Abstract
Currently, the primary challenges in entity relation extraction are the existence of overlapping relations and cascading errors. In addressing these issues, both CasRel and TPLinker have demonstrated their competitiveness. This study aims to explore the application of these two models in the context of entity relation extraction from Chinese medical text. We evaluate the performance of these models using the publicly available dataset CMeIE and further enhance their capabilities through the incorporation of pre-trained models that are tailored to the specific characteristics of the text. The experimental findings demonstrate that the TPLinker model exhibits a heightened and consistent boosting effect compared to CasRel, while also attaining superior performance through the utilization of advanced pre-trained models. Notably, the MacBERT + TPLinker combination emerges as the optimal choice, surpassing the benchmark model by 12.45% and outperforming the leading model ERNIE-Health 3.0 in the CBLUE challenge by 2.31%.
Introduction
As one of the key tasks in a knowledge graph, entity relation extraction is of great significance and affects the quality of the knowledge graph. Entity relation extraction aims to identify specific associations between two entities and construct relation triples. For example, according to the disease diagnosis text, we can extract relations between “COVID-19”, “fever” and “cough” as (COVID-19, symptom, fever) and (COVID-19, symptom, cough). At present, the common method is neural network models such as Convolutional Neural Networks (CNN), 1 Recurrent Neural Networks (RNN), and Long Short-Term Memory (LSTM). These models are more advantageous in terms of autonomous learning features and good adaptability. With the development of natural language processing (NLP) technology, advanced methods such as neural network model and attention mechanism fusion, 2 tagging framework improvement, 3 and triple generation order adjustment 4 have successively achieved state-of-the-art performance on entity relation extraction.
In the field of medical text, medical entity relation extraction has been gradually taken seriously since the Informatics for Integrating Biology & the Bedside (i2b2) released a relevant task in 2010. However, due to the large differences between Chinese and English, the entities and entity relation types of Chinese texts are more special, so the effect of related techniques in processing Chinese texts may be less effective. Meanwhile, due to the privacy of medical information, the professionalism of the medical field, and the high standards and requirements of medical research, there is still much room for improving the effect of entity relation extraction for Chinese medical texts.
In addition, in the entity relation extraction of Chinese medical texts, there are often problems such as entity nesting and overlapping relations. For example, COVID-19, as shown in the previous example, can be used as the subject of a relation triple corresponding to different objects, thus forming multiple relation triples. At the same time, the traditional entity relation extraction adopts pipeline approaches, divides the entity extraction and the relation extraction into two subtasks, and regards the relation extraction as a classification problem, which leads to cascading errors. Therefore, this paper focuses on the entity relation extraction task of Chinese medical text in Chinese Biomedical Language Understanding Evaluation (CBLUE). 5 CasRel 6 and TPLinker 7 are selected to solve the problems of overlapping relations and cascade error in entity relation extraction, and pre-training models are added to improve the effect of entity relation extraction.
Related works
Pre-trained model
Pre-trained models are generally considered an efficient method to improve NLP tasks,8–11 especially after BERT was proposed. BERT can be fine-tuned to fit multiple downstream tasks, which accelerates and improves NLP tasks. 12
It is well known that BERT includes two main tasks, the masked language model (MLM) and next sentence prediction (NSP), which differ from traditional pre-trained models and make BERT a state-of-the-art model. Later, numerous derivative models were proposed to improve these tasks. e.g., the RoBERTa model is an enhanced BERT model with several improvements; it changes static masks to dynamic masks in MLM, removes the NSP task, and adopts a larger level of byte-pair text encoding and other changes in parameters and the volume of training data. 13 In addition, transforming the MLM task in BERT from the original random masking to the whole word masking also derived the BERT-wwm model and RoBERTa-wwm model. This kind of model is superior to the general BERT model and ERNIE model in tasks such as machine reading comprehension, natural language inference, and sentence pair matching. 14
However, it is still necessary to consider the applicability of different pre-training models in specific fields or tasks. e.g., Hofer et al. 10 analyzed the comparison of five optimization improvement measures of named entity recognition task in the case of a handful of tagged text for special corpora, such as medical texts and electronic medical records, and found that the best improvement effect could be achieved by using pre-training methods similar to the target field.
Joint extraction of entities and relations
Pipelined and joint learning are two main approaches in entity relation extraction. Pipeline extraction identifies named entities in sentences from a given dataset and then learns the relation between entities. The pipeline method is easy to conduct and flexible because entity extraction and relation extraction are operated separately, and datasets can be different. However, it also has some disadvantages. Errors in entity identification will be passed to the relation extraction task, thus affecting the following steps (i.e., the cascading error). Moreover, possible connections and dependencies between two highly decoupled tasks might be ignored. The task of pairing two entities can cause redundant information between two unrelated entities, which raises the calculation complexity and leads to errors.
Joint extraction combines entity and relation extraction to mitigate possible errors and error accumulation in the pipeline method. Some studies also show that joint extraction obtains better performance than pipeline extraction.15–17 Joint extraction can be divided into multitask learning based on shared parameters and structured prediction based on joint decoding according to the difference in model design.
Multitask Learning Based on Shared Parameters
In this approach, the state of input features and internal hidden layers in entity and relation extraction are joined together by sharing the same coding layer of the model. Another strategy is to strengthen the interaction between two tasks that apply encoders separately and finally obtain the optimal parameters, such as dependency enhancement such as reinforcement learning, 18 and risk minimization. 19 Miwa and Bansal 20 and FENG et al. 18 both take a “BiLSTM + Tree-LSTM” structure to achieve joint extraction through parameter sharing, but the latter applies reinforcement learning to relation classification, which leads to better results than the former. Zheng et al. 21 proposed a BiLSTM-ED method for entity recognition where Bi-LSTM parameters are shared in encoders and achieved superior results compared with Miwa and Bansal 20 and other models.
