Abstract
The aim of this study was to determine how genetic variants contribute to warfarin dosing variability when non-genetic factors are controlled. Thirty healthy subjects were subjected to a warfarin dosing algorithm with daily international normalized ratio (INR) measurements to INR ≥ 2.0, then off warfarin to INR ≤ 1.2. The primary outcome was the cumulative dose required to achieve INR ≥ 2.0 for 2 consecutive days. CYP2C9 (p=0.004) and VKORC1 (p=0.02) variant carriers required lower cumulative doses, and CYP4F2 carriers required higher doses (p=0.04). Subjects with variants in both CYP2C9 and VKORC1 required fewer days to reach INR ≥ 2.0 than wild-type subjects or those with variants in CYP2C9 or VKORC1 (p=0.01). Genetic contribution to dose variability (~62%) was greater than previously reported, suggesting that uncontrolled clinical variables influence the effect of these variants. In conclusion, genotype-guided warfarin-dosing algorithms may rely more on genetic variables in healthier individuals than in patients with clinical confounders.
Introduction
Warfarin, an oral anticoagulant widely prescribed for the prevention and treatment of thromboembolism, 1 has a narrow therapeutic window and variable dose–response influenced by genetic variation. Adverse events associated with warfarin therapy are common, 2 and bleeding complications are particularly frequent during the initial dose-finding phase.3,4 Typically, anticoagulation is achieved through empiric dose-titration with consideration of clinical variables and frequent assessment of the international normalized ratio (INR), a standard measure of anticoagulation intensity based on the prothrombin time. Despite these precautions, conventional dosing strategies are associated with therapeutic anticoagulation intensity only about half the time. 5 The mutable dose–response remains an inherent limitation of warfarin since insufficient and excessive anticoagulation are associated with elevated risks of thrombosis and bleeding, respectively. 6 Previous studies have reported up to 40% dose variability attributable to common variants in the cytochrome P450-2C9 (CYP2C9) and vitamin K epoxide reductase complex 1 (VKORC1) genes among patients with multiple clinical confounders, including concomitant medications, compliance and co-morbid conditions7,8 The genetic contribution in the absence of these clinical variables has not been adequately assessed, and may provide insight into the selection of patients who benefit most from pharmacogenetically guided warfarin dosing.
Warfarin dosing algorithms that include both genetic and clinical variables have been developed to estimate warfarin dose requirements,9–12 and these are more effective than dosing either empirically or according to the information packaged with warfarin tablets.11,13 Variant CYP2C9 alleles with reduced activity result in delayed warfarin elimination, lower therapeutic dose and greater risk of bleeding7,14 The VKORC1 enzyme is the direct target of warfarin, and common haplotypes resulting in reduced gene expression have been associated with increased warfarin sensitivity and lower therapeutic doses.10,11,15–18 Genome-wide association studies have identified a variant in the CYP4F2 gene (c.1297G>A; p.V433M) that contributes to warfarin dose variability, albeit less than CYP2C9 and VKORC1.19–21 Despite advances in warfarin pharmacogenetics and evaluation in prospective trials,22–26 the independent contribution of allelic variation to the anticoagulant response is incompletely understood.
To address potential confounding of clinical variables on warfarin dosing, we measured the impact of allelic variation on dose–response in healthy subjects to test the hypothesis that genetic variation in CYP2C9, VKORC1 and CYP4F2 explains a greater proportion of the variability in response when non-genetic factors influencing warfarin pharmacokinetics and pharmacodynamics are controlled.
Methods
Subjects
Participants were healthy, non-smoking volunteers, identified and recruited through posted advertisements and email solicitation. Eligible subjects were 18–74 years old, not taking warfarin, able to provide informed consent and available for the 3-week duration of data collection. Women of childbearing potential were included if pregnancy was excluded and abstaining from sexual activity or using barrier contraception during the study. We excluded subjects taking (within 2 weeks) medications known to interact with warfarin, aspirin or clopidogrel, those with thrombotic disorders requiring anticoagulation, thrombophilia or coagulopathy identified by history or a screening INR ≥ 1.2, activated partial thromboplastin time (aPTT) more than twice the upper limit of normal (ULN), serum Protein C or S deficiency, family history of thrombophilia or coagulopathy, prisoners or wards of the state, scheduled surgery within 1 month of participation, active liver disease based on history or serum transaminase elevation (> 2× ULN), premenopausal women on oral contraception, and non-English-speaking individuals. Qualifying subjects were financially compensated, and were assigned unique identifying codes to track samples and data throughout the study. The protocol was approved by the Institutional Review Board governing research involving human subjects (ClinicalTrials.gov/NCT01520402) and monitored by a Data Safety Monitoring Committee chaired by an independent coagulation specialist.
