Abstract
With the vast amount of medical data being generated on an electronic basis, to ensure that individual patient information is properly linked across multiple sources, efficient patient matching algorithms are critical. We describe a novel naturalistic approach that is a hybrid of both deterministic (exact) and probabilistic (“close”) systems that allows for manual adjudication of cases where necessary, but whose basis stems from the thought processes that a rational individual would follow in the comparison of medical records. A validation of this algorithm using large databases from two disparate sources demonstrates that the naturalistic approach can largely eliminate false positives (false matches) and false negatives (false mismatches).
Keywords
Introduction
Accurately identifying a patient allows health systems to construct a longitudinal view of each patient’s health history. This is essential to protecting patient safety yet very difficult with the multitude of electronic systems used to track and report medical data. This problem, known as “record linkage,” relates to the task of finding and linking records that should be associated to the same individual across different data sources. 1
In the past, record linkage was performed manually by a well-informed person who could adjudicate which records belonged to which patient. Typical matching processes were based on exact matching of demographics values such as first name, last name, date of birth (DOB), and US social security number (SSN). This type of exact matching method became to known as “deterministic” matching. In an effort to find a more standardized approach to record linkage, Newcombe and Kennedy 2 introduced the first method of matching based upon the weighted probability that two particular values would be the same. This idea was expanded upon by Felliegi and Sunter with mathematical models which enabled a more flexible approach to record linkage. This approach provides the ability to link based upon “close” matches of values that are similar enough based upon probability. This method of record linkage became the basis for today’s “probabilistic” matching methods.1,3
The widespread use of computer systems for capture and retention of demographics records has made the process of record linkage more complex and the large number of transactions involved requires decisions to be made automatically by high-speed computer systems. 4 In the United States, medical laboratories routinely receive and generate HL7-based order, result and admission/discharge/transfer (ADT) messages in their laboratory information systems (LISs), billing systems, and (in the case of hospital-based laboratories) patient registration and hospital information systems. These systems often use different technology platforms that have been configured to organize and store data in different ways, and the lab typically does not control the administration of all of these systems. As a result, there is usually no single reliable identifier (PatientID) that can be used to identify unique patients across all systems and all transaction types. 5 This problem is exacerbated when the laboratory is connected to customers and stakeholders outside of the boundaries of its enterprise, for example, physicians using electronic medical record (EMR) systems and other platforms to generate lab orders. (Due to the nature of the survey data used, the scope of this article is necessarily focused on a limited set of patient matching issues that arise in medical laboratories in the United States. However, strong anecdotal evidence suggests that the patient matching issues described are prevalent in other diagnostics modalities such as radiology/imaging, cardiology, and other specialized diagnostics procedures.) Given these challenges, some healthcare institutions fail to consistently match patient records accurately. 6
Purpose of patient matching algorithms
In general, medical laboratories in the United States receive and process requests to perform specific tests and deliver the associated results as isolated transactions. In order for medical laboratories to ensure patient safety and address a wide variety of business needs, it is necessary to use a comprehensive matching algorithm to automatically associate these separate transactions with the appropriate uniquely-identified individual patients. This allows the laboratory to perform their clinical responsibilities and facilitate workflow in a “patient-centric” rather than a “transaction-centric” framework, even in the absence of a universally defined patient identifier. To address this problem, medical laboratories implement systems to analyze the data contents of each transaction and then apply patient matching algorithms to that data in order to associate the transaction with a unique patient.
Data fluctuations as a hindrance to matching algorithms
A significant problem when applying patient matching algorithms is the quality of the data that is being transmitted. Medical transaction files often contain inconsistencies in the data due to transcription errors, misspellings, missing data, and changes in data. These variations in the data can hinder the accuracy of patient matching systems. Commonly seen issues include the following categories and examples:
Coverage—Individual data elements may or may not be completed on particular transactions.
Quality—Data may include misspellings and typographical errors.
Consistency—A patient’s first name may be represented at times by a nickname or by the use of a middle initial.
Mutability—A patient may change his or her last name.
Reliability—The owner of source data may change systems which can have unexpected effects such as the renumbering of locally assigned patient identifiers.
Encoding—Identical addresses may be represented differently.
Strength of a name—Match based on the commonness or rarity of the name.
Representation of filler data—A DOB of 01/01/1900 may be used to populate a required field when the DOB is unknown.
Application of matching algorithms
Within any patient matching system, as the matching algorithm assesses each transaction, it will use the potentially flawed data it receives to either associate the transaction with a previously identified patient or determine that the patient represented in the transaction is previously unknown to the system and is to be added to the patient record list. Additionally, the system that employs a patient matching algorithm may, or may not, include a user interface to allow a user to review and manually adjudicate near matches or near non-matches.
Traditional types of patient matching algorithms
Traditionally, systems have relied upon one of two types of algorithms to address patient matching needs; deterministic and probabilistic matching. The former uses one or several sets of fields to match exactly (or with a fixed number of leading characters) the transaction message and the system’s representation of the patient. 7 These algorithms are simple to understand and configure but typically do a poor job at accounting for the types of data fluctuations described above and usually do not include a user interface to manually adjudicate near matches and near non-matches.
Probabilistic patient matching uses the configurable assignment of weights to particular data fields along with a measure of the closeness of the match between any two instances in a given data field. This information is used to create an overall metric for the degree of matching that is compared to some predefined threshold. If the value on this metric is greater than or equal to this threshold, the match is made. 1 These algorithms account for some (but not necessarily all) of the types of data fluctuations previously mentioned. However, probabilistic algorithms have the disadvantage of being complex to configure and not always intuitive in their automated adjudication of particular transactions.
Deterministic matching is typically used by applications that receive data from external systems where the receiving and the sending systems have their own local unique patient identifiers that are not shared between the systems. Examples include a physician office-based EMR system receiving results from a hospital’s LIS. In these cases, when the EMR receives data from the LIS, it may link the incoming results data to existing patient records (i.e. identify the EMRs unique patient identifier) by exactly matching the received data’s physician, full patient name, DOB, sex, and zip code. Failing to find a match with those data elements, the EMR may try to match based on physician office code which identifies a particular physician’s office within the EMR system (aka ClientID), DOB, SSN, and last name. When no match is made, the system may assume that the new results’ data represent an unknown patient and flag the result data for manual review.
