Abstract
Deep learning models are necessary in the field of healthcare for the diagnosis of cardiac rhythm diseases since the conventional ECG classification is based on hand-crafted feature engineering and traditional machine learning. Nevertheless, CNN and BiLSTM architectures provide automatic feature learning, enhancing ECG classification accuracy. The current research work puts forward a framework integrating CNN with CBAM and BiLSTM layers for the purpose of extracting valuable features and classifying ECG signals. The model classifies heartbeats according to the AAMI EC57 standard into 5 categories: normal beats (N), supraventricular ectopic beats (S), ventricular ectopic beats (V), fusion beats (F), and unknown beats (Q). To tackle uneven class distributions, SMOTE synthesizes new samples, making the model more robust. Evaluation on MIT-BIH arrhythmia database yields remarkable results with 99.20% accuracy, 97.50% sensitivity, 99.81% specificity, and 98.29% mean F1 score. Deep learning methods have great potential to alleviate clinicians’ workload and improve diagnostic accuracy of cardiac diseases.
Introduction
Cardiovascular diseases (CVD) according to the World Health Organization (WHO) account for 32% of total human mortality worldwide in 2019, which represents over 85% of the figure of World Health Organization. 1 The traditional diagnosis relies on clinical history and physical examination, using a uniform set of quantifiable medical criteria for the classification of disease according to Zhang et al., 2 Haberl et al., 3 and Du and Jose 4 But this technique appears to be conventional as it entails a detailed investigation of various information, which can take more than a single doctor as stated by Ebrahimi et al. 5 Poor countries are particularly challenged as they do not have adequate healthcare professionals and facilities in hospitals. Thus, there is a need for an effective and low-cost computer-aided diagnosis system (CADS) that can perform adequately in environments where resources are limited. CADS functions by reading physiological signals to monitor and evaluate organ function based on Du and Jose 4 and Ebrahimi et al. 5 CADS is needed to provide affordable healthcare options, especially in poor communities. Deriving useful and reliable information from ECG records is the most significant problem CADS face. Each cardiac cycle has individualized electrical depolarization-repolarization patterns, which all add to the various electrical activities of the heart based on Zhang et al. 2
Deciphering these complex patterns is crucial in identifying abnormalities and making corresponding clinical decisions. The complexities of understanding how such electrical events interact within the cardiac cycle add a further level of complexity. Hence, there is an acute need to increase efficiency and performance of cardiology’s computerized diagnostic equipment by developing sophisticated algorithms capable of extracting valuable information from ECG signals with utmost precision. Normal heart rhythm features are significantly variable in morphology from individual to individual and even within the same individual under different conditions, where QRS complex configuration and R-R interval changes may be noted as revealed by Karpagachelvi et al. 6 The classification of arrhythmias based on the ECG signal remains challenging due to the presence of high intra-class variability and inter-class proximity among different cardiac rhythm abnormalities. Some arrhythmias exhibit very small waveform differences and are therefore challenging to classify as distinct from normal heartbeats. The addition of noise, artifacts, and individual physiological fluctuations further complicates the automation of the classification. Ex-perienced doctors utilize the variation in rhythmic pulse shape throughout a heart rhythm analysis to spot abnormal heart activity. However, automating the process is difficult for computerized systems due to interference from external noise. Unlike human clinicians, computerized systems must struggle to maintain accuracy and validity amidst numerous exogenous influences and imbalances between data classes, and hence the classification of anomalies in heart rate patterns becomes a daunting task.
Therefore, there is a need to improve the performance of automatic systems in identifying abnormalities in cardiac functions. An ECG, as a dynamic physiological signal depicting the complex electrical activity of the heart, conveys important information through its PQRST waveform according to Karpagachelvi et al., 6 Abdul Jamil et al., 7 and Bogdanov et al. 8 Sinus rhythm, a sign of coordinated cardiac activity, and P wave, QRS complex, S wave, T wave amplitudes, and intervals (RR, PR, QT, and QRS complex) are crucial indicators to ascertain the heart strength as Du and Jose 4 emphasize in their study. Any abnormality from these normal values can suggest an abnormal cardiac cycle, which means that there is something wrong with the heart. Since each type of arrhythmia manifests itself by specific alterations in these waveforms and intervals, an effective classification model must be able to accurately distinguish normal and abnormal ECG signals based on their morphological and temporal characteristics. These models can thus help physicians quickly make a reliable diagnosis, especially in emergency settings such as intensive care units (ICUs). The current research follows the development of an effective and lightweight automated diagnosis system that may support cardiologists by introducing a smart deep-learning method that is both swift and cost-effective and simultaneously minimizes the number of incorrect arrhythmias diagnosed. The proposed model that includes convolutional neural networks (CNN) together with recurrent neural networks (RNN), namely long-term memory (LSTM), for general classification of various cardiac arrhythmias. By utilizing deep learning, the approach is supposed to enhance the accuracy of arrhythmia classification and achieve real-time, scalable, and efficient ECG-based diagnosis. The ECG recordings used in this study were obtained from the MIT-BIH Arrhythmia Database.
Although several previous studies have explored the combined use of CNN-BiLSTM networks with attention mechanisms for arrhythmia classification from ECG signals, these approaches still exhibit structural and practical limitations that hinder their real-world clinical applicability. On the one hand, most of these models are designed and evaluated on controlled datasets such as MIT-BIH, without incorporating robust techniques for handling class imbalance. This often results in significantly reduced performance when dealing with rare or minority classes. On the other hand, the attention mechanisms employed are typically global and poorly suited to capturing the fine temporal and spatial complexity of ECG signals, particularly when identifying local multichannel interactions and a typical morphological variation. Moreover, these architectures often sacrifice computational efficiency to achieve higher accuracy, thereby limiting their deployment on real-time or embedded systems. To address these shortcomings, the model proposed in this study adopts a synergistic strategy that integrates multi-scale CNN blocks to effectively extract local features at different resolutions, a dual attention mechanism (spatial and channel-based) for finer contextual weighting, and a BiLSTM layer to model long-term temporal dependencies. In addition, the use of the Synthetic Minority Oversampling Technique (SMOTE) helps to effectively mitigate class distribution asymmetry, ensuring more balanced learning and better model generalization. This novel combination aims to overcome the limitations of existing approaches by simultaneously enhancing robustness, accuracy, and computational efficiency.
In summary, the main contributions and innovations of this article can be briefly described as follows:
(i) Using convolutional neural networks coupled with channel attention mechanisms and long-term memory networks has led to an exemplary contribution to the classification of electrocardiogram signals.
(ii) The approach uses the synthetic minority oversampling technique (SMOTE), which generates synthetic samples of a minority class aimed at balancing the skewed distribution of classes that characterize most ECG datasets, making the model competent to handle these problems.
The structure of this paper proceeds as follows: Section II undertakes an in depth review of the relevant literature, contextualizing the proposed system within the existing body of work. Section III delves deeper into the nuances of the system architecture, providing a detailed exploration of its design and operational features. Going further, Section IV presents the experimental results, making comparisons with literature results to verify the effectiveness of the proposed approach. Section V discusses the limitations of the current study, addressing the challenges encountered and the potential areas for improvement in future research. The concluding Section VI summarizes the current state of the proposed technique, elucidating its advantages and disadvantages, thereby providing a holistic perspective on the study results.
Literature Review
In recent years, CNN has gained extensive attention in biomedical signal processing, especially for electrocardiogram (ECG) signal classification. The stability and effectiveness of CNN in image analysis tasks in these studies9,10 have encouraged their adaptation to ECG signal analysis, acknowledging their ability to detect intricate patterns associated with various cardiac abnormalities. Hierarchical feature extraction ability of CNNs has been utilized in numerous research papers to learn automatically discriminative features from ECG signals without any preprocessing, and this has been beneficial in producing effective and correct classification models such as Porumb et al. 11 and Baloglu et al. 12 Use of deep architectures and the ability to use several layers in CNNs has also proved to be very useful, allowing the network to learn the complex hierarchical temporal pattern representations from the ECG signals effectively. 13 Proposed a comprehensive 34-layer end-to-end deep neural network to differentiate 12 categories of rhythms. Their network was extensively evaluated using a new large single-lead ECG database and reported satisfactory results with an impressive area under the curve (AUC) of 97%. Similarly, Zhang et al. 14 proposed a 1-dimensional CNN of 12 layers tailored for classifying a single 1-lead heartbeat signal into 5 heart disease classes. Validation of their method using the MIT/BIH arrhythmia database yielded remarkable performance metrics like 97.7% positive predictive value, 97.6% sensitivity, and an F1 score of 97.6%. Further, Zhang et al. 15 presented a comprehensive review of deep residual learning for ECG signal classification and described the superiority of CNNs with fewer layers in extracting suitable features. Moreover, techniques such as transfer learning have also enhanced the accuracy of CNNs for ECG signal classification issues as shown in the study of Cote-Allard et al. 16 However, there are still concerns like the need for large collections of labeled data, interpretability of deep models, and overfitting risk as illustrated by the research of Liu et al. 17 But studies continue to advance and refine deep CNN structures for ECG signal classification to contribute to the development of automated diagnostic tools in cardiology. On the contrary, more recent studies have especially weighed the resilience associated with the blend of CNNs with LSTMs in ECG signature classification. 18 Study showed utmost accuracy when they employed hybrid CNN-LSTM to detect arrhythmia and is better at identifying spatial and temporal features of signal ECG compared with employing just LSTM. But Mukhametkaly et al. 19 took that even further in detecting anomalies, specifying the coherence between spatial features which are learned through CNN and temporal dependencies embedded within LSTM. For instance, other recent studies like Alamatsaz et al. 20 and Yildirim et al. 21 have demonstrated the efficiency of CNN-LSTM based on ECG readings with better accuracy and responsiveness for a variety of heart ailments. Specifically, the experiments by Warrick and Homsi 22 used multimodal approaches that demonstrate the effectiveness of joint usage of CNN and LSTM networks to detect Cardiac arrhythmia. The research by Siami-Namini et al. 23 is a comparison of BiLSTM and LSTM RNN structures to determine the performance impacts of additional training layers. The result is that BiLSTM trained with more data outperforms normal LSTM and ARIMA models in capturing complex temporal patterns. Otherwise, Wang and Li 24 present an automated framework that combines CNN and BiLSTM for the classification of ECG signals. Their cascade model achieves a higher weighted F1 score of 0.82, which is very effective in classifying heterogeneous ECG signals. Cheng et al. 25 also propose a heartbeat network that integrates a 24-layer DCNN and BiLSTM framework. Their model achieves a high accuracy rate of 89.3% and an F1 score of 0.891, showing its effectiveness in classifying heart rhythms from ECG signals. The above development validates the suitability of CNN-BiLSTM combination for a comprehensive solution to ECG signal classification complexities. In parallel, the integration of attention mechanisms such as CAM or CBAM with CNNs has revolutionized signal processing by enabling focus on significant features within complex data sets. The work of Woo et al. 26 and Wang et al. 27 confirmed the effectiveness of this approach for medical signal analysis, offering an optimal framework for anomaly detection. Our approach builds upon these foundations while overcoming the limitations of competing models. Unlike Chorney 28 model, which neglects local spatial relationships and class imbalances, we integrate sophisticated attention mechanisms that effectively capture significant features. Our architecture avoids the excessive computational complexity of Pandey et al. 29 model and resolves the overfitting problems identified by Yuan 30 through a nested framework combining the spatial extraction capabilities of CNNs and the temporal modeling of BiLSTMs.
Unlike the M2MASC integration by Pandey et al., 31 our architecture maintains optimal efficiency while effectively addressing the data imbalances that limit generalization in the study by Xu et al. 32 In summary, our innovative application combining CNNs, attention mechanisms, and BiLSTM networks constitutes an exemplary contribution to ECG signal classification. Our nested framework effectively recognizes complex patterns through the spatial extraction capabilities of CNNs, the attentional enrichment from CBAM, and the temporal dependency modeling by BiLSTM.
Methodology of Arrhythmia Classification
In this section, we will elucidate our data preprocessing methodology provide a concise overview of the database employed and finally apply our model for ECG classification.
ECG Dataset
This study uses the MIT-BIH arrhythmia dataset to implement and validate the proposed methodology. The MIT-BIH includes 48 readings obtained by recording the cardiac activity of 47 subjects (25 men aged 32-89 years and 22 women aged 23-89 years). Each patient contributes to both readings. It lasts approximately 30 minutes carefully selected from the numerous 24-hour recordings as Goldberger et al. 33 and Alqudah 34 consisting of twice Lead ECG, Limb Lead II, and (often) V5. The sampling rate of ECG signals is 360 Hz and an annotation for each beat occurs during a reading. The dataset marks 2 types of beats, but we will limit our attention to 1 type, subdivided into 5 classes, following the AAMI recommendation. These categories are N (normal), S (premature supraventricular beat), V (premature ventricular contraction), F (merge of ventricular and normal beat), and Q (unclassifiable/unknown beat). The detailed dataset presented in Table 1 provides useful information for research and investigation into the classification of heart diseases and arrhythmia.
Annotations in the MITBIH dataset labeled following the AAMI standard, with consolidated classes including N, S, V, F, and Q and several beats used in each class.

