Abstract
Background:
Circular RNAs (circRNAs) have been implicated in the pathogenesis of atherosclerosis (AS) and the migration and proliferation of vascular smooth muscle cells (VSMCs) under oxidized low-density lipoprotein (ox-LDL). Here, we defined the exact action of human circ_0007478 in VSMC migration and proliferation induced by ox-LDL.
Methods:
Human VSMCs (HVSMCs) were exposed to ox-LDL. Circ_0007478, microRNA (miR)-638, and rho-associated protein kinase 2 (ROCK2) levels were gauged by quantitative real-time PCR (qRT-PCR) and western blot. Cell viability and proliferation were assessed by MTT and EdU assays, respectively. Transwell assays were used to detect cell migration and invasion. Dual-luciferase reporter and RNA immunoprecipitation (RIP) assays were used to evaluate the direct relationship between miR-638 and circ_0007478 or ROCK2.
Results:
Our data indicated that circ_0007478 expression was augmented in AS serum samples and ox-LDL-treated HVSMCs. Depletion of circ_0007478 attenuated HVSMC proliferation, migration, and invasion induced by ox-LDL. Mechanistically, circ_0007478 targeted miR-638 by directly pairing to miR-638. Reduction of miR-638 reversed the effects of circ_0007478 depletion on ox-LDL-evoked proliferation, migration, and invasion in HVSMCs. ROCK2 was a direct miR-638 target and miR-638-mediated inhibition of ROCK2 relieved ox-LDL-evoked HVSMC proliferation, migration, and invasion. Furthermore, circ_0007478 was identified as a competing endogenous RNA (ceRNA) for miR-638 to modulate ROCK2 expression.
Conclusion:
Our present study establishes an undescribed ceRNA regulatory network, in which circ_0007478 targets miR-638 to upregulate ROCK2, thereby contributing to ox-LDL-induced proliferation and migration in HVSMCs.
Introduction
Atherosclerosis (AS) is a slowly progressing inflammatory disease of medium and large-sized arteries that is the underlying pathological cause of the majority of cardiovascular diseases.1,2 The pathological mechanism of AS is highly complex, and the accumulation of lipids, the infiltration of immunocytes, as well as the formation of a fibrous cap by vascular smooth muscle cells (VSMCs) are considered to develop the atherosclerotic plaque. 1 Oxidized low-density lipoprotein (ox-LDL) has established a central role in AS by acting on various cells, such as VSMCs, macrophages, and endothelial cells. 3 Numerous studies have documented that ox-LDL can induce VSMC migration and proliferation and thus contributes to the formation of atherosclerotic plaque.4,5 Vital modulators of ox-LDL-evoked VSMC migration and proliferation, including circular RNAs (circRNAs) and proteins, are under exploration for the moment.4,6,7 Elucidating the functions of these regulatory molecules will provide a new opportunity for developing better interventions in AS.
Covalently closed circRNAs are natural RNA circles that are engendered mostly by back-splicing of exons. 8 Some circRNAs inhibit microRNA (miRNA) activity by pairing to miRNAs and thus function as competing endogenous RNAs (ceRNAs). 9 Recent reports have implicated circRNAs and their ceRNA activity in the pathogenesis of AS.10,11 Moreover, several circRNAs, including circUBR4 and circCHFR, can participate in ox-LDL-evoked VSMC migration and proliferation by miRNA competition.12,13 Huang et al. also highlighted that silencing endogenous circ_0029589 in VSMCs under ox-LDL hindered cell growth and migration through miR-214-3p-dependent regulation of stromal interaction molecule 1. 14 Wang and colleagues unveiled that circ_0002984 operated as a miR-181b-5p ceRNA to exacerbate ox-LDL-triggered VSMC migration and proliferation by elevating vascular endothelial growth factor A. 15 Circ_0007478 (also called circRNA transmembrane 7 superfamily member 3 [circTM7SF3]), produced by exonic circularization of TM7SF3, is highly upregulated in serum samples from patients with AS and it works as a contributor to ox-LDL-evoked oxidative stress, inflammation, and apoptosis in THP-1-derived macrophages. 16 The document also uncovered the ceRNA activity of circ_0007478 to regulate aspartyl (asparaginyl) β-hydroxylase (ASPH) via miR-206 competition. 16 Nevertheless, the exact action of circ_0007478 in VSMC migration and proliferation induced by ox-LDL remains to be clarified.
MiRNAs have regulatory functions in VSMC migration and proliferation during the developmental process of AS.17,18 Reduced miR-638 expression has been reported in the serum of patients with AS,19,20 and miR-638 can impede VSMC migration and proliferation induced by platelet-derived growth factor.21,22 Rho-associated protein kinase 2 (ROCK2), a serine/threonine kinase, has established a role in promoting the process of AS.23,24 ROCK2 actively participates in VSMC migration and proliferation,25,26 and its inhibitors have been highlighted to have a therapeutic value for AS. 24 However, whether miR-638 and ROCK2 are involved in the regulation of circ_0007478 is largely unclear.
In the current work, we sought to define the function of circ_0007478 in VSMC migration and proliferation induced by ox-LDL and addressed its underlying ceRNA mechanism by miRNAs.
Materials and methods
Human serum specimens
Thirty-three subjects with AS (mean age: 54.32 ± 7.57; 15 males and 18 females; 20 cases at coronary ischemia stage, 13 cases at fibrosis stage) and the same number of age- and sex-matched healthy donors (mean age: 56.25 ± 5.89; 17 males and 16 females) were recruited at First Affiliated Hospital of Bengbu Medical College from October 2019 to June 2020, with documented informed consent. We harvested serum samples from these participators as described 19 and transferred them to a ‒80°C refrigerator for storage. Subjects with tumors, hepatic disease, renal failure, recent infections, and trauma were excluded from the current work.
Human experimental protocols were granted by the institute’s Committee for Ethics at First Affiliated Hospital of Bengbu Medical College. The present study was approved by the ethical review committee of First Affiliated Hospital of Bengbu Medical College. Written informed consent was obtained from all enrolled patients, who agreed to participate in this work.
