Abstract
General practice records present a comprehensive source of data that could form a variety of anonymised or pseudonymised research databases to aid identification of potential research participants regardless of location. A proof-of-concept study was undertaken to extract data from general practice systems in 15 practices across the region to form pseudo and anonymised research data sets. Two feasibility studies and a disease surveillance study compared numbers of potential study participants and accuracy of disease prevalence, respectively. There was a marked reduction in screening time and increase in numbers of potential study participants identified with the research repository compared with conventional methods. Accurate disease prevalence was established and enhanced with the addition of selective text mining. This study confirms the potential for development of national anonymised research database from general practice records in addition to improving data collection for local or national audits and epidemiological projects.
Keywords
Introduction
While health researchers in Scotland may have access to disease-specific databases, identifying and recruiting participants to research studies remains difficult, 1 especially for trials with more complex inclusion and exclusion criteria. As a result, failure to return feasibility information for research in a timely fashion coupled with research projects experiencing difficulties with recruitment leads to delay and reduced access to academic and commercial research trials. Epidemiological or observational studies regarding population health are also disadvantaged.
General practice (GP) records present a rich and comprehensive source of regularly updated data that could aid efficient identification and equitable recruitment of participants for observational and recruitment studies regardless of location, thus forming the basis of well-populated research databases. However, access to information requires physical data extraction by staff within the practice setting, and there is no system to link all GP record systems. All GP practices in Scotland use either Egton Medical Information Systems (EMIS) PCS or the InPS Vision 3 as their clinical system for recording and storage of their patient medical records. Currently, the EMIS- and InPS-centralised data solutions cannot stream data held in the other suppliers’ GP practice system, and reports cannot be run against a single, complete set of GP patient records. 2
A centrally managed General Practice Extraction Service (GPES) 3 has been set up in England with both the EMIS and InPS GP systems having been developed to work within this data extraction service. This system was developed for movement of data for the purpose of sharing rather than data mining. GPES now contributes to care.data 4 mandated by the Health and Social Care Act in England. In Wales, the Audit+ 5 system has been in use for a number of years with recent commissioning of a primary care data extraction system to contribute to the Secure Anonymised Information Linkage (SAIL) 6 system, and for practices that use Vision systems, a similar process to Quality Outcome Framework (QOF) is in place to send anonymised patient data to the General Practice Research Database/Clinical Practice Research Datalink (GPRD/CPRD) 7 and The Health Improvement Network (THIN). 8
The UK Government has recognised these emerging markets and is promoting the CPRD as the national research repository for life science and pharmaceutical research. This is nationally funded by National Health Service (NHS) National Institute for Health Research (NIHR) and Medicines and Healthcare products Regulatory Agency (MHRA) and operates in England.
A research alliance has evolved comprising three partners: academic, clinical (NHS) and industry to facilitate a process for transforming data from various GP databases to information for the benefit of recruitment to research studies. This is founded on a primary care data pump, able to extract a complete range of READ codes developed by a local Small/Medium Enterprise company specialising in primary care informatics and data extraction.
Aim
To form a pseudo/anonymised research database (ARD) based on a subset of practices within a health board area and assess the following:
Identification of eligible participants for research study;
Speed of research study feasibility testing;
Accuracy of data extraction from records.
Methods
Practice recruitment
The data pump software has been established within practices for several years under an extraction contract with the local health board. The contract endorses due regard to patient confidentiality and adheres to the board information governance guidelines. Specific permission to extract data for the purposes of the proof-of-concept study by the ‘trusted third party’ (TTP), working to International Organisation for Standardisation (ISO 9000) quality standards, was provided by the Local Medical Sub-Committee, and individual written consent was obtained from participating practices.
An audit trail for data extraction was established with a separate record of data used. Access to anonymised data extracts was provided through an analytical platform and restricted to two parties (S.M. and J.F.) for the purpose of data cube formation and descriptive analysis by password-controlled remote desk top. All data sets were deleted on completion of the data analysis.
Information about the project was provided to practices prior to participation, and patient leaflets were made available in GP surgeries outlining the process for those participating practices. Data for 57,680 patients were available from 15 practices spread throughout the region.
Data extraction
This was carried out in a three-staged process (Figure 1):
Practice information was filtered prior to extraction to exclude workforce or performance information and then assembled into a standard format for transporting to the repository.
After the initial individual extractions, each data set was loaded into a pre-processing repository (safe haven) where all identifiable information relating to practice, clinician or patient was removed. Patient unique identifiers (Community Health Index) were hived off into a separate secure store and the original replaced with a single generated repository number to maintain record linkage.