Data collection and follow-up
At the screening visit, subjects meeting inclusion criteria completed an intake questionnaire including demographic information (age, height, weight, race, ethnicity, smoking history, history of liver disease, cardiovascular disease, bleeding problems or family history of liver disease, cardiovascular disease or bleeding problems) and current medications, which were assessed for possible interaction with warfarin. Participants underwent blood sampling for genotyping, coagulation testing, measurement of serum Protein C and S activity, and hepatic function tests. Genotyping of CYP2C9, VKORC1 and CYP4F2 was performed blinded to clinical data.
Subjects were educated regarding the vitamin K content of foods and instructed to follow a fixed diet (120 μg/day for men and 90 μg/day for women). Each maintained a log of food intake and refrained from over-the-counter medications or alcohol for 1 week before data collection began. Only subjects adhering to the prescribed diet (based on review of dietary logs) and avoiding medications that potentially interfere with warfarin metabolism qualified for the dose–response testing phase. These subjects took daily oral doses of warfarin (Coumadin® tablets; Bristol-Meyers Squibb, Princeton, NJ, USA) based on INR measurements made with the CoaguChek-S Coagulation Meter (Roche Diagnostics, Germany). INR measurements obtained using the meter were compared with values processed in the clinical laboratory during the screening visit (accepting INR variability ± 0.1). The initial warfarin dose for all subjects was 5 mg daily; if the INR was <2.0 after 4 days, the dose was increased to 7.5 mg daily, and subsequently increased by 2.5 mg every 3 days until the INR reached ≥ 2.0 for 2 consecutive days to a maximum of 15 mg daily. Warfarin was then stopped and INR measurements continued daily until the value fell below 1.2.
Genotyping
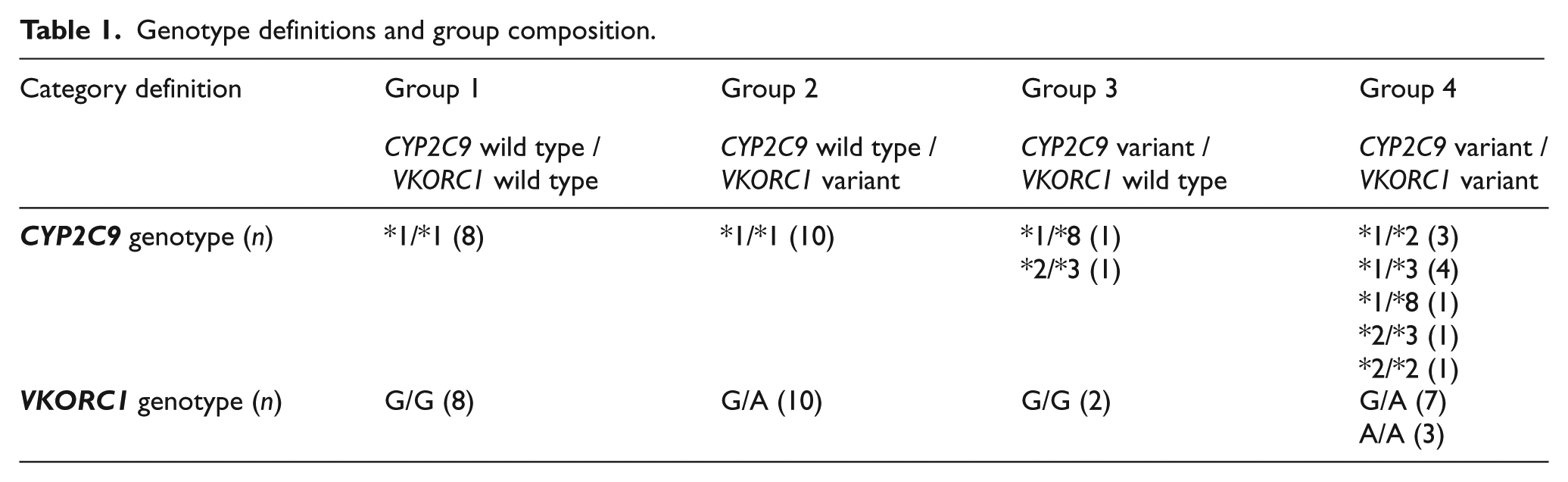
Genotyping of CYP2C9 (*2, *3, *4, *5, *6, *8, *11, *13), VKORC1 (-1639G>A and selected coding region mutations [p.V29L, p.A41S, p.V45A, p.R58G, p.V66M, p.L128R]), and CYP4F2 (rs2108622 [p.V433M]) was performed using the Tag-It™ Mutation Detection Kit (Luminex Molecular Diagnostics, Toronto, ON, Canada) and PCR-restriction fragment length polymorphism assays, as previously described.12,27,28 The wild-type CYP2C9 allele (*1) was assigned in the absence of other detectable variant alleles. Extensive metabolizers (EMs) were defined as *1/*1; intermediate metabolizers (IMs) as *1/variant; and poor metabolizers (PMs) as variant/variant. Subjects were also grouped into four categories based on the CYP2C9 and VKORC1 genotype profile (Table 1): Group 1 (CYP2C9 wild type and VKORC1 wild type), Group 2 (CYP2C9 wild type and VKORC1 variant), Group 3 (CYP2C9 variant and VKORC1 wild type), and Group 4 (CYP2C9 variant and VKORC1 variant).