On the other hand, probabilistic matching is typically performed by specialized Electronic Master Patient Index (EMPI) systems that serve to broker–patient identity between multiple systems that do not share unique patient identifiers. So, in the example above, if the EMR and LIS both had access to an EMPI system with probabilistic matching, they could use that system to cross-reference the EMR and the LIS patient identifiers so that the patient identifier can be translated into the receiving system’s version when data are passed from one system to another. Alternately, the EMPI may generate a new unique patient identifier using data from both the EMR and the LIS. This “Master Index” then can be used by both the EMR and the LIS systems to reliably match received data to patient records.
ATLAS Naturalistic™ matching algorithm
This article explores the accuracy of a third type of matching algorithm. This algorithm employs a heuristic approach and fuzzy logic to account for the data fluctuations described above. This methodology inherently solves many data quality and inconsistency issues encountered in medical transaction files. Like deterministic algorithms, the naturalistic algorithm is relatively intuitive and easy to understand. Moreover, like probabilistic algorithms, the naturalistic algorithm accounts for the complexity caused by data fluctuations that otherwise decrease the accuracy of deterministic algorithms. This enhanced type of algorithm is termed “naturalistic” because it is designed to match transactions using the same steps that a careful, well-informed person would take.
Atlas Medical’s HealthCentric® EMPI (Atlas Development Corporation, Calabasas, CA) employs the ATLAS Naturalistic patient matching algorithm and also provides a user interface to flag near matches and near non-matches for manual review and adjudication. The methodology is described below. The following diagram outlines the naturalistic matching algorithm logic flow:
Methods
The naturalistic algorithm allows for matching rules to be configured based upon the combination of selected data elements. A data element is a string or number that represents a discrete piece of information within a transaction record that can inform the record linkage process (e.g. first name, last name, DOB, and phone number). When comparing the respective values of a given data element across two transactions, it can
Provide strong evidence that two transactions are associated with the same patient. For example, if both transactions contain the same patient phone number, it is strong evidence that the two represented patients live in the same household and may in fact be the same patient.
Provide strong evidence that the two transactions are not associated with the same patient. For example, if both transactions contain significantly different dates of birth, it is likely that the two represented patients are in fact different from each other. However, significantly different patient phone numbers add little information because patient phone numbers readily change.
Provide weak evidence that those two transactions are or not associated with the same patient. For example, if both transactions have the same last name, this information is evidence that the two patients are in fact the same. However, if the last names differ, it does not exclude the possibility.
As the number of data elements that match (or nearly match) between two transactions increases, the evidence that the two transactions are associated with the same patient also increases. The naturalistic algorithm is based on the selection of various combinations (rule sets) of these data elements. The rule set definitions also include logic that determines the matching of similar, but not identical values (aka Relaxation described in Table 2). Additionally, other override logic (to be described below) can be applied that restricts the applicability of particular data sets. If all the data elements in any one rule set match (or nearly match) and there are no restrictive overriding logic, then the patients represented in the two transactions are considered to be the same.
The matching validation was performed with a set of configurations used to determine if two transactions match to the same unique patient. Each row in Table 1 represents a set of rules used by the naturalistic algorithm to perform patient matching across different transactions within a given database. Each cell in each row represents a data element and/or a matching rule. Each row of Table 2 details the various values of the cells in Table 1. This approach is tuned to approximate the steps that a person deciding to match a transaction to a candidate patient record would perform. When determining whether a transaction belongs to a particular candidate patient record, the algorithm evaluates each individual rule within a given set of rules. If all of the individual rules in any set are true, then the algorithm matches the transaction to the candidate patient record. If no candidate patient is found to match, the algorithm assigns the transaction to a newly created patient record.
Summary of matching configuration rules.
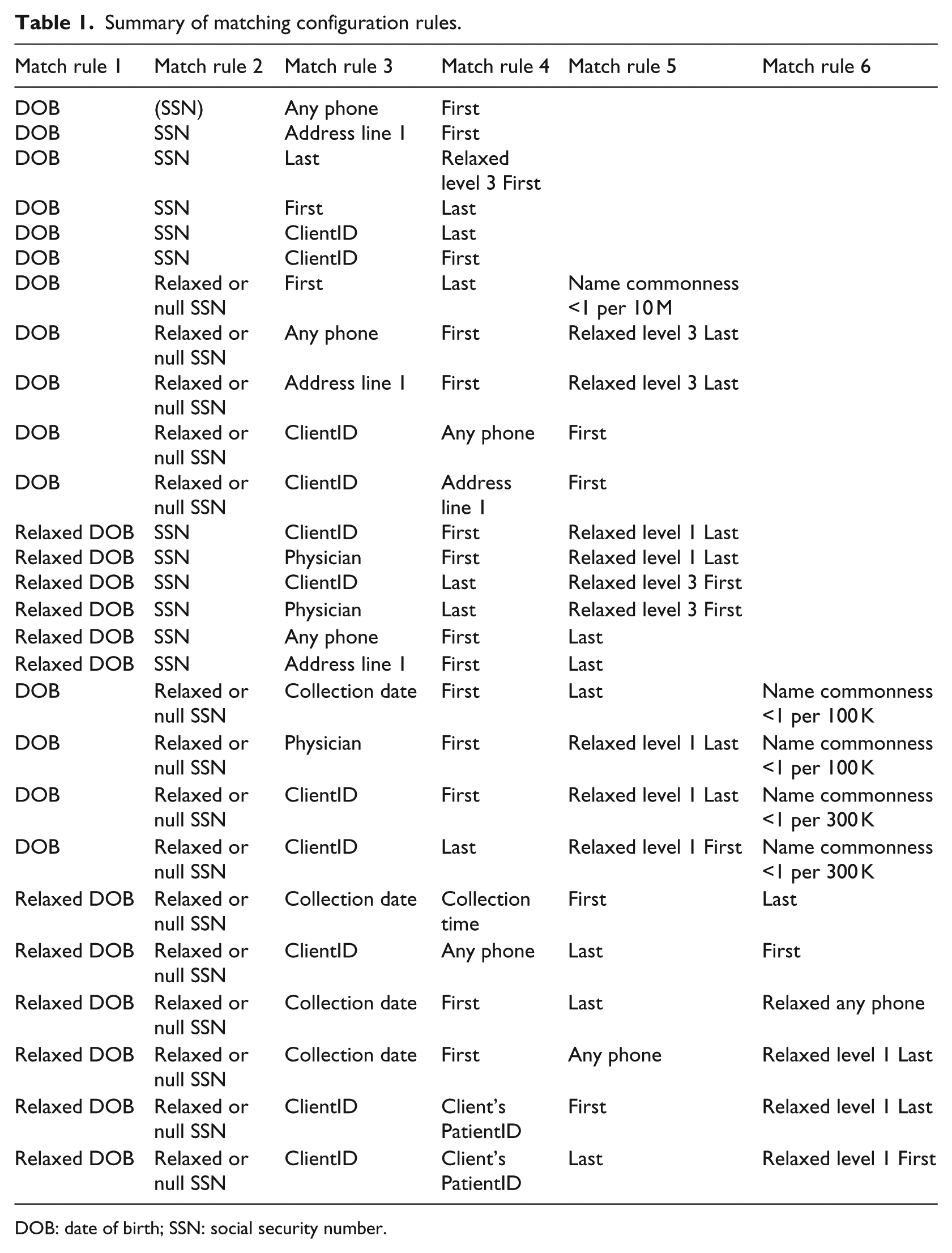
DOB: date of birth; SSN: social security number.