Cell line and oxidized LDL (ox-LDL) stimulation
We obtained human VSMCs (HVSMCs) from ATCC (Manassas, VA, USA) and propagated them with the VSMC Growth Kit from ATCC at 5% CO2 at 37°C. Ox-LDL was procured from Sigma-Aldrich (Sydney, Australia). For ox-LDL stimulation, cultured HVSMCs were exposed to ox-LDL at a concentration of 100 µg/mL for 24 h. For the dose course experiment of ox-LDL stimulation, cultured HVSMCs were plated at 2.5 × 106 cells per 100-mm dish before stimulation with ox-LDL for 24 h at 0, 25, 50, and 100 µg/mL concentrations.27,28
RNA expression analysis by quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA from 200 µL of serum specimens and cultured HVSMCs was extracted using the RNeasy Mini Kit and following the suggestion of the manufacturers (Qiagen; Basel, Switzerland). For circ_0007478 and ROCK2 mRNA detection, cDNA was randomly primed from 1 to 2 µg of RNA using Roche AMV reverse transcriptase (Tokyo, Japan); diluted cDNA was subjected to qRT-PCR with the primer sets shown in online Supplementary Table 1 using SYBR Green (TaKaRa; Dalian, China). For miR-638 detection, 200 ng of RNA was reverse transcribed using miR-638-specific stem-loop primers and miScript RT Kit (both from Qiagen); miRNA-cDNA was amplified by using a miScript SYBR Green Kit (Qiagen) and miR-638-specific primer sets shown in online Supplementary Table 1. Relative RNA expression was scored by comparative cycle threshold (2–ΔΔCt) with GAPDH (for circ_0007478 and ROCK2) or U6 (for miR-638) as the reference. Three duplicated wells were used for each sample, and a total of three independent experiments were conducted for this assay.
Expression plasmids, oligonucleotides, and transfection of human VSMCs (HVSMCs)
The following oligonucleotides were used in this study. The siRNA specific to circ_0007478 (si-circ_0007478, 5’-AAAGGAGAGUCUAGGGAAGCA-3’) and si-NC negative control (5’-AUACGUGUGACAACGCAUGGA-3’) were procured from Geneseed (Guangzhou, China). MicrON human miR-638 mimic, micrOFF miR-638 inhibitor (anti-miR-638), and the corresponding miR-NC and anti-miR-NC negative controls were procured from Ribobio (Guangzhou, China). Human ROCK2 expression plasmid pcDNA3.1-ROCK2 that expresses ROCK2 coding sequence without the 3’UTR was generated by inserting human ROCK2 coding sequence (synthesized by BGI; Shenzhen, China) into the pcDNA3.1(+) vector (Life Technologies; Paisley, UK). As control, the nontargeting pcDNA3.1 vector was employed. Oligonucleotides (200 nM siRNA, 50 nM miRNA mimic, and 50 nM miRNA inhibitor) or/and expression plasmids (200 ng) were transiently transfected in 2.5 × 105 HVSMCs using RNAiMAX or Lipofectamine 3000 reagent (both from Life Technologies), respectively, as per the manufacturing guideline. Ox-LDL stimulation and expression experiments were done 24–72 h posttransfection. We harvested HVSMCs after transfection and ox-LDL stimulation for further analyses as below.
MTT cell viability assay
For viability assay, we plated HVSMCs after transfection as indicated and ox-LDL stimulation at 3 × 103 cells/well in 96-well dishes and added 3-(4,5-dimethythiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) reagent (10 µL, 12 mM; MedChemExpress; Shanghai, China) to each well 48 h later. At 2–3 h after addition, 100 µL of SDS-HCl solution (Yeasen; Shanghai, China) was used to dissolve the produced formazan. We read the optical density at 570 nm with a Bio-Rad iMark reader (Sundbyberg, Sweden).
EdU incorporation assay
For proliferation assay, we plated HVSMCs after transfection as indicated and ox-LDL stimulation at 3 × 103 cells/well in 96-well dishes with 5-ethynyl-2’-deoxyuridine (Edu) solution (10 µM; Life Technologies) for 48 h. For EdU visualization, the Click-iT EdU Imaging Kit with Alexa Fluor 594 dye was used as recommended by the manufacturers (Life Technologies). Following nuclei staining with DAPI (Yeasen), we determined the percent of EdU positive cells relative to DAPI-stained cells under IX73 fluorescence microscopy (Olympus; Markham, ON, Canada).
Transwell migration and invasion assays
Following transfection and ox-LDL stimulation, we seeded HVSMCs in serum-free media in 24-well, 6.5 mm internal diameter transwell cluster plates with polycarbonate membranes (8 µm pore size; Corning Costar; Bodenheim, Germany) precoated with (at 2 × 105 cells/well, for invasion assay) or without (at 5 × 104 cells/well, for migration assay) Matrigel (Corning Costar). The lower chamber was filled with standard growth media. Plates were then incubated for 24 h at 37°C. We stained the cells migrating or invading through the inserts with crystal violet (0.2%; Yeasen) and counted their number by microscopy (100× magnification; Olympus).
Wound healing migration assay
After transfection and ox-LDL stimulation, HVSMCs were seeded into 24-well plates at a density of 5 × 104 cells per well, and serum-free culture medium was added to culture cells. The surface of cells was scratched to create a wound using a sterile pipette tip. The distance of the wound was quickly recorded by light microscopy (40× magnification). After culturing cells for 24 h, the distance of the wound was recorded again.
Western blot
Whole-cell lysates were prepared and immunoblotting was carried out as described. 29 Primary antibodies (Life Technologies) for immunoblotting were against ROCK2 (rabbit polyclonal, 1:5000, #PA5-78291), PCNA (mouse monoclonal, 1:1000, #13-3900), Bax (mouse monoclonal, 1:100, #MA5-14003), Bcl-2 (rabbit polyclonal, 1:100, #PA5-20068), and GAPDH (mouse monoclonal, 1:1000, #437000). The anti-rabbit and anti-mouse IgG-HRP antibodies (1:200‒1:500, #PA128786 and#32430; Life Technologies) were employed as secondary antibodies. Immunoreactivity signals were analyzed using GelQuant software (DNR Bio-Imaging; Jerusalem, Israel).
Prediction of the targeted microRNAs (miRNAs) of circ_0007478 and miR-638 targets
For prediction of the targeted miRNAs that potentially bind to circ_0007478, we used the computational tool circInteractome. 30 For prediction of miR-638 target genes, we employed the target-prediction program TargetScan (http://www.targetscan.org/vert_72/).
Dual-luciferase reporter assay
The fragments of circ_0007478 and ROCK2 3’UTR encompassing miR-638 putative target sequence or mutated target sites were inserted downstream of the Renilla luciferase cassette in psiCHECK-2 vector (Promega; Charbonnières, France). The indicated psiCHECK-2 wild-type or mutant plasmids (200 ng) were introduced into 2.5 × 105 HVSMCs along with miR-638 mimic (50 nM) or mimic control. Renilla luciferase was gauged using a Standard Luciferase Assay Kit (Promega) after 48 h and normalized to firefly luciferase to control for differences in transfection efficiency.
