Abstract
Introduction
Pituitary adenomas are one of the most common intracranial tumors, occurring in the intracranial human endocrine center, the pituitary gland, with a high prevalence reported up to 20%. 1 In the general population, approximately 1 in 1100 individuals suffers from clinically evident pituitary adenomas. 2 Moreover, they mostly occur in young and middle-aged adults, and seriously damage patient’s growth, development, labor capacity, fertility, etc., resulting physical, mental, economic and social burden.
Clinically, the comprehensive assessment of patients with pituitary adenoma faces the challenges of complex clinical manifestations and high pathological heterogeneity, limiting the health management, diagnosis and treatment plan recommendations, and recurrence risk prediction of such patients. Clinical notes contain rich diagnostic and treatment information that can be used to characterize pituitary adenomas. 1 However, since these notes are recorded in natural language, useful sematic information and features needs to be identified and annotated from the free-text narratives as machine-readable and computable structured knowledge of characterization.3–5
Several excellent corpora have been present based on specialized diseases with different application objectives. To evaluate future machine learning (ML)-based NLP methods, Chaturvedi et al. 6 manually developed a pain-related corpus based on mental health electronic health records (EHRs). Given the urgent need for scientific and knowledge on rare diseases, Martínez-deMiguel et al. 7 developed a gold standard corpus through a dictionary-based approach for extracting medical information. Additionally, facing the growing demand for non-English clinical corpora, Frei et al. 8 created a small annotated text dataset to train a medical named entity recognition (NER) model on German text, Oliveira et al. 9 developed a semantically annotated corpus for Portuguese clinical NLP tasks, and Cai et al. 10 annotated the clinical diagnostic criteria for premature ovarian decline based on Chinese medical records. Nevertheless, existing annotated corpora are still mainly in English.
Corpus annotation is a labor-intensive task, especially for purely manual annotation. ML techniques facilitate automated processing of texts, and a series of excellent models and methods have emerged. Among them, active learning11,12 has been shown to be effective in reducing the cost of annotation compared to the typical passive framework based on random selection of samples. Recently, pre-trained language models have greatly improved the performance of NLP, but they still need to be combined with domain knowledge to support the underlying understanding and reasoning of specific tasks. 13 To demonstrate the use of case reports for data construction and pandemic surveillance, Raza et al. 14 created a gold standard annotation dataset using limited COVID-19 case reports to train BERT during the few-shot learning process. In a study on the annotation of a large multi-cancer genomic dataset, Kehl et al. 15 trained deep NLP models to extract clinical outcomes based on limited manual medical record annotations derived from separate retrospective cohort studies.
To improve the efficiency and reduce the labor and time cost of annotation, this study presents a semi-automatically clinical text corpus construction framework with a combination of ML and NLP approaches, and characterizes pituitary adenomas by annotating sematic information and constructing a clinical texts corpus. Furthermore, experiments on emerging advanced large language models (LLMs)16–18 demonstrate the practicality of pituitary adenoma characterization and its value for medical artificial intelligence (AI). To the best of our knowledge, this is the first clinical text corpus study dedicated to characterizing pituitary adenomas.
The main contributions of this work are as follows: (1) a semi-automatic framework that combines ML and NLP methods was designed to build a corpus; (2) a novel clinical text annotation guideline was developed, according to which the text corpus of pituitary adenomas (TCPA) was constructed, and common clinical information related to pituitary adenomas were identified; and (3) the fine-tuning and prompting experiments were conducted, and the practical applicability of this study in LLMs era was verified.
Material and methods
Dataset
Statistics of data distribution.
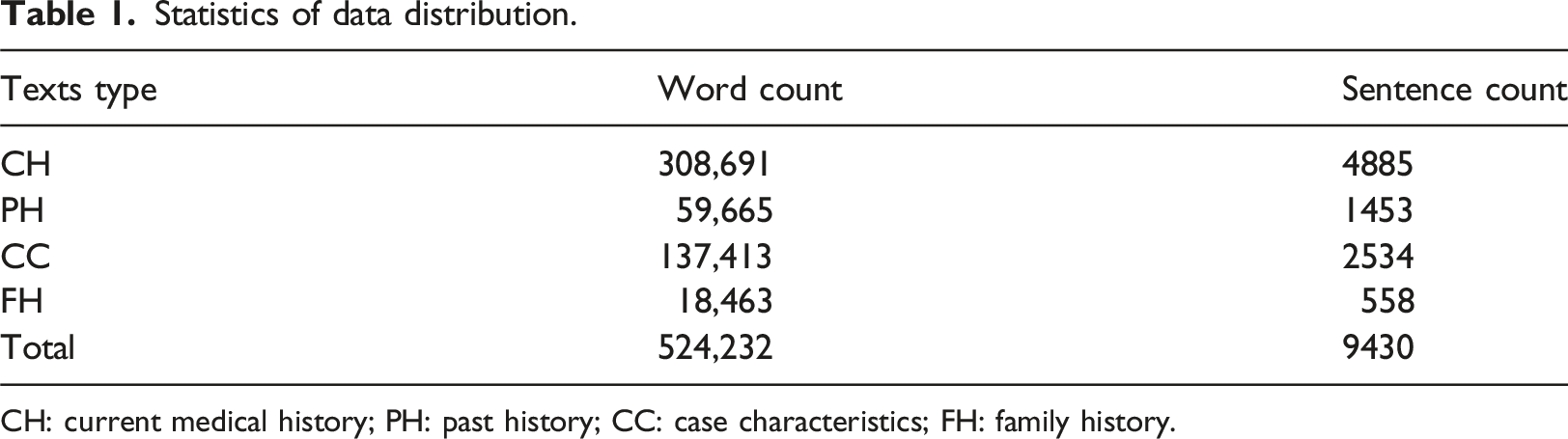
CH: current medical history; PH: past history; CC: case characteristics; FH: family history.