Summary of field fuzzy logic rules to account for matching with data content variability.

DOB: date of birth; SSN: social security number.
The authors determined the relaxation rule logic (described in Table 2), the manual review logic (described in Table 3), and the “tuned” matching rules (described in Table 1) prior to the start of this study. This determination was made through the experience of sequentially analyzing the matching performance of unrelated, data sets in isolation. These “training” data sets were obtained from four separate laboratories and these data do not overlap the data used in the study below. Additionally, the authors reviewed the findings with staff from each laboratory to better understand the data and tune the logic. These data set sizes ranged between approximately 200,000 and 10,000,000 transactions.
Summary of rules to flag near matches and near non-matches for manual review.
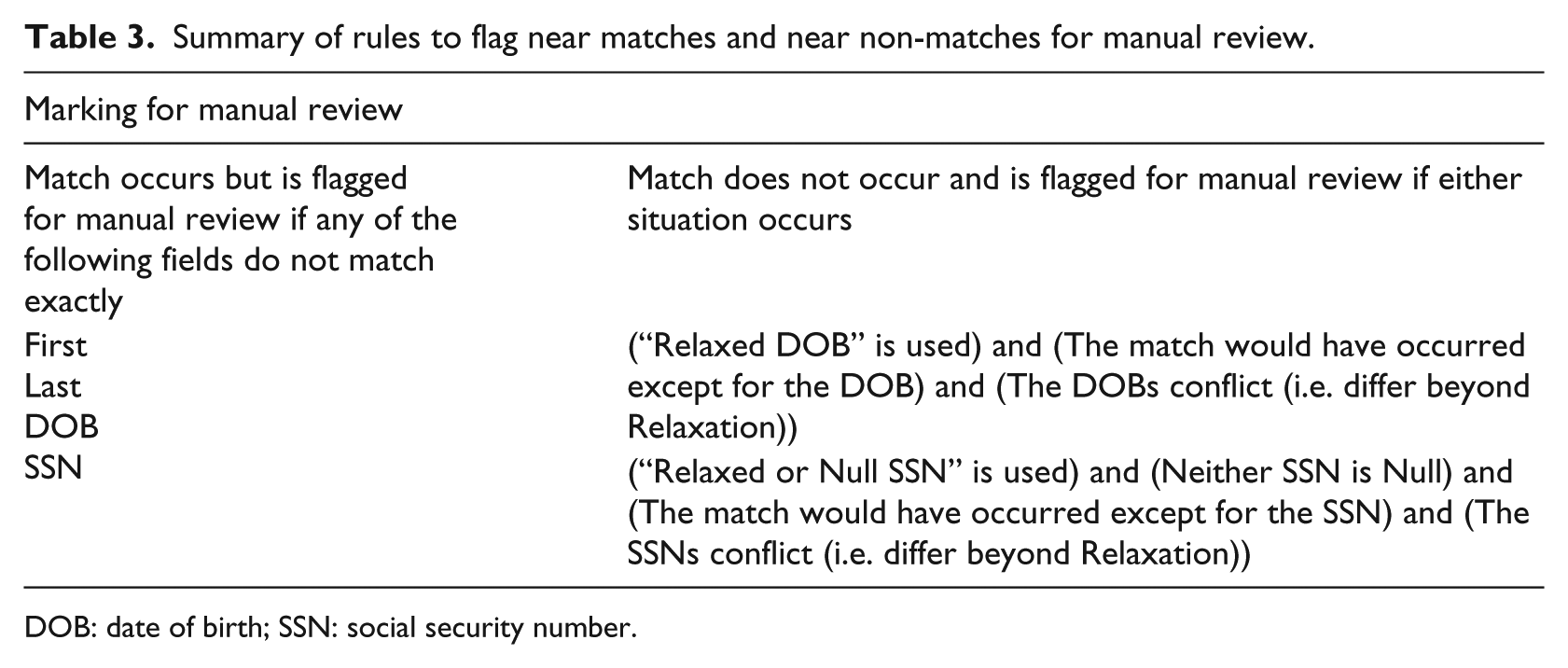

DOB: date of birth; SSN: social security number.
Although the authors performed the analysis using this particular set of rules, since the rule sets are configurable, this approach allows the matching algorithm to be tuned so as to approximate the record matching logic that any particular person or organization desires to implement. In the authors’ experience, this function is typically performed by the health system’s department that is responsible for patient data quality.