RNA immunoprecipitation (RIP) assay
Lysates of HVSMCs extracted with RIPA buffer (Yeasen) were added with Protein A+G Agarose (Roche; Shanghai, China) previously conjugated with the anti-Ago2 (rabbit polyclonal, 1:500, #PA5-117725; Life Technologies) or anti-IgG (rabbit monoclonal, 1:500, #ab172730; Abcam; Cambridge, UK) antibody. After addition, a 6-h incubation was allowed at 4°C. We harvested RNA bound to beads to perform qRT-PCR for quantification of circ_0007478, miR-638, and ROCK2 mRNA enrichment levels.
Statistical analysis
The GraphPad Prism 8.0 software (La Jolla, CA, USA) was used for statistical analysis. Data were expressed as mean ± SEM of separate experiments (n ⩾ 3). Differences were compared using ANOVA with Tukey’s multiple comparisons test for more than two groups or using Student’s t-test for two groups. Statistical significance was regarded as p < 0.05. For gene expression correlation, we used Pearson’s correlation coefficients.
Results
Circ_0007478 expression is elevated in AS serum samples and HVSMCs under ox-LDL
As shown in Figure 1A, circ_0007478 was a 622-nucleotide single-stranded circRNA that was produced by back-splicing of exons 3–6 of the linear pre-mRNA of TM7SF3 located on chromosome 12 (p11.23). We analyzed the expression of circ_0007478 in the serum samples from 33 patients with AS and 33 normal donors. The circ_0007478 level was remarkably augmented in AS serum samples as compared with normal controls (Figure 1B). We then stimulated HVSMCs with different doses of ox-LDL for 24 h and examined circ_0007478 expression. As expected, ox-LDL led to a dose-dependent elevation in circ_0007478 expression in HVSMCs (Figure 1C). Furthermore, when HVSMCs were treated with ox-LDL for 24 h, cell viability was dose-dependently increased (Figure 1D). Taken together, these results indicate the upregulation of circ_0007478 in AS serum samples and ox-LDL-treated HVSMCs.
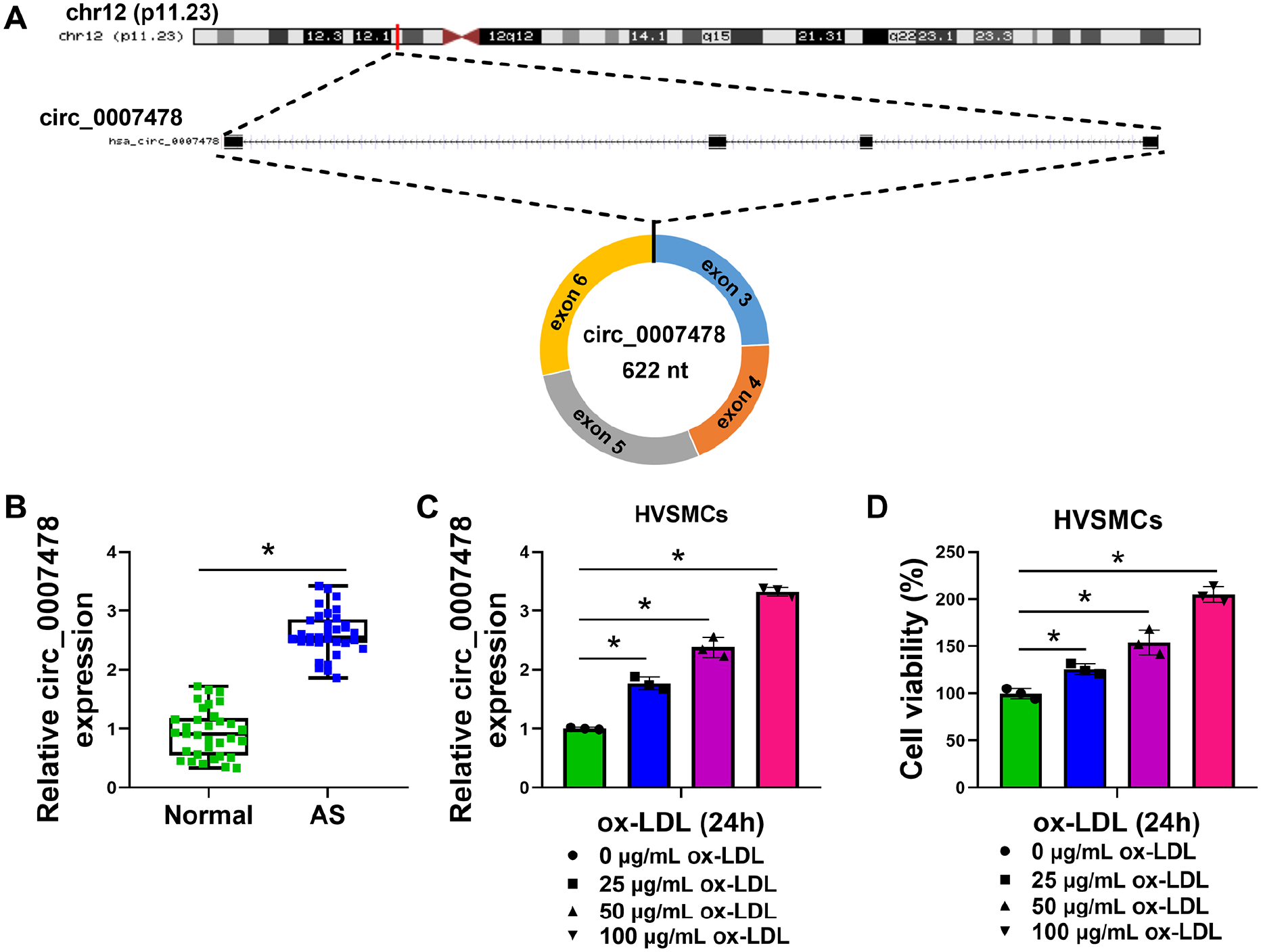
Circ_0007478 is overexpressed in serum of patients with AS and ox-LDL-stimulated HVSMCs.