The final stage involved transfer of the anonymised records into a single anonymised research repository (ARD). This was the outward facing research database with no access or linkage to any of the previous stages.
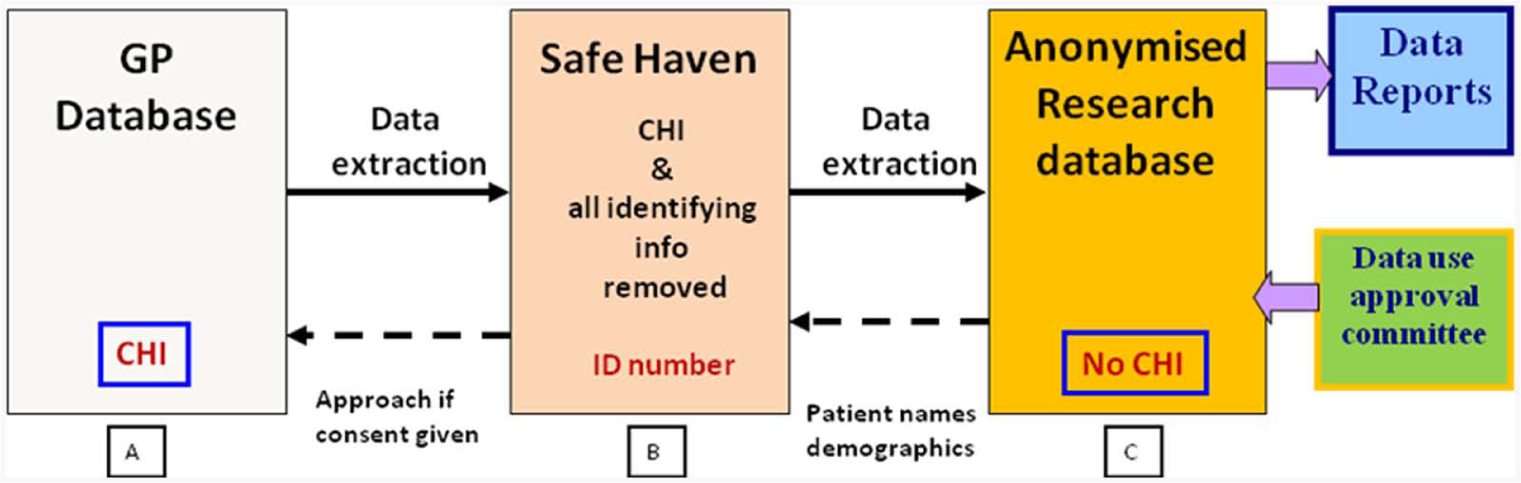
Pseudo/anonymised research database, data extraction process and potential link for re-identification.
Proof-of-concept studies
Identification of eligible participants for a research study and speed of research study feasibility testing
We used the example of a recently conducted feasibility study searching for patients with complex multi-morbidity to compare numbers of individuals identified by inclusion/exclusion criteria from the ARD retrospectively to those classified from the original feasibility assessment. A data cube based on READ codes for the relevant disease area was prepared. Time to achieve final numbers was also estimated.
A second retrospective comparison with ARD was carried out to evaluate numbers identified and time taken to achieve this using a study that recruited from primary care. The original approach was facilitated by a research coordinator visiting practices to specifically search individual practice databases for eligible participants.
Testing accuracy of data extraction from records
We used a simple assessment of disease surveillance to compare period prevalence figures extrapolated from ARD with accepted prevalence figures based on a serological test for a condition diagnosed mainly in primary care (Lyme disease).
Results
Over a 2-month period of time, data were extracted manually from a sample of 15 practices covering the breadth of the largest health board area in Scotland with data items relating to 57,680 patients.
The results of the three different types of study chosen to test the proof of concept are shown below.
Feasibility study
Using a dedicated research database of 1400 individuals to identify potential patients for a diabetes and cardiovascular disease outcome study took in order of 100 h of research nurse time spread over 2 weeks. This comprised creation and submission of data extraction queries; initial review of patient lists returned; cross-reference with hospital results system and finally manual review of case notes. This exercise resulted in 58 subjects being identified as potentially suitable for this particular study. With relevant READ codes matched to the same inclusion and exclusion criteria, search of the PRD database of 57,680 individuals from 2004 onwards resulted in the need for 5 h of programmer time which included 0.5 h to create the READ code–based repository (Figure 2) and 4.5 h to identify the cohort data set; 368 potential participants were identified through this method.