The algorithm allows for “relaxed” matching of fields in the candidate database. Table 2 provides a set of definitions based on fuzzy logic that clarifies what is meant by the term “relaxed.” This allows for the equating of two values or names that are close but not exact, which improves the quality of the matching outcome. This improvement is due to the ability of the fuzzy matching technique to address typographical errors and variations in the data. 8
Finally, Table 3 provides a set of rules for determining when a potential match or near non-match should be flagged for manual review. Flagged near matches may indicate receipt of an incorrect piece of data (e.g. sex error) or a legitimate data fluctuation such as a change of last name. The flagging of near non-matches is used to identify transactions that would be linked based on an agreement of data elements if not for a significant disagreement between values of an immutable data element (e.g. DOB or SSN). This situation can indicate either a data error in one of the non-matching immutable data elements or a more systemic error in the other matching data. In either case, the system flags this near non-match as an unexpected situation that should be investigated and corrected.
Data sets used for validation
The data used for the validation was made available by two medical laboratories that have implemented the naturalistic patient matching algorithm using the methods listed above. Olympic Medical Center (OMC) is a hospital located in Port Angeles, WA, with an active community physician outreach program. To service these outreach physicians, OMC provides an order entry and results delivery portal that allows for manual and electronic order entry, results review, results auto-printing, and physician EMR integration. Each HL7 result transaction that OMC forwards to this portal contains a patient identifier determined by OMC’s EPIC system’s registration module which is part of its Hospital Information System. This data set consists of 137,470 HL7 result transactions generated by OMC’s LIS between 18 July 2013 and 23 June 2014. These transactions included 84,458 separate accessions. OMC’s EPIC system’s registration module employs a traditional probabilistic matching to validate record linkage after the manual registration process has occurred.
Diagnostic Laboratory Services (DLS) is an independent laboratory located in the State of Hawaii, Guam, and Saipan that serves community physicians as well as the Queen’s Medical Center health system. As with OMC, DLS’s LIS forwards HL7 result transactions to an order entry and result delivery portal. These transactions include results destined for community physicians as well as results delivered to Queen’s Medical Center. Since DLS is an independent, rather than a hospital-based laboratory, the results generated by its LIS do not contain a patient identifier that is created through a hospital registration process. This data set consists of 1,134,406 HL7 result transactions generated by DLS’s LIS during a 3 month period that overlapped the OMC data timeframe. These transactions included 505,225 separate accessions.
General validation approach
Two separate validation steps were performed in order to test whether the naturalistic algorithm is susceptible to either
Results
Given the nature of the imperfect and incomplete information available (data fluctuations and inconstancies due to data entry errors, name changes, alternative name usage, etc.), any algorithm will produce either false negative and/or false positive errors. For clinical purposes, false positive errors are much more damaging and are avoided if at all possible. Therefore, implemented algorithms tend to make some false negative errors. To measure the naturalistic algorithm’s propensity to make false negative errors, the authors compared its results with results from an established probabilistic algorithm implementation for identical data sets. It was not practical for the authors to compare the naturalist algorithm with the exact “truth” by cross referencing other data sources such as phone records because of the author’s inability to violate patient privacy.