Circ_0007478 depletion reduces HVSMC proliferation, migration, and invasion induced by ox-LDL
Having demonstrated elevation of circ_0007478 expression in ox-LDL-stimulated HVSMCs, we sought to evaluate the regulatory role of circ_0007478. A siRNA targeting circ_0007478 (si-circ_0007478) was used to silence circ_0007478 in ox-LDL-treated HVSMCs. Transfection of si-circ_0007478, but not the si-NC control, strikingly reduced circ_0007478 expression induced by ox-LDL in HVSMCs (Figure 2A). In function, circ_0007478 depletion strongly attenuated ox-LDL-driven proliferation and viability enhancement of HVSMCs (Figures 2B and 2C). Western blot results also showed that ox-LDL in HVSMCs resulted in increased levels of proliferating maker PCNA and antiapoptotic protein Bcl-2 and decreased expression of pro-apoptotic factor Bax, whereas knocking down circ_0007478 abrogated these effects (Figure 2D). Additionally, circ_0007478 loss of function led to suppressed migration and invasion rates of ox-LDL-stimulated HVSMCs by transwell assays (Figures 2E and 2F). Wound healing assay also presented that ox-LDL-enhanced HVSMC migration was attenuated by circ_0007478 downregulation (Figure 2G). These data establish that depletion of circ_0007478 protects HVSMCs against ox-LDL-induced proliferation and migration.
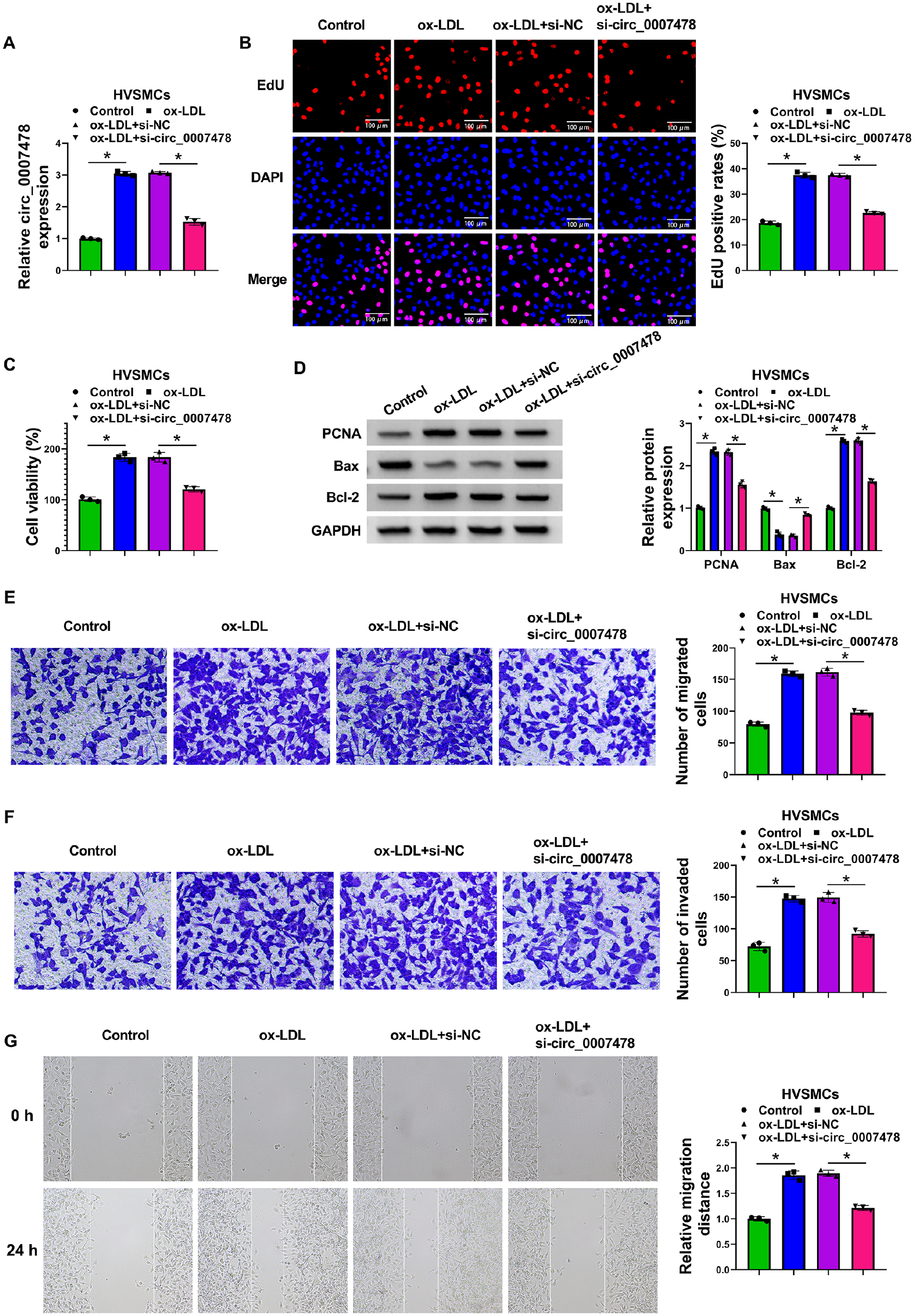
Circ_0007478 depletion mitigates HVSMC proliferation and migration induced by ox-LDL. HVSMCs were introduced with or without si-circ_0007478 or si-NC and then stimulated with 0 or 100 µg/mL of ox-LDL for 24 h.
Circ_0007478 directly targets miR-638
To search specific targeted miRNAs of circ_0007478 with potential relevance in the pathogenesis of AS, we used the computational tool circInteractome 30 and found a putative pairing site for miR-638, a pivotal regulator in AS pathogenesis and VSMC proliferation and migration,19,21,22 within circ_0007478 (context + score percentile: 96; Figure 3A). To establish the direct circ_0007478–miR-638 relationship, we inserted the circ_0007478 sectional sequence encompassing the predicted target site into a dual-luciferase reporter vector and transfected it into HVSMCs along with miR-638 mimic. The miR-638 increased efficacy of miR-638 mimic transfection was confirmed by qRT-PCR (Figure 3B). Remarkably, the luciferase expression of this reporter plasmid appeared to be suppressed by miR-638 (Figure 3C). We next constructed mutations in the target site and found that the reporter carrying a mutated binding site was refractory to repression by miR-638 (Figure 3C). MiRNAs direct gene expression in the form of miRNA ribonucleoprotein complexes that also harbor Ago2, the core component of the RNA-induced silencing complexes (RISCs). 31 To test whether circ_0007478 associates with RISCs, we performed RIP experiments using an anti-Ago2 antibody on extracts of HVSMCs. In agreement with miR-638 enrichment, circ_0007478 was substantially enriched in Ago2-containing RISCs compared with IgG immunoprecipitates (Figure 3D). Analysis of the serum samples revealed the decrease in miR-638 expression in AS serum samples relative to normal controls (Figure 3E). Moreover, there was a significant and inverse correlation between miR-638 and circ_0007478 levels in AS serum samples (Figure 3F). We also found that ox-LDL repressed miR-638 expression in HVSMCs in a dose-dependent manner (Figure 3G). It was important to note that we observed a striking elevation in the expression of endogenous miR-638 in circ_0007478-silenced HVSMCs (Figure 3H). Collectively, we conclude that circ_0007478 negatively regulates miR-638 expression via a binding site.
