Abstract
Objectives: This study aims to identify necessary adjustments required in existing oncological datasets to effectively support automated patient recruitment. Methods: We extracted and categorized the inclusion and exclusion criteria from 115 oncological trials registered on ClinicalTrials.gov in 2022. These criteria were then compared with the content of the oBDS (Oncological Base Dataset version 3.0), Germany’s legally mandated oncological data standard. Results: The analysis revealed that 42.9% of generalized inclusion and exclusion criteria are typically present as data fields in the oBDS. On average, 54.6% of all criteria per trial were covered. Notably, certain criteria such as comorbidities, pregnancy status, and laboratory values frequently appeared in trial protocols but were absent in the oBDS. Conclusion: The omission of criteria, notably comorbidities, within the oBDS restricts its functionality to support trial recruitment. Addressing this limitation would enhance its overall effectiveness. Furthermore, the implications of these findings extend beyond Germany, suggesting potential relevance and applicability to oncological datasets globally.
Introduction
Over the past few decades, rapid advancements have been witnessed in the field of oncology, leading to significant improvements in cancer patient treatments. Clinical research has played a pivotal role in driving this progress. 1 Participating in oncological studies offers patients the opportunity to receive early treatment using new and innovative methods. However, the current approach of relying on doctors, tumor boards, or interdisciplinary oncological councils to recruit patients for study participation poses numerous challenges. Even well-informed doctors typically lack knowledge of all ongoing studies and their inclusion and exclusion criteria. Therefore, a limited availability of potential participants does not necessarily stem from a lack of interest or refusal on the part of patients. 2 Other factors contributing to low recruitment numbers may include inadequate organizational structures and ethical conflicts, which may also vary depending on the locality.3–6 Despite advancements in medical and information technology, patient recruitment in Germany continues to rely heavily on direct patient contact, with limited utilization of available technologies. 7
Though, the concept of automating patient recruitment to alleviate the associated challenges has been contemplated for some time. 7 For instance, the German Medical Informatics in Research and Care in University Medicine (MIRACUM) consortium 8 is developing an automated recruitment tool that utilizes patient data to determine their eligibility for clinical trials. 9 However, a significant hurdle lies in the increasing complexity of patient disease profiles, requiring more comprehensive documentation. In the field of oncology, initiatives such as the German Network for Personalized Medicine (DNPM) strive to provide personalized therapy for complex medical cases. 10 This paper examines the relevant inclusion and exclusion criteria in current oncological studies and evaluates the extent to which existing enforced oncological standards, in particular, the German oncological base data set - version 3.0 (oBDS), can support automated patient recruitment. 11 The selection of the oBDS dataset for analysis is based on its status as a legally mandated dataset in Germany and its integral role as a major inspiration in numerous large-scale oncological projects. As an example, the oBDS serves as the foundation for the clinical data catalogue of the National Network Genomic Medicine (nNGM), which draws extensively from the oBDS. 12 Additionally, the oBDS serves as a blueprint for the expansion module of the medical informatics core dataset, aiming to connect various medical domains, including e.g. oncology and radiology, with the oBDS dataset as its cornerstone within the oncology domain. 13 While the oBDS may not encompass all necessary data categories for all clinical trials, it remains the nationwide standard for tumor documentation in Germany. 11 Similarly, oncological datasets exist in other countries like the United States, England or others.14,15 While their structures differ, these datasets are similar in complexity and general data contents among each other, suggesting that the findings in this study may have relevance beyond Germany.
This paper analyzes the potential utilization of enforced oncological datasets, specifically the oBDS dataset, to support automated patient recruitment in clinical trials. It assesses the extent to which enforced oncological datasets can fully cover studies and discusses the need for dataset expansion.
Methods
The first part of this work was the detection of relevant studies. The source of the study collection is the website ClinicalTrials.gov. It is provided by the U.S. National Library of Medicine and serves as the world’s most important registry which provides access to information about clinical trials. 16
The initial detection of studies was performed on January 31 of 2022. The first filter set was the subject area of studies; in this case, oncological studies were required. There was no further specification regarding the cancer variations or the organ entities. To narrow it down it was necessary that the study was either recruiting or not yet recruiting. Additionally, it needed to be an interventional study, which contained a study protocol. The study protocol was fundamental since it holds information about the inclusion and exclusion criteria. Furthermore, the studies must have taken place between January 1 of 2019 and the day of the initial detection, January 31 of 2022.
