Abstract
Background:
Over 1.5–2 million tuberculosis deaths occur annually. Medical professionals are faced with a lot of challenges in delivering good health-care with unassisted automation in hospitals where there are several patients who need the doctor’s attention.
Objective:
To automate the pre-laboratory screening process against tuberculosis infection to aid diagnosis and make it fast and accessible to the public via the Internet. The expert system we have built is designed to also take care of people who do not have access to medical experts, but would want to check their medical status.
Methods:
A rule-based approach has been used, and unified modeling language and the client–server architecture technique were applied to model the system and to develop it as a web-based expert system for tuberculosis diagnosis. Algorithmic rules in the Tuberculosis–Diagnosis Expert System necessitate decision coverage where tuberculosis is either suspected or not suspected. The architecture consists of a rule base, knowledge base, and patient database. These units interact with the inference engine, which receives patient’ data through the Internet via a user interface.
Results:
We present the architecture of the Tuberculosis–Diagnosis Expert System and its implementation. We evaluated it for usability to determine the level of effectiveness, efficiency and user satisfaction. The result of the usability evaluation reveals that the system has a usability of 4.08 out of a scale of 5. This is an indication of a more-than-average system performance.
Conclusion:
Several existing expert systems have been developed for the purpose of supporting different medical diagnoses, but none is designed to translate tuberculosis patients’ symptomatic data for online pre-laboratory screening. Our Tuberculosis–Diagnosis Expert System is an effective solution for the implementation of the needed web-based expert system diagnosis.
Introduction
Mycobacterium is the microbe that is the causative agent of tuberculosis (TB) and infects humans and other animals. This pathogen consists of seven species that cause the disease, and they include the following: Mycobacterium tuberculosis, Mycobacterium bovis, Mycobacterium canetti, Mycobacterium africanum, Mycobacterium pinnipedi, Mycobacterium microti, and Mycobacterium caprae. 1 We define TB as an infectious disease of both animals and humans caused by tubercle bacillus bacteria, which is contracted by airborne or by droplet contact with exudates, and symptomatic with characteristic cough and bloody sputum. It is believed that TB affects over 2 billion people or one-third of the world’s population with an estimate of 1.5–2 million deaths annually. 2
We present an expert system designed to automate the pre-laboratory screening process against tuberculosis infection to aid diagnosis and make it fast and accessible to the public via the Internet.
Trends in automated medical diagnosis
An important breakthrough in the field of medical expert systems was realized in 1976 when Shortliffe published MYCIN used to assist physicians to administer blood infection treatment from diagnosis of infectious blood diseases to subsequent recommendation of antibiotics, with the dosage prescription based on the patient’s body weight. 3 Other software produced during this early period included Present Illness Program (PIP)—a system for renal disease by Pauker et al. 4 —and in 1978, Close Air Support Network (CASNET), an ophthalmology advisor that was implemented to recommend management for patients with glaucoma 5 as shown in Table 1. ONCOCIN was developed in the early 1980s with the aim of assisting physicians in the management and treatment of cancer patients. 6 INTERNIST-1 is a system for diagnosing complex general internal medicine cases and was created in 1982 by Miller and others. 7 A hybrid medical expert system called XBONE was developed in 1997, and it uses a hybrid representation formalism to integrate rules and an Adaline artificial neural unit for bone disease diagnosis using patients’ clinical, demographic, and scintigraphic (nuclear medicine image (NMI)) data. 8 EpiMAN-TB developed by McKenzie et al. 9 is a decision-support system using spatial information for the management of TB in cattle and deer, which works by prediction of possum TB hot spots using farm boundary, vegetation, and slope information.