Genotype definitions and group composition.
Main outcome measures
To assess the effect of CYP2C9, VKORC1 -1639G>A and CYP4F2 genotypes on the anticoagulant response to warfarin, the primary outcome was the cumulative dose required to reach an INR within the usual clinical therapeutic range 2.0–3.0) for 2 consecutive days. The secondary outcome was the time required for the INR to return to baseline (≤ 1.2) after warfarin cessation. Subjects were monitored for adverse clinical events, including skin necrosis, hospitalization and major or minor bleeding (e.g. bruising). All authors had access to primary clinical trial data and contributed to analysis and interpretation of the results.
Statistical analysis
Analyses were performed using SAS/STAT® software, version 9.1 (SAS Institute, Inc., Cary, NC, USA). Associations between the primary outcome measure (cumulative warfarin dose to achieve therapeutic INR) and genotype (CYP2C9, VKORC1, CYP4F2 or CYP2C9/VKORC1 genotype group) were assessed using the Kruskal–Wallis and Wilcoxon rank-sum tests. The proportion of variance (R2) explained by each predictor was calculated using multivariate regression analysis adjusted for age, sex and reported race, with outcome values logarithmically transformed. The study was powered to detect R2 > 20%, and significance was accepted at p ≤ 0.05.
Results
Study population
Thirty-five subjects responded to email and printed solicitations for the study, and five were excluded due to medication conflicts, unwillingness to take warfarin, or medical conditions precluding participation. Table 2 summarizes the baseline characteristics of subjects grouped by genotype. Approximately 60% were male in all genotype groups except Group 1 (38%). Subject age ranged from 24 to 55 years; the mean age (32 ± 8 years) and calculated body mass index (BMI) (24 ± 3 kg/m2) did not differ between groups. In all genotype groups except Group 3, the majority of subjects identified themselves as white. Overall, the median cumulative therapeutic warfarin dose was 39 mg for all subjects (interquartile range [IR]: 27–52 mg), but sensitivity varied considerably from 10 to 110 mg, corresponding to 2–13 days of warfarin dosing.