After a thorough selection of the studies, the review of each study protocol with regard to their inclusion and exclusion criteria began. The review of the study protocols started out with the identification of each inclusion and exclusion criteria of every study. Using Microsoft Excel 2016, the transcription of the study protocol’s criteria into a database was carried out. During the transfer, no attention was paid to whether it was an inclusion or exclusion criteria, but rather that it was a criteria in the general sense as all exclusion criteria can be formulated as negated inclusion criteria.
This process was followed by a generalization of each study criteria. The generalization serves to group multiple criteria into fitting categories. To demonstrate the idea of the generalization the following example can be given: Extracted criteria regarding the age, in this case for example ‘18+ (NCT04342429)’ and ‘Adult 21+ (NCT04745754)’ were transcribed into a newly generalized category called ‘age’. This process served to generalize criteria that fundamentally share a common goal. The new categories were not established based on existing standards, for example, those derived from the EHR4CR project or other sources.17–19 Instead, they were empirically formed by extracting all inclusion and exclusion criteria in their raw form and grouping them based on their similarities. This approach was chosen to remain open to potential new categories, such as COVID-19, and also to provide a specific focus on oncology, as exemplified in the case of molecular markers.
While the primary author undertook the direct extraction of raw criteria from the trial protocols, discussions on generalization were conducted in collaboration with senior staff at the local Comprehensive Cancer Center. Additionally, insights were sought from the center’s physicians, tumor documentalists, and medical informatics practitioners