In the joint extraction method of shared parameters, most experiments use the encoding of BiLSTM to achieve parameter sharing and then incorporate models such as the attention mechanism and text span representation to enhance semantic association,2,3,17,22,23 followed by classification mapping of relations through the identified entities. In terms of the execution order of the two subtasks of entity recognition and relation extraction, there are also some experiments in which relations are extracted first and then mapped to entities through relations24–26, or the head entity in the relation triad is mapped to relations and tail entities.6,27,28
Structured prediction based on joint decoding
In the multitask learning method based on shared parameters, entity recognition, and relation extraction are still completed separately in a certain order, which cannot avoid generating redundant information. The joint decoding method will further strengthen the interaction between the two subtask models for global optimization. There are two main solutions: one is to use sequence tagging to obtain relation triples,3,7 and the other is to use the sequence-to-sequence method to re-encode and decode.29–32 In the entity relation extraction method of joint decoding, more attention is given to solving the problem of overlapping. According to the different types of overlapping, it can be divided into the following three categories: ①Normal, that is, no overlap; ②EPO (Entity Pair Overlap) or relations overlapping, that is, there are multiple relations in an entity pair; ③SEO (Single Entity Overlap) or entity overlapping, that is, different triples share the head entity or tail entity.
Zeng et al. 29 first proposed the classification of overlapping and designed an end-to-end model based on sequence-to-sequence learning of the replication mechanism, which has two ways of one decoder and multiple decoders in the decoding stage. According to the experimental results, regardless of whether one decoder or multiple decoders are used, the F1 value of the model is slightly worse than that of the optimal NovelTagging model 3 in the case of no overlap. However, in the case of EPO and SEO, the one-decoder and multiple-decoder modes are improved by 7.8% and 31.1%, respectively, and the multiple-decoder mode achieves the highest F1 value. Yuan et al. 25 proposed the relation-specific attention network (RSAN) model to specifically solve the overlapping problem and had the best performance in solving SEO and EPO types. The F1 value on the EPO type reaches the same performance as the current best in normal cases. In addition, there are other optimization approaches, such as supervised multi-head self-attention mechanism, 2 position attention mechanism, 16 and hybrid dual pointer network. 33
Methodology
CasRel
CasRel is a general algorithmic framework proposed by the team of Jianlin Su to solve the overlapping problem. Unlike other approaches regarding relations as discrete labels on entity pairs, CasRel models relations as a function of subject mapping to objects. The reason is that if the relations of entity pairs are discrete, the model will obtain unevenly distributed data, and for overlapping labels, if the training data are insufficient, the accuracy rate of the classifier will be greatly reduced.
Relation extraction in CasRel identifies sentence subjects first and then identifies all possible relations and objects for each subject. As shown in Figure 1, the model consists of three modules: BERT-based encoder, subject tagging, and relation-specific object tagging. Subject tagger adopts two binary classifiers to predict the head and tail index of subjects. Classifiers calculate the subject start or end position probability of each word. The corresponding token will be tagged as one when the probability exceeds a given threshold and otherwise as 0, thus predicting all subjects. The relation-specific object tagger takes the encoded vector of the subject into account. Vectors are passed to the process of conditional layer normalization, where the input data with different dimensions of features are normalized. All the relations in the dataset are traversed once to obtain objects corresponding to this subject under each relation. The structure of CasRel.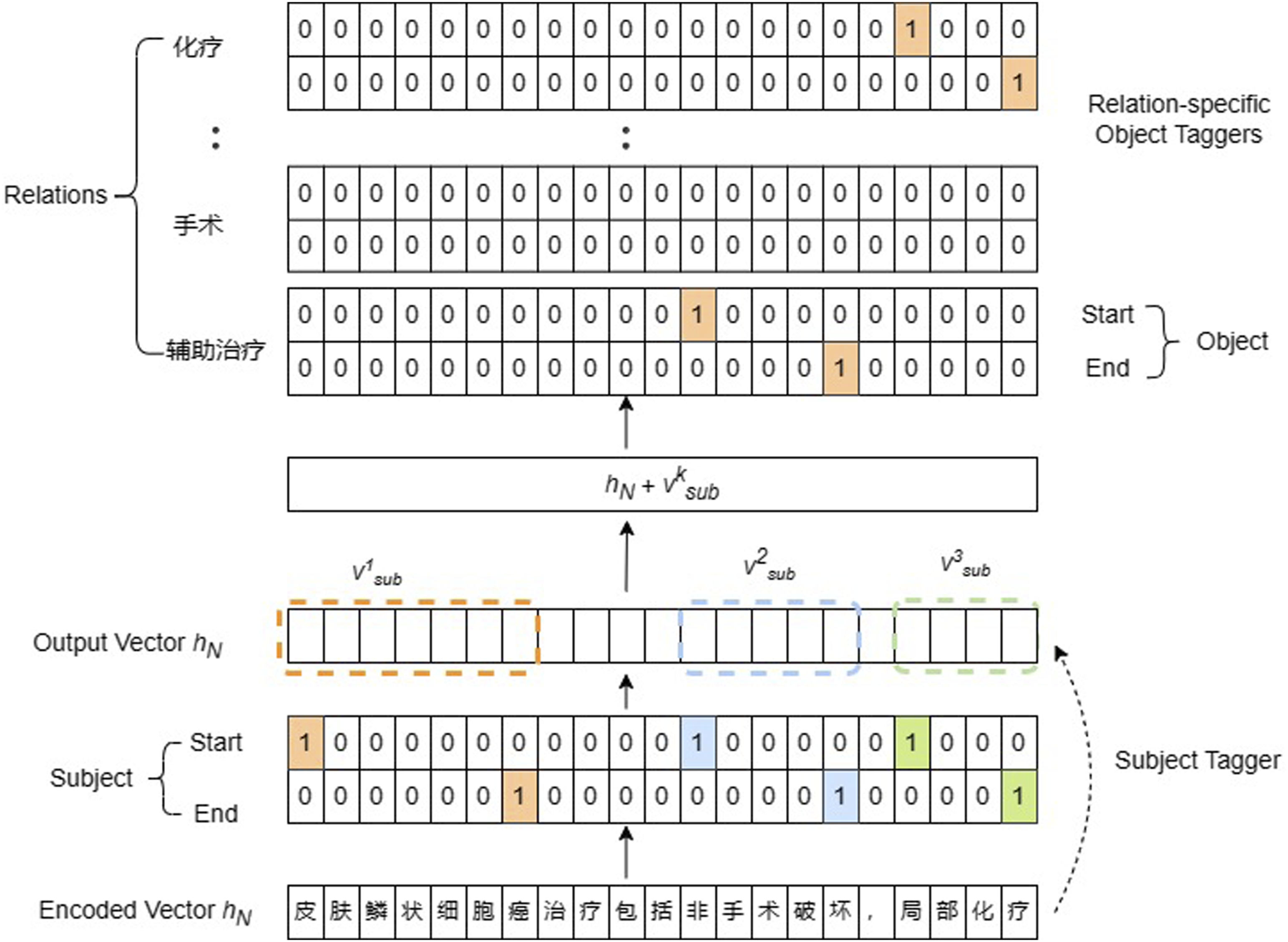
TPLinker
CasRel is an effective solution to the overlapping problem, but exposure bias still exists during training and inference, while TPLinker tackles overlapping and exposure bias based on the Muti-head selection (MHS) 15 mechanism, which appears to be more complete than the former.