Automated diagnosis systems. This shows selected and existing automated diagnosis system from 1976 to 2011.
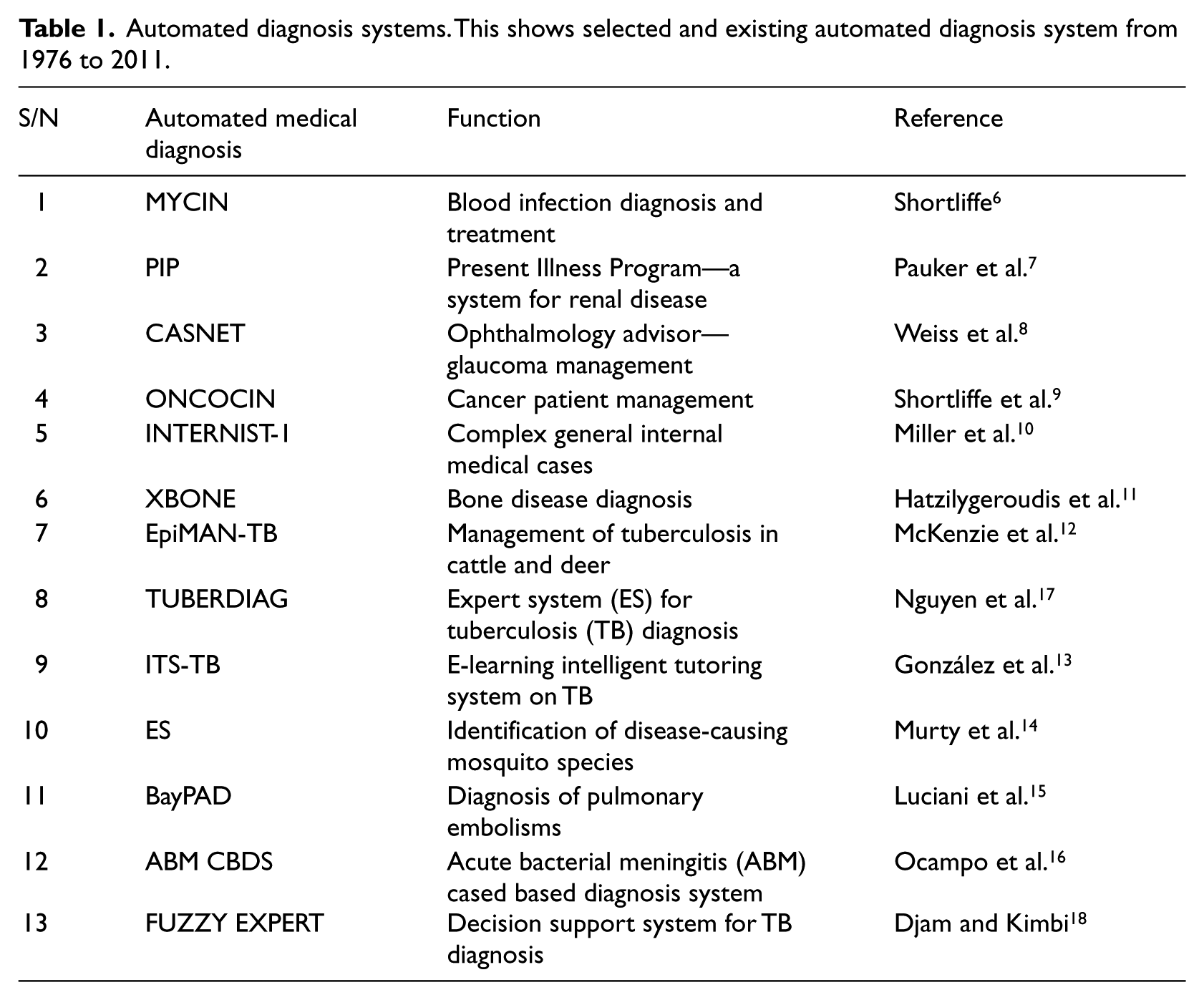
In 2005, attention temporarily shifted from diagnosis to public enlightenment when González et al. 10 presented ITS-TB, an e-learning Intelligent Tutoring System for TB by integrating multiagent systems and the case-based reasoning (CBR) paradigm. Murty et al. 11 described the use of the ID3 algorithm in the development of an online rule-based expert system for the identification of disease-causing mosquito species. A Bayesian probabilistic model-based system called BayPAD was developed by Luciani et al. 12 to perform diagnosis of pulmonary embolism. Based on the symptoms and signs that a patient presents, a Bayesian clinical decision support system (CDSS) uses Bayesian inference engine to propose differential diagnoses with their corresponding degree of certainty. Recently, Ocampo et al. 13 engineered the construction of diagnosis support tools for acute bacterial meningitis (ABM) using a CBR inference engine and also reported a comparative assessment of the quality of a CDSS resulting from the application of CBR to that of system developed using a Bayesian CDSS. TUBERDIAG 14 was developed in 1999 as a fuzzy logic and approximate reasoning system. In 2011, Djam and Kimbi 15 developed a fuzzy expert system for TB diagnosis, which provides decision support platform to TB researchers, physicians and other health-care practitioners in tropical medicine.
Methodology
Basic concepts of rule-based model
The basic model of a rule-based expert system is composed of sets of rules that are regularly applied to the sets of facts in the database. While facts represent circumstances that describe a certain situation in the real world, rules consist of an “if” and a “then” portion or “if-then” statement which represents heuristics that define a set of actions to be carried out in a given situation or circumstance.16,17 The “If” portion is on the left-hand side (LHS) and is called premise or predicate, while the “then” portion is on the right-hand side (RHS) and is either a conclusion or action.
- Action or set of actions represent the operations of the rule that determine the response of the system.
- Condition or set of conditions is expressed as a (simple or complex) Boolean expression which represents the situation under which the actions of the rule are enabled, in case the specified events occur. 19
Algorithmic rules in the TB–Diagnosis Expert System (TB-DES)
Where TB is suspected
The presence of cough with bloody sputum and swollen lymph is probably an indication of Mycobacterium infection, which is captured in the following algorithmic rules.
If <symptom is cough AND
(NOT symptom is headache)
AND (symptom is bloody sputum)
AND symptom is swollen lymph/neck/joint>
then <Notify (Patient), “Tuberculosis is very likely, please go and see your doctor”>.
Where TB is not suspected
When cough is not observed and sputum occurrence is none or minimal, and where it occurs, it is not bloody, then TB is not likely. This is depicted in the following algorithmic rules.
If <symptom is cough AND
(NOT Symptom is bloody sputum)
AND (NOT symptom is swollen lymph/neck/joint>
then <Notify (Patient), “Tuberculosis is not likely but ordinary cough”>.
Design of TB-DES
To support the design of TB-DES, the unified modeling language (UML) was applied to model the system and facilitate easy implementation. 18 Use case (Figure 1), and Sequence diagrams (Figure 2) were created to describe the system. The architecture of TB-DES shown in Figure 3 consists of a rule base, knowledge base, and patient database. Patient data is acquired over the Internet via a user interface.