Baseline characteristics of all subjects by genotype category.
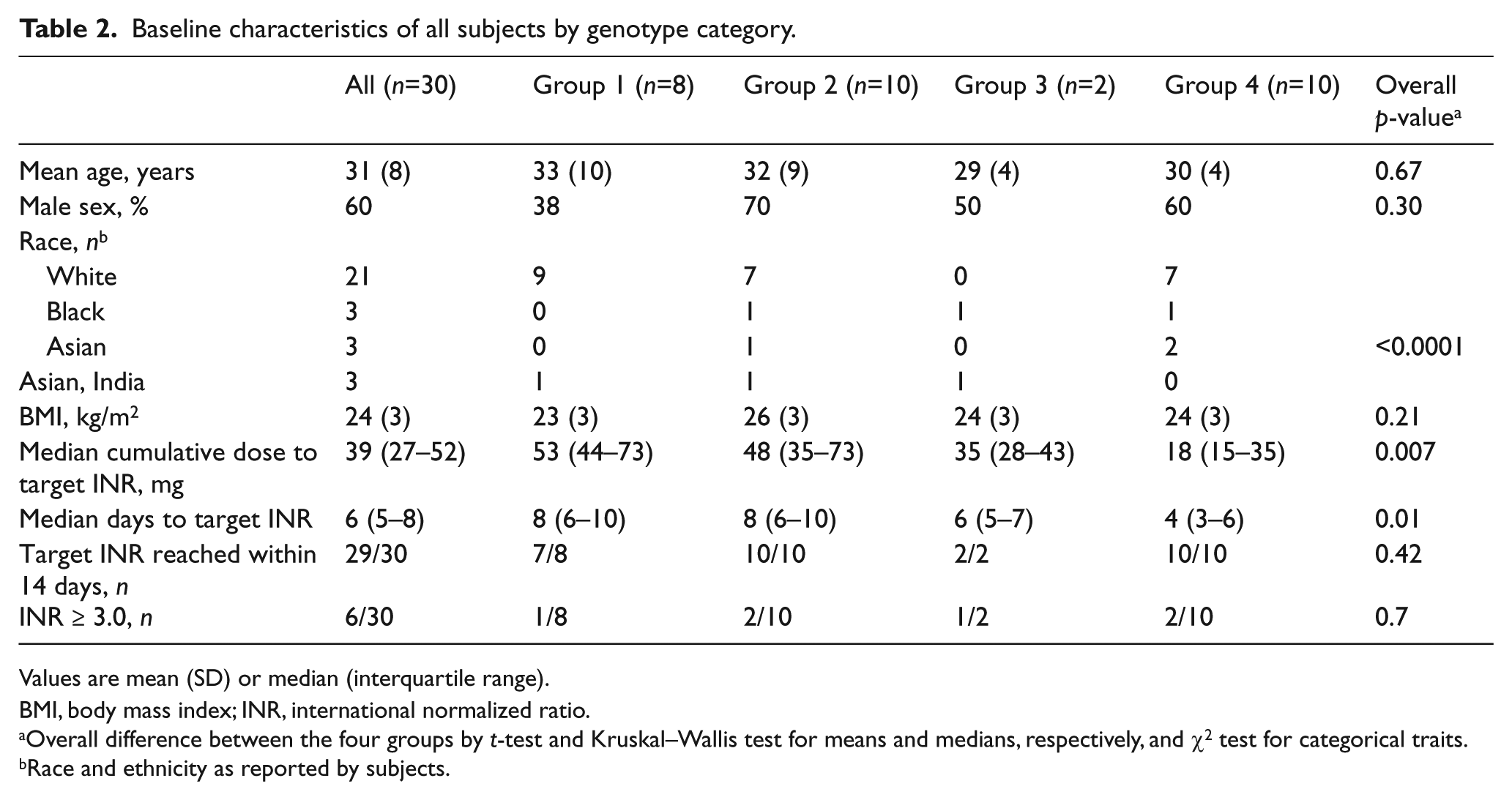
Values are mean (SD) or median (interquartile range).
BMI, body mass index; INR, international normalized ratio.
Overall difference between the four groups by t-test and Kruskal–Wallis test for means and medians, respectively, and χ2 test for categorical traits.
Race and ethnicity as reported by subjects.
One subject failed to reach a therapeutic INR despite 13 days of warfarin and the maximum dose of 15 mg daily, limited by protocol. This individual was wild type for both CYP2C9 (*1/*1) and VKORC1 (G/G) and did not carry any of the tested ‘resistant’ VKORC1 mutations. During the course of the study, the INR exceeded 3.0 in six subjects, and remained ≥ 3.0 for more than 1 day (2 days) in one subject. All possible combinations of wild-type and variant CYP2C9 and VKORC1 alleles were represented in these six subjects, as defined by the genotype groups (Group 1 = one subject, Group 2 = two subjects, Group 3 = one subject, and Group 4 = two subjects). The time required for recovery from the target (≥ 2) to baseline INR (≤ 1.2) varied from 1 to 9 days (mean 5 ± 2 days) among all subjects and did not differ significantly between groups. Subjects adhered to the vitamin K restricted diet until return to baseline INR. No adverse reactions and minor or major bleeding events occurred.
CYP2C9 and VKORC1 genotype groups
A significant difference in time required to reach an INR ≥ 2.0 for 2 consecutive days was identified between the four genotype groups, with subjects carrying variants in both CYP2C9 and VKORC1 (Group 4) requiring the fewest days to reach target anticoagulation intensity (median=4 days; overall p=0.01) (Figure 1). As illustrated in Figure 2, target anticoagulation intensity was achieved after a median cumulative warfarin dose of 52 (IR 44–72) mg in Group 1 subjects, 42 (IR 35–72) mg in Group 2 subjects, 35 (IR 27–42) mg in Group 3 subjects, and 15 (IR 15–35) mg in Group 4 subjects (overall p=0.007). This corresponded to median warfarin dosing periods ranging from 4 (IR 3–6) days in Group 4 subjects to 8 (IR 6–10) days in Group 1 subjects.