to ensure a comprehensive and well-informed approach. After these preparatory steps, a comparative data analysis was carried out with the goal to identify the previously generalized categories in data fields of the oBDS dataset. This comparative data analysis additionally aimed to identify the extent to which the oBDS dataset can provide information usable in an automated patient recruitment process. It also allows identifying those frequently occurring generalized categories that are missing in the oBDS.
To alleviate the effort of interpretation, a descriptive analysis was conducted using the free software programming language R (version 4.2.1), developed by the R Core Team. 20 The goal was to evaluate which of the categories have the highest impact when it comes to patient recruitment, either in terms of individual studies or more generally the overall impact.
Results
On the day of the initial detection, 402,624 studies were registered on the clincaltrial.gov website. First, the filter “oncology” was set, which resulted in an inclusion of 98,880 studies. Then, another filter was set regarding the start date, which reduced the number of studies to 23,985. The filters “recruiting and not recruiting”, “interventional” and “study protocol” reduced the number of studies to 506. The last filter was the end date, which was the date of the initial detection, and reduced the size to 145 studies. Later on, another 30 studies were excluded. This was due to some trials being labeled as non-oncological trials as well as some trials being deleted from the clinialtrial.gov website. The final study collection contains 115 studies. The exact number of studies selected after each subsequent filter can be seen in Figure 1. Process of study selection.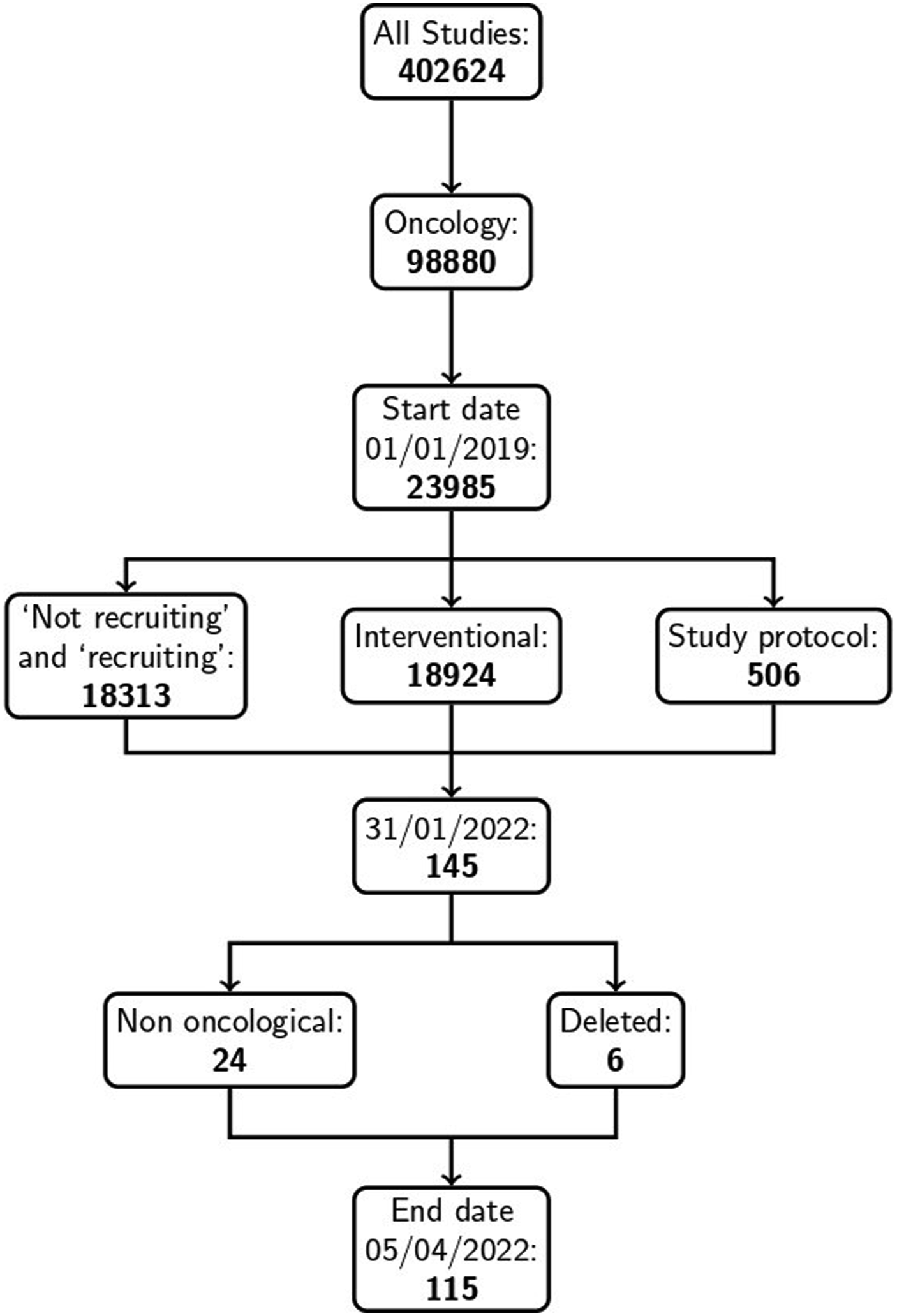
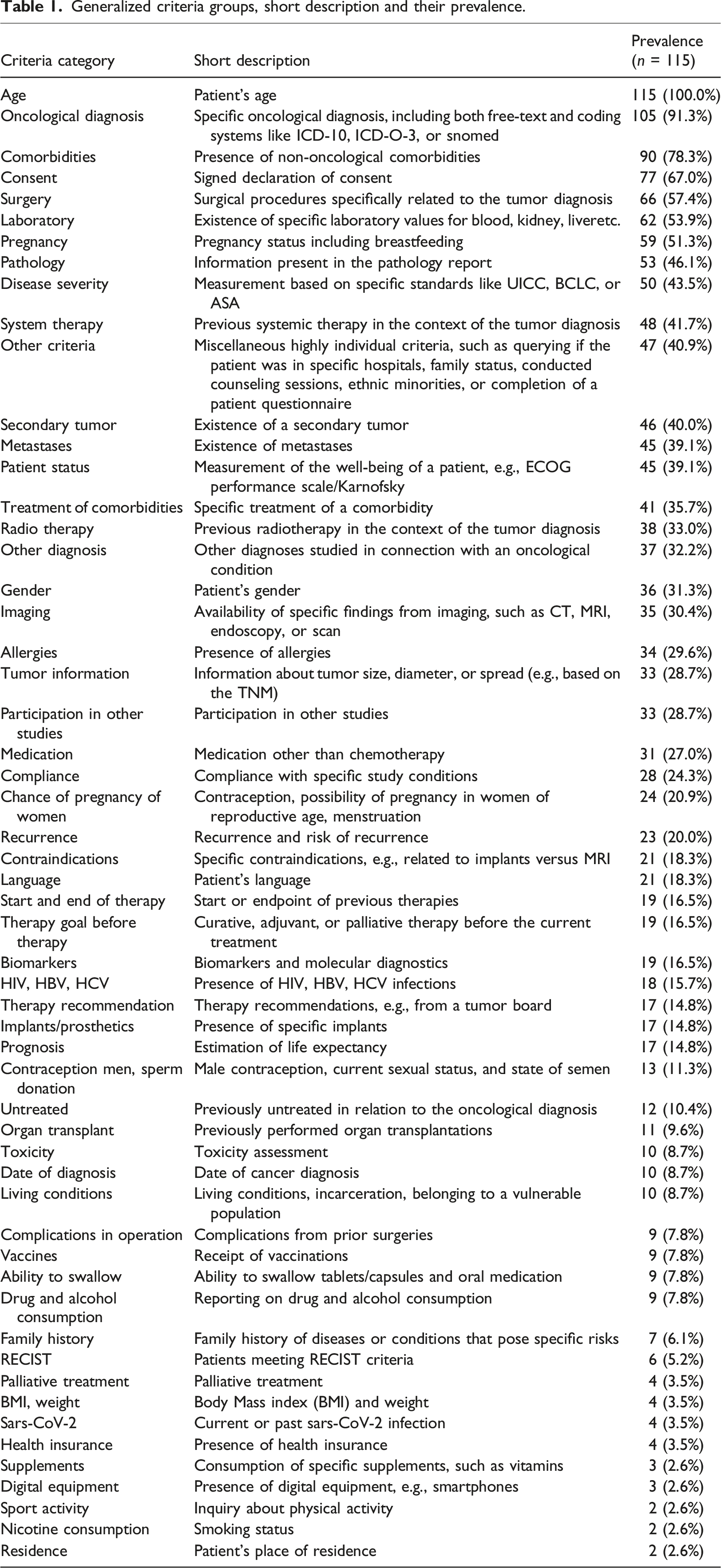
Generalized criteria groups, short description and their prevalence.