The naturalistic patient matching process was compared to OMC’s registration processes matching algorithm by first considering the following:
Identified the percentage of instances and the matching determinations were identical (agreement with a strong independent patient matching process).
Identified the percentage of instances, and reasons why the OMC registration process identified a single patient while the naturalistic algorithm identified multiple individuals (potential for false negative matches).
Identified the percentage of instances, and reasons why the naturalistic algorithm identified a single patient while the OMC registration process identified multiple individuals (potential for false positive matches).
Second, to continue the evaluation of these two approaches, any patients that were determined to reside in both the OMC and the DLS data sets were verified that they were the same patients (potential for false positive matches)
In order to perform this comparison, the OMC registration PatientID was removed from the OMC HL7 result transaction and then the transactions were processed through the naturalistic algorithm with matching rules defined below. This approach grouped transactions into sets that shared the same patient, identified near match transactions that were associated with a single patient, but had enough data differences resulting in a flag for manual review, and identified near non-match transactions that were not associated with a single patient, but had enough data in common resulting in a flag for manual review.
Table 4 summarizes the results of the naturalistic algorithm’s transaction grouping compared with that from the OMC registration process. The high degree of agreement between the two independent matching algorithms largely validates the veracity of both approaches. That said, by closely examining the remaining 0.35 percent of records for which there was a disagreement, we can gain more insight into the likelihood of false positive or false negative errors made by either algorithm.
Summary comparing the NA transaction and OMC registration process groupings.
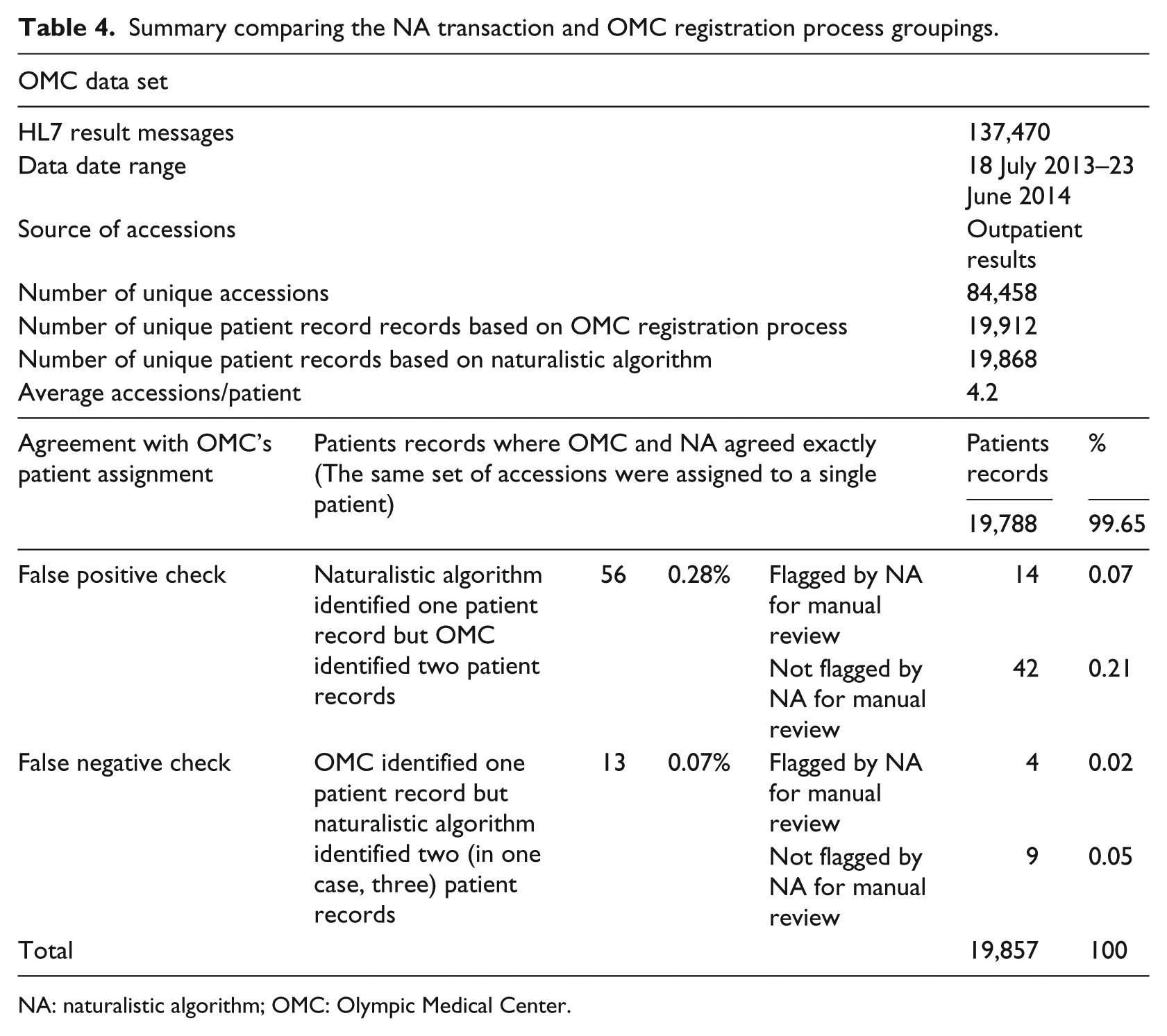
NA: naturalistic algorithm; OMC: Olympic Medical Center.
False positive check analysis
For the 56 cases where the naturalistic algorithm identified a single patient but OMC identified two separate individuals, 14 were flagged for manual review. Of those 14 sets of transactions, 9 had last names that did not match exactly, 4 had SSNs that did not match, and 1 had a first name that did not match exactly. Aside from these minor variations, the transactions had many other fields in common such as the remainder of the name, the phone number, the ClientID, the address, and so on. Upon detailed user examination, these transactions were undoubtedly determined to be from the same patient.
A total of 42 records were not flagged for manual review. Of these, 35 had no data differences that might disqualify a match and instead had numerous field values in common, and 7 had transactions where some of the SSNs were populated with either 999-99-9999 or 000-00-0000. This misleading information may have resulted in different patients being created by the OMC registration process. Again, since many other fields matched in these cases, the transactions undoubtedly belonged to the same patient. Based on this analysis, there is no evidence that the naturalistic algorithm created false positive matching errors. The details of these set variances are listed in Tables 5 and 6 in the “Detailed matching information” section.
Fourteen sets of OMC transactions that ATLAS determined to be same patient and flagged for review while EPIC determined the pairs to be different patients.
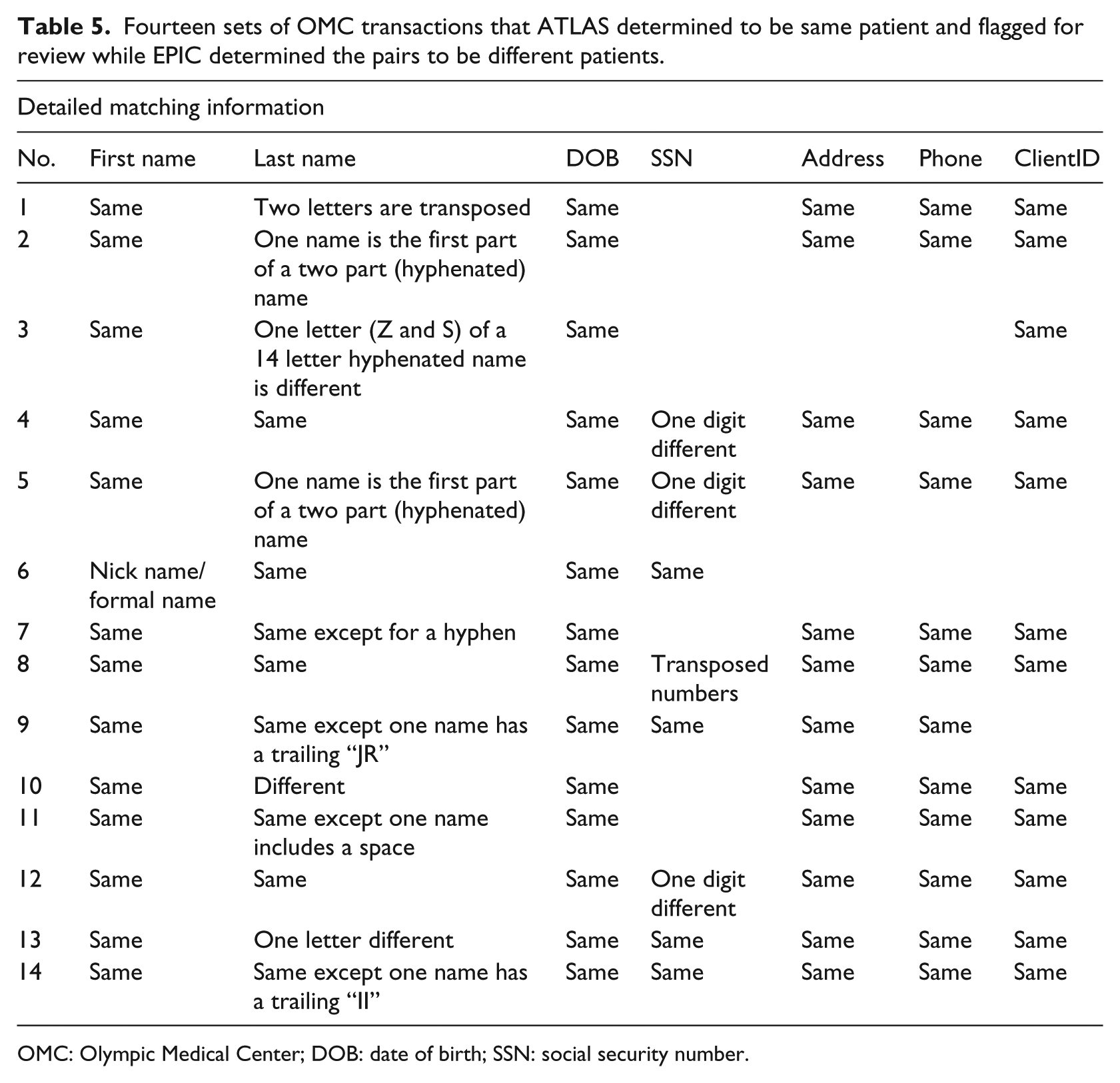
OMC: Olympic Medical Center; DOB: date of birth; SSN: social security number.