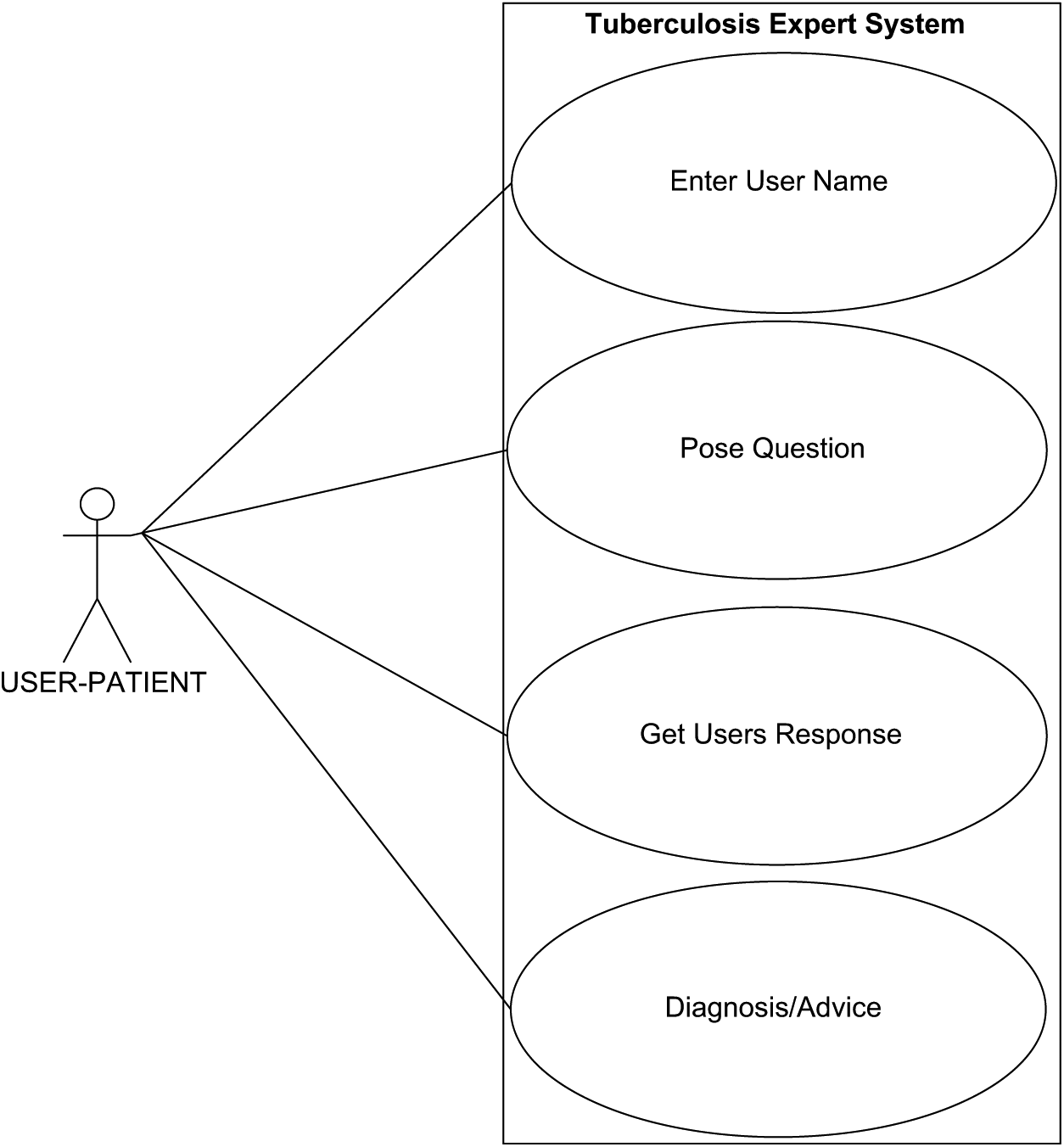
The use case diagrams show actors and their relationships to provide an overview of the functionality. An actor represents a role that a user plays when interacting with TB-DES, as indicated by the patient actor.
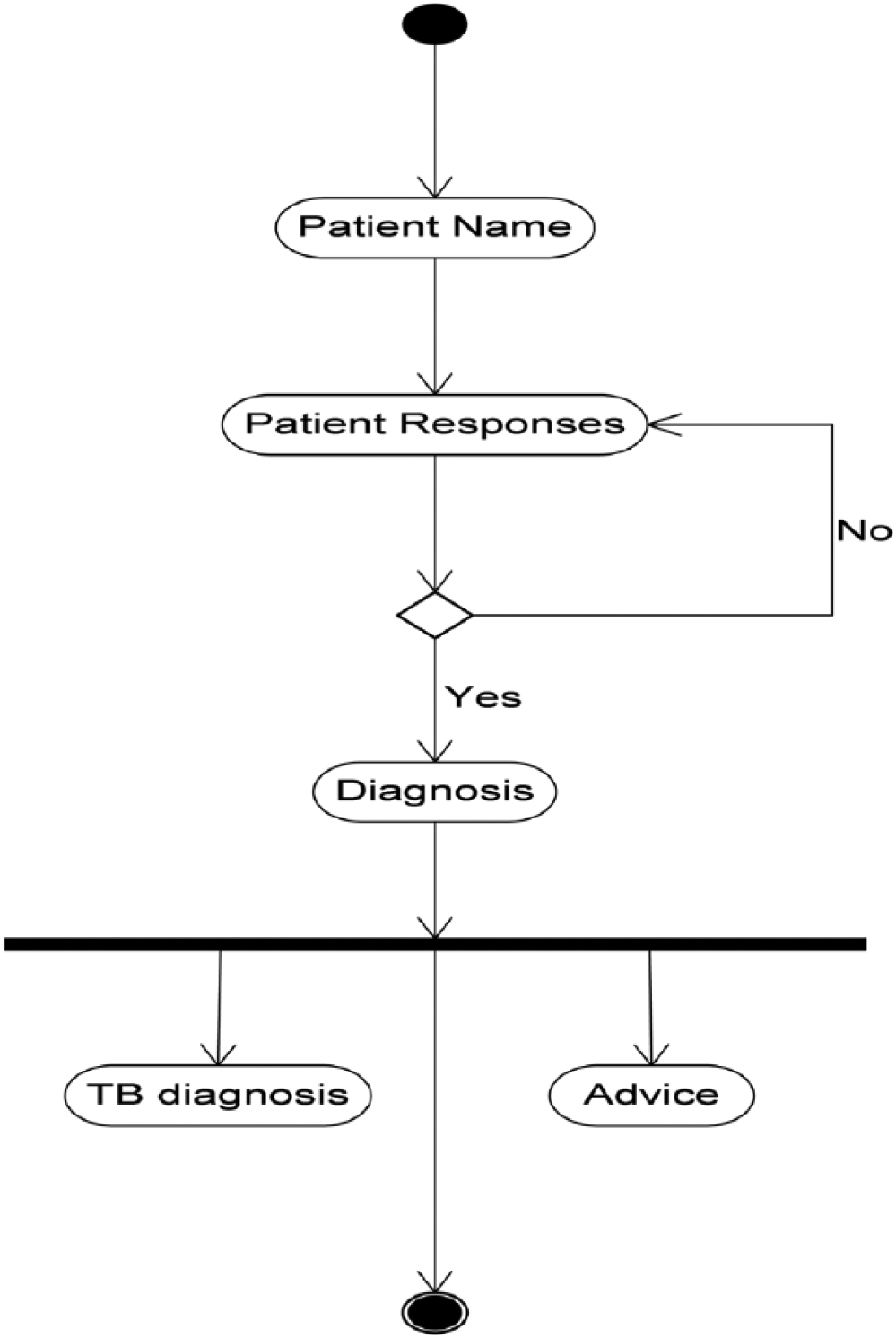
The activity diagram represents the transaction processes of TB-DES.