The color difference on the bar chart in Figure 2 is intended to show coverage by the oBDS dataset in addition to the prevalence of the criteria. The lighter bars show the criteria found in the oBDS dataset while the darker bars show which criteria were not found in the oBDS dataset. Prevalence of the generalized criteria groups and it’s occurence in the oBDS dataset.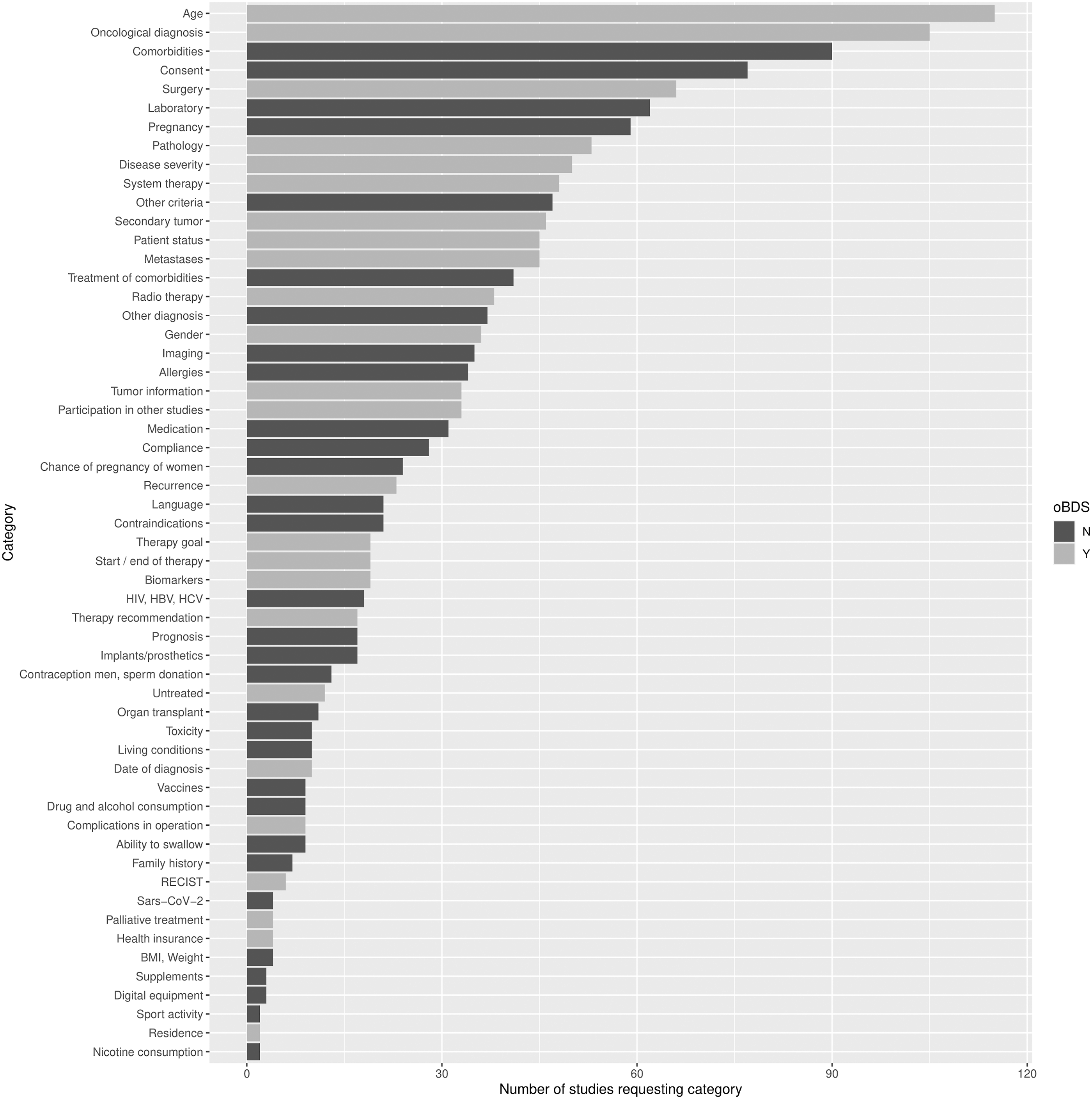
The results of the comparative analysis show that 24 out of 56 identified criteria groups from the ClinicalTrial.gov study protocols are listed in the oBDS dataset. Out of the five most common required criteria three were covered by the oBDS dataset.