Methods
To characterize pituitary adenomas in clinical notes, TCPA is constructed following a semi-automatic framework based on machine-assisted manual annotation. Then, fine-tuning and prompting experiments are conducted to demonstrate the practical applicability of this study in the LLMs era. The workflow of our approach is shown in Figure 1. Workflow overview of our approach. CCNLP: Chinese clinical natural language processing; CNPL: clinical natural language processing; TCPA: text corpus of pituitary adenomas; LLMs: large language models.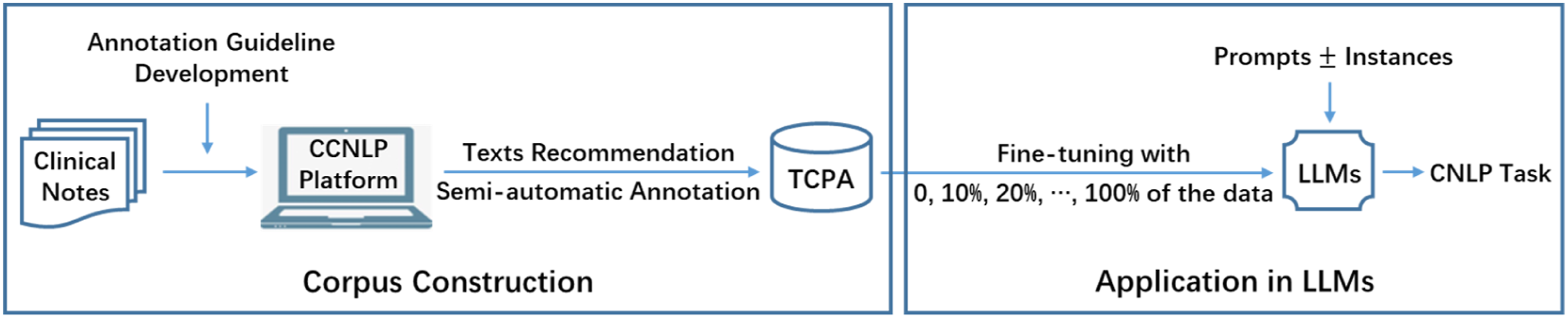
Specifically, the annotated corpus construction framework of TCPA is illustrated in Figure 2, consisting the following steps: (1) guideline development, (2) unannotated texts recommendation, and (3) corpus construction. An initial guideline is drafted based on the discussion of domain experts, including the clinicians and information scientists, as well as referring to the relevant annotation specifications. In view of the special language expression of clinical notes, several typical medical records are selected, and the annotation specifications are discussed and revised by analyzing the language and structural characteristics of notes. The annotators are trained on the annotation guidelines to manually pre-annotate part of the texts until the consistency satisfies the stability requirements. To improve the efficiency of annotation, the active learning method is used to query the most valuable texts from the clinical texts to be annotated, and the top N texts are recommended for machine-assisted annotation. Then, the ML algorithm is used to automatically annotate the recommended texts. Thus, annotators can easily make additions, deletions and modifications based on the automatic annotation results. Different annotators may have different understandings of the annotation contents, so the multi-round annotation mode is adopted to ensure the corpus quality. Besides, a senior clinician is added as reviewer to unify the annotation results being aggregated into the final corpus. Diagram illustrating the corpus construction framework. CCNLP: Chinese clinical natural language processing.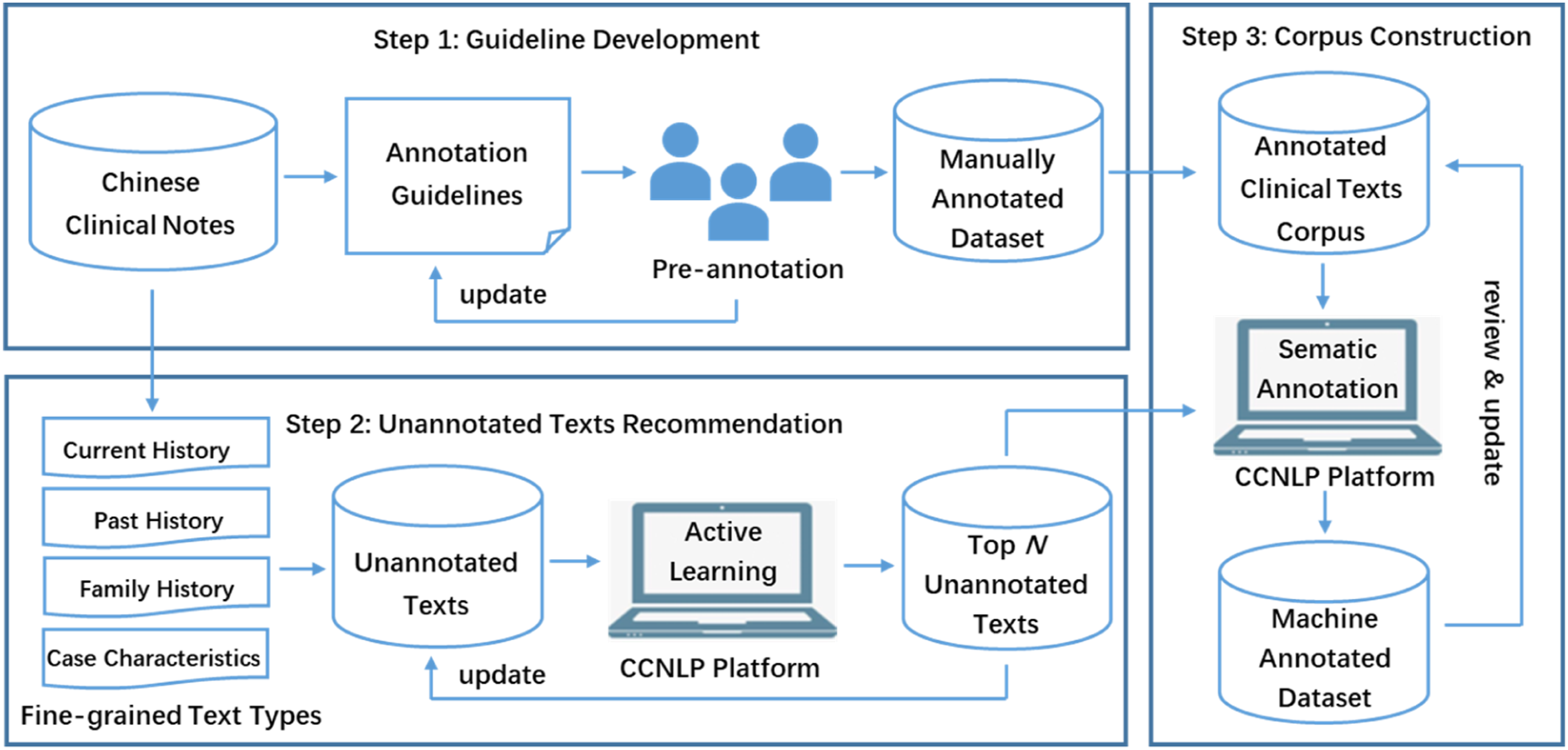
Annotation guideline
Concepts and their assertions of interest in this study.