Preprocessing
Noise Filtering
Noise reduction in electrocardiogram (ECG) signals represents a crucial preprocessing step aimed at enhancing the accuracy of diagnostic procedures. The utilization of the Discrete Wavelet Transform (DWT) provides an effective approach, enabling a time-frequency decomposition of signals to identify and eliminate unwanted noise components while preserving essential cardiac information. This method has proven effective in attenuating various types of noise, including baseline wander and muscle artifacts like in Aqil et al. 35 study. Comparative studies with other noise suppression techniques, such as finite impulse response (FIR) filters, adaptive filters, and empirical mode decomposition (EMD), have underscored the superiority of wavelet transform in preserving signal characteristics while effectively removing noise demonstrated by Rout et al. 36 The notable work conducted by Ali 37 highlighted the efficacy of DWT in noise elimination, making it a preferred choice in the preprocessing pipeline for accurate ECG signal analysis. The integration of DWT with a customized wavelet like the “db5″ wavelet and a decomposition level of 4 holds significant promise. The “db5” wavelet, known for its ability to preserve signal features while suppressing noise, ensures robust feature extraction essential for accurate classification such as Sawant and Patil. 38 This balanced approach, aiming to preserve both global and local signal characteristics, strengthens the effectiveness of ECG signal classification algorithms, providing clinicians with more reliable diagnostic information, and facilitating rapid interventions in patient care. Figure 1 illustrates the ECG signal before and after applying preprocessing techniques with DWT.