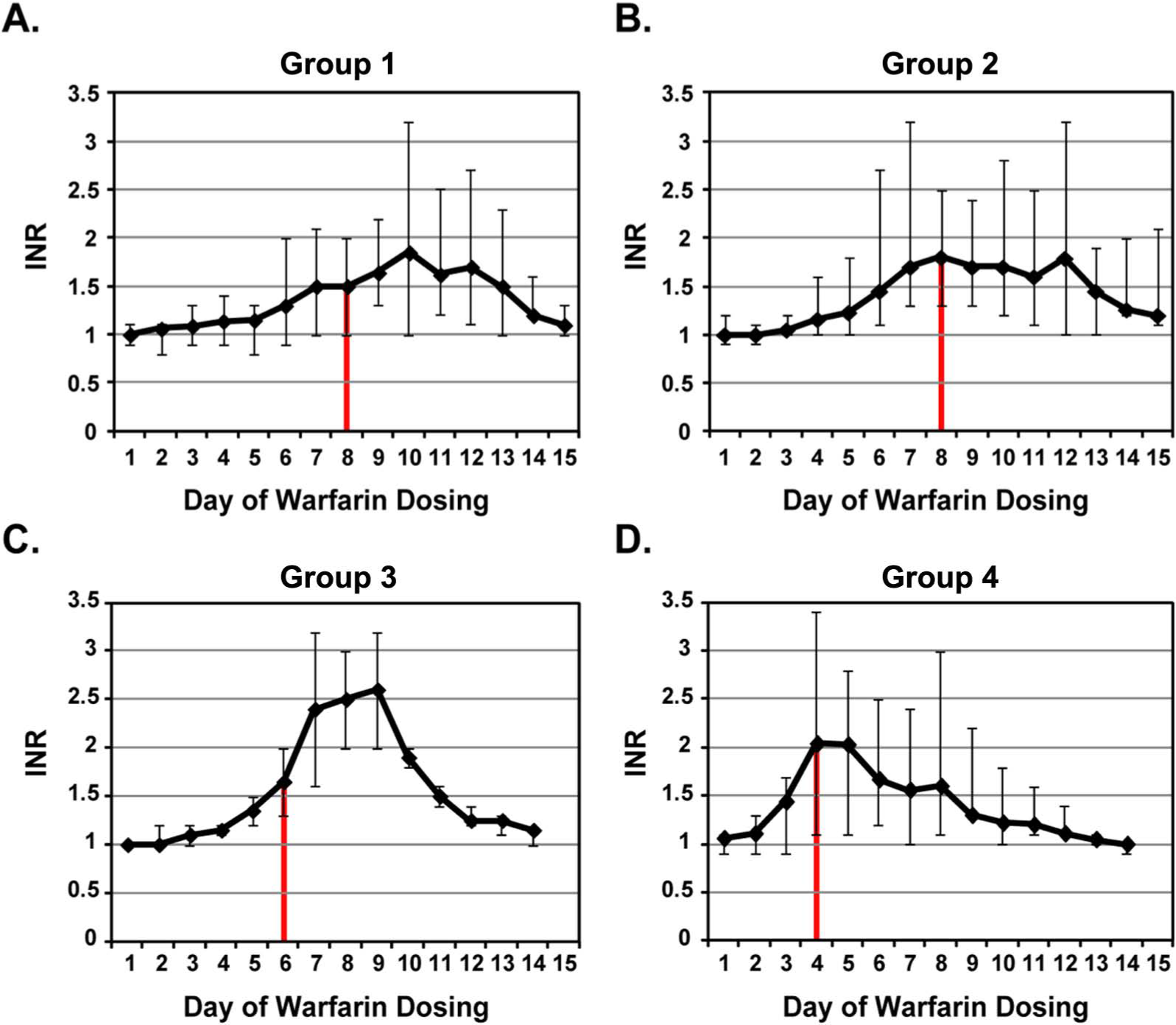
Time-based mean dose-response to warfarin by genotype group. The median days required to achieve an INR ≥ 2.0 for two consecutive days for (
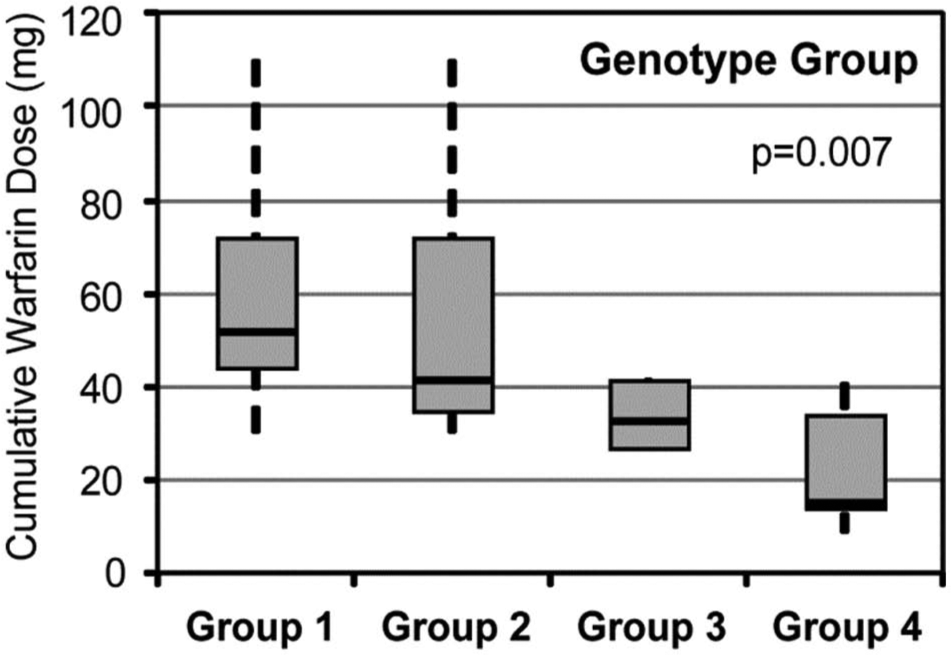
Median cumulative warfarin dose requirements by genotype group. Cumulative warfarin dose requirements are shown according genotype group. Median doses are indicated by the solid black bars; boxes indicate the interquartile range of warfarin doses; whiskers represent the range of observed doses falling within 1.5 times the interquartile range.