The differences of the annotation guidelines between 2010 i2b2/VA and our work include: (1) the concept categories of medical problem and treatment defined in 2010 i2b2/VA are fine-grained into disease, symptom, drug, and surgery, (2) since the examination and test information have been structured and stored in the dedicated structured databases, concept test is not annotated rationally, (3) three concept categories are added to improve the refinement of annotation, that is, body, disease course, and family history, and (4) the assertion annotation guideline of 2010 i2b2/VA 14 is simplified from a practical perspective as following: positive concepts in clinical texts need to be identified; negative disease, symptom and drug providing auxiliary reference for clinical diagnosis and treatment19,20 need to be analyzed; and especially, alleviated symptom is valuable to complete clues for disease progression.
Annotation platform
To support the annotation and analysis of clinical information, we developed a web-based platform named Chinese clinical natural language processing (CCNLP).
21
CCNLP provides online and incremental learning, and the active learning algorithm used in Step 2 (detailed in Appendix 2) and automatic text annotation algorithm used in Step 3 (detailed in Appendix 3) are both integrated to facilitate the visualization of man-machine interaction. Figure 3 gives a screenshot of the CCNLP interface. Screenshot of the configuration interface for the automatic machine-assisted annotation in CCNLP. The English explanations corresponding to the functions displayed on the platform interface are provided in orange font.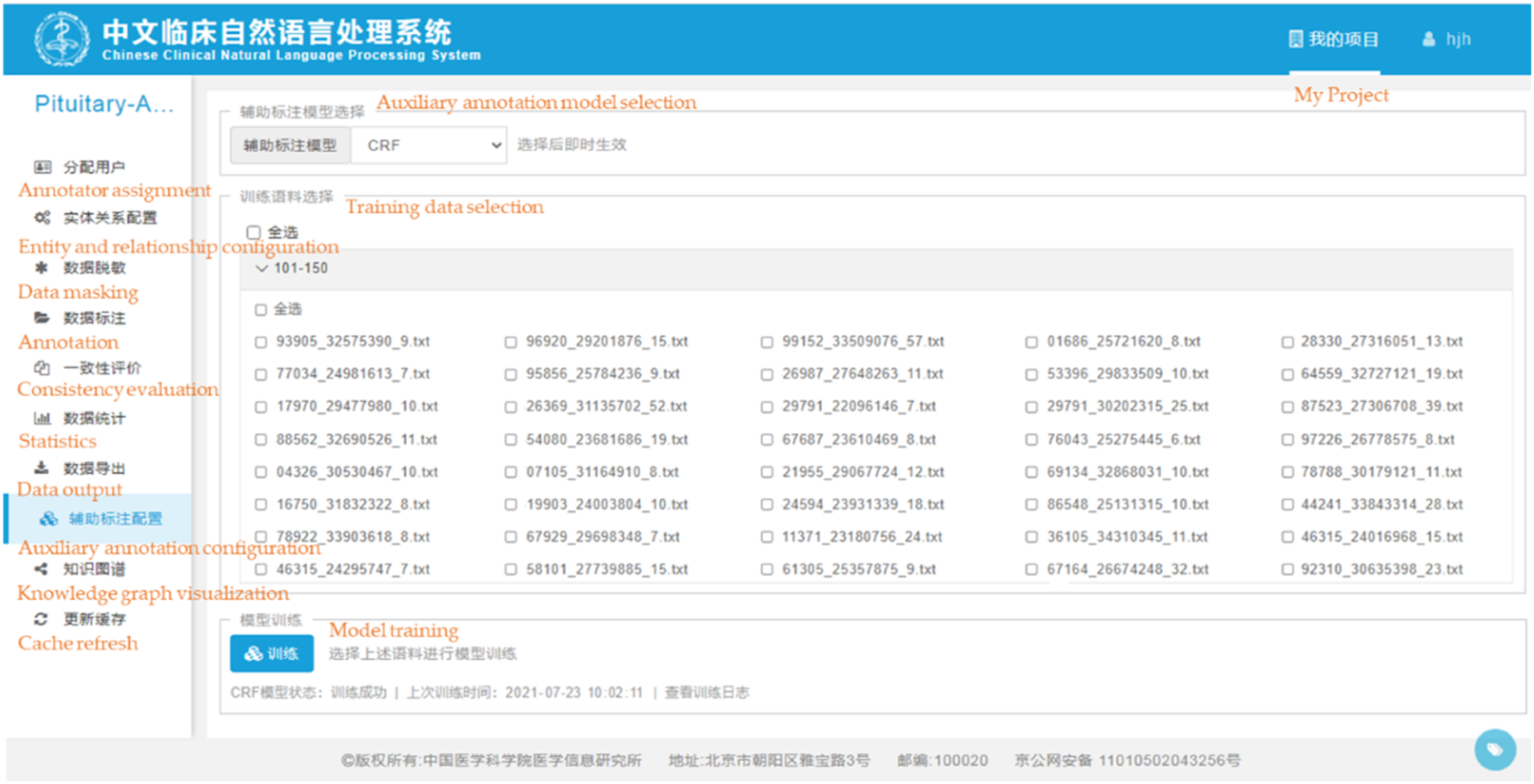
In CCNLP, the personalization of medical concepts and their assertions is supported. It also provides convenient annotation operations and user-friendly keyboard shortcuts for quick annotation. Figure 4 shows an example of the annotation of Chinese clinical notes using CCNLP. An annotation example of Chinese clinical notes in CCNLP. Different medical concepts are distinguished by different colors. CCNLP: Chinese clinical natural language processing.
Corpus application
In the field of clinical NLP, the fine-tuning of LLMs for specific tasks such as clinical NER22–24 is a pivotal area of research. A novel approach in this domain involves the integration of human-annotated data with Low-Rank Adaptation (LoRA) 25 to enhance the model’s performance in identifying and classifying named entities within text. Our approach takes advantage of TCPA, which serves as a gold standard, providing the model with precise examples of named entities to learn from. Subsequently, the LoRA model, which introduces low-rank matrices into the pre-trained LLM’s architecture, is employed to