ECG signal of patient 100: (a) before and (b) after denoising.
Segmentation
However, before performing ECG classification, each heartbeat must be separated from the overall ECG signal. The identification should be pinpoint accurate on the QRS wave and the corresponding fiducial point for each heartbeat pulse. The Pan-Tompkins algorithm created by Pan and Tompkins 39 is a commonly used method for detecting QRS complexes in electrocardiogram (ECG) signals. Additionally, Suresh et al. 40 introduced SeismoNet, a fully convolutional deep neural network designed to precisely identify R peaks for reliable SCG signal monitoring. SeismoNet undergoes end-to-end training to accurately detect R peak positions even in the presence of noise. Although several exact (99%) methods of QRS beat and fiducial point setting can be found in many papers, in this paper no attempt was made to provide new insights on the issue.
However, instead, the annotated R-peak position serves as the fiducial element, and the ECG gets split into a series of beats. Such an approach makes it easy to rate one’s work in direct comparison to other existing works. Fixed-size ECG signal (300 samples) with 99 samples before R-peak and 201 samples following it. The selection of these particular sample points is such that they describe the essential aspects of a heartbeat. Segmentation is depicted in Figure 2.
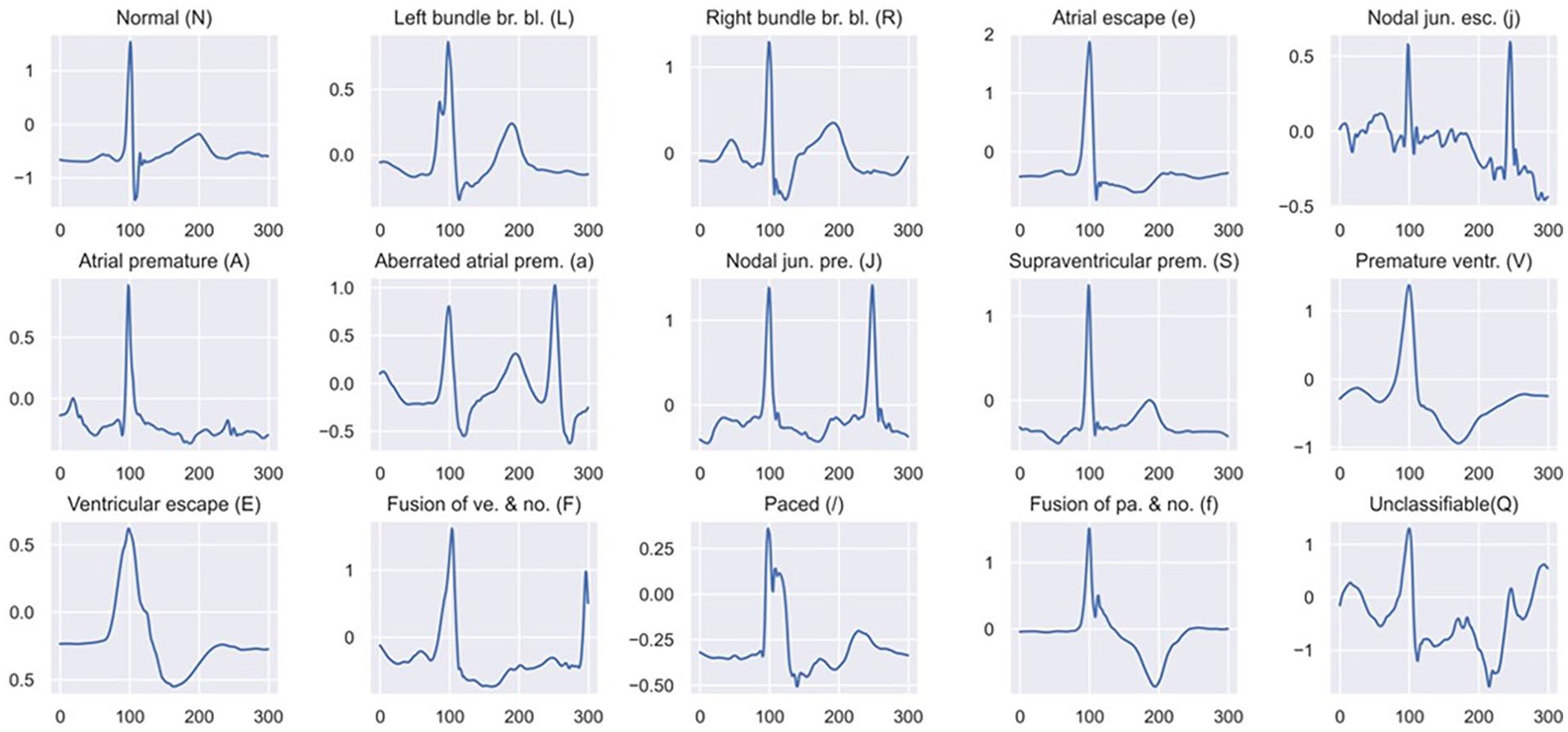
Segmented ECG beat samples for each heart type in the MITBIH.
Through the segmentation process, a total of 109 305 ECG beats were effectively extracted. The distribution of ECG beats across various classes is comprehensively presented in Table 1. These beats were divided into a training set comprising 80% of the beats and a test set containing the remaining 20% of segmented raw ECG signals. This partitioning ensures a balanced distribution of different arrhythmia classes, facilitating robust model training and evaluation. The distribution of ECG beats across various classes with both training and testing sets is comprehensively presented in Figure 3.