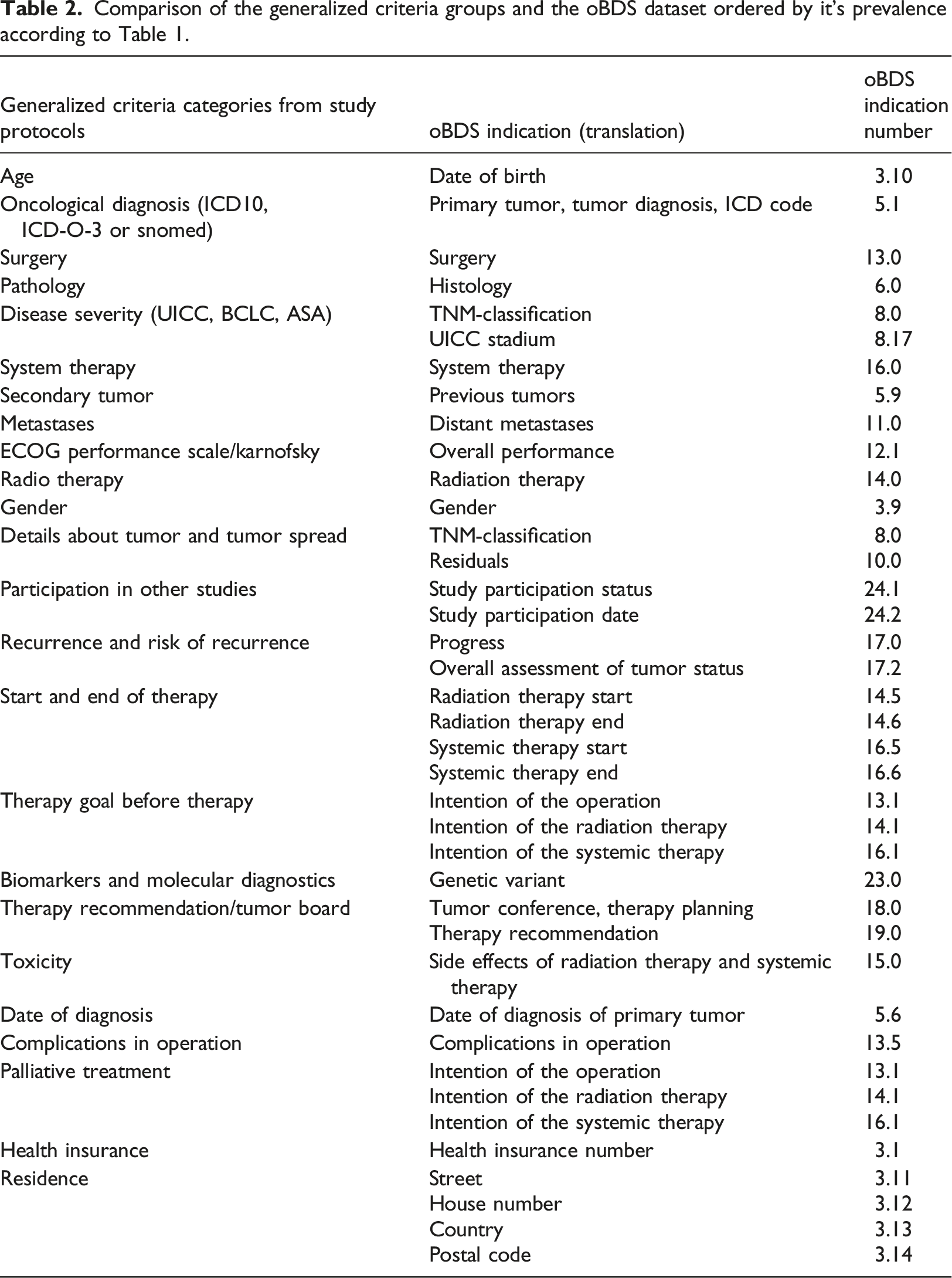
Comparison of the generalized criteria groups and the oBDS dataset ordered by it’s prevalence according to Table 1.