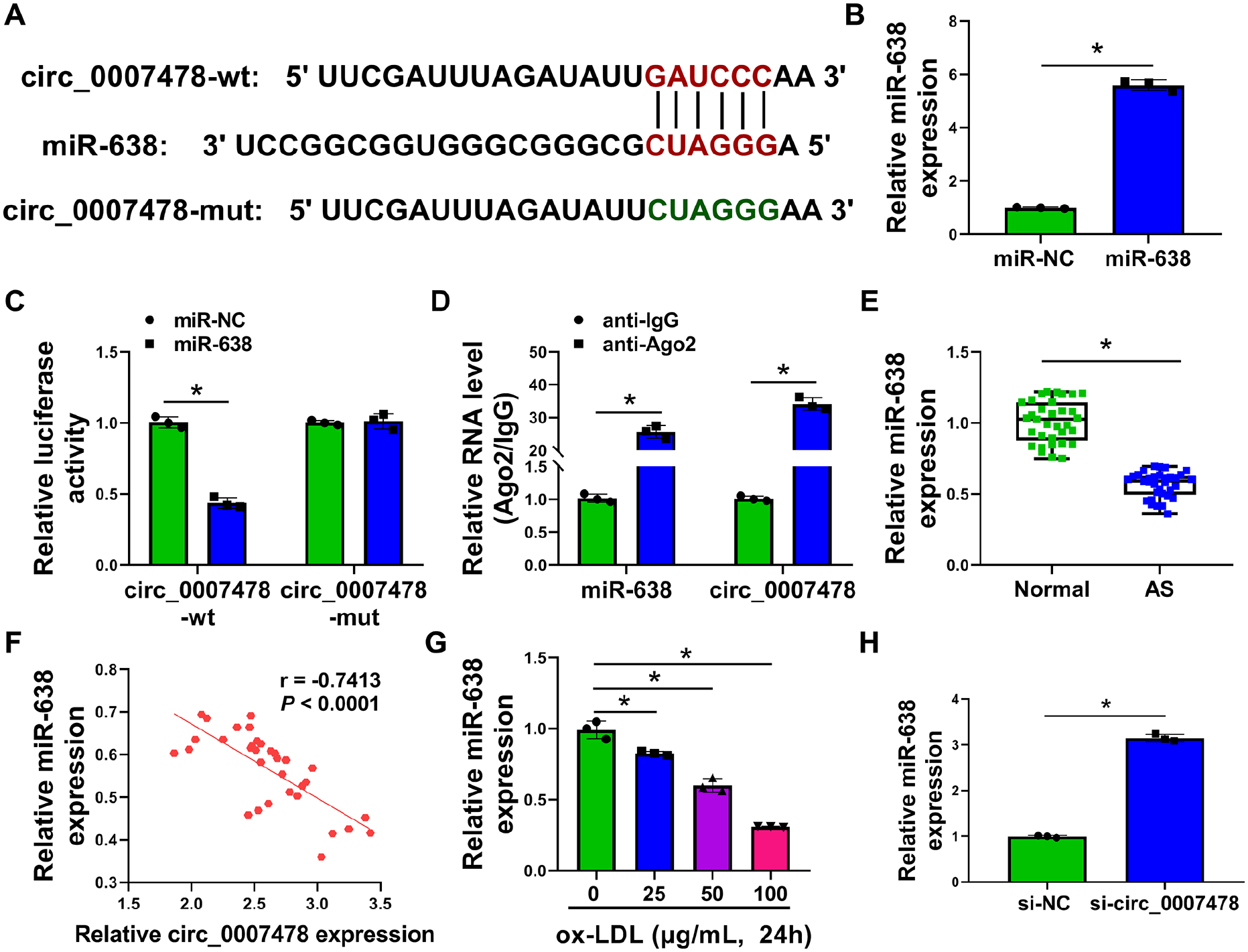
Circ_0007478 directly targets miR-638.
MiR-638 is responsible for the regulation of circ_0007478 in ox-LDL-induced HVSMC proliferation, migration, and invasion
Next, we determined whether circ_0007478 depletion mitigates ox-LDL-evoked proliferation and migration by the upregulation of miR-638. An inhibitor of miR-638 (anti-miR-638) was used to reduce miR-638 expression in circ_0007478-silenced HVSMCs, and the efficiency of anti-miR-638 transfection was validated by qRT-PCR (Figure 4A). Notably, reduction of miR-638 counteracted circ_0007478 depletion-driven repression of proliferation and viability of HVSMCs (Figures 4B and 4C; online Supplementary Figure S1A). Reduced expression of miR-638 also abrogated circ_0007478 knockdown-mediated alteration in the expression levels of PCNA, Bax, and Bcl-2 in HVSMCs (Figure 4D). Furthermore, miR-638 downregulation in HVSMCs clearly reversed the antimigration and anti-invasion effects of circ_0007478 depletion (Figures 4E–4G; online Supplementary Figures S1B–S1D). These findings establish that miR-638 represents a molecular mediator of circ_0007478 regulation in ox-LDL-evoked HVSMCs.