fine-tune the model. Unlike traditional fine-tuning that adjusts all parameters and risks overfitting, LoRA strategically modifies only a small subset of the model’s weights, preserving the general knowledge while adapting to the nuances of the NER task. This approach not only enhances the model’s ability to generalize across different contexts but also efficiently leverages the human-annotated data, resulting in a more accurate and robust system for NER.
Since the study data is in Chinese, here we use the Chinese-friendly ChatGLM3-6B 26 as the base model of LLM. The experimental environment and parameter settings are listed in Appendix 4.
In addition, prompting is an important method for embedding LLMs with clinical information extraction pipeline. But the prompt designs are generally hard in clinical domain in terms of the LLMs lack of medical knowledge. In our dataset annotation work, we not only conducted the high quality annotated dataset, but also provide the prompting instances recommended by clinical experts (see Appendix 5).
As shown in Figure 1, we randomly selected samples from the corpus, analyzed the performance of the fine-tuned model with different proportions of corpus participation, and compared the differences between fine-tuned LLMs prompting without instances and with instances.
Statistical analysis
All statistical analyses in this study were performed in Python (version 3.10.13) with the following libraries: NumPy (version 1.23.4) for numerical calculations, pandas (version 2.2.2) for data processing, and SciPy (version 1.14.0) for scientific computing. For the deep learning framework, PyTorch (version 2.1.0) was chosen with the Hugging Face’s transformers library (version 4.43.4). In addition, the following libraries were also used: datasets (version 2.20.0) for processing and loading datasets, DeepSpeed (version 0.14.0) for accelerating model training, and flash-attn (version 2.6.3) for optimizing the computational efficiency of the attention mechanism. The experimental environment and parameter settings are available in Appendix 4.
Results
Annotation consistency
The quality of annotated corpus is related to the efficiency and depth of clinical information application. To ensure the annotation quality, the principle of multi-person annotation is strictly followed, and the annotation consistency is evaluated by calculating the inter-annotator agreement (IAA) value using the measure of F1-sorce.27,28
Denote 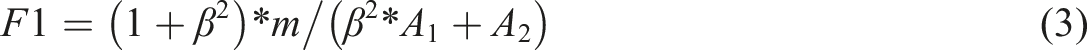
IAA value for the corpus construction.