CYP2C9
The CYP2C9 extensive (EM), intermediate (IM), and poor (PM) metabolizer frequencies in the tested population were 63%, 27%, and 10%, respectively. CYP2C9 EM (*1/*1) subjects required a median cumulative therapeutic warfarin dose of 52 mg (IR 35–72 mg) to reach the target INR, as compared to CYP2C9 variant allele carriers (i.e. IMs and PMs) who required 17 (IR 15–42) and 27 (IR 15–35) mg, respectively (p=0.004; Figure 3A). These corresponded to median durations of 8 (IR 6–10), 3 (IR 3–7), and 5 (IR 3–5) days of warfarin to reach the target INR, respectively.
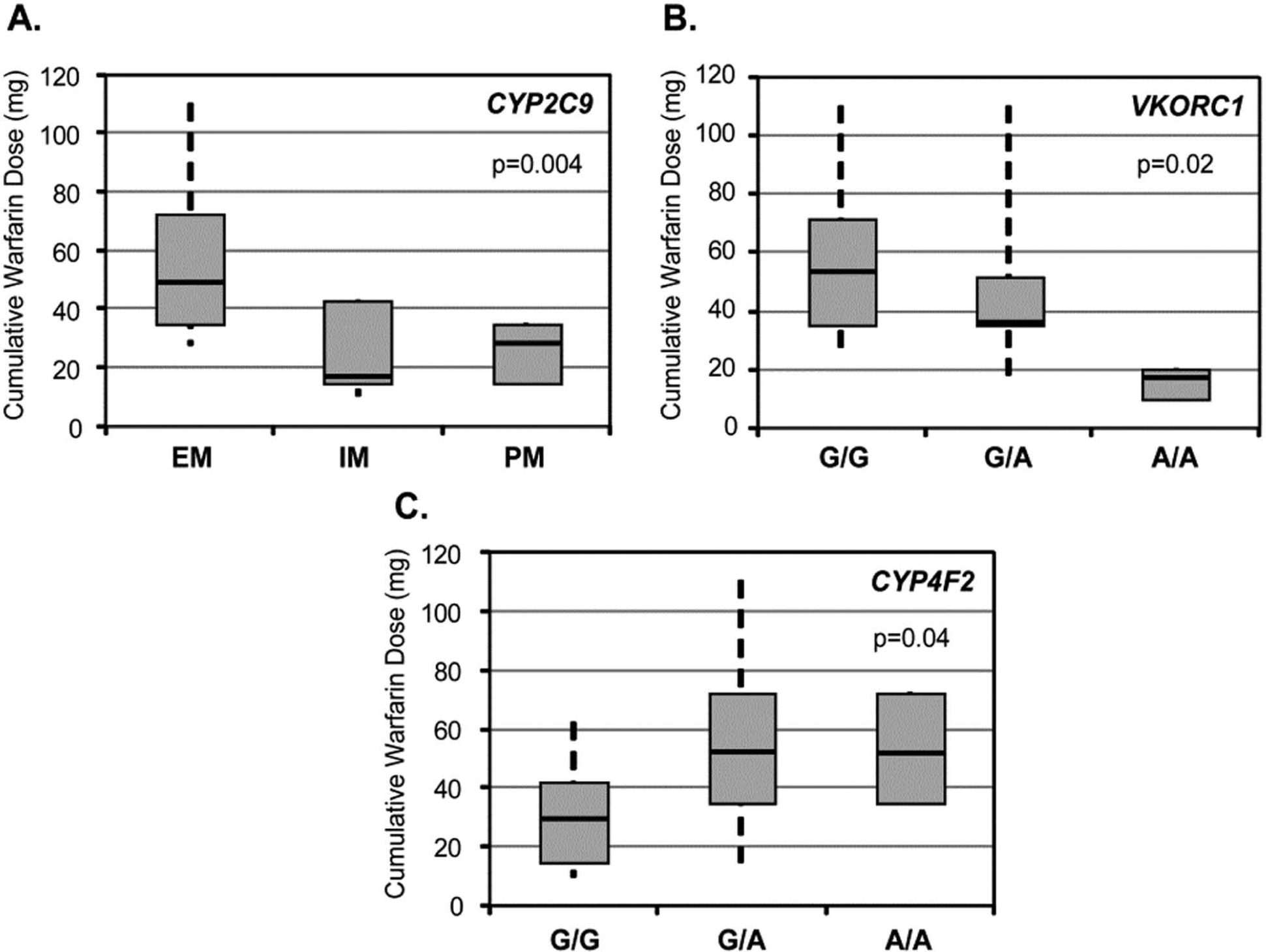
Median cumulative warfarin dose requirements by individual genotypes. Cumulative warfarin dose requirements are shown according to (
VKORC1
The VKORC1 -1639 G/G, G/A, and A/A genotype frequencies were 33%, 57% and 10%, respectively. No carriers of the ‘resistant’ VKORC1 coding region mutations were identified. VKORC1 -1639 G/G subjects demonstrated a median cumulative therapeutic dose requirement of 52 (IR 35–72) versus 35 (IR 35–53) mg in G/A subjects and 15 (IR 10–20) mg in subjects with the A/A genotype (overall p = 0.02; Figure 3B), corresponding to 8 (IR 6–10), 6 (IR 6–8) and 3 (IR 2–4) days of warfarin to reach the target INR, respectively.