Another focus of this investigation was how many criteria, regardless of whether they were inclusion or exclusion criteria, were required by the study protocol for each study and how many of these could be supported by the oBDS dataset.
Two studies were fully covered by the oBDS dataset. The study with the ClinicalTrials.gov study ID NCT04424758 requires three criteria which were included by the oBDS dataset and the study with the ClinicalTrials.gov study ID NCT04743999 requires six criteria which were included by the oBDS dataset. The study protocol that required the most criteria has the ClinicalTrials.gov study ID NCT04357873. This protocol requires 35 (out of 56) criteria with 15 (42.9%) being covered by the oBDS dataset. The bar chart in Figure 3 visualizes the number of required criteria per study protocol and whether they are covered by the oBDS dataset. Required criteria in each study and to what extend they are covered by the oBDS dataset. (Y = yes, N = no).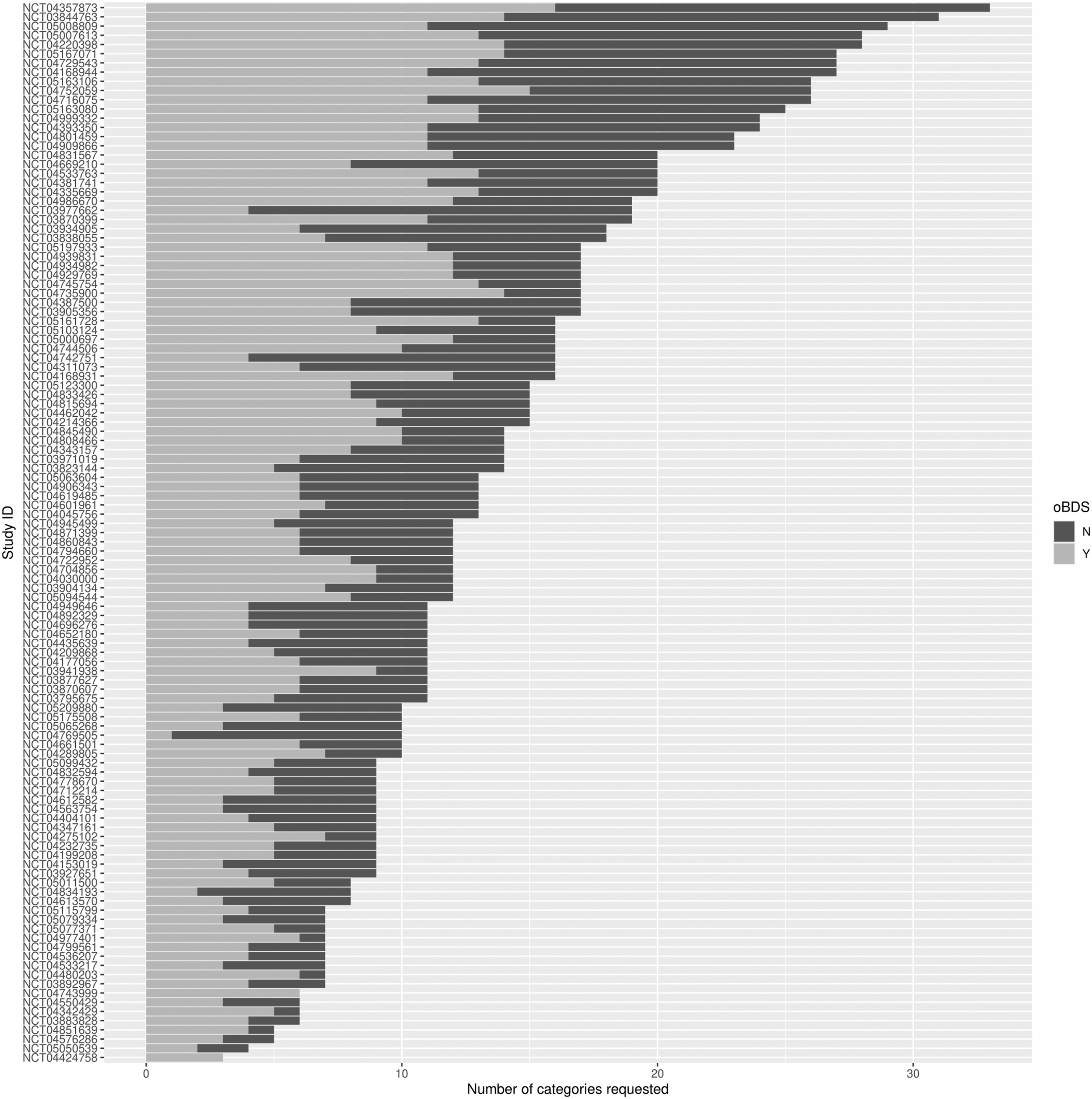
Discussion
At the outset of the study, three primary questions were posed: To what extent can the oBDS dataset comprehensively cover various studies? Which categories prove useful to expand the oBDS in context of automated trial recruitment? To what degree can tumor documentation data already be used to support automated trial recruitment?