Exposure bias is caused by the inconsistency of training input and inference input. Each word input is ground truth in the training process, but the inference input is from the output of the last word depending on the model prediction, which causes errors to accumulate.
The tags in TPLinker include 0, 1, and 2, where one indicates the relation from subject to object and two from object to subject. As shown in Figure 2(a), given a sentence example “小明在北京 (Xiaoming is in Beijing)”, the cross position of the row of “小” and the column of “明” with tag one means that the sequence fragment “小明” is an entity. The cross position of the row “小” and the column “北” has a tag of 1, which means a relation between two entities starting with “小” and “北”, respectively; that is, there is a correlation between “小明” and “北京”. In this way, entity extraction and relation extraction can be performed in the same phase, without the need to decouple the task as in CasRel. An example of tagging in TPLinker.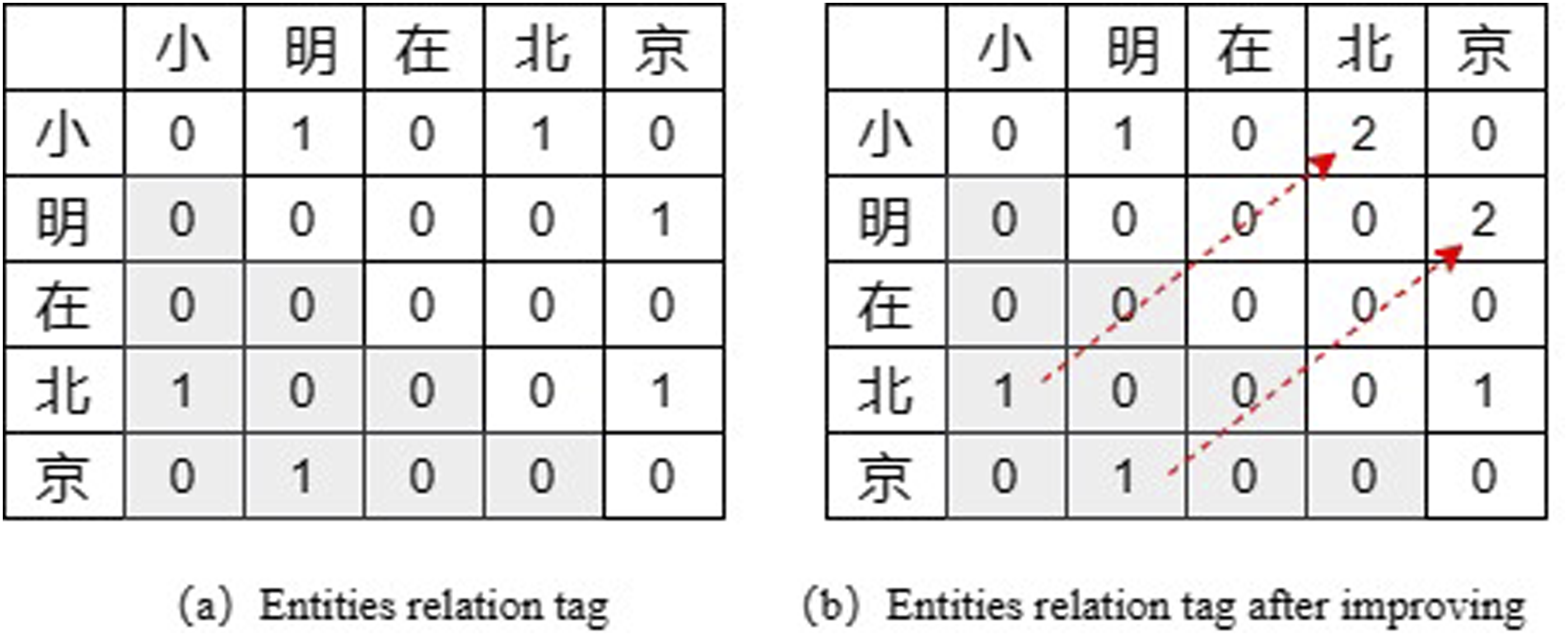
There are a large number of tag 0 in the lower left triangle region due to the logical order of the entity head and tail, which will lead to an enormous waste of memory and increase the calculation amount of the whole model. However, because the order between the object and subject is not certain, deleting the lower triangle directly will also lead to the loss of some relations. In this regard, the TPLinker model proposes a new solution by mapping the tag “1” in the lower left triangle to the upper right triangle and then relabelling it as “2”. The final tag result is shown in Figure 2(b). In the end, the remaining parts of the triangle are flattened into a sequence for calculation.