CYP4F2
The CYP4F2 c.1297 G/G, G/A, and A/A genotype frequencies were 46%, 46% and 8%, respectively. Subjects with a CYP4F2 G/G genotype demonstrated a median dose requirement of 31 (IR 15–42) mg versus 52 (IR 35–72) and 54 (IR 35–72) mg among G/A and A/A subjects, respectively (overall p=0.04; Figure 3C). This corresponded to 5 (IR 3–7) days of warfarin for G/G subjects, compared to 8 (IR 6–10) days for G/A and A/A subjects.
Multivariable analyses
Table 3 summarizes the multivariate regression analyses. Dose variability attributable to demographic variables was small and not statistically significant (overall model adjusted R2 = 0.03; p=0.98). The cumulative variance significantly increased when the CYP2C9 genotype was included in the model (adjusted R2 = 0.57; overall p=0.004). After inclusion of the VKORC1 genotype, the dose variability attributable to both CYP2C9 and VKORC1 adjusted for age, race and sex was 64% (overall model p=0.006). The c.1297G>A (p.V433M) CYP4F2 variant contributed 1% to the variation in warfarin dose, bringing the total variation attributable to demographic and genetic factors to 65% (overall model p=0.017). None of the demographic and genetic predictors for the secondary outcome (time to restoration of baseline INR) reached statistical significance.
Explained variation in combined therapeutic warfarin dose models.
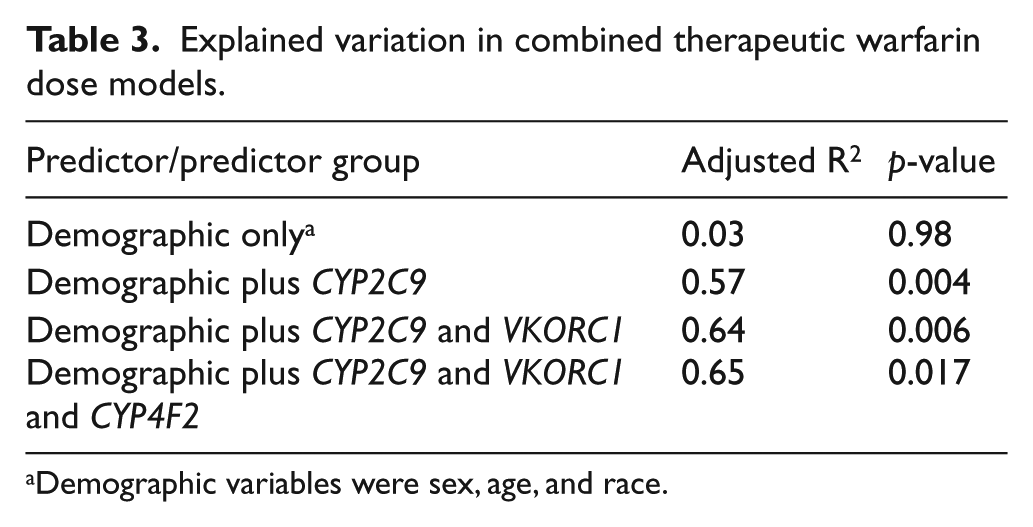
Demographic variables were sex, age, and race.
Discussion
Genotype-based warfarin dosing can improve the time in therapeutic range to 80%, 29 prompting recommendations that physicians consider genotyping at the inception of therapy.30,31 Beyond the early phase, the influence of CYP2C9 and VKORC1 genotypes is largely captured by INR measurements.32,33 The clinical utility of genotype-guided dosing is under investigation in the Clarification of Optimal Anticoagulation through Genetics (COAG) trial24,34 targeting the same INR level in patients with clinical indications for warfarin. The aim of this study was to determine the genetic contribution to warfarin dose variability when non-genetic variables are eliminated. We included 30 healthy volunteers, allowing for optimal control of clinical variables (diet, medications, comorbid conditions) that influence warfarin pharmacokinetics and pharmacodynamics. In the absence of these confounders, common CYP2C9 and VKORC1 variants lowered by nearly half the warfarin dose required to reach the anticoagulation intensity most commonly prescribed for clinical conditions.