Regarding the first question about a possible coverage by the oBDS dataset: It can be stated that barely any study can be fully covered by the oBDS, in fact only 2 (1.7%) studies can. Also only 24 of the 56 categories are supported by the oBDS dataset (42.9%). The listed names of the criteria in the study protocols displayed high heterogeneity but could be generalized into similar groups.
The most commonly asked for criteria, such as age or diagnosis, are indeed present in the oBDS dataset, however, other criteria such as pregnancy, informed consent, and comorbidities are not present in the oBDS dataset, despite frequent demand. Age was the only criteria that was needed in every analyzed study, thus it was requested 115 (100%) times. This is not an unexpected result, as the age information is already mandatory when submitting the study protocol to the ethics committee for the assessment of the clinical trial. 21 A question about possible comorbidities appeared in 90 (78.3%) study protocols. Despite its high frequency in the trial protocols, this field is yet not included in the oBDS dataset. This field provides a good example that regardless of a possible automated patient recruitment, it would be useful for research intentions to include comorbidities in the oBDS dataset. In fact, some cancer related comorbidities might potentially be available alongside some of the international tumor set equivalents like the British Cancer Outcomes and Services Data (COSD). In this case the National Disease Registration Service (NDRS) compiled comorbidity data by using Hospital Episode Statistics (HES) and other data sources but the usage of this data is not recommended.14,22 Similarly, comorbidities might also be abundant in other datasets like the German MI core dataset, documentation of this data is not enforced yet, hence, its completeness might be lacking and lead to new issues.
Incorporating laboratory values and pregnancy status into the oBDS dataset could also offer improvements. However, it’s crucial to note that these values may fluctuate between the time of documentation and the screening phase for an oncological trial, thus, their inclusion presents only potential benefits depending on the time of documentation.
The frequency with which consent was listed as an inclusion criterion (77/67.0%) was noteable, given that consent is inherently obligatory, particularly in interventional studies. Consequently, including this criterion in the oBDS, even if it appeared frequently, does not enhance the dataset’s content.
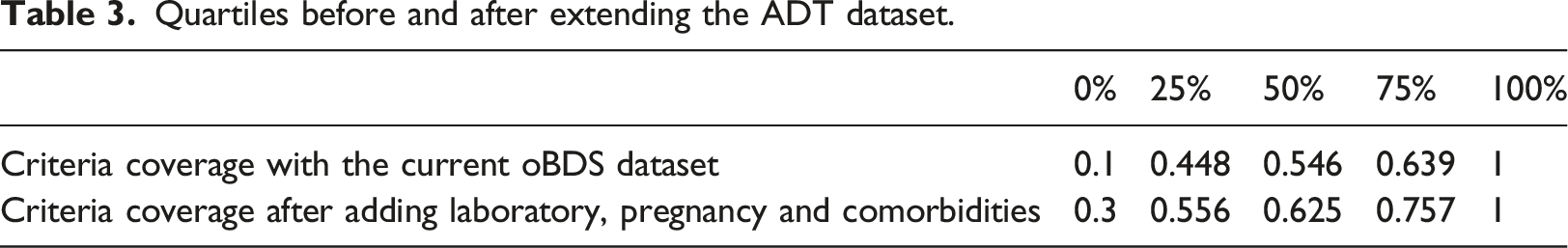
Quartiles before and after extending the ADT dataset.