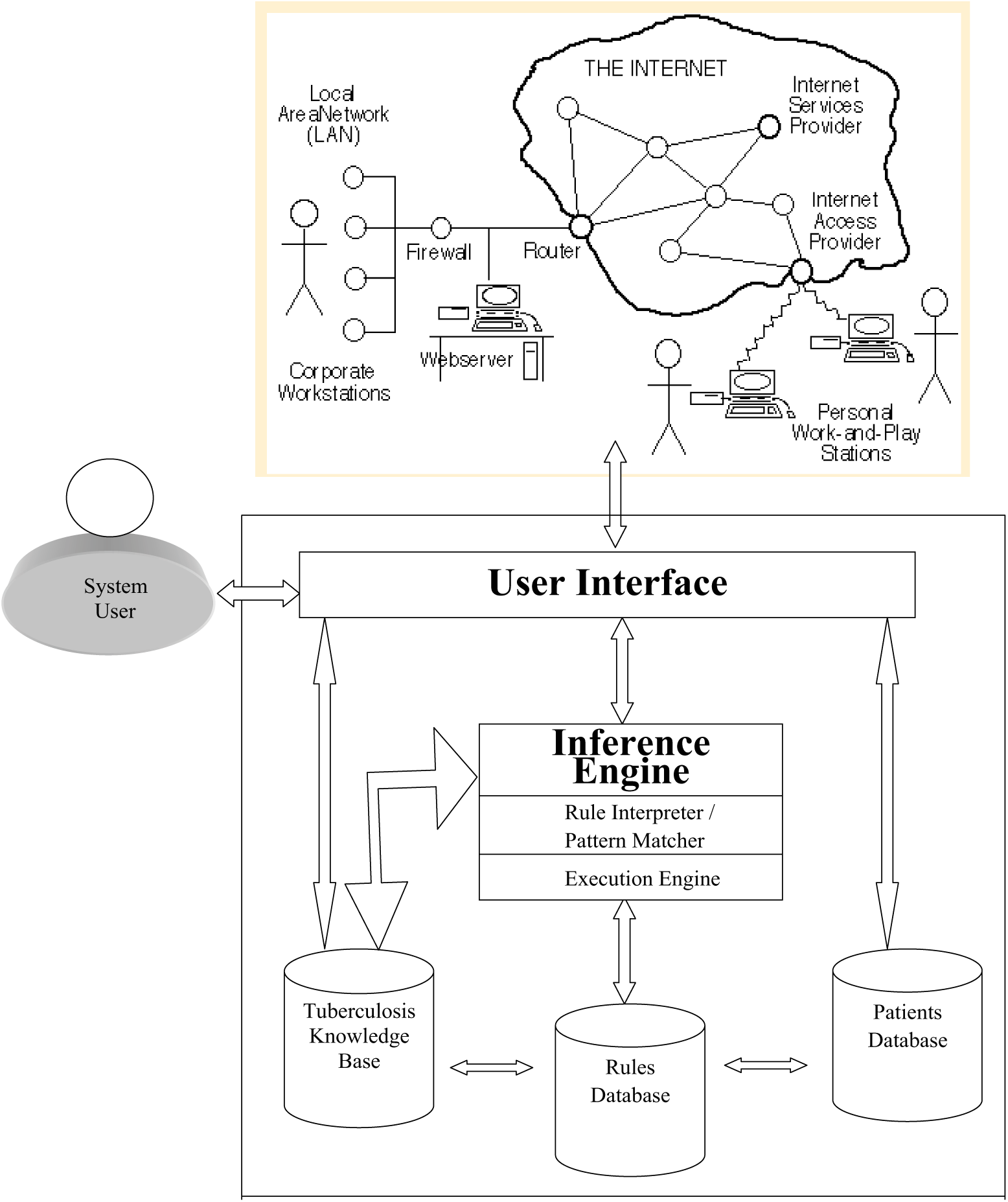
The TB-DES architecture consists of a rule base, knowledge base, and patient database. These units interact with the inference engine that receives patient data through the Internet via a user interface.
Systems implementation
TB-DES was implemented using Java NetBeans Integrated Development Environment (IDE) version 6.0 hosted on the Windows Vista operating system. NetBeans was chosen because it is a robust IDE suitable for rapid and reliable development of enterprise Java web applications. Java was chosen because of its multiplatform and concurrency support.
The various modules as described in Figure 3 are integrated and implemented using Java Server Pages (JSP), Apache Tomcat web server, Apache Derby Version 2.0, and Java NetBeans IDE. JSP technology enabled us to put snippets of Servlet Code directly into a text-based document. A JSP page is a text-based document that contains wireless markup language (WML), extensible markup language (XML) and JSP elements, which determine how the page constructs dynamic content. Apache Tomcat is the Servlet Container used in the official reference implementation for the Java Servlet and JSP technologies. The Java Servlet and JSP specification are developed by Sun under the Java Community Process.
Apache Derby is an open source relational database implemented entirely in Java and available under the Apache license version 2.0. Derby provides an embedded JDBC driver that lets one embed Derby in any Java-based solution. Derby also supports the more familiar client/server mode with the Derby Network Client JDBC driver and Derby Network Server. In other words, JDBC is responsible for initiating and maintaining the connection between the interface client and the Derby relational database. Java NetBeans IDE provided the total programming environment where all development took place. The system was tested with varying symptomatic input. The system was able to navigate through various prompted questions relevant to the preclinical screening and finally terminated with a conclusion that defines the infectious or noninfectious status of TB. In some cases, it also includes additional advisory information to help the patient improve his or her health. The sample screen shots of the implemented system are shown in Figures 4 and 5.