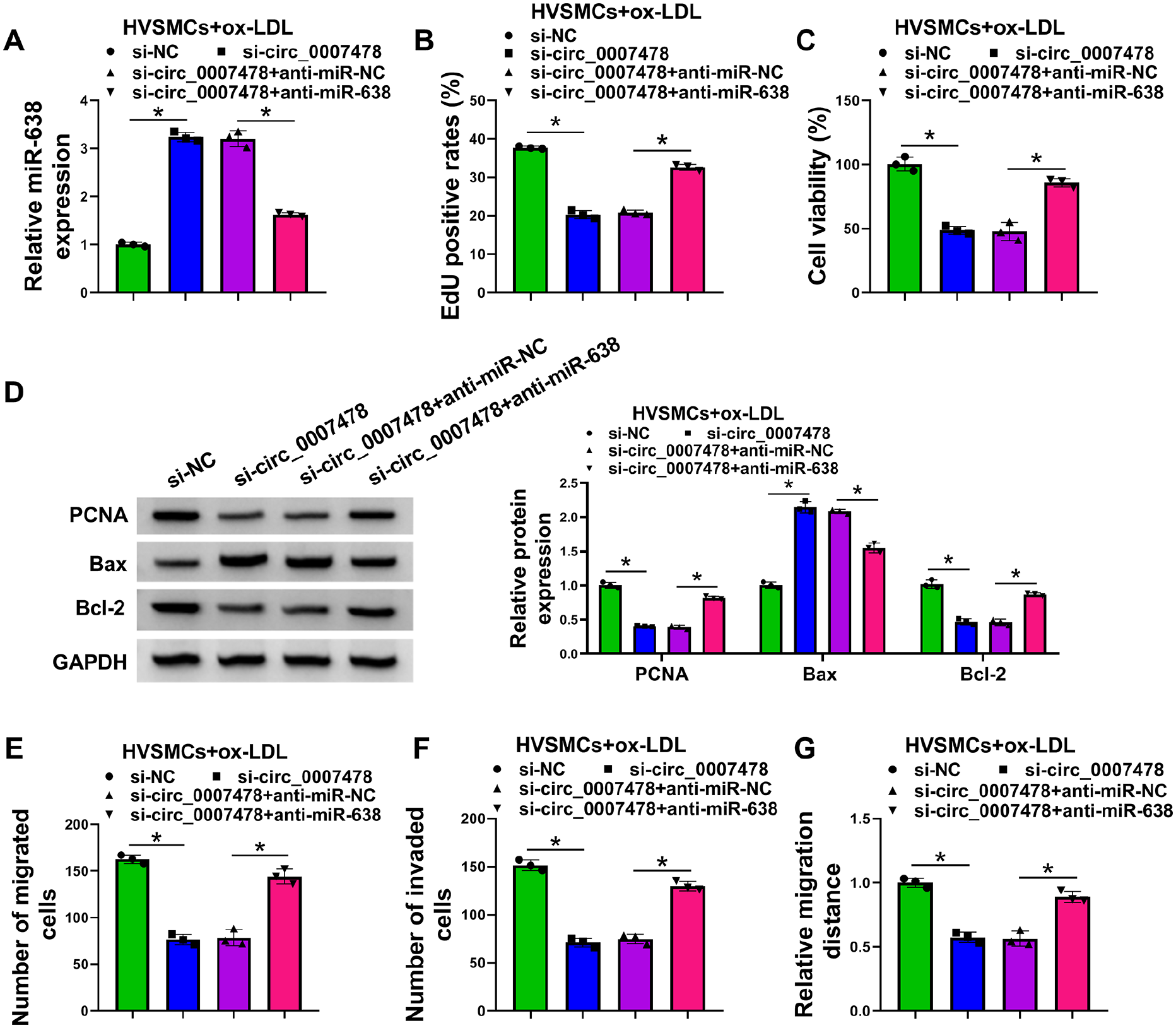
Depletion of circ_0007478 relieves ox-LDL-evoked HVSMC proliferation and migration via the elevation of miR-638 expression. HVSMCs were transiently transfected with anti-miR-638 + si-circ_0007478, anti-miR-NC + si-circ_0007478, si-circ_0007478, or si-NC before stimulation with 100 µg/mL of ox-LDL for 24 h.
ROCK2 is a direct miR-638 target and it is regulated by circ_0007478 via miR-638
To identify specific miR-638 targets, we interrogated the target-prediction program TargetScan and found a putative miR-638 complementary site within the 3’UTR of ROCK2 (Figure 5A). Besides, ROCK2 expression was decreased most in HVSMCs overexpressing miR-638 compared to other predicted genes (online Supplementary Figure S2). The 3’UTR of ROCK2 with the wild-type, but not with a mutant, miR-638-binding site conferred suppression by miR-638 to a luciferase reporter (Figure 5B). RIP assays also validated the association between ROCK2 and the RISCs (Figure 5C). We also found a striking upregulation of ROCK2 expression in AS serum samples and ox-LDL-stimulated HVSMCs (Figures 5D–5F). It was important to note that in AS serum samples, ROCK2 mRNA expression negatively correlated with miR-638 level, while the expression of ROCK2 and circ_0007478 showed a positive correlation (Figures 5G and 5H). The negative regulation of miR-638 in ROCK2 expression further supported the targeting of ROCK2 by miR-638 (Figure 5I). Intriguingly, we observed an obvious reduction in the expression of ROCK2 protein in HVSMCs after transfection by si-circ_0007478; this reduction was dramatically reversed by miR-638 knockdown (Figure 5J), pointing to that circ_0007478 functions as a ceRNA for miR-638 to modulate ROCK2 expression.