Certain studies exhibited an unusually high number of inclusion and exclusion criteria, exemplified by the clinical trial with the ID NCT04357873, which necessitated the inclusion of 35 (out of 56) distinct criteria. There is a trend that shows that studies that require a larger number of criteria are proportionally less likely to be covered by the oBDS dataset. In most cases, there are more than 50% of categories that are not included in the oBDS dataset. Though, 100% compliance with the oBDS dataset does not seem desirable, because extending the oBDS dataset with all these fields would move it away from its original purpose. An extension consisting of the most frequently occurring fields would be desirable in the sense of a beneficial extension in terms of patient recruitment. Striving for a 100% inclusion of trial criteria within the oBDS may not be essential, especially when dealing with time-sensitive variables like laboratory values or pregnancy status, as previously mentioned. Instead, a preliminary selection of studies can serve as a decision support system for physicians, underscoring that it complements rather than supplants the doctor’s role by suggesting potential study candidates. In this context, prioritizing the most commonly used criteria can prove advantageous. This approach aligns with the findings of Gulden et al., which also acknowledge the potential for errors in the formulation of inclusion and exclusion criteria or in the documentation of this information. 23
In order to answer the question regarding the possible support of patient recruitment by tumor documentation, it is important to consider possible limitations. Overall, it should be noted that tumor documentation in Germany is usually delayed by multiple months and therefore data may not be available in time. 24 This might prompt the question of why even consider tumor documentation data for trial recruitment. However, due to being manually curated, a tumor dataset inherently comprises structured information of comparatively high data and content quality, setting it apart from typical Electronic Health Record (EHR) entries. As an example, diagnosis codes found in medical billing are often inaccurate. It is worth considering whether interfaces to subsystems such as a chemotherapy system could reduce the necessary time for documentation. Some tumor documentation systems already do this for at least some subsystems (e.g., CREDOS directly pulls surgery data). 25
A potential, forward-thinking approach to mitigate documentation time is the utilization of natural language processing (NLP) for the processing of medical reports. Given the substantial volume of data within unstructured free text in healthcare, employing machine-learning algorithms to cleanse and integrate this data into tumor documentation systems emerges as a possible solution. 26 In that regard, the commercial text mining system Averbis Health Discovery has a cooperation with the cancer registry of the federal state of Baden-Württemberg, the registry of Rhineland-Palatinate and the registry of Lower-Saxony. It aims to improve and speed up the data extraction from free text fields. 27 In general, natural language processing could also be a possible option to support digital trial recruitment by applying it on the study protocols and automatically parsing the inclusion and exclusion criteria towards a fixed catalogue.17–19
Perhaps other datasets (aside from the oBDS) could also be considered as the base for a similar project, such as the national core dataset of the Medical Informatics Initiative. 13 In this paper the oBDS dataset was used as an example for the previously mentioned reasons. Another option to focus on would, as an example, have been the Observational Medical Outcomes Partnership (OMOP) Common Data Model. 28 Though, while this model potentially covers all medical domains, it is, at least in Germany, not as widespread as compared to the oBDS dataset. It can be questioned whether there are oncological oriented datasets in other countries that could be considered for such projects.
As of the year 2023, the exclusive reliance on automated trial recruitment based on obligatory collected tumour data may prove unattainable until certain essential prerequisites are satisfied. These conditions, amongst others, include the necessity to expand existing datasets, for instance, by incorporating additional variables such as comorbidities, enhancing the quality of documented data in terms of completeness and accuracy, diminishing the temporal lag between the act of documentation and the occurrence of the associated events, such as laboratory values, and standardizing the criteria for inclusion and exclusion in alignment with an internationally or nationally accepted common framework, as established within study protocols and registries.
Conclusion
To assess how useful common oncology datasets are for automatically selecting patients for clinical trials, we examined 115 oncology trial protocols and compared them with the German oBDS dataset. Within the present oBDS dataset, on average, 54.6% of all investigated categories can be accommodated. However, it is worth noting that the majority of studies do not achieve full coverage. Moreover, the outcomes of this study underscore that the ability of the oBDS dataset to encompass an expanding array of inclusion and exclusion criteria in clinical studies is notably enhanced when certain categories, in particular comorbidities, are incorporated. Consequently, the current utility of the oBDS lies in its capacity to function as a tool for the preliminary selection of potential trial recruits. In this sense it might be viable as a minimum set for trial recruitment, aside from its lack of comorbidities. Nevertheless, it is important to emphasize that the complete automation of trial recruitment, as of the year 2023, remains an unattained objective.
Footnotes
Acknowledgements
We would like to express our gratitude to the staff of the Comprehensive Cancer Center Munich, supporting this project.
Author contributions
DN conceptualized the project, while MLM oversaw the data collection and preparation process. DN, SK, SM, and TF brought their senior expertise to the project, particularly in the realm of category generalization. AK played a crucial role in analytics and plot creation. All contributors collaborated on manuscript writing, with DN taking the lead in this aspect.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical statement
ORCID iDs
Data availability statement
Information about the selected trials can be requested by the corresponding author.