According to the tagging strategy of TPLinker, the method transforms the task of joint entity relation extraction into a Token Pair Linking (TPLinker) problem, thus proposing three different linking methods, namely, head-to-tail linking of the same entity (EH-TO-ET), head-to-head linking of different entities (SH-TO-OH), and tail-to-tail linking of different entities (ST-TO-OT).
14
All token pairs are enumerated and tagged through matrices for the convenience of obtaining entities. In decoding, for the type EH-TO-ET, all entities can be extracted from the sentence and stored in the dictionary. For each type of relationship, decoding ST-TO-OT and SH-TO-OH yields the tail position set of the subject and object entity pairs and the start position set of the subject and object entity pairs, respectively. Then, TPLinker traverses the entity dictionary using the start position set corresponding to the entity pair and checks the tail position set. If the tail exists, an SPO triple is successfully extracted. In Figure 3, the text “故宫博物院坐落于中国首都北京” is an example showing the Token Pair Linking process of TPLinker. An example of a token pair link through TPLinker.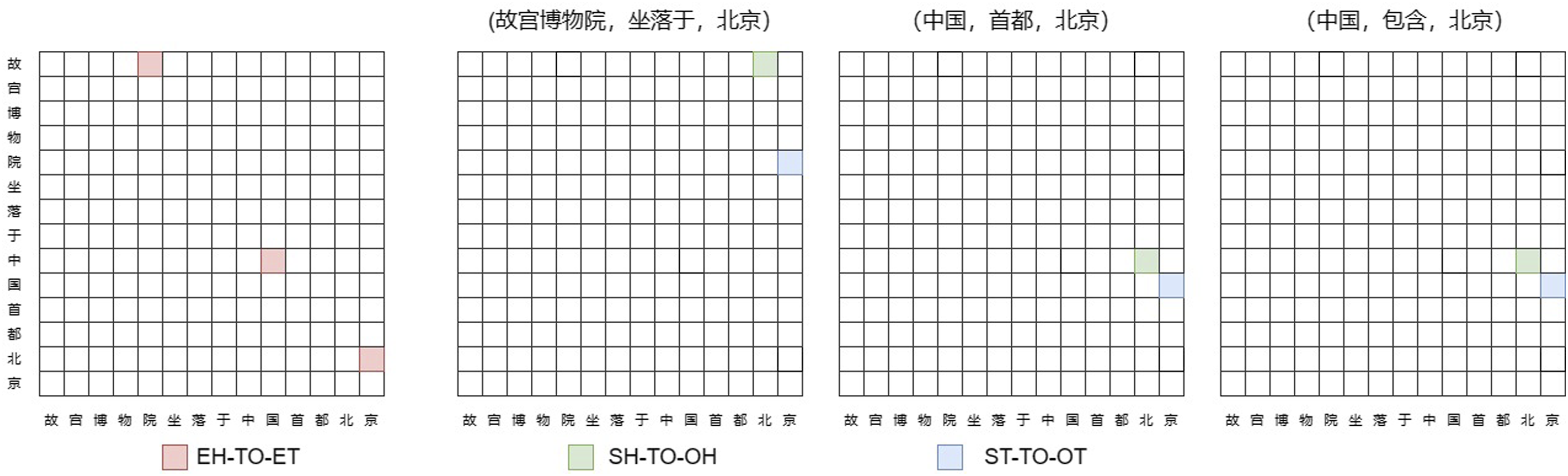
Experiments
Dataset
The dataset used in this study is CMeIE provided by the Chinese Biomedical Language Understanding Evaluation (CBLUE). 5 CMeIE contains 14,339 pieces of data from the open training set, 3585 from the validation set, and 4482 from the test set, involving a corpus of 518 pediatric diseases and 109 common diseases, all from medical textbooks and medical texts of clinical practice, with a total of approximately 75,000 triples, 28,000 disease statements, and 53 defined schemas. There are 10 types of head entities and 11 types of tail entities in entity categories, one synonym sub-relation, and 43 other subtypes in relationship categories, for a total of 53 schemas.