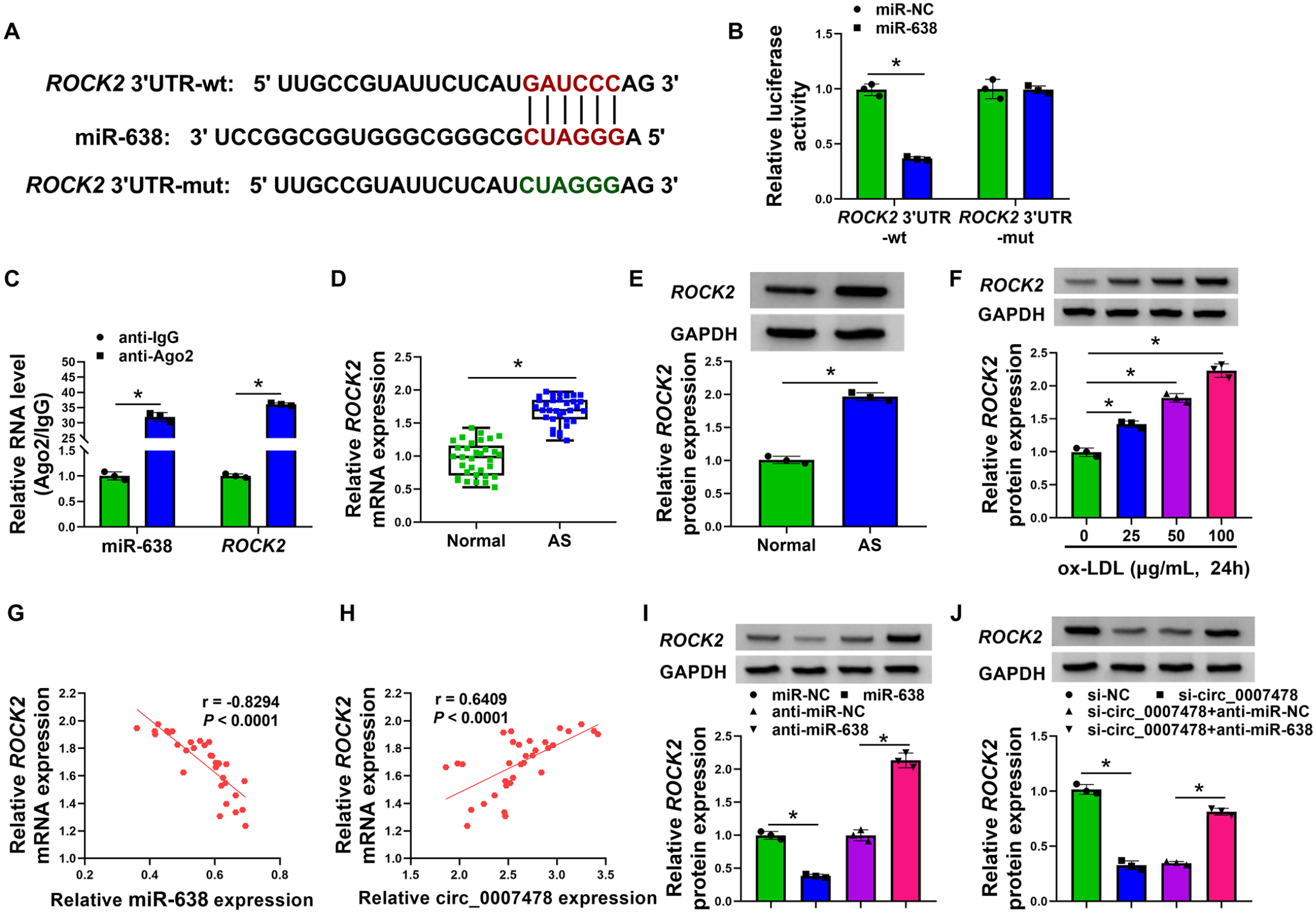
Circ_0007478 involves the expression regulation of ROCK2 by acting as a ceRNA for miR-638.
MiR-638-mediated inhibition of ROCK2 relieves HVSMC proliferation, migration, and invasion induced by ox-LDL
Our above data indicated that ROCK2 is directly targeted and inhibited by miR-638. ROCK2 has been well established as a potent contributor to AS.24,32 We therefore hypothesized that miR-638-mediated targeting of ROCK2 may impact ox-LDL-evoked proliferation and migration in HVSMCs. To test this hypothesis, we transfected HVSMCs with miR-638 mimic alone or along with a plasmid construct expressing ROCK2 coding sequence without the 3’UTR, and therefore insensitive to miR-638, before ox-LDL stimulation. In ox-LDL-stimulated HVSMCs, the ROCK2 plasmid construct significantly abolished miR-638-mediated inhibition of ROCK2 (Figure 6A). Overexpression of miR-638 in ox-LDL-stimulated HVSMCs suppressed cell proliferation (Figure 6B and 6C; online Supplementary Figure S3A) and elevated Bax expression, as well as decreased the levels of PCNA and Bcl-2 (Figure 6D); however, re-expression of ROCK2 reversed these effects (Figures 6B–6D). Moreover, overexpression of miR-638 led to suppressed motility and invasion rates of ox-LDL-stimulated HVSMCs, which was remarkably counteracted by re-expression of ROCK2 (Figures 6E–6G; online Supplementary Figures S3B–S3D). These findings collectively demonstrate that miR-638 protects HVSMCs from ox-LDL-induced proliferation and migration by inhibiting ROCK2.
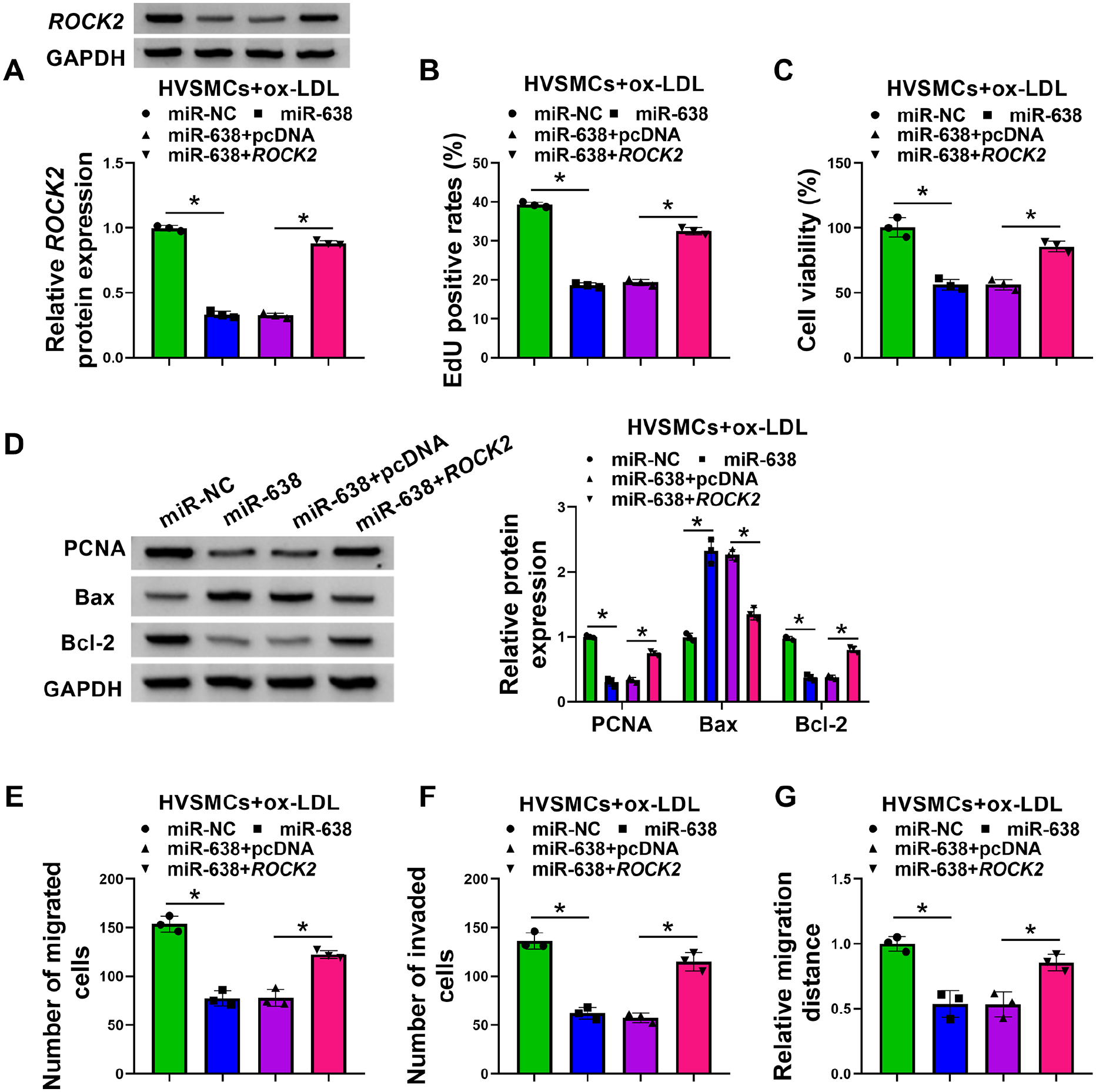
MiR-638 overexpression relieves ox-LDL-induced HVSMC proliferation and migration by inhibiting ROCK2. HVSMCs were introduced with miR-638 mimic + ROCK2, miR-638 mimic + pcDNA vector, miR-638 mimic, or miR-NC mimic before stimulation with 100 µg/mL of ox-LDL for 24 h.