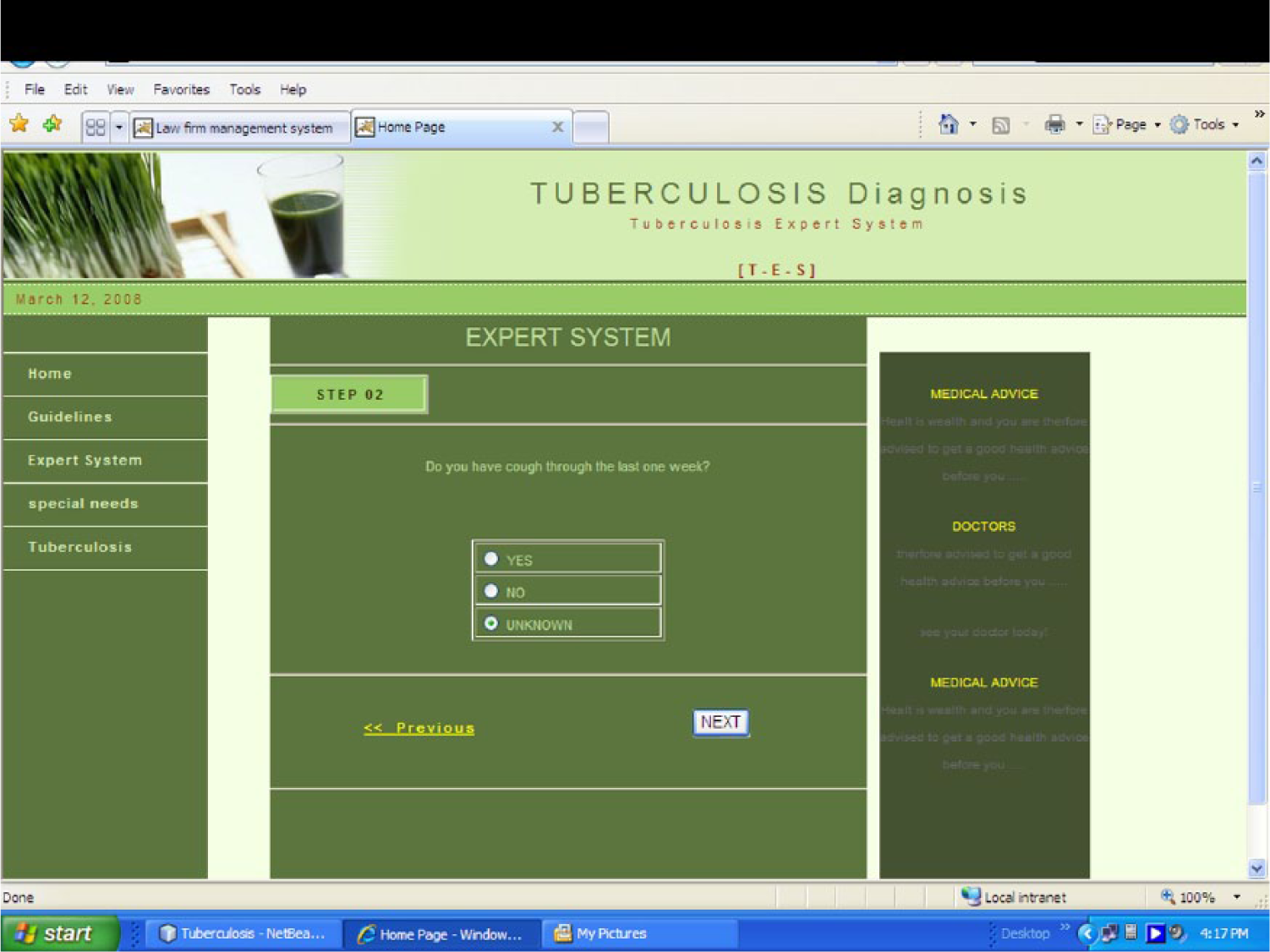
Question about any recent cough symptoms. The user is asked whether he or she had cough during the past week; the user selects an option and then moves to the next page.
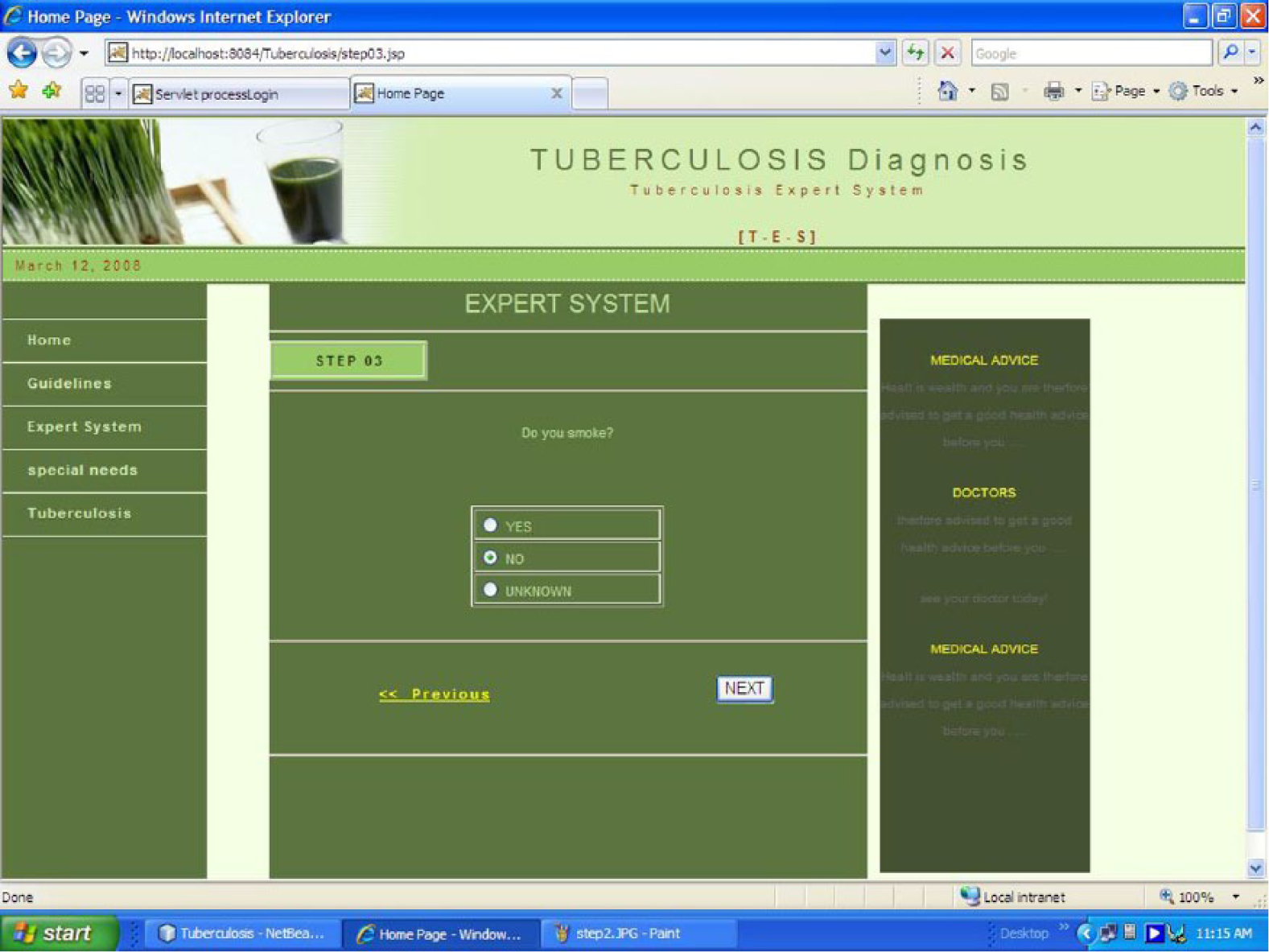
Smoking habits. The user is asked whether he or she smokes, because such behavior may aggravate TB; the user selects an option and then clicks on the next button to move to the next page..