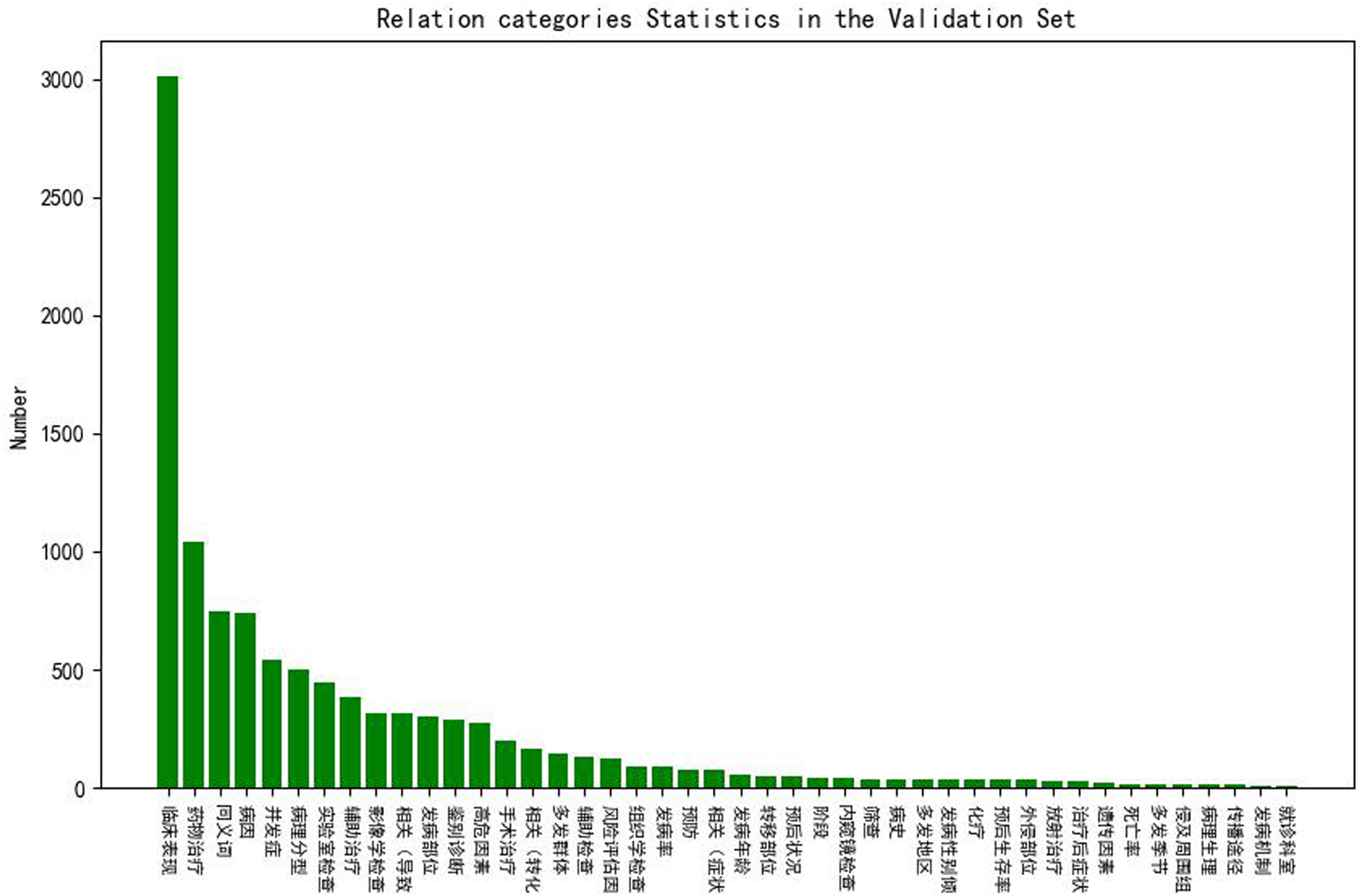
Figures 4 and 5 show the statistics of the relation categories in the training and validation sets. From the figures, we can see that the distributions of the two datasets are similar, and the most numerous relation categories are “临床表现 (clinical manifestation)”, “药物表现 (drug manifestation)”, “同义词 (synonym)” and “病因 (etiology)”. However, the distribution of relation categories is not balanced. We can see that the most frequent category “临床表现 (clinical manifestation)” is more than twice as frequent as the second most frequent category “药物表现 (drug manifestation)” and 200 times more frequent than minor categories in the training set. Relation categories statistics in the training set. Relation categories statistics in the validation set.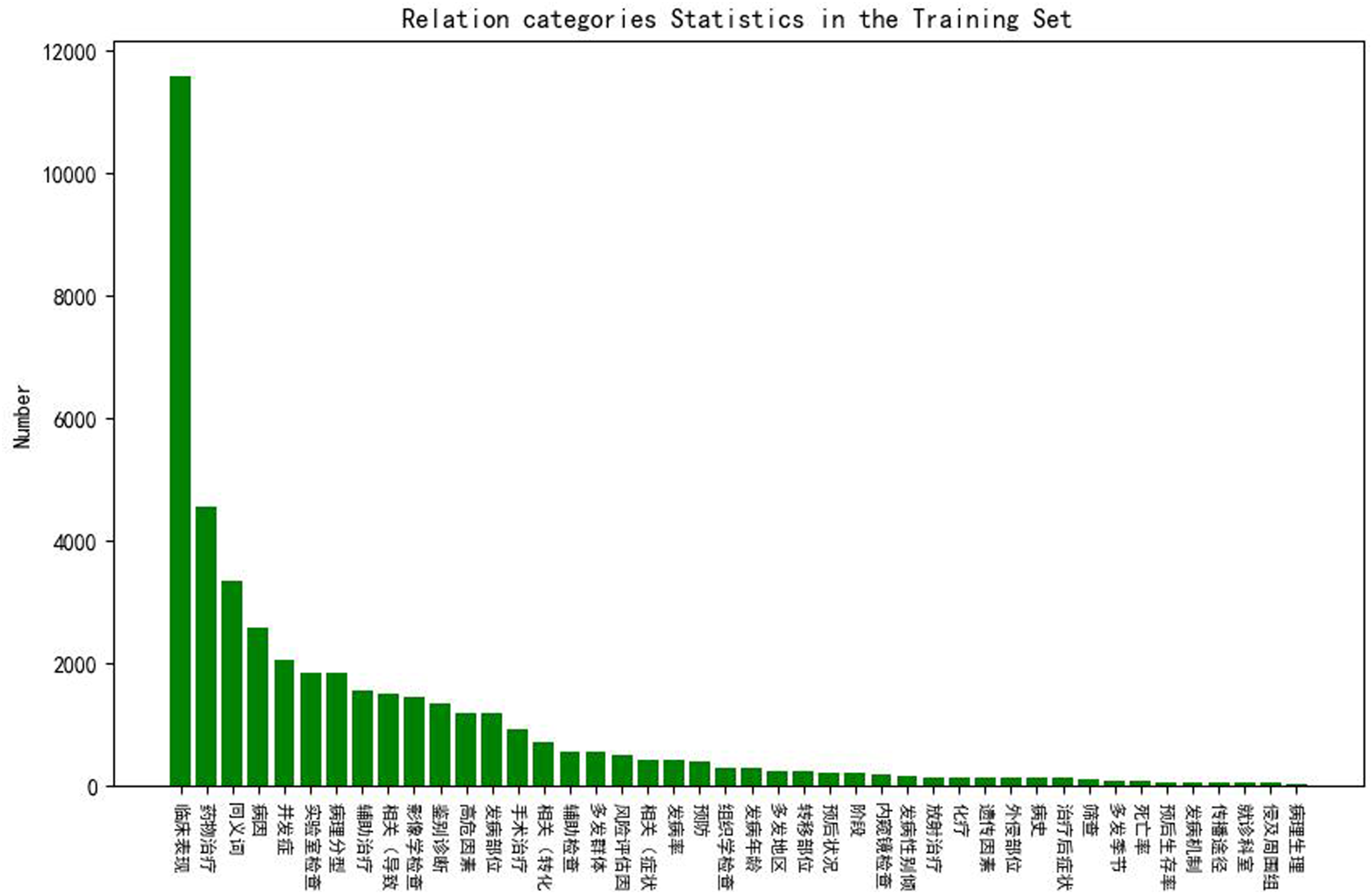
Evaluation and baseline
The operating system used in our experiments is Linux Spuer-HR 4.15.0-189-generic, Intel(R) Xeon(R) Gold 6139 M CPU @ 2.30 GHz CPU, and NVIDIA GeForce RTX 2080Ti GPU.