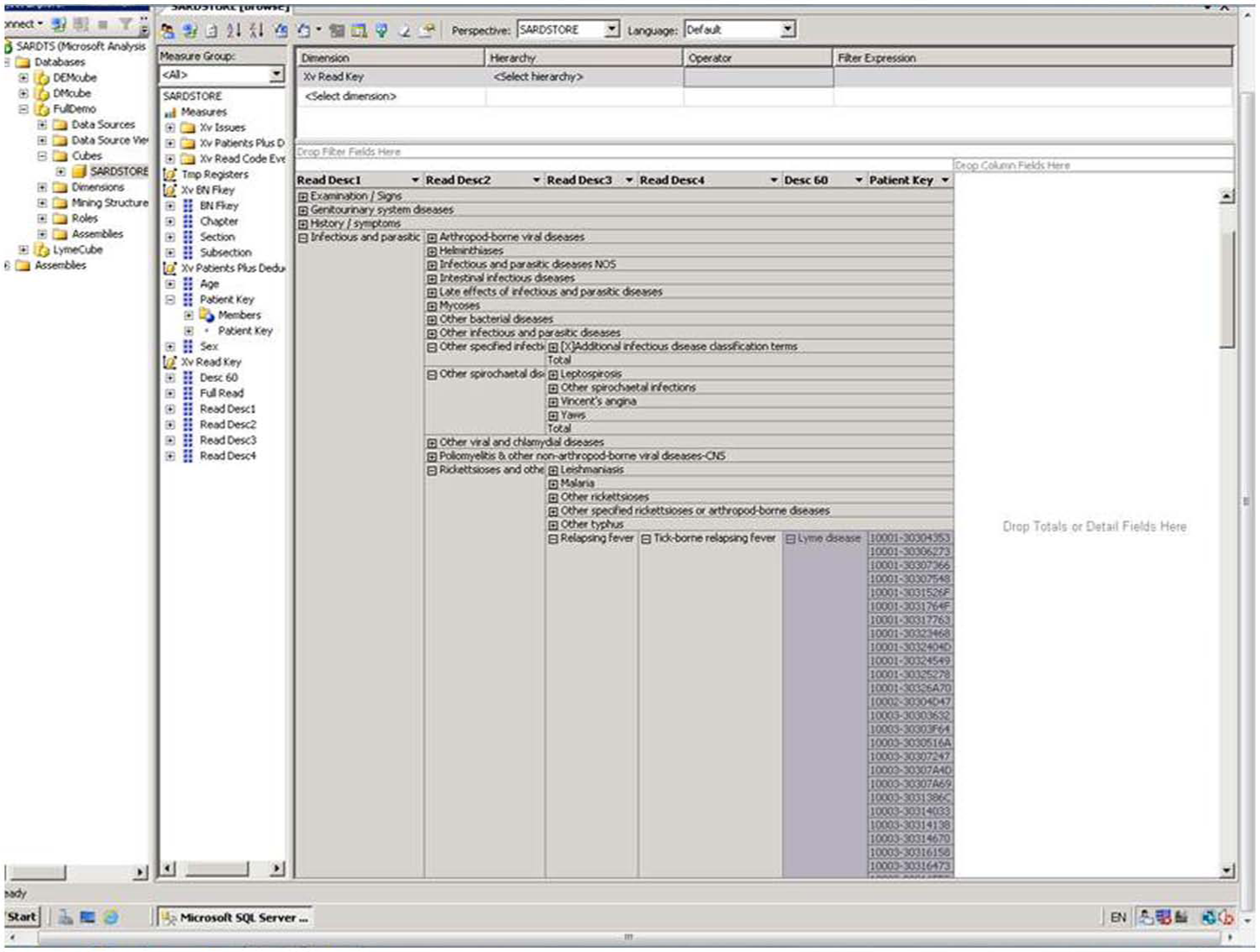
Example of SQL data cube formed from research repository.
Direct primary care recruitment study
Recruiting for a primary care paediatric study involved a coordinator visiting 18 practices during 2011 and building a data set for each practice to enable the search of records for children between the ages of 6 and 16 years. This resulted in 847 patients being identified as potentially suitable for study participation. Time taken was in the order of 5 months. Using the 15 practices and PRD research repository, 607 patients matching the same age span were identified within 0.5 h.