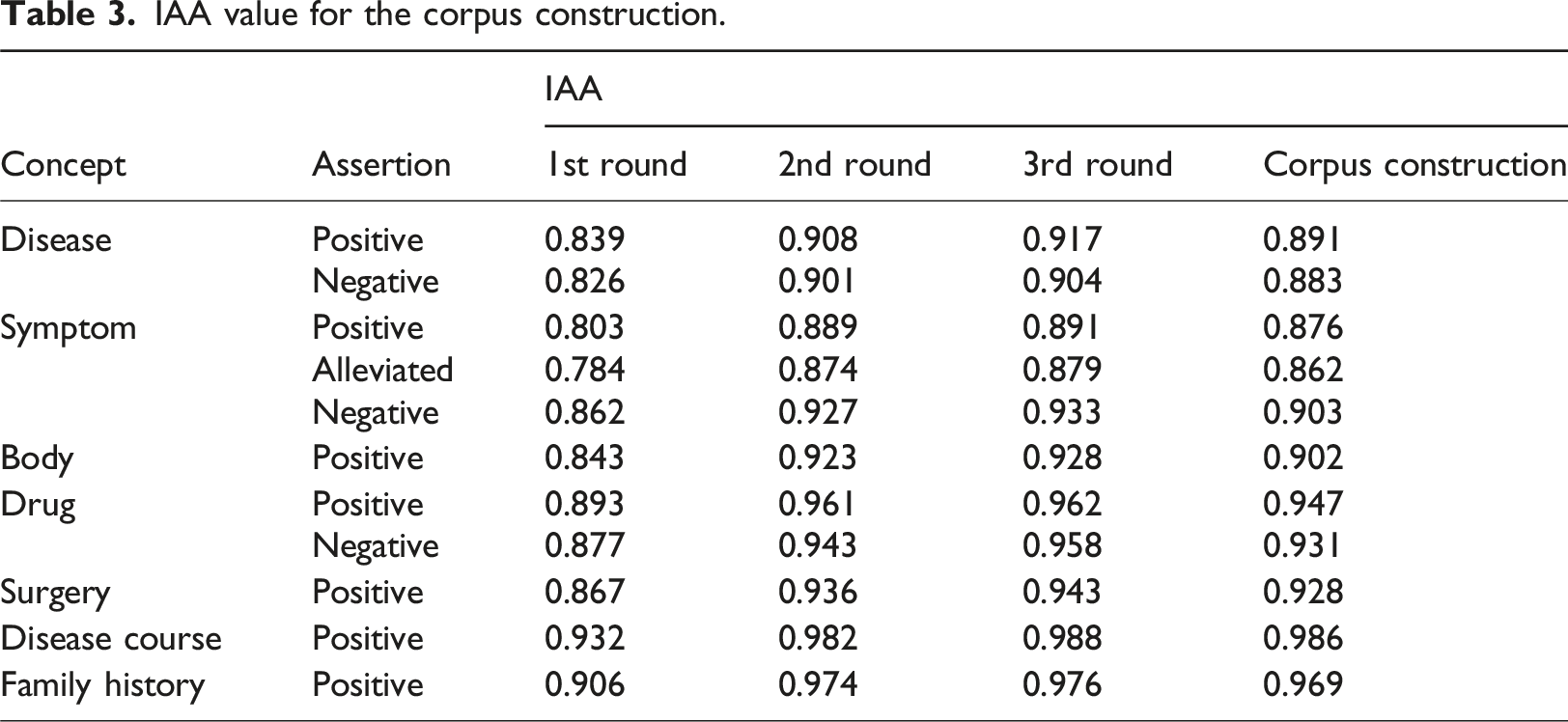
Corpus statistics
Concept distribution in our annotated clinical texts.
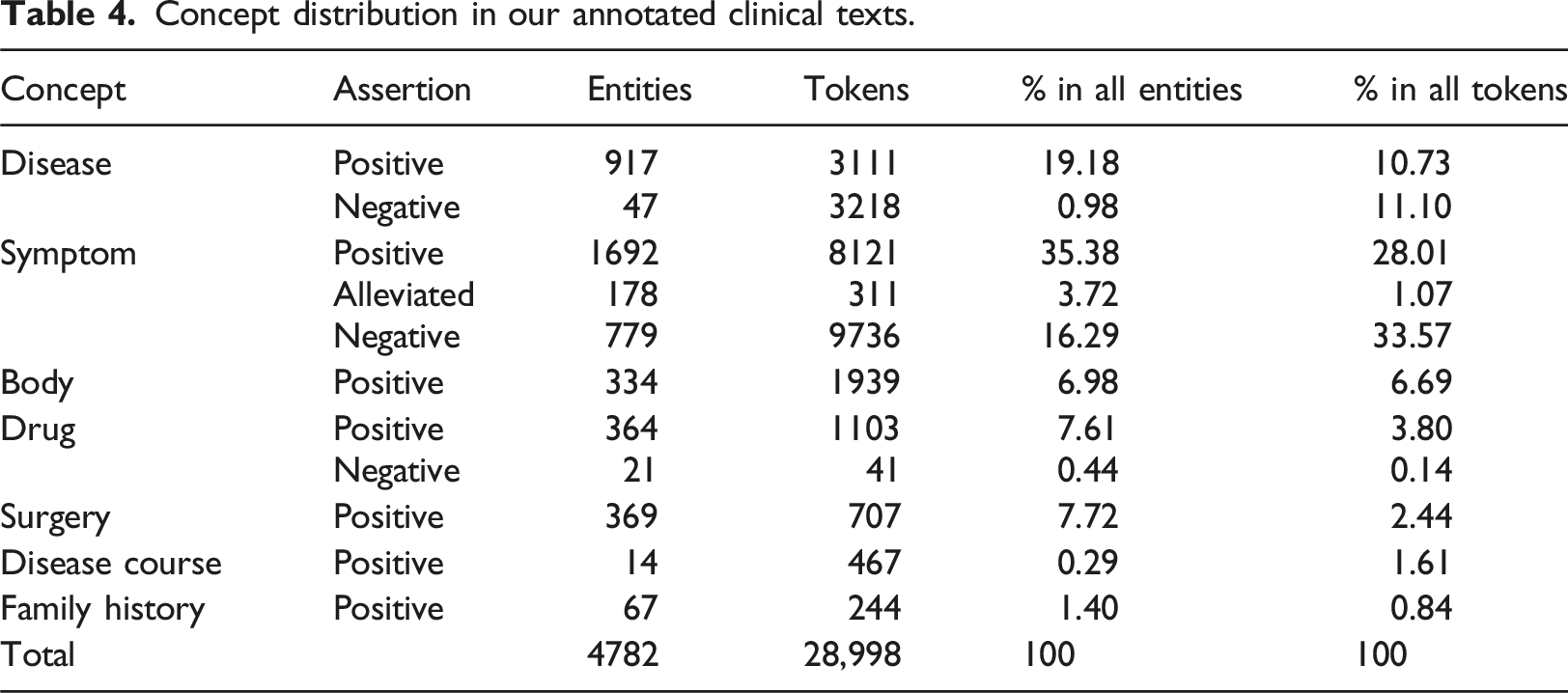
Top-10 tokens for each fine-grained medical concept.