The CasRel model in our experiments is adopted from Wei et al. 6 and implemented in TensorFlow. TPLinker is adopted from the optimized version TPLinker_plus by Wang et al, 7 implemented in PyTorch. In our experiment, we use exact matches to evaluate whether entity relations are correct. We report three metrics, Precision(P), Recall(R), and F1 value, which are commonly used in NLP tasks.
Both models were previously conducted on two English datasets, NYT and WebNLG, for the entity relation extraction task, and both BERT and BiLSTM encodings were tested. 7 From the experimental results, the TPLinker model has a 2.3% advantage over the CasRel model when using BERT encoding on the NYT dataset; the two models have the same effect when using BiLSTM encoding. On the WebNLG dataset, the TPLinker model has only a 0.1% advantage over the CasRel model when using BERT coding; the TPLinker model has a 6.8% advantage over the CasRel model when using BiLSTM coding. Overall, the TPLinker model has a slight advantage over the CasRel model in handling the entity relations extraction task for the English dataset, but the advantage is not stable due to the difference in dataset and encoding method.
Comparison of CasRel and TPLinker with BERT-base, Chinese.

From the experimental results, the precision, recall, and F1 values of the TPLinker model are higher than those of the CasRel model in the face of entity relationship extraction of Chinese medical text. The recall rate of the former exceeds that of the latter by 8.6% and the F1 value by 6.01%, and the advantage of the TPLinker model is more significant.
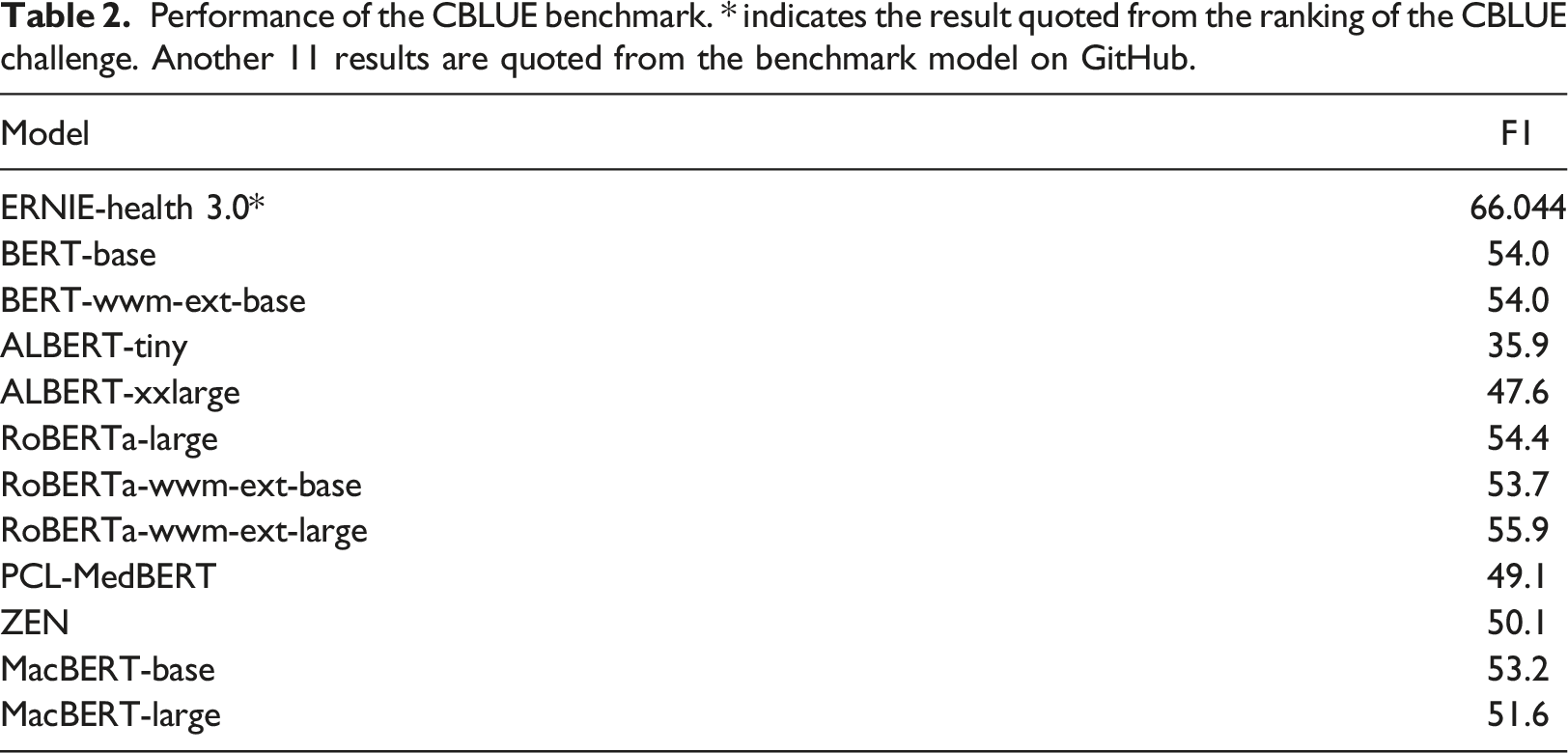
Performance of the CBLUE benchmark. * indicates the result quoted from the ranking of the CBLUE challenge. Another 11 results are quoted from the benchmark model on GitHub.

