced LOS in our analysis, clinicians require clear explanations of how and why these medications influence outcomes—especially for patients with specific comorbidities. Techniques like SHAP (Shapley Additive Explanations) and LIME (Local Interpretable Model-Agnostic Explanations) could bridge this gap by quantifying feature contributions 42 thereby transforming our models into clinically adoptable tools. Our ML-based algorithms have been rigorously designed to account for and neutralize the effects of potential confounding variables. This was achieved through a combination of robust feature selection, cross-validation, and evaluation on an independent test dataset. The performance metrics across these datasets demonstrate the model’s generalizability and stability, even in the presence of confounding factors. Additionally, we employed techniques such as to further mitigate bias. The comprehensive evaluation confirms that the model maintains high predictive accuracy without being systematically influenced by confounders under standard experimental conditions.
Since our dataset spans multiple pandemic waves, it likely includes a heterogeneous mix of clinical practices, treatment protocols, and circulating viral strains, all of which could influence the observed associations between treatments and outcomes. Although our current analysis offers valuable insights, it does not explicitly stratify or adjust for these evolving factors due to limitations in the available data. Nevertheless, it is noteworthy that ML models employed during this period demonstrated a substantial ability to predict current patient conditions despite these variations. Their high accuracy indicates their practical applicability in real-time clinical settings. Furthermore, with ongoing updates and expansion of the dataset, these models could serve as efficient and adaptable tools for identifying effective treatments even as new variants emerge. Retraining these models with more recent data is expected to enhance their robustness and generalizability, thereby increasing their utility for future clinical decision-making and rapid response to the evolving landscape of the pandemic.
Future research should consider incorporating temporal variables, such as treatment protocol changes and dominant circulating variants, to better contextualize their effects. Additionally, employing ML models that are periodically retrained with updated datasets could enhance their effectiveness and utility in managing ongoing and future public health challenges.
Generalizability to other provinces or countries may be limited. It is also possible that differences in treatment protocols, hospital resources, or clinical decision-making strategies across regions could influence the outcomes. Future multicenter studies involving diverse geographic locations and healthcare settings are recommended to improve the external validity of the findings.
Conclusion
This research has used ML to predict the outcome of COVID-19 drugs on the LOS by surmounting the complex challenges posed by the pandemic. This research employs SVM, kNN, and ANN to assess the treatment strategy, taking advantage of a large patient dataset for robust prediction. This study shows that all the ML algorithms studied herein were very effective in assessing drug efficacy for personalized COVID-19 treatment and thus open ways for developing better strategies. Our analysis suggested that early introduction of some immunosuppressive medications such as tocilizumab (Actemra) and bromhexine were associated with a reduction in the LOS in patients with COVID-19. LOS reductions were more obvious in patients with limited major comorbidities. This may mean that early introduction of treatment delayed or prevented the progression of the illness in otherwise healthy patients suffering from COVID-19. Such findings demonstrate the utility of decision support systems for personalized treatment. While mentioning certain limitations of this approach, given that the database is linked to two COVID-19 referral centers, results are highly applicable to healthcare professionals and policy makers, helping toward resource optimization and improving outcomes for patients. The present study illustrates that AI may be a useful component in clinical decision-making and could fertilize this event to future events in health crises. For further investigation, hybrid models for LOS and mortality predictions are considered. Such a hybrid model should include diverse populations and be validated with real-world sets of patients. Such efforts will be able to bridge gaps between treatment efficacy and patient outcomes, hence refining clinical decision-making related to COVID-19 and beyond. Independent testing and practical application are crucial to realizing these models’ full potential in clinical practice.
Supplemental Material
Suppplemental Material - Impact of treatment protocols on hospital length of stay for COVID-19 patients: A machine learning analysis of cases in Khuzestan province, Iran
Suppplemental Material for Impact of treatment protocols on hospital length of stay for COVID-19 patients: A machine learning analysis of cases in Khuzestan province, Iran by Soheila Shamouni Sayaei, Amir Jamshidnezhad, Javad Zarei, Maryam Haddadzadeh Shoushtari, Mohammad Reza Akhound in Health Informatics Journal
Footnotes
Acknowledgement
The authors would like to acknowledge Ahvaz Jundishapur University of Medical Sciences and the Vice-Chancellor for Research Affairs at Ahvaz Jundishapur University of Medical Sciences for their financial and administrative support in undertaking this project. Research Grant No: U-010.15. Research Ethics Code: IR. AJUMS.REC.1401.010. The authors declare that AI tools were used for manuscript proofreading in some parts of the manuscript. The authors would also like to express their gratitude to the COVID-19 Registry System in Khuzestan province for providing access to the data.
Ethical considerations
The present prospective study has been approved by the local ethical committee (Quality Management Unit) in Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran. The need to obtain informed consent was waived by the local ethical committee. The reference number is not applicable and not available.
Author contributions
A.J. conceived of the idea. A.J. and J.Z. designed the study. S.Sh.S. developed the models. S.Sh.S. analyzed the data and achieved the outcomes. MR.A. reviews the statistical experiments. M.H.S. reviewed the clinical factors. All authors discussed the outcomes. S.Sh.S. wrote the first draft with contributions. S.Sh.S reviewed and commented on the manuscript, as well as all responsible for the content of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the Key Supporting Scientific Research Projects of Ahvaz Jundishapur University of Medical Sciences (Grant Number U-010.15).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data that support the findings of this study are available from Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran. However, restrictions apply to the availability of these data, which were used under license for the current study and are not publicly available. Data are available from the author, Mrs. Soheila Shamouni Sayaei, upon reasonable request via the following email, and with permission from Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran:
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