Discussion
Recent evidence has identified circRNAs that contribute to the pathology of AS through the regulation of VSMC migration and proliferation via ceRNA activity.10,11 Considering the stimulative role of circ_0007478 in ox-LDL-induced macrophage apoptosis and inflammation during AS, 16 we set out to define the function and ceRNA activity of circ_0007478 in VSMC migration and proliferation induced by ox-LDL in vitro. Such investigation is limited at present by the lack of in vivo analyses using AS animal models.
In line with the findings reported by Wang et al., 16 our data also discovered the overexpression of circ_0007478 in AS serum, hinting the significance of circ_0007478 as a potential marker for AS diagnosis. Because of the elevation of circ_0007478 level in HVSMCs under ox-LDL, we used the siRNA silencing experiments to identify that circ_0007478 depletion protects HVSMCs against ox-LDL-triggered migration and proliferation in vitro. The data implied that the targeted inhibition of circ_0007478 might be a strategy for AS therapy development in the future.
We have also demonstrated that circ_0007478 targets miR-638 by directly pairing to miR-638. MiR-638 is an established antitumor factor in gastric cardia adenocarcinoma and endometrial carcinoma,33,34 and it functions as a potential driver in renal cell carcinoma. 35 Dysregulation of miR-638 has also been illustrated in diabetic nephropathy and it can impact inflammatory response. 36 Additionally, miR-638 possesses antimigration and antiproliferation activity in human abnormal airway smooth muscle cells. 37 The underexpression of serum miR-638 is tightly linked to vulnerable atherosclerotic plaques in patients with advanced carotid stenosis. 19 Moreover, miR-638 is the capacity of repressing VSMC migration and proliferation induced by platelet-derived growth factor21,22 and endothelial cell proliferation induced by ox-LDL. 20 Our study first unveils the repressive effect of miR-638 on ox-LDL-evoked HVSMC proliferation and migration. Importantly, we identify, for the first time, miR-638 as a molecular mediator of circ_0007478 regulation in ox-LDL-evoked HVSMCs.
ROCK2 functions as a strong contributor in AS,16,23 and inhibition of ROCK2 may be a promising therapeutic method for AS. 24 Here, we provide definitive evidence for the notion that ROCK2 is a direct miR-638 target and the regulation of miR-638 in ox-LDL-evoked HVSMCs depends, at least in part, on miR-638-mediated inhibition of ROCK2. Based on the shared binding site for miR-638, we first highlight the ceRNA activity of circ_0007478 to modulate ROCK2 expression through miR-638 competition, suggesting the circ_0007478/miR-638/ROCK2 ceRNA crosstalk in ox-LDL-evoked HVSMCs and providing a previously unknown regulatory network in AS. Similarly, the findings reported by Wang et al. pointed to the circ_0007478/miR-206/ASPH axis in an ox-LDL-stimulated AS cell model in vitro. 16 There may be other ceRNA crosstalks that remain to be clarified in the function of circ_0007478 in AS pathogenesis.
Study limitations
In terms of the limitations of present study, the data showing the levels of circ_0007478, miR-638, and ROCK2 in HVSMCs of the atherosclerotic plaque are lacking. Future work should conduct co-immunostaining of plaques or vessels in an animal model of AS or in excised human tissues to address these issues. In addition, the strategy for circ_0007478 siRNA-associated targeted therapy should focus on the potential off-target side effects, which needs to be further studied.
Conclusion
To conclude, our study has demonstrated an undescribed ceRNA regulatory network (online Supplementary Figure S4), in which circ_0007478 targets miR-638 to upregulate ROCK2 expression, thereby contributing to ox-LDL-induced proliferation and migration in HVSMCs. Importantly, this network has potential targets that can be explored for therapeutic interventions against AS.
Supplemental Material
sj-pdf-1-vmj-10.1177_1358863X221137617 – Supplemental material for Regulation of oxidized LDL-induced proliferation and migration in human vascular smooth muscle cells by a novel circ_0007478/miR-638/ROCK2 ceRNA network
Supplemental material, sj-pdf-1-vmj-10.1177_1358863X221137617 for Regulation of oxidized LDL-induced proliferation and migration in human vascular smooth muscle cells by a novel circ_0007478/miR-638/ROCK2 ceRNA network by Zeyu Guan, Ran Lu, Yong Sun, Xiaogao Wang, Chaowen Yu and Tao Song in Vascular Medicine
Supplemental Material
sj-pdf-2-vmj-10.1177_1358863X221137617 – Supplemental material for Regulation of oxidized LDL-induced proliferation and migration in human vascular smooth muscle cells by a novel circ_0007478/miR-638/ROCK2 ceRNA network
Supplemental material, sj-pdf-2-vmj-10.1177_1358863X221137617 for Regulation of oxidized LDL-induced proliferation and migration in human vascular smooth muscle cells by a novel circ_0007478/miR-638/ROCK2 ceRNA network by Zeyu Guan, Ran Lu, Yong Sun, Xiaogao Wang, Chaowen Yu and Tao Song in Vascular Medicine
Footnotes
Availability of data and materials
The analyzed data sets generated during the present study are available from the corresponding author on reasonable request.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplementary material
The supplementary material is available online with the article
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