Optimization
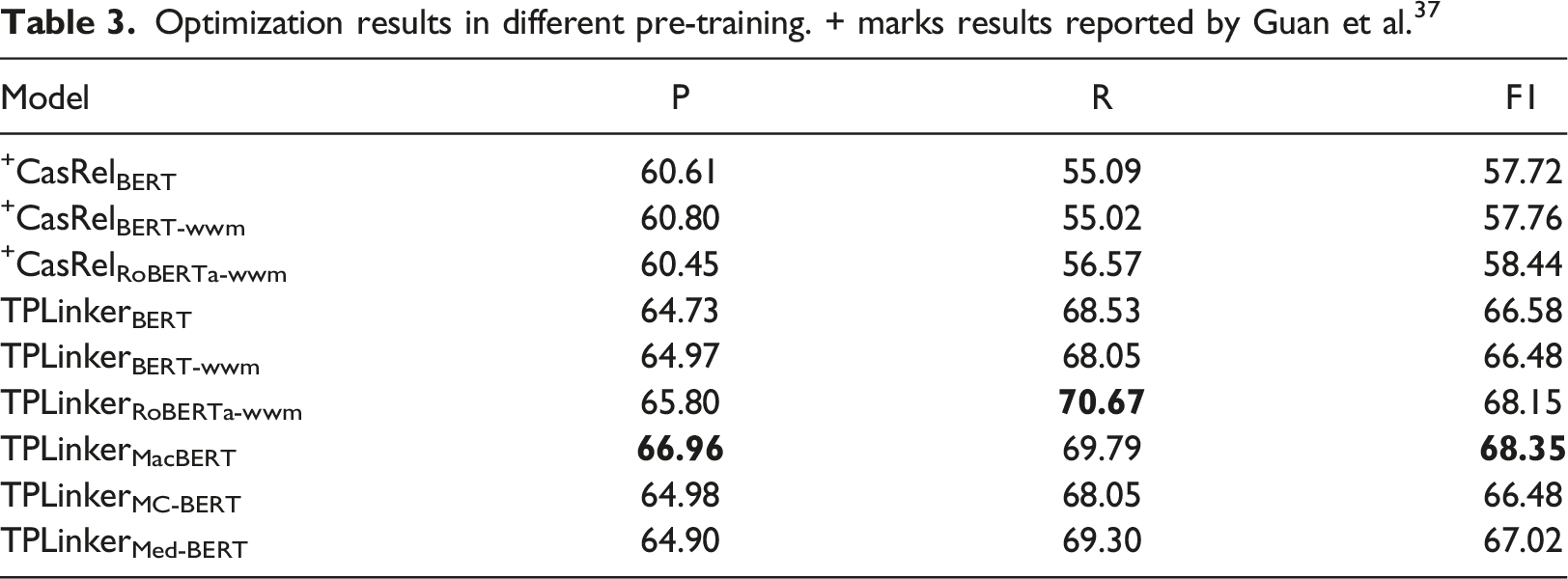
Optimization results in different pre-training. + marks results reported by Guan et al. 37
Discussion
Model comparison for joint extraction
Based on the findings presented in Table 3, the experimental results indicate that, for the first six sets of experiments, the TPLinker model consistently exhibits a more stable improvement advantage over the CasRel model when processing the Chinese medical dataset CMeIE, irrespective of the encoding method employed. In terms of encoding methods, the RoBERTa-wwm model demonstrates superior support compared to both BERT and BERT-wwm-ext-base.
The CasRel model employs an entity mapping relation approach to facilitate joint extraction. Within this model, the extraction of relations relies on the correspondence between subject entities and relation-specific object entities. Consequently, if the object entities are either untagged or incorrectly tagged, the extraction of the triple is impeded or may result in erroneous extraction. Conversely, the TPLinker model generates a span matrix by considering token pairs during the tagging phase. After decoding the span matrix, the corresponding triples can be obtained based on the entity tag and relations tag types. Furthermore, entity relations can be acquired through a unified encoding and decoding process. In contrast, the TPLinker model maintains consistency throughout the tag, encoding, and decoding stages, thereby yielding superior outcomes and demonstrating enhanced suitability for Chinese medical text.
Optimization comparison of the pre-training model
During the optimization process of the TPLinker model, it was observed that the pre-trained models MacBERT, RoBERTa-wwm, and MedBERT exhibited superior performance. In comparison to the original model utilizing BERT, these models demonstrated increases in F1 scores by 1.77%, 1.57%, and 0.44% respectively. However, BERT-wwm and MC-BERT experienced slight decreases in performance. Notably, the F1 value achieved by the optimized structure “MacBERT + TPLinker” reached 68.35%, surpassing the F1 score of the top model Ernic-Health 3.0 in the CBLUE challenge by 2.31%. The findings demonstrate that the utilization of pre-trained models trained on a corpus resembling Chinese medical text yields superior model outcomes. However, the magnitude of improvement is contingent upon the source of training data, the volume of training data, and the parameter configurations of the pre-training models.