After adjustment for sex, age and race, the variation in warfarin dose–response attributable to both CYP2C9 and VKORC1 (61%) exceeded that reported in clinical studies (30–40% dose variability).7,8 The contribution of CYP4F2 (p.V433M) to dose variability (R2 = 1.6%), accounting for CYP2C9 and VKORC1 status, was consistent with previous clinical reports.19,35 The contribution of demographic variables, including age and body weight, was no greater (3%) in our population than reported in patients with clinical indications for warfarin (15–22%; p=0.98 [NS]).10,16 This suggests that CYP2C9 and VKORC1 variants play a larger role in warfarin dose variability in patients with fewer comorbid conditions and medications and non-genetic factors contribute less.
There were two unexpected findings: (1) CYP2C9 and VKORC1 variant alleles were not associated with differences in time to normalization of INR after withdrawal of warfarin; and (2) CYP2C9 influenced warfarin dose–response variability at least as much as VKORC1; this conflicts with earlier observations suggesting that VKORC1 better predicts the early warfarin dose–response.17,36 Specifically, CYP2C9 EMs and VKORC1 -1639G/G subjects were anticoagulated after a mean cumulative dose of 60 mg. CYP2C9 IMs and PMs required half the dose of EMs (overall p=0.004), as did the heterozygous VKORC1 -1639A carriers (overall p=0.02) when compared to wild-type individuals.
The effect of genetic variation on INR recovery has not been directly studied previously, although the effects of CYP2C9 on warfarin clearance and VKORC1 as the direct target of warfarin suggest that variants may influence INR correction with administration of vitamin K. 37 Less common variants not adequately represented in our population or unidentified genetic factors could influence the pharmacokinetics and pharmacodynamics of warfarin. One subject carried the CYP2C9*8 allele, present in some 10–12% of African Americans and associated with lower warfarin dose requirement.27,38 Another subject did not reach the goal INR despite 13 days of warfarin and the maximum dose of 15 mg daily. This individual was wild type for both CYP2C9 (*1/*1) and VKORC1 (-1639G/G), the group with the highest cumulative dose requirement, but unidentified genetic factors may also contribute to the warfarin dose–response.
These observations must be interpreted cautiously because of the limited sample size, impeding power to distinguish differences in warfarin sensitivity. The small number of subjects, particularly in Group 3 (n=2), impeded detection of modest effects. Measuring the contribution of CYP2C9 and VKORC1 to warfarin dose–response in healthy subjects also has limited generalizability to patients commonly treated with warfarin. The primary outcome of this assessment (INR ≥ 2 for 2 consecutive days) does not reflect a steady state. Future studies should include measurements of serum vitamin K and warfarin metabolite levels to elucidate the mechanisms by which genetic variants influence warfarin pharmacology in the absence of thrombotic disorders. We relied on reported compliance with diet and avoidance of confounding medications, which may influence time to INR recovery. We did not include markers of ancestry to control for population stratification beyond self-reported race. Future research should include under-represented genotypes and broader ethnic representation, particularly since allele frequencies vary across ethnic groups. 28
The implications of this study are that clinical confounders mask the phenotypic effect of genotype and the value of genotype-guided dosing algorithms that input both genetic and clinical information varies across the spectrum of patients receiving warfarin. Dosing may rely more heavily on genetic than demographic factors, particularly for healthier patients taking few medications.
Footnotes
Acknowledgements
The authors thank Dr Louis M Aledort for his role as chair of the Data and Safety Monitoring Board for this project.
Declaration of conflicting interest
Dr Desnick serves as a site Principal Investigator for Clarification of Optimal Anticoagulation through Genetics (COAG) trial. Dr Scott is a member of the COAG Genotyping Committee. Dr Halperin has received consulting fees from Bayer AG HealthCare, Boehringer Ingelheim (serving on the Executive Steering Committee of GLORIA-AF Registry), Daiichi Sankyo, Johnson & Johnson, Ortho-McNeil-Janssen Pharmaceuticals, Sanofi-Aventis (having served on Clinical Trial Operations Committee for the PALLAS trial), Biotronik, Inc. (as co-Chair of the Steering Committee for the IMPACT clinical trial), and Medtronic (as a member of the Steering Committee for the REVEAL-AF study), and is an investigator in the COAG trial.
Funding
This work was supported by the Icahn School of Medicine at Mount Sinai [GCO# 08-1442].