Arrangement of both training and testing data within distinct classes.
Data Augmentation
To achieve an even class distribution in the dataset, a multi-dimensional method is used with caution, making use of a combination of under-sampling, and the Synthetic Minority Oversampling Technique (SMOTE). SMOTE is a significant contributor toward balancing the inherent imbalances in the dataset such as Pandey and Janghel 41 and Kummer et al. 42 Specifically, for Class N (normal), a statistically significant, conscious random sample of 50 000 instances is maintained, which gives an effective representation of this majority class. However, realizing the importance of equitable representation of all classes, further attempts are made to balance Classes S (supraventricular premature beat), V (premature ventricular contraction), F (fusion of ventricular and normal beat), and Q (unclassifiable/unknown beat). To do so, the SMOTE process is run systematically and diligently oversampling each of the aforementioned minority classes until they have all reached the standard number of 50 000 data points.
The very tangible outcome of this diligently balancing procedure is actually shown in Figure 4 here, providing visual illustration of class-balanced distribution. This strategic strategy transcends straightforward numerical balancing: its final end is to work toward a better representative and balanced presentation of each class in the dataset. Counteracting the errors that are invariably associated with imbalanced datasets, such as model learning bias and unbalanced predictions, this approach aims to ensure an environment conductive to augmented model generalization. The enhanced class balance results in a more robust and stable training set for machine learning algorithms, thereby enhancing their performance and predictability. Such refined method considers diversity and richness in the dataset while aiming to perfect the process of learning and fortify the collective power of machine learning models during implementation in filtered data.