The minimum hardware requirements for the TB-DES to work at an optimal level are Pentium III (600 MHz) processor or higher, 128 MB of RAM or more, with hard disk size of 2 GB and above, and Operating System Software (Windows XP, 2000 and above).
Results and discussion
The result of the integration of the various parts of the system modeled in Figures 1 to 2 gave rise to the complete architecture as shown in Figure 3. The core of the architecture consists of the patient database, rules base, inference engine and web client–server interface. The patient database contains facts that are the smallest piece of information supported by the inference engine. The rule base contains rules in the form of if-then statements, which represent the knowledge provided by the user and/or an expert in the problem domain. The inference engine matches facts in the patient database against rules in the rule base, and it determines which rules are applicable according to the reasoning method adopted by the engine.
The user who is ready to undertake the TB diagnosis pre-laboratory prediction connects through the Interface layer via a browser client, sending his or her personal (patient) data and associated symptoms as prompted by the system. The web server handles the requests by sending it to the expert system through same Interface layer. The required diagnostics prediction is processed by the expert system and delivered to the client. This is made possible by the internal working of the inference engine, which matches the facts (the symptoms supplied by user) in the patient database against the rules defined and sourced by the TB expert and placed in the knowledge base.
The TB-DES starts by asking series of questions from patients and data submitted to the system are taken as symptoms, for example, cough, loss of appetite, or any other symptoms. These symptoms may be associated with several illnesses like common cold, asthma, and so on, but if the patient is predicted to have TB, then lab confirmation is required. Possible symptoms of TB recognised by TB-ES are shown in Table 2.
Symptoms of tuberculosis.
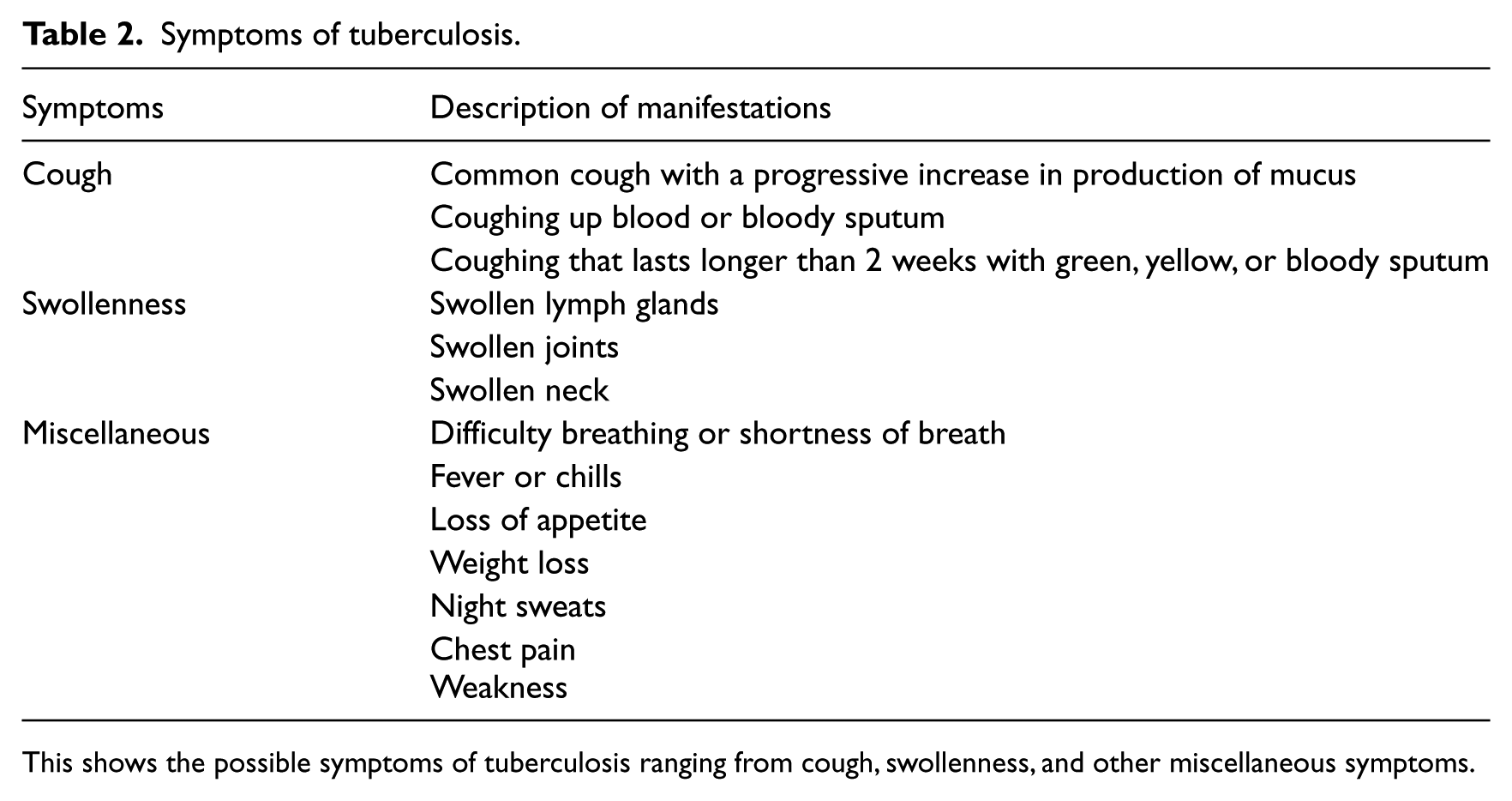

This shows the possible symptoms of tuberculosis ranging from cough, swollenness, and other miscellaneous symptoms.