Forty-two sets of OMC transactions that ATLAS determined to be same patient and NOT flagged for review while EPIC determined the pairs to be different patients.
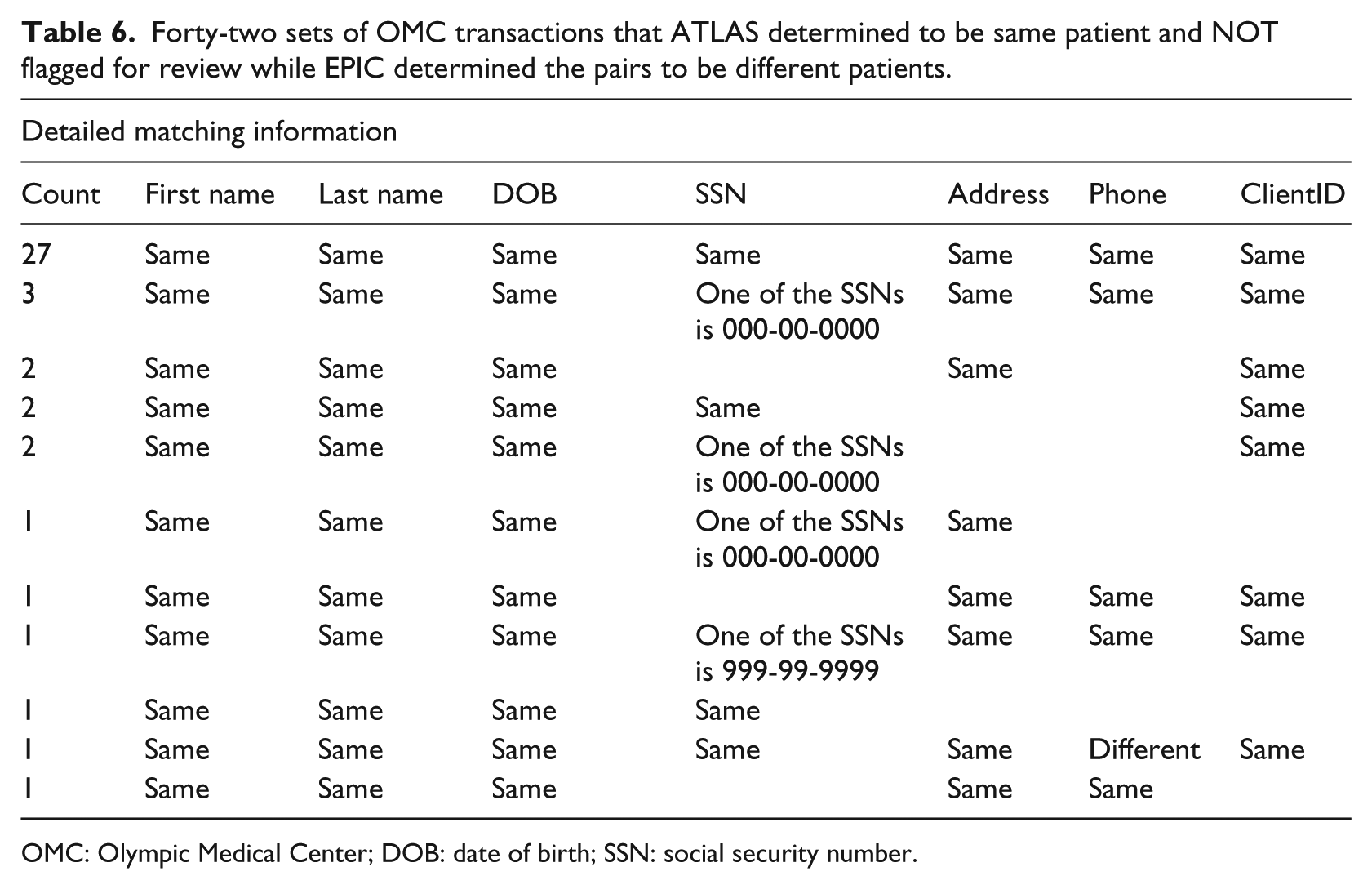
OMC: Olympic Medical Center; DOB: date of birth; SSN: social security number.
False negative check analysis
There were 13 transactions where OMC identified a single patient record, but the naturalistic algorithm identified two (in one case, three) patient records. Four were flagged for manual review. Two of those sets had transactions that contained completely different SSNs, one set contained transactions where the SSNs were different by two digits and one set had transactions where the “DOB” differed by two digits. Although these differences may or may not have led to four false negative errors, the reasons for caution with the non-matches are clear and highlighted appropriately for manual review.
Of the nine remaining cases, one set contained clearly different patients and so the naturalistic algorithm performed appropriately, one set contained both DOBs and SSNs that differed between transactions, one set contained transactions that had slightly different first names with no way to rule out that twins were represented on the separate transaction, three sets contained transactions where the first name was only an initial, which limited the sets of rules that were available to make a match, one set contained first names that were completely different, one set contained transactions that only had matching name and DOB with an uncommon name, and one set contained a first name and the last name where neither matched exactly. Although a case can be made that some of the above nine instances may represent false negative errors, the cost of the alternative scenario making false positive errors is much higher. Given this risk trade-off, the naturalistic algorithm appears to have performed this risk trade-off in a way consistent with a rational person closely examining the data.