Training data balanced by SMOTE.
Proposed Model Architecture
One of the latest and most advanced artificial intelligence technologies is deep learning which rose in tandem with the proliferation of big data sets like Porumb et al. 11 and Pandey and Janghel 41 Deep learning differs with its particular structure made up of various consecutive layered structures intended to provide for progressively more advanced input processing.2,5 Mimicking the human brain in its complex structure through the innovative 43 approach. Deep learning is very powerful because it can automatically learn hierarchical representations that enable the detection of intricate patterns and features within enormous and complex datasets. Therefore, deep learning has established itself as an essential tool in numerous disciplines, including pattern understanding in pictures, speech comprehension, and most importantly in the examination of biomedical data, such as ECGs, wherein its potential to highlight features and categorization has proven very beneficial.
CNN
Initially developed in the 1980s, CNNs gained significant traction with improved deep learning theories and computational resources in the 2000s, with notable architecture like LeNet-5 paving the way for advancements like Lecun et al. 44 Despite being traditionally applied to 2-dimensional image data, CNNs have also found utility in processing 1-dimensional signals, 45 such as electrocardiogram (ECG) data utilized in this study. In adapting CNNs to 1-dimensional inputs, we capitalized on their ability to automatically learn spatial features without explicit feature selection. However, since ECG signals exhibit strong temporal characteristics, a simple CNN architecture may struggle to capture temporal features effectively. To address this limitation, we used a deep CNN architecture, comprising four 1-dimensional convolution layers followed by 3 pooling layers (2 max pooling layers and 1 average pooling layer). By incorporating pooling layers after the convolutional layers, we sought to reduce the dimensionality of the feature maps.
LSTM
LSTM represents a type of RNN particularly adept at processing time sequences similar to Greff et al. 46 RNNs are designed to assess current information based on preceding data, making them effective for timing-related tasks. However, they are susceptible to the vanishing gradient problem as the network grows deeper. LSTM addresses this issue by enhancing memory retention and selectively preserving pertinent information while mitigating the problem of gradient disappearance and explosion inherent in RNNs corresponding to Gers et al. 47
The internal structure of an LSTM memory block, depicted in Figure 5, comprises Ct and Ct−1 representing the current and previous neuronal states, respectively. Additionally, ht and ht−1 denote the output of the unit at the current and previous times, respectively, with xt serving as the network input. The LSTM forget gate, ft, regulates the forgotten information via the sigmoid function.
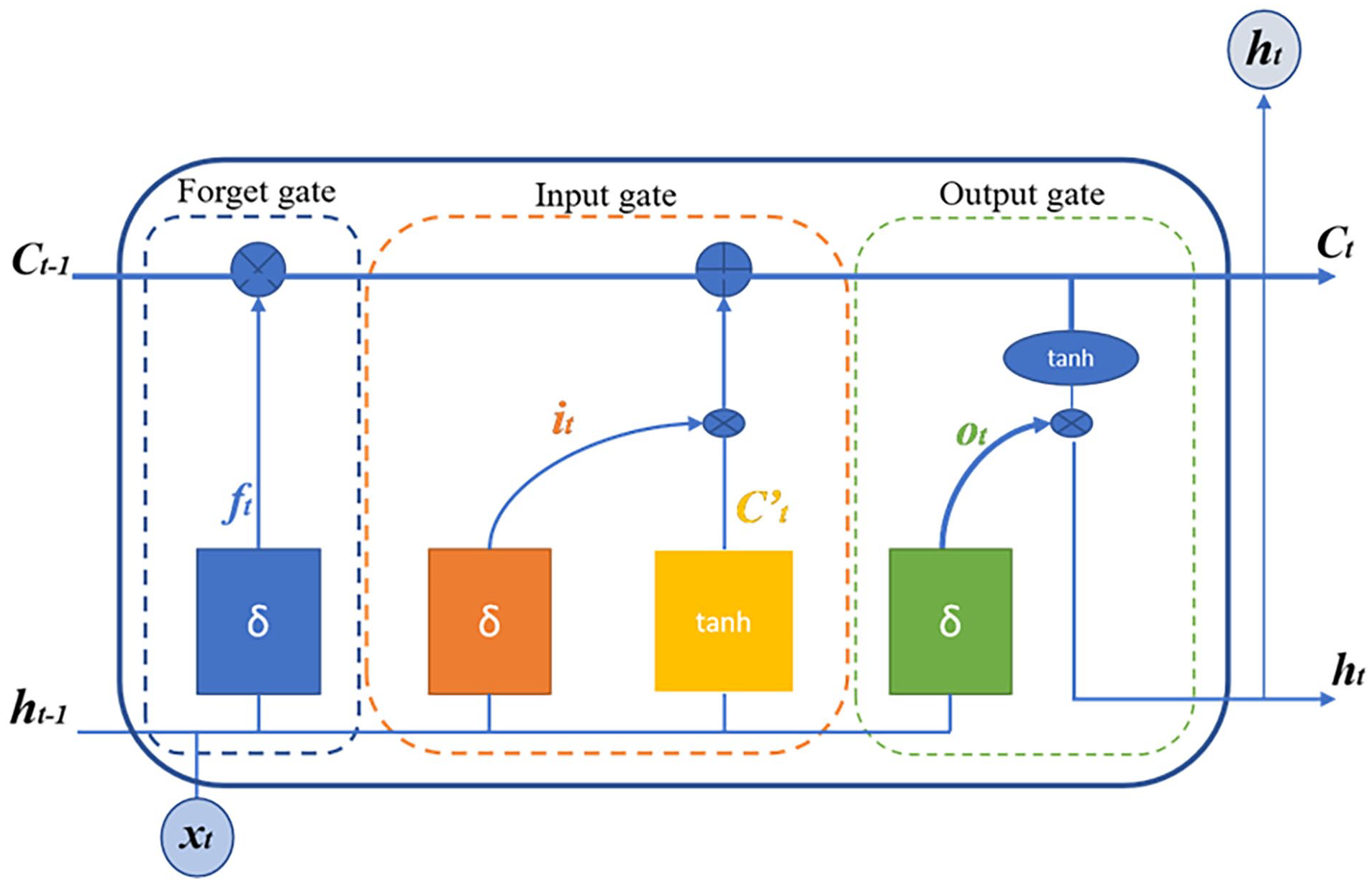
Architectural composition of an LSTM unit.
At the same time, it acts as the input gate, setting the threshold and utilizing the tanh function to determine the neuron state. Lastly, Ot is the output gate, controlling the output information through the sigmoid function, with corresponding mathematical expressions governing their operations.
The forget gate equation:

This equation defines how much of the previous memory should be retained or forgotten by applying the sigmoid function to a weighted sum of the previous output ht−1 and the current input xt, with a bias term Rf .
The input gate equation:

This equation regulates the intake of new information by using the sigmoid function, determining how much of the new information should be incorporated into the memory based on the previous state ht−1 and the current input xt.
The output gate equation:

Here, the output gate equation defines how much of the memory will be output to the next layer, controlling the flow of information using the sigmoid function.
The candidate neuron state equation:

This equation computes the candidate neuron state
In the LSTM framework, the weight matrices Lf, Li, Lo, and Lc are attributed to the forget gate, input gate, output gate, and neuron state matrix, respectively, while Rf, Ri, Ro, and Rc denote the offsets for each gate. These matrices play pivotal roles in determining the flow of information within the LSTM unit, facilitating memory management and information retention. Specifically, Lf regulates the forgetting of unnecessary information, Li governs the intake of new information, Lo controls the output of information, and Lc manages the neuron state. Additionally, the expressions for the current state of the neuron and the output of the cell are as follows:

This equation updates the current neuron state Ct by combining the previous memory Ct−1, weighted by the forget gate ft, and the candidate state
The output of the LSTM cell is computed as:

This equation calculates the output ht by applying the output gate Ot to the current neuron state Ct, with the tanh function ensuring that the output remains bounded. In the proposed architecture, 1 Bidirectional Long Short-Term Memory (BiLSTM) layer is utilized to capture not only the context preceding a given point but also the context following it, allowing a more complete understanding of temporal data.
CBAM
The attention mechanism, initially inspired by the study of human vision, has been gradually integrated into the field of deep learning algorithms. Its operation relies on adjusting the network parameters through the generation and allocation of weights, thereby efficiently allocating computational resources to more important tasks like in Shan et al. 48 study. This mechanism plays a crucial role in focusing on specific parts of input data, enhancing model performance by emphasizing the most relevant information for the task at hand. In this study, the Convolutional Block Attention Module (CBAM) is employed to enhance the performance of the arrhythmia classification model. CBAM, introduced by Woo et al., 26 is a multi-attention mechanism combining channel and spatial attention modules. CBAM considers both channel and spatial attention simultaneously, resulting in improved outcomes, unlike attention mechanisms that target only one aspect. The architecture of CBAM is depicted in Figure 6.

The comprehensive structure of the convolutional block attention module (CBAM): (a) channel-attention module (CAM) and (b) spatial-attention module (SAM).
The structure of the channel attention module, illustrated in Figure 6a, begins with subjecting the input features to global maximum pooling and global average pooling operations. These pooled features are then simultaneously input into a multi-layer perceptron network.
Following this, the output features are successively summed and activated using the sigmoid function to derive the channel attention feature weight. Finally, this weight is applied to the input features through multiplication to generate the features essential for the spatial attention module. The spatial attention module’s architecture, depicted in Figure 6b, begins with subjecting the input feature map to channel-based global maximum pooling and global average pooling operations. The resulting feature map undergoes channel concatenation. Subsequently, after convolution and sigmoid activation operations, the spatial attention feature weight is produced. This weight is then multiplied by the input feature map to yield the final feature representation. In the Convolutional Block Attention Module (CBAM), the channel attention mechanism plays a crucial role in enhancing the model’s performance, especially in the context of 1D signals such as electrocardiogram (ECG) data. Considering the inherently 1-dimensional nature of ECG signals and their typical single lead recording setup, it’s conceivable that the spatial attention module within CBAM might not offer significant relevance or effectiveness in such a context. Therefore, the focus is primarily on leveraging the channel attention mechanism to capture and emphasize important features within the ECG signal. By selectively attending to specific channels or features across the input signal, the channel attention mechanism enables the model to adaptively adjust its attention to relevant information, enhancing its ability to discriminate between different arrhythmia classes, or other relevant diagnostic categories. Consequently, by leveraging the channel attention component of CBAM tailored to the characteristics of ECG signals, the model can effectively extract discriminative features and improve its classification performance for tasks related to ECG analysis and diagnosis.
The hybrid CNN-BILSTM model
This paper proposes a novel deep learning method that combines convolutional neural networks (CNNs) with channel and spatial attention mechanisms, and then bidirectional long short-term memory (BiLSTM) structures with the main aim of improving the diagnostic accuracy of various cardiac arrhythmias. The holistic diagnostic model ensures complete coverage, highlighting the model’s ability to produce correct estimates of a variety of cardiac abnormalities.
As shown in Figure 7, the schematic illustration of the proposed model presents its architecture as follows: starting with an input layer for ECG signal processing, having a channel allocated to receive sequences of 300 data points. The model integrates sequential convolutional layers, the first one composed of 16 filters with 21 × 1 convolution kernels and augmented with channel attention mechanisms (CAMs) to provide finer channel features. Additional max pooling operations with a 3 × 1 kernel and stride of 2 are used to perform further downsampling, and finally, a second convolutional layer consisting of 32 filters with channel attention mechanisms. Further feature extraction occurs via another max pooling, a third convolutional layer with 64 filters, and then the next average pooling. Inserting a fourth convolutional layer with 128 filters and adding attention mechanisms introduces varied kernel sizes, providing more flexibility. BiLSTM layers are also followed, effective in recognizing sequential dependency in temporal space. Dropout layers are inserted at critical locations to avoid overfitting, with a dense layer with 128 nodes and a rectified linear unit (ReLU) activation refining the extracted representations. The model ends with a dense layer with softmax activation, producing a multi-class classification output. In terms of trainable parameters, the architecture achieves an optimal balance between modeling capacity and overfitting risk. The model contains exactly 1 575 717 trainable parameters, with the majority concentrated in the dense layer and deep convolutional layers. This parametrization enables robust feature extraction while maintaining manageable computational complexity. For hyperparameter initialization, a meticulous approach based on empirical trials and rigorous cross validation was adopted. The convolutional layer weights are initialized using the He et al. 49 method, particularly suited for ReLU activations, while BiLSTM cells are initialized using a uniform distribution to facilitate convergence. An initial learning rate of 0.001 with the Adam optimization algorithm was found to be optimal for model convergence. Dropout rates are set at 0.2, following a progressive strategy that preserves critical information while effectively preventing overfitting. This novel architecture, integrating convolutional and LSTM layers with different attention and kernel sizes, enables robust classification of the ECG signal, which reflects its efficiency, and adaptability in discriminating complex physiological patterns.