The symbolic representation of expert knowledge in a computer can be represented using a number of schemes, but the rule-based approach is perhaps the most straightforward. Rules are conditional “if-then” statements that indicate circumstances under which conclusions can be drawn or actions taken, while categorical rule may be cause-to-effect, effect-to-cause, associational, or definitional rule. 3 In some cases, knowledge represented in expert systems may include indications of the degree of certainty associated with the rule.
A usability study was carried out using the specification in ISO 9241–11 19
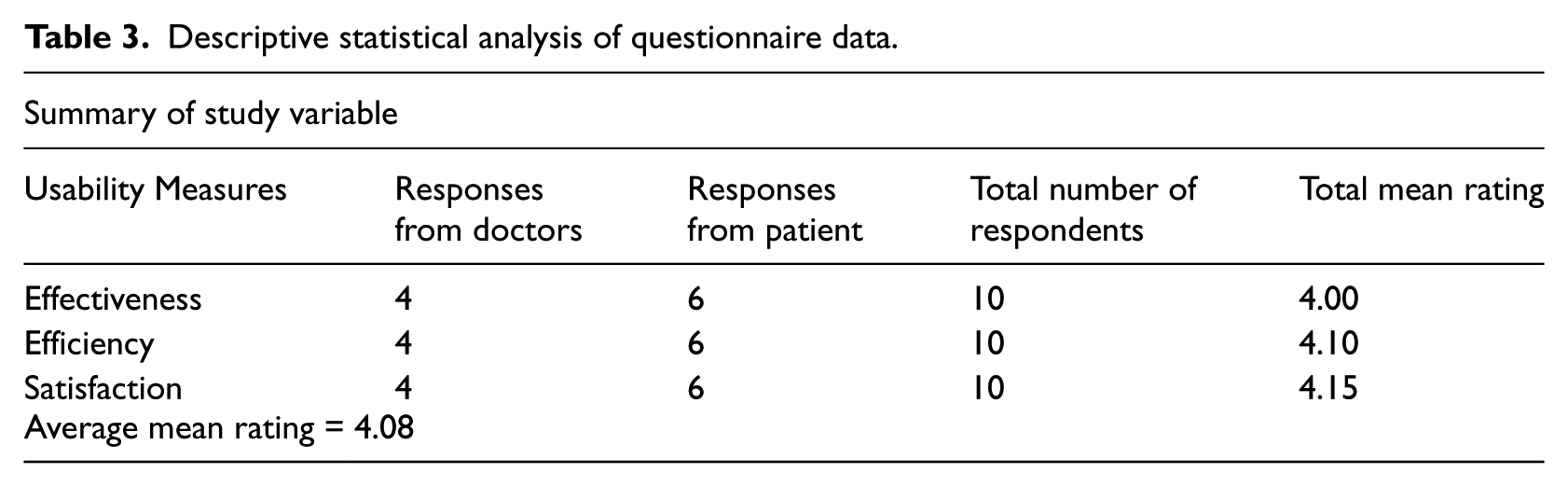
To conduct the study, a questionnaire was designed and administered to doctors and patients. The survey questionnaire was divided into five sections, namely, background information, effectiveness of the system, efficiency of the system, and user satisfaction. Each section on effectiveness, efficiency, and user satisfaction contains four questions represented by Q1, Q2, Q3, and Q4. A total of 12 questionnaires were administered, but only 10 responses were received, analyzed and reported (see Table 3). The questions were designed using a 5-point Likert scale, where 1 = strongly disagree, 2 = disagree, 3 = undecided, 4 = agree, and 5 = strongly agree.
Descriptive statistical analysis of questionnaire data.
The average ratings, standard deviation (SD) and variance (VAR) are presented in Table 4. In terms of questions such as “Would you support the use of computer diagnosis for TB in your hospital?,” 6 users said “yes” and 4 said “no.” Responses to Q4 on effectiveness – “I feel comfortable using the application” – were lower compared to other questions. Notwithstanding, the user survey results were encouraging, particularly, the responses to questions on user satisfaction. It was proposed by Sauro and Kindlund 20 that “Good Usability” should have a mean rating of 4 on a 1–5 scale and 5.6 on a 1–7 scale. Therefore, it can be concluded that the application has “Good Usability” based on the average total rating of 4.08. The usability analysis graph for each survey variable tested is shown in Figure 6 and Kindlund32 that “Good Usability” should have a mean rating of 4 on a 1–5 scale and 5.6 on a 1–7 scale. Therefore, it can be concluded that the web-based application developed has “Good Usability” based on the average total rating of 4.08. The usability analysis graph for each survey variable tested is shown in Figure 7.
User survey results N = 10.