Disease surveillance
The estimated prevalence of Lyme disease in Highland region is 39.2–49.7/100,000, 9 a figure which is based on positive serology testing. Extrapolation using the ARD data set of all records with no age restriction over a 12-month period in 2011, matching the time frame of the reference paper, provided an estimated period prevalence figure of 46.8/100,000 which accords well with the anticipated prevalence. We also used specific defined text to search in the clinical notes in the ‘freetext’ field of GP records to capture additional cases where serology was not undertaken. When added to the original search criteria the terms erythema chronicum migrans, tick bite + rash or tick bite + antibiotic prescription and Lyme disease without ‘not’ in the text, the figure increased substantially to 62.4/100,000.
Discussion
It is imperative that in the interests of data protection, General Medical Council (GMC) guidelines are adhered to GMC 10 with regard to data collection schemes and that once collected, de-identified data are retained securely within a ‘safe haven’ repository and only retained for a period of time defined by the study requirements.11,12 These vital steps have been successfully achieved under the guidance of a research alliance. This has involved intensive discussions with the GP community as custodians of individuals’ medical records.
We have established a pseudo/ARD using safe-haven technology with pseudonymisation of the data extraction from 15 primary care data sets dispersed throughout a wide geographical area. Our proof-of-concept study has demonstrated that data sets can be prepared for a variety of research questions across several specialty areas. Time for preparation and searching is considerably shorter than when using conventional methods by factors of up to 20 times faster even with a disease-specific research database. When compared to a more time-consuming method of searching for research subjects that involves physically visiting a GP surgery and extracting data from individual practice data sets, the difference in time required for searching and identification of eligible subjects utilising the research database is considerable. The two main factors contributing to the time difference are removal of the need to set up a mutually convenient time to visit the practice and the requirement to build a search data set for each practice on site.
One of the key benefits of this type of multi-condition repository is the ability to search for individuals with several co-morbidities, thus providing a greater return on searches, for example the sixfold increase in the return of eligible subjects when searching for patients with multiple medical problems such as type 2 diabetes and cardiovascular complications. In our feasibility study, we used a disease-specific research register for comparison with ARD, and while the former produced a relatively higher percentage of potential recruits per population surveyed using the same search criteria, such registers are rare and absolute numbers, as highlighted, are likely to be lower. One advantage of a research register is the explicit expression of interest in research participation and when used in tandem with an ARD would constitute a powerful tool for enhanced study recruitment.
While normal practice has been to not extract any freetext to the repository, for data quality and information governance reasons, there is advantage in adding specific defined text to an extraction using READ codes. This aims to identify cases where the data have been recorded only in the freetext but had failed to also add a READ code. 13 In this study, the data pump searched for the pre-defined search terms only on text extracted from clinical comments with no patient identifiable information and with outputs limited to positive instances alone without displaying other freetext.
A key aim and component of the repository will be to design a system that will be accessible for research, through a local safe haven, by skilled academics, health-care developers and the clinical community without the need for advanced information technology (IT) or clinical data mining skills. The repository will allow them to interpret and present the information in a meaningful way. Access to repository data could be controlled via a GP consortium model and adherence to national data governance principles. 14
The research governance framework prevents researchers from making direct contact with potentially eligible subjects; 15 however, this work complements a parallel project that has piloted and continues to develop a comprehensive Scottish Health Research Register (SHARE). 16 These systems when combined could facilitate a national or, indeed, international unified system by linking the SHARE register of volunteers to their GP records to allow a match with relevant studies as well as facilitating local or national audit and epidemiological projects. In addition, data sets may be linked through the electronic Data Research and Innovation Service (eDRIS) and the Scottish Health Informatics Programme (SHIP) model to other national health and social care data sets.
The model is now complete and ready to be implemented as a health board wide demonstrator project to test the protocols, procedures and technology established in the pilot studies. Scaling up the extraction process to health board and thereafter national level would enable rapid turnaround of study feasibility data, significant reductions in recruitment times and potentially increased overall recruitment numbers. Furthermore, representative epidemiological data could be collected efficiently and accurately from a national perspective.
Conclusion
We have demonstrated through a proof-of-concept study that it is possible to establish a secure pseudo/anonymised repository of primary care data to form a research repository which yields fit-for-purpose extraction of data sets which profile individuals over multiple disease areas. Moreover, we have shown that highly significant reductions in time to obtain relevant data are possible using the database; these would translate into important cost savings in terms of research personnel time, enhanced recruitment to studies and support for public health initiatives. Expansion of the text mining facility adds a further important dimension to improving data extraction.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by funding provided by Highlands and Islands Enterprise. Data Pump and extraction provided by Albasoft Ltd, Scotland.