Design of the envisioned deep learning model.
Results and Discussion
Our proposed model is discussed in detail, covering both its architecture and configuration. A comprehensive analysis of its performance parameters follows, highlighting its unique qualities. To ensure relevance, we compare the results of our model against others to assess its suitability for the task.
Results
To judge any system with AI, it is important to compare its performance against new data. The performance of the proposed models is measured by comparing the original annotations of ECG beats against the annotations predicted by the models. The accuracy, sensitivity, accuracy, and specificity are then calculated from these annotations, and this provides information about the correctness of the diagnosis. The calculation of such measures needs true positive (TP), false positive (FP), false negative (FN), and true negative (TN) in order to properly evaluate the diagnostic capabilities of the system.
The accuracy equation:

This equation calculates the overall accuracy of the system by measuring the proportion of correct predictions (TP and TN) out of all predictions made.
The sensitivity equation:

Sensitivity, also known as recall, measures the proportion of actual positive cases (TP) that were correctly identified by the system, indicating how well the system detects positive instances.
The specificity equation:
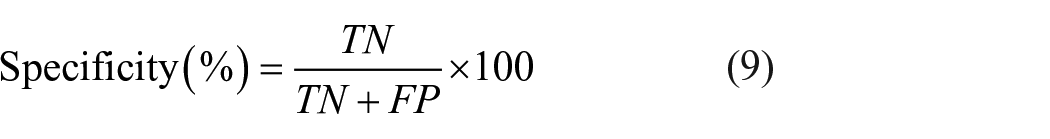
Specificity measures the proportion of actual negative cases (TN) that were correctly identified by the system, indicating how well the system avoids FP.
The F1 score equation:

The F1 score is the harmonic mean of precision and sensitivity, providing a balance between the 2 metrics, especially in situations where there is an uneven class distribution.
This ECG signal classification study was conducted using Python, Keras, and TensorFlow on a laptop with an Intel Core i7 processor, 8 GB RAM, and an NVIDIA K80 GPU. The model was trained using a momentum optimizer with adaptive backpropagation, with a batch size of 128 over 500 episodes of 30 epochs each. The dataset was split into 80% training and 20% testing using stratified sampling. The model was evaluated using cross-entropy loss, which minimizes loss values during training by adjusting model weights. Figures 8 and 9 shows a decline in Accurancy and loss over 30 epochs, indicating effective model performance.

Progress charts illustrating the performance of accuracy during training.
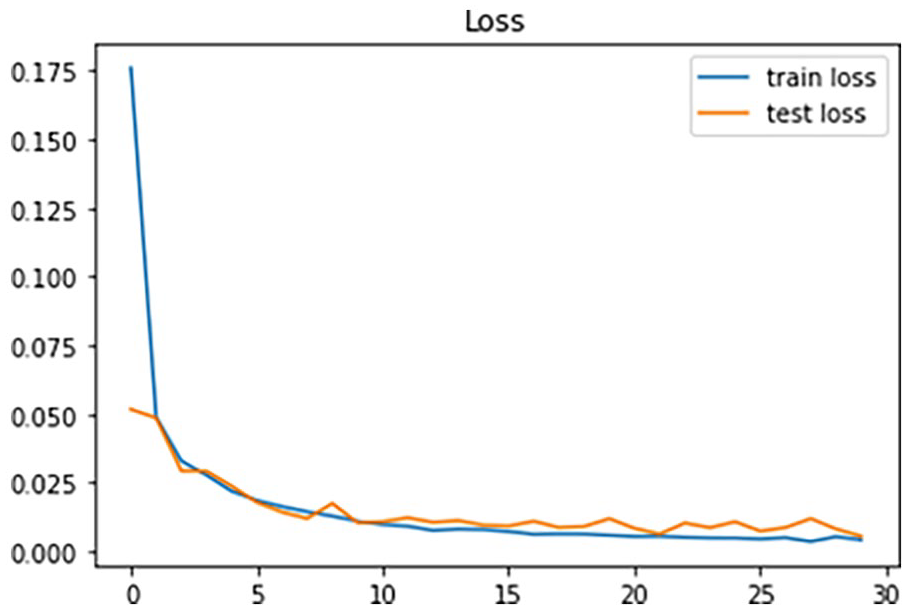
Progress charts illustrating the performance of loss during training.
In particular, Figure 8 revealed model’s superior accuracy of more than 99% to detect 5 types of arrhythmias. Furthermore, Figure 9 showed the effective error of the model during training, with a small number of 1.24%. This general approach highlights the robustness and effectiveness of the model in ECG signal segmentation. The confusion matrix, displayed in Figure 10 serves as a critical visualization tool for evaluating the classification algorithm’s performance.

Confusion matrix depicting the predictions generated by the proposed methodology.
Its representation offers a clear insight into the model’s ability to accurately classify instances, with the numbers along the main diagonal indicating true positives a direct measure of the model’s proficiency in correctly identifying instances within each class. Confusion matrix depicting the predictions generated by the proposed methodology. Moreover, the matrix provides a comprehensive snapshot of the model’s performance across various classes, revealing an impressive total accuracy of 99.2%. Notably, class N demonstrates particularly high accuracy, with 18 147 instances correctly classified. However, challenges arise in distinguishing between classes S and V, as evidenced by observed misclassifications. Despite these hurdles, the model demonstrates commendable performance across most classes, reflecting its robustness and effectiveness in accurately classifying instances and instilling confidence in its predictive capabilities.
In this study, we evaluate the performance of our proposed model against existing methods that utilize attention models. Our model, which integrates advanced techniques such as attention mechanisms and LSTM networks, demonstrates significant improvements across various metrics compared to the literature benchmarks, as illustrated in Table 2. Notably, our model achieves superior accuracy, sensitivity, specificity, and F1 score in classifying cardiac arrhythmias, showcasing its robustness and efficacy, particularly in detecting subtle abnormalities. For instance, while existing methods like CNN-LSTM-SE by Zhang et al. 50 and CNN-LSTM-attention gate by Jin et al. 51 achieve high accuracy, our model outperforms them in terms of sensitivity and specificity. Furthermore, compared to attentional CNN (ABCNN) in the Liu and Zhang 52 study, our model exhibits higher precision, recall, and F1 score. Additionally, our model’s performance surpasses that of models like CNN-LSTM Attention illustrated by Sakr et al., 53 CSA-MResNet demonstrated by Wang et al., 54 and MA1DCNN proven by Wang et al., 55 across various evaluation metrics. Specifically, our model achieves an accuracy of 99.20%, sensitivity of 97.50%, specificity of 99.81%, and an F1 score of 98.29%, highlighting its effectiveness in accurately diagnosing and classifying cardiac arrhythmias. These findings underscore the potential of our proposed model as a reliable tool for enhancing patient care and clinical decision-making in real-world scenarios.
Evaluation of methods incorporating attention mechanisms.