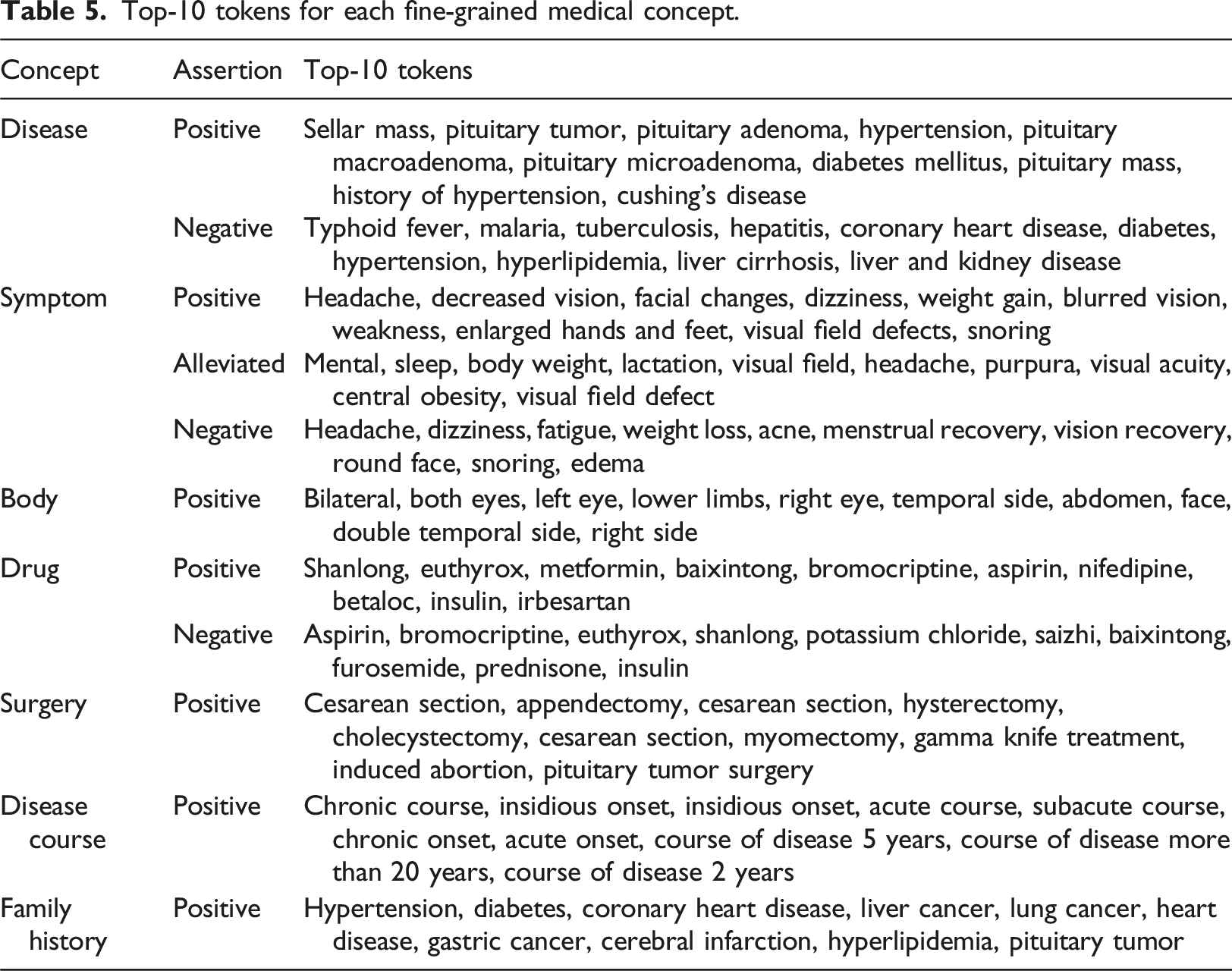
LLMs experiments
In clinical information extraction, the human expert annotated corpus are crucial in training a robust deep learning model for related information extraction. In LLMs experiments, we conduct the evaluation on how the annotated corpus and prompts benefit the clinical NLP task in the LLMs era. The experimental results are shown in Figure 5. Performance changes of LLMs fine-tuned with different percentages of the corpus. LLMs: larger language models.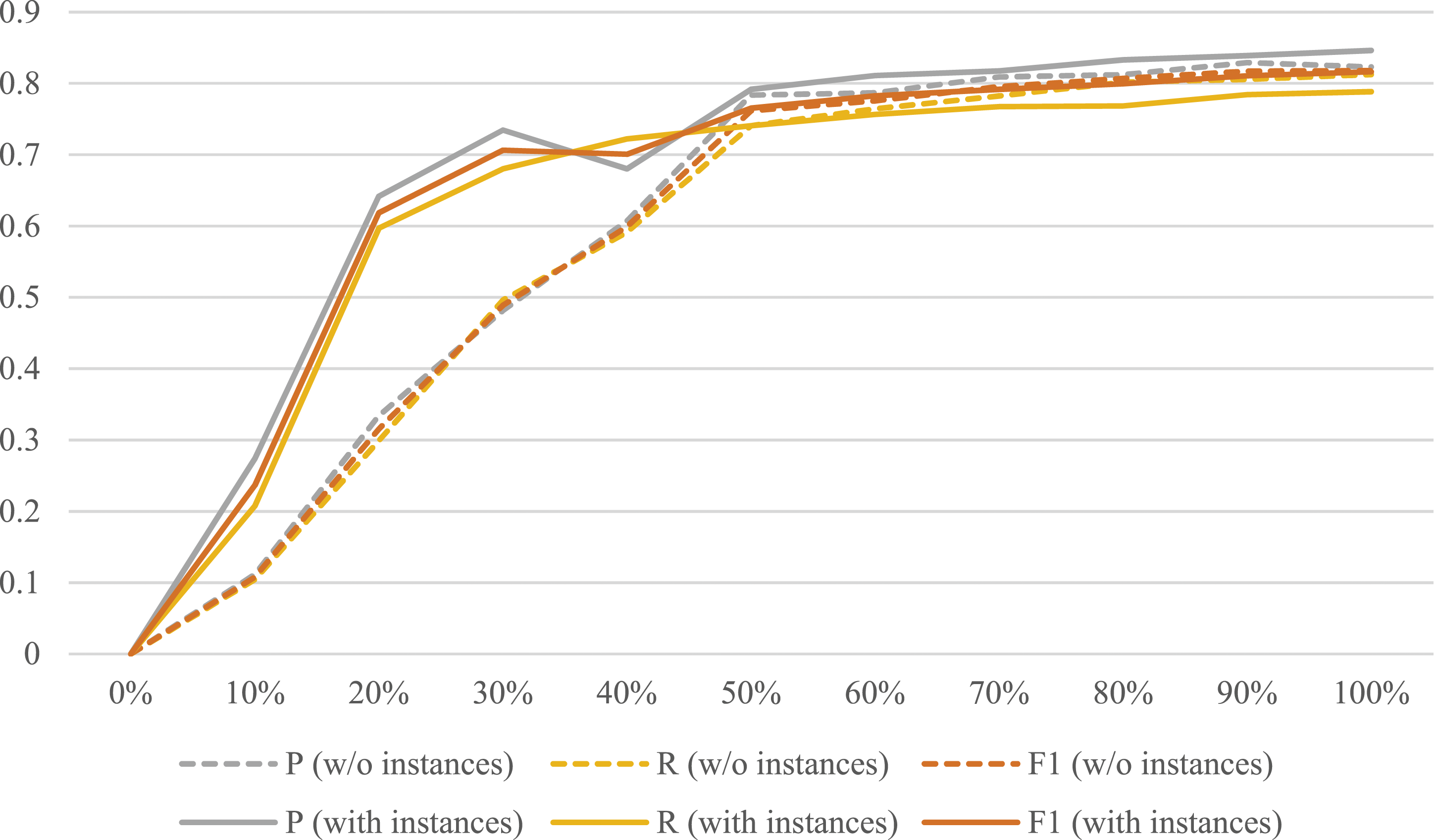
The experimental results show that when TCPA is not used for fine-tuning, that is, when 0% of the corpus is used, the F1 score is 0 regardless of whether it is prompting with instances or without instances. This indicates that it is difficult for a general LLM to understand the semantics of clinical concept entities without fine-tuning with domain data.
Considering semantics of entities are hard for LLMs to understand in this task, TCPA was used and the F1 score increased to 0.8179 (orange dotted line in Figure 5). Thus, it is verified that TCPA can significantly improve the performance of LLMs in understanding and processing pituitary adenoma-related texts, which is of great significance for clinical natural language scenarios with highly specialized language structures and terminology.
Further prompting engineering experiment show that instances provided by domain experts are beneficial to performance improvement (solid line in Figure 5). The LLM fine-tuned on 20% of the annotated data and prompted with instances provided by experts can achieve the same effect as that fine-tuned on 45% of the dataset and prompted without instances. Thus, the same performance can be achieved with less annotated data, thereby reducing labor costs.