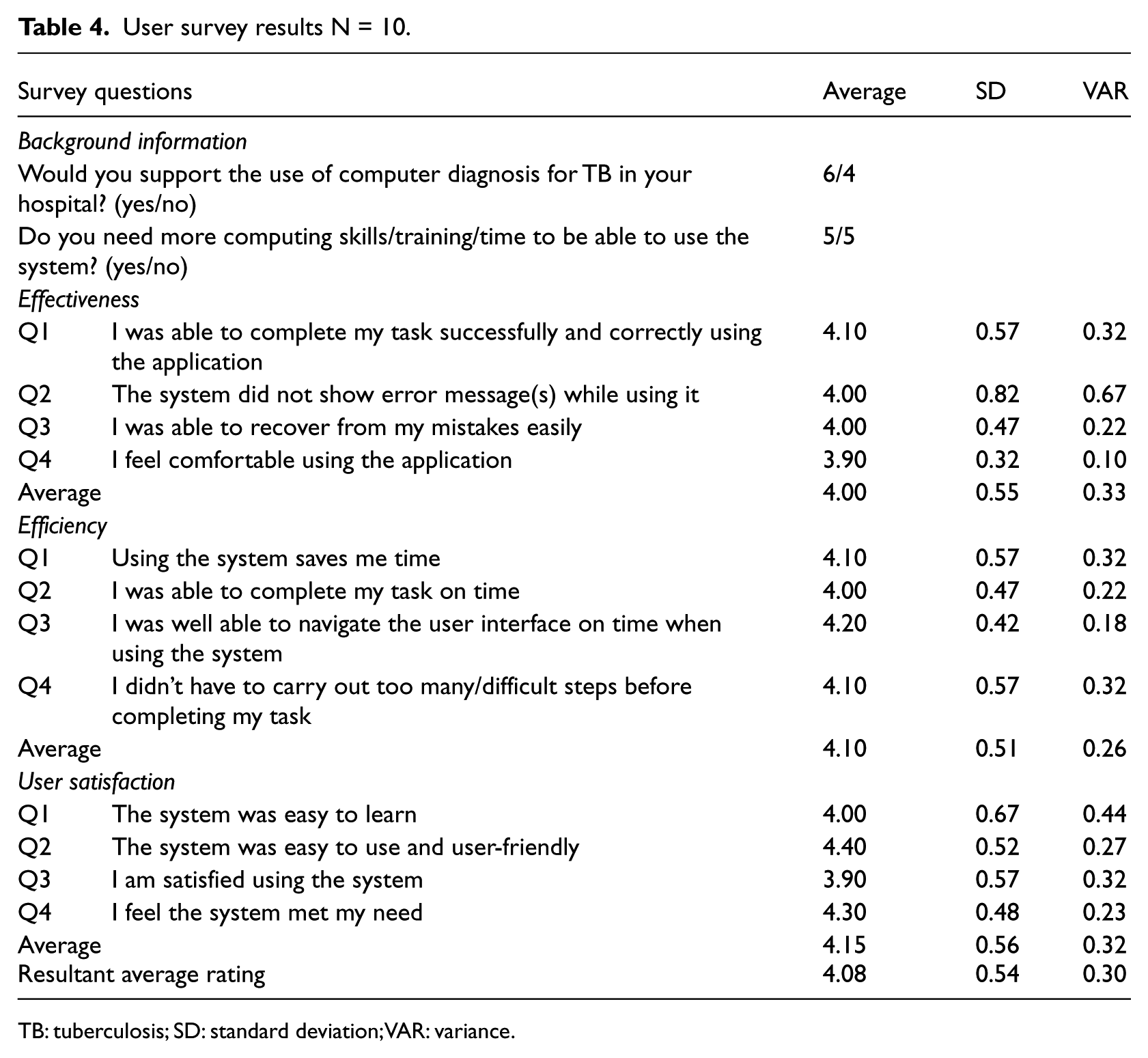
TB: tuberculosis; SD: standard deviation; VAR: variance.
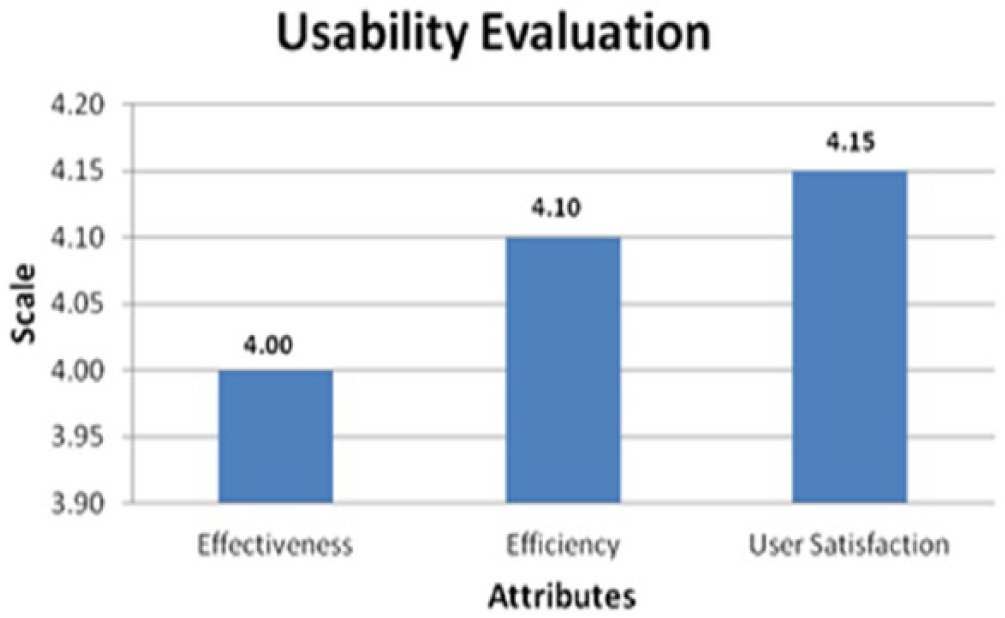
Usability analysis. The effectiveness, efficiency and user satisfaction with the system
Conclusion
Following the vast information harvested from the clinical biology of Mycobacterium sp., we applied a rule-based algorithm for the development of the TB-DES TB prediction system. Our knowledge base is inspired largely by the depth of information gathered on the clinical biology of Mycobacterium sp. A rule-based approach has been applied and knowledge stored as rules and facts to help medical practitioners reduce patient–doctor ratio in hospitals and ultimately consultation time in the long run. This system further aids in acquiring patients’ medical information via web for use in TB diagnosis, thereby ensuring medical consultation without walls.
The result of the usability evaluation reveals that the prototype application developed has “Good Usability” rating of 4.08 out of 5. This shows that the application will not only complement existing manual TB diagnosis systems but will be of benefit to doctors and patients.
Footnotes
Declaration of conflicting interests
There is no conflict of interest for this work.
Funding
This work was carried out using the authors’ own funds