In addition to its remarkable performance, the computational analysis highlights that our model is well suited to handle computational constraints. ECG signal preprocessing, including filtering, resizing to 300 samples, and data balancing, is computationally inexpensive and runs quickly. The 1D convolution layers, optimized for GPU parallelism, ensure efficient feature extraction without incurring significant computational costs. The inclusion of CBAM modules, which enhance spatial and channel attention, adds minimal overhead, maintaining the model’s efficiency. However, the BiLSTM layer, due to its sequential processing nature, remains the most computationally expensive component in the pipeline. Despite this, the complete training process over 30 epochs, with a batch size of 128 and an NVIDIA K80 GPU, was completed in approximately 1 hour and 30 minutes. When compared to other state of the art models, our approach strikes an optimal balance between complexity and performance, offering significant computational efficiency alongside exceptional accuracy in cardiac arrhythmia classification.
Discussion
Previous studies utilizing the MIT-BIH Arrhythmia Database have enhanced machine learning and neural networks for classifying cardiac rhythm abnormalities (arrhythmias). Recently, deep learning techniques have been employed for arrhythmia categorization. Our proposed method outperforms previous approaches using the MIT-BIH, achieving higher overall accuracy and specificity in arrhythmia classification. Our model maintains a reasonable number of parameters, allowing for efficient signal classification without becoming overly complex. It is specifically designed to extract features from ECG signals and use them to classify rhythms according to AAMI criteria. This is made possible by the inclusion of a multiscale convolutional module and an attention module, which enhance the model’s ability to capture important signal characteristics while benefiting from the bidirectional LSTM module. Addi-tionally, the use of the SMOTE technique for data balancing contributes to improving the accuracy of arrhythmia classification. Despite making progress, there are still challenges to address. The first involves optimizing the performance of deep learning models through hyperparameter tuning. Automating this tuning process, using techniques such as the Orthogonal Array approach, could enhance efficiency and parameter optimization as reported in the study by Zhang et al. 56 Furthermore, although many models already achieve high performance, researchers continue to pursue architectural innovations with the goal of approaching 100% accuracy, as even small gains can significantly impact clinical safety, particularly in detecting rare but life threatening arrhythmias. This pursuit is also motivated by the need to minimize diagnostic errors and improve generalization across diverse patient populations. Despite high performance in theoretical contexts, implementing AI based clinical diagnosis for cardiac arrhythmias faces practical challenges. It is crucial to ensure that offline performance translates into real world use. Overcoming these obstacles is essential to create a reliable and stable diagnostic system, making it a significant focus of future research.
Limitations
Some assumptions were made in this work to guide the model development, with some constraints added. The fixed length of 300 samples was used for ECG signal processing, making it easier to standardize and perform parallel processing. However, this assumption can alter some meaningful features, especially when real signals have varying lengths in acquisition systems. Training was also performed using public databases, which are frequently utilized in the literature because they are available. However, these data may not necessarily reflect the pathophysiological heterogeneity one observes in real clinical practice. Further, the experiments were conducted on a specific hardware configuration (NVIDIA K80 GPU), which may limit reproducibility of performance on other platforms. Finally, the model evaluation was conducted using a static test set, which is not able to reflect all the data variability. An experimental setup or a cross-validation procedure based on real clinical data would thus serve to confirm the strength of the result. Such implications thus open the door to fairly many potential improvements toward improving the model’s generalization in practice.
Conclusion and Future Work
With the growing incidence of cardiovascular diseases, there is a pressing need for rapid and accurate electrocardiogram (ECG) signal classification. In fulfilling this need, we have developed a lean neural network that is particularly tailored for ECG classification. This lean architecture aims to accelerate the process of diagnosing cardiovascular disease, thereby enhancing the effectiveness of healthcare intervention. By synergistic use of the CBAM attention mechanism, convolutional network, and LSTM network, our model offers a complete solution to the ECG classification problem. Incorporating convolutional layers enables automatic feature extraction from raw ECG signals, and this increases the model’s ability to identify finer patterns indicative of cardiac diseases. By extensive testing on the MIT-BIH database, our model exhibits state of the art performance metrics like improved accuracy and robustness against traditional benchmarks. These results demonstrate the efficacy and potential clinical utility of our proposed method in accurate identification and classification of cardiac arrhythmias.
Looking ahead, several exciting avenues for future work are proposed to further enhance the model’s applicability and performance in clinical settings. First, we plan to extend the model to handle multichannel signals, such as 12-lead ECG, to address more complex clinical scenarios. Second, integrating a self-supervised learning strategy will allow the model to exploit unannotated data, expanding its usability in real world environments where labeled data may be scarce. Additionally, we aim to compress the model for implementation on embedded devices, supporting edge computing and enabling real-time analysis in resource-constrained settings. Finally, a multicenter evaluation of the model with real clinical data will be essential to assess its generalizability and robustness across diverse patient populations and clinical conditions.
Footnotes
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