Considering the DLS data, we note that OMC and DLS are geographically separated. When examining the patient populations of both data sets together during overlapping periods of time, it would be reasonable to expect that very few individual patients would appear in both data sets. If the naturalistic algorithm was susceptible to false positive matches, this would not be the case. Table 7 provides a summary of the matching results for the DLS and OMC databases.
Summarizes the matching results for the DLS data set along with the overlap with the OMC data set.
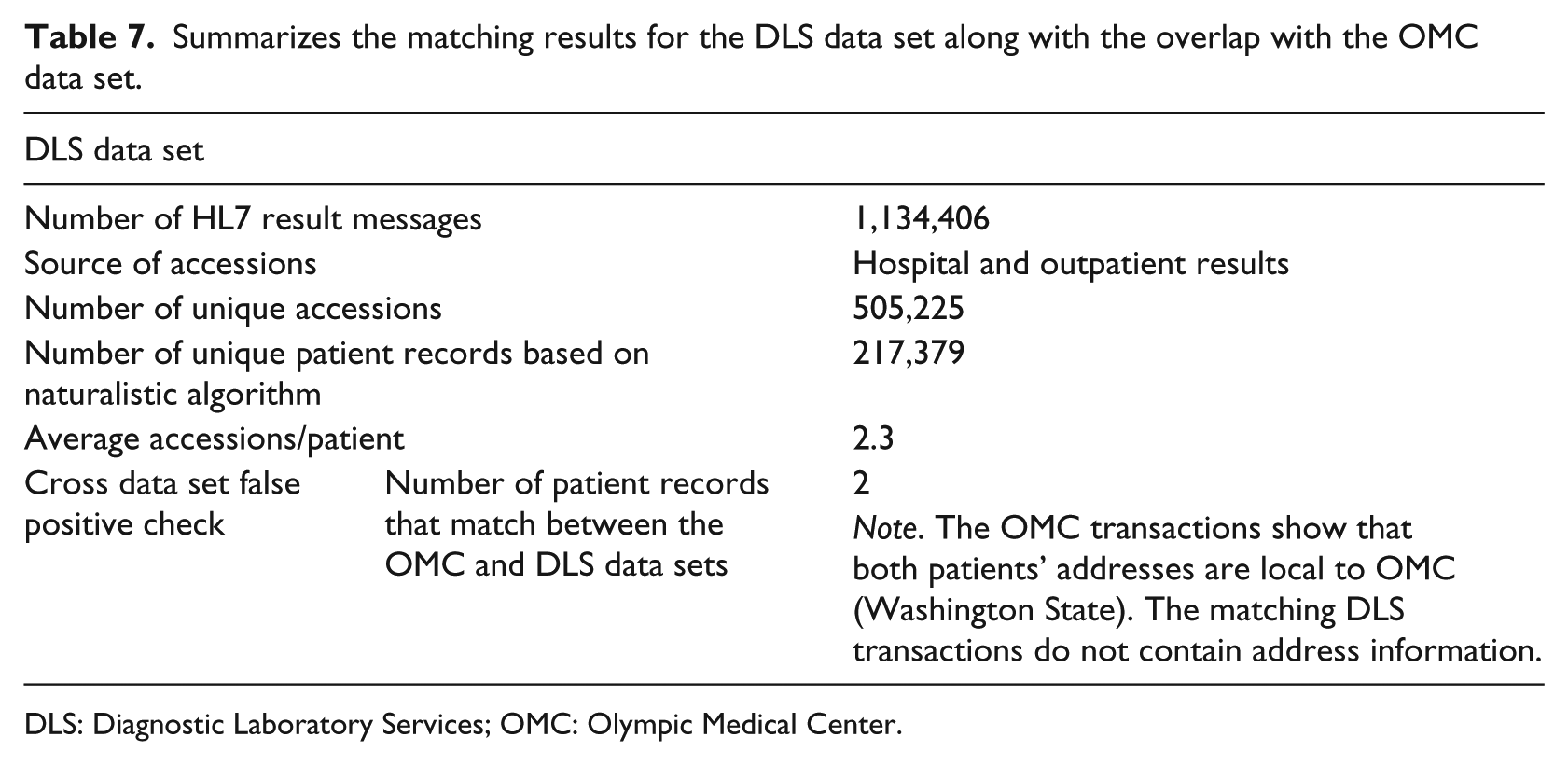
DLS: Diagnostic Laboratory Services; OMC: Olympic Medical Center.
Of the 19,788 patient records identified in the OMC data set, two were also identified in the DLS data set by the naturalistic algorithm. Both records were matched using only one rule set:

In both cases, the SSN was null in the DLS transactions and there was no other identifying information available besides first name, last name, and DOB. In both cases, the DLS transactions were associated with the Queen’s Medical Center (as opposed to outreach orders). However, by looking at the physician portal systems that DLS and OMC utilize, it was possible to find additional patient information that was not included in the HL7 result transactions. This information was used to either help confirm that the matching patient records are truly the same individual or examples of false positive matches.
In the first instance, examination of DLS’s order entry and result reporting portal system revealed that this patient’s DLS stored phone number matches the phone number on the OMC transactions. This data match, along with the matching name and DOB, provided conclusive evidence that the patient represented by DLS transactions and the patient represented by OMC transactions were in fact the same person.
For the second subject, the evaluation of DLS’s order entry and result reporting portal system revealed that this patient’s DLS stored middle initial matches the middle name in OMC order entry and results review portal. This data match, along with the matching name and DOB, was evidence that the patient represented by DLS transactions and the patient represented by OMC transactions are in fact the same person.
Thus, the two pairs of patients found in both the OMC and the DLS data sets, almost certainly represented single instances of individuals, accurately matched across the data sets. This provided compelling evidence that the naturalistic algorithm is not susceptible to making false positive matching errors.
Discussion
Properly matching patient records in an environment in which information overload is a reality provides an extreme challenge to healthcare data systems. The stakes are high in terms of patient safety and the incipient costs associated with potential errors. Because of the size of the typical patient database (both in terms of number of individuals and their related numerous data sources) for which the matching is required, manual or simple search algorithms are not feasible in a real-time environment even with high-speed computing hardware and software. Methods for overcoming these problems have included deterministic and probabilistic approaches. The former takes a simple approach based on the premise that a certain number of fields need to match in order to declare that two records come from the same individual. This, of course, does not allow for slight variations in key fields, for example, names, DOB, or SSNs, that naturally occur in the process of entering large volumes of data. On the other hand, a probabilistic algorithm incorporates these data fluctuations into a decision-making process based on the probability of a match rather than an exact agreement in the deterministic algorithm. However, these methods are necessarily more complex, can lead to multiple decision-making processes, and, as a result, are not easily understood. It is important to note that in neither approach is there an allowance for manual adjudication of cases that might require it.