RoBERTa employs a significantly larger dataset (160G) during the pre-training phase, resulting in an extended training duration. It is worth mentioning that the RoBERTa-wwm variant incorporates a whole word mask mechanism, effectively addressing Chinese semantic complexities and enhancing its applicability in Chinese text-mining endeavors. In contrast, MacBERT is a perpetually refined model built upon the integrated BERT model and its enhanced iterations. It employs a blend of synonymous and random words for masking during the MLM task, thereby mitigating the disparity between the pre-training and fine-tuning stages. 36
Both the MC-BERT and MedBERT models are pre-trained models derived from BERT models, specifically designed for the biomedical field. The MC-BERT model is trained using Chinese Biomedical Community Question Answering, Chinese Medical Encyclopedia, and Chinese Electronic Health Records (EHR) as data sources, with a training data volume of approximately 20 M. Moreover, this model incorporates explicit medical knowledge by employing whole entity masking and whole span masking methods. 34 The MedBERT model utilizes Cerner Health Facts, a structured electronic health record (EHR) database, as its training data source, which consists of approximately 20 million data points. To pre-train, MedBERT employs the International Classification of Diseases (ICD) encoding and serialized word embedding. 35 However, it is worth noting that incorporating explicit medical knowledge as input restricts the model’s learning capacity. In contrast, the adoption of a generalized serialized word embedding learning approach proves more suitable for the practical implementation of medical texts.
Comparison with ChatGPT
The triplet results were extracted by ChatGPT.

We provide ChatGPT with sample data and all available relationship types for learning and then input sample data and request ChatGPT to output triples in (subject, predicate, object) format. 42 In Table 4, Sample data represents the data input into ChatGPT, ChatGPT results are the triplets extracted by ChatGPT, Standard results are the triplets in the validation dataset, and Overlap or not means the triplet overlap type.
According to the results in Table 4, ChatGPT has the functions of entity identification and triplet extraction. For simple text (e.g. Sample data one and Sample data 4), the extraction results can be close to the verification set results, but they do not meet the strict standards. In the context of text extraction involving compound structure (e.g. Sample data two and Sample data 3), ChatGPT demonstrates an inability to accurately generate triples. Furthermore, it is crucial to acknowledge that despite the progress achieved in the development of ChatGPT, there remains a necessity for comprehensive training and expert guidance to effectively employ the model for unfamiliar tasks. Additionally, it is noteworthy that the output produced by the model may not consistently align with accuracy. The experimental findings of Qin et al. 43 further illustrate the formidable obstacles ChatGPT encounters when addressing sequence annotation tasks, including named entity recognition. Our model is primarily designed to tackle this particular problem, focusing on the simultaneous extraction of entity and relation while also considering relation overlap in complexly structured text. As a result, our model offers distinct advantages in addressing this issue.
Limitations
It is important to acknowledge that the findings of this experiment reveal certain limitations in the performance of CasRel and TPLinker. For instance, when considering the CasRel model, it achieved notable improvements of 17.5% and 30.2% on the English datasets NYT and WebNLG, respectively, surpassing the previous best results. However, in the present experiment, the disparity in results between CasRel and the benchmark is merely 4.59%, with CasRel’s performance falling short of the result obtained by the CBLUE challenge ERNIE-Health 3.0. The TPLinker achieved a significant advancement, as evidenced by the superior performance of all optimized pre-training models compared to the result obtained from the ERNIE-Health 3.0 model in the CBLUE challenge. The potential factors contributing to this success are comprehensively examined as follows: (1) The Chinese medical text dataset contains a substantial quantity of lengthy text and cross-sentence SPO triples, with an uneven distribution of triples across different relation categories. The dataset exhibits a prominent long-tail problem, as the “clinical manifestations” relation category is frequently represented, whereas the “pathogenesis” category is less prevalent. (2) The absence of any subsequent fine-tuning for the pre-trained model is a significant limitation. Fine-tuning plays a crucial role in enhancing the performance of pre-trained models, particularly in the realm of natural language processing tasks. By allowing pre-trained models to adapt to specific downstream tasks, fine-tuning not only enhances their performance and efficiency in processing new data and tasks but also capitalizes on their pre-existing knowledge and representations, thereby mitigating the necessity for extensive training on task-specific data.43–46
Conclusion
This study centers on the extraction of entity relations in Chinese medical text. To tackle the issues of overlapping relations and cascading errors, we employ CasRel and TPLinker to compare their performance in Chinese medical text and determine the most suitable pre-trained model for optimization. The findings indicate that TPLinker exhibits a more consistent improvement compared to CasRel and achieves superior results when supported by an advanced pre-trained model. The optimal model, MacBERT + TPLinker, surpasses the top-performing model, Ernic-Health 3.0, in the CBLUE challenge by 2.31%, and exceeding the benchmark model by 12.45%. Additionally, we conducted supplementary experiments using ChatGPT, which revealed its capability to generate triples. However, when applied to Chinese medical texts characterized by intricate structures, ChatGPT exhibited limitations in generating logical triples and addressing the issue of relationship overlap.
Footnotes
Acknowledgements
Our work is supported by the Basic Project of the Science and Technology Program of Guangzhou, China (NO.202002020036). The results disclosed in this work are obtained using the high-performance computing infrastructure and resources of Sun Yat-Sen University and the Reporting System of China Chest Pain Center Certification (Website:![]() ).
).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Our work is supported by the Basic Project of the Science and Technology Program of Guangzhou, China (NO.202002020036)