Discussion
Clinical findings
The pituitary gland is the most important endocrine gland in the body, locating at the base of the brain, and pituitary adenomas may cause significant morbidity or mortality.1,2 Early diagnosis, timely treatment and effective prognosis can significantly improve the life and health of patients with pituitary adenomas. However, the characteristics of pituitary adenomas are only partly understood. 31 The Pituitary Society recently recommended the integration of various types of information on pituitary-related tumors, including clinical information, bioinformatics, etc. at the international Pituitary Neoplasm Nomenclature (PANOMEN) workshop. 32
This study confirmed our hypothesis that rich information about pituitary adenomas is available in clinical notes and can be standardized and automated for annotation and identification based on NLP and ML. The most common disease, symptom, body, drug, surgery, disease course and family history of pituitary adenomas were identified and analyzed. The study found that visual field-related symptoms are representative in the pituitary adenomas notes, and most of body concepts are related to vision, such as binocular, bilateral, temporal, etc. These observations are consistent with previous research findings that visual field defects are typical symptoms of pituitary adenoma.33,34 Additionally, fine-grained annotated data such as unilateral or bilateral and temporal visual field defects in this study have reference value and positive effects in assisting clinical diagnosis and treatment.
The annotated corpus constructed in this study initiatively provides clinical characteristics knowledge of pituitary adenomas, which is quality-assured and can serve as a reliable source for future integration of multi-source and multi-type information resources. In addition, the characteristics identified are computer-readable and computable, which accelerates the future development and application with large-scale biomedical knowledgebase such as gene ontology (GO) 35 and DrugBank. 36
Technical significance
Annotation of clinical data is a labor-intensive task, and how to reduce the labor costs and improve the efficiency is being studied. The semi-automatic corpus construction framework proposed is generally applicable to medical natural language scenarios. With the growing demand for non-English clinical corpora,8–10 the annotation guideline jointly discussed and revised by domain experts can provide a reference for tasks such as annotation, information extraction, data integration, and knowledge fusion of pituitary adenoma-related texts in other languages. The characterization idea is applicable to the related research of other diseases. Additionally, the recommendation and annotation models have been integrated into the CCNLP platform accessible online.
The constructed corpus not only achieves the characterization of pituitary adenomas, but also facilitates secondary utilization for clinical NLP due to its good computer readability and computability. LLMs have demonstrated powerful learning and reasoning capabilities in general fields. But for clinical natural language scenarios where language structure and terminology are highly specialized, its performance is worrisome and even faces hallucination challenges.37,38 Some studies39–41 have raised concerns that the clinical NLP performance of LLMs may not be as good as existing deep learning models. Consistent with previous studies, our study shows that the performance of LLMs without fine-tuning is indeed unsatisfactory. In the meanwhile, an impressive finding is that after fine-tuning with TCPA, the performance has been greatly improved, which suggests that the disease characteristic knowledge annotated in the domain corpus is conducive to enhance the clinical utility effect of LLMs.
Limitations and future work
Although we have characterized pituitary adenomas and applied them on LLMs, our study was limited to Chinese clinical notes, and the study cohort was not fully representative of the patient population with pituitary adenomas. For future work, with the development of deep neural networks and generative language models, the constructed text corpus would be combined with larger and more diverse data such as laboratory examinations and tests for the joint multimodal clinical data analysis and mining. The integrated application of the identified clinical knowledge with large-scale biomedical ontologies is also our next research focus. In addition, this study preliminarily explored the feasibility of using clinical knowledge of pituitary adenomas to guide LLMs for automated information extraction. In view of the positive experimental results, the use of LLMs for the construction of a larger medical corpus is also the next direction of effort to accelerate the secondary use of clinical NLP and develop the huge potential of AI technology in clinical auxiliary diagnosis and treatment.
Conclusions
In this paper, we constructed an annotated corpus and characterized pituitary adenomas in clinical notes. To improve the efficiency of text annotation, we proposed a corpus construction framework and developed CCNLP platform with integration of the text recommendation algorithm and the automatic annotation algorithm. An annotation guideline defining medical concepts and their assertions of clinical notes was developed, and fine-grained text types were identified to be annotated with the comprehensive consideration of the language, structure and content characteristics of clinical texts. Based on the method proposed in this paper, an annotated clinical text corpus of pituitary adenomas has been constructed. The rich clinical information about disease, symptom, body, drug, surgery, disease course and family history of pituitary adenomas were identified and analyzed. Additionally, this study applied the constructed TCPA to fine-tune LLMs and manifested the benefit of internal domain knowledge to LLMs in medical applications.
The constructed corpus has good machine-readable and computability, and can be further used in AI-aided medical applications such as multi-dimensional risk assessment, postoperative recurrence prediction, and patient health management. The corpus construction framework proposed in our work is general for clinical free-texts and can provide a reference for the characterization of other diseases.
Supplemental Material
Supplemental Material - Characterizing pituitary adenomas in clinical notes: Corpus construction and its application in LLMs
Supplemental Material for Characterizing pituitary adenomas in clinical notes: Corpus construction and its application in LLMs by Jiahui Hu, Jin Fu, Wanqing Zhao, Pei Lou, Ming Feng, Huiling Ren, Shanshan Feng, Yansheng Li and An Fang in Health Informatics Journal.
Footnotes
Acknowledgements
The author expressed gratitude to the relevant contributors.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research was supported by the CAMS Innovation Fund for Medical Sciences (CIFMS) (Grant No. 2021-I2M-1-056), the National Natural Science Foundation of China (Grant No. 72074222), and the National Social Science Foundation of China (Grant No. 21CTQ016).
Ethical statement
ORCID iDs
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