The authors chose not to synthetically generate data sets to simulate real world data. To do this in a meaningful way would require a profoundly detailed understanding of the real world variation for each particular data element based on patients’ region, ethnicity, sex, and so on. For example, the degree to which data elements fluctuate varies by patient age. 9 Additionally, data content will vary based on the type of source system from which the data originate. For example, medical laboratories which enter orders from handwritten requisitions will likely have less accurate and less rich data than laboratories which accept orders through electronic interfaces. Gaining such a detailed understanding of these variations and then programmatically generating data based on that understanding was deemed unnecessary, given the author’s access to actual data sets.
The naturalistic algorithm combines aspects of both these processes. Exact matches are, of course, one outcome of this algorithm, but close matches can also be considered (the “relaxed” portion of the methodology). However, when there is uncertainty in the decision concerning specific records, the approach can trigger a manual adjudication of these data to arrive at a final conclusion regarding the match. Fortunately, the number of these manual reviews is minimized by the automated algorithm.
Deterministic, probabilistic, and naturalist algorithms are all actually “deterministic” in the sense that given particular data inputs, the behavior is predetermined and non-random. Probabilistic algorithms are “probabilistic” only in the sense that they can match data elements across transactions that are literally different (because of fluctuation) but have a “probability” of representing the same patient because they are similar or related. The naturalistic algorithm likewise performs this matching despite data fluctuation.
The goal of the current evaluation and validation of the naturalistic algorithm was to demonstrate a virtual absence of false positives (false matches) and false negatives (false mismatches) using the naturalistic algorithm. With the available representative data sets, and using the HealthCentric EMPI system configuration described above as a comparator, the analysis was unable to show any evidence that the naturalistic algorithm leads to either false positive errors or undesired false negative errors. The matching agreement of 99.65 percent with OMC’s EPIC-based registration process is strong evidence that this approach yields results almost identical to other sophisticated and validated algorithms. In fact, examination of the cases representing the 0.35 percent disagreement between the two systems supports the assertion that the naturalistic algorithm can lead to more accurate matches. Furthermore, evaluation of large databases is underway to buttress this finding.
The original attempt to formalize an algorithm for record linkage was provided by Newcombe and Kennedy. Their work utilized a weighted probability approach in order to determine whether two records matched based on a threshold likelihood of agreement. This idea was extended by Felliegi and Sunter, who provided a more rigorous mathematical approach that put probabilistic matching on a sound analytical footing. They developed a probabilistic decision rule that was shown to be optimal when the comparison attributes are conditional independent. Many of the modern probabilistic linkage methods are based on the Felliegi–Sunter technique. Of course, these methods are only as good as the data provided and may not be particularly accurate given the data fluctuations that were described earlier. Moreover, there is no allowance for manual adjudication.2,3
More recently, Victor and Mera 1 reported on a method that combines data standardization (to address some of the data fluctuation issues), weight estimation (which provides a basis for evaluating the discriminating power of a given variable), and a matching approach that combines both deterministic and probabilistic methods. In a sense, this methodology is conceptually closer to the naturalistic algorithm described in this article. However, it may not be able to overcome all of the potential data fluctuations without the use of manual adjudication that is part of the naturalistic process. Moreover, the inclusion of a multistep algorithmically complex method can provide difficulties in exposition and may not be as computational efficient as the authors indicate. Thus, the naturalistic algorithm has the multiple advantages of simplicity, utilization of both probabilistic and deterministic approaches, and the ability to handle data fluctuations by including careful selection of cases that require manual adjudication.
Table 1 summarizes the rules used to match patients across transactions. Each row in the table consists of a set of rules. If all the rules in any one row are true, then the patients are considered to be the same and they are “matched.” Each of the separate rules is defined as follows:
DOB—exact match of the patients’ DOB excluding trivial values such as “01/01/1900” which are typically used to populate a required date field when the DOB is unknown. Invalid DOBs are not considered.
SSN—exact match of the patients’ SSN excluding trivial values such as “999-99-9999” which are typically used to populate a required field when the SSN is unknown. Invalid SSNs are not considered.
Any phone—exact match of any of the patients’ phone numbers. There may be more than one phone number stored in a given transaction. Invalid phone numbers are not considered.
First—exact match (exclude letter case) of the patients’ first names. Null or one letter names are not considered.
Address line 1—exact match (exclude letter case) of the first line of an address (e.g. “123 Oak Street”). Null or one letter addresses are not considered.
Last—exact match (exclude letter case) of the patients’ last names. Null or one letter names are not considered.
Relaxed 3 First—a relaxed, level 3, match (exclude letter case) of the patients’ first names (see Table 2 for details). Null or one letter names are not considered.
ClientID—exact match of the identification of the laboratory client. This identifier typically represents a physician office.
Relaxed or null SSN—(see Table 2 for details).
Name commonness <1 per “N.” The algorithm calculated commonness of the first name/last name combination is less common than 1 in “N” where N is listed (e.g. 10 M = 10,000,000).
Relaxed 3 Last—a relaxed, level 3, match (exclude letter case) of the patients’ last names (see Table 2 for details). Null or one letter names are not considered.
Relaxed 1 Last—a relaxed, level 3, match (exclude letter case) on the patients’ last names (see Table 2 for details).
Relaxed DOB—(see Table 2 for details). Invalid or trivial DOBs are not considered.
Physician—exact match (exclude letter match) of the physician name.
Collection date—exact match of the transactions’ collection date excluding trivial values such as “01/01/1900” which are typically used to populate a required date field when the collection date is unknown. Invalid collection dates are not considered. This represents two transactions being ordered the same day.
Client’s patient ID—match of the client’s patient identifier as defined for transactions containing the same ClientID (i.e. the same physician’s office).
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was entirely funded by the Atlas Development Corporation.
