Abstract
Background:
The production of inflammatory factors in periodontium is increased by LPS, particularly from P. gingivalis, and the damage to periodontal tissues is exacerbated. Exosomes from periodontal ligament stem cells change regeneration and repair brought on by bacterial LPS. MiRNAs are carried by exosomes to recipient cells to affect epigenetic functions. Thus, this study aims to utilize deep learning algorithms to uncover novel micro-RNA biomarkers in bacterial LPS-exposed PDLSC stem cells to understand the activation pathway.
Methods:
Using NCBI GEO DATA SET GSE163489, the most differentially expressed micro RNAs were found to differ between healthy and LPS-induced PDLSC cells. Deep learning analysis, employing a Random Forest, Artificial Neural Network c, a Support Vector Machine (SVM), and a Linear Regression model implemented within the orange data mining toolkit, identified novel microRNA biomarkers. The orange data mining toolkit was utilized for deep learning analysis of microRNA expression data, providing a user-friendly environment for machine learning tasks like classification, regression, and clustering.
Results:
Random Forest emerged as the superior model, achieving the highest R2 score (.985) and the lowest RMSE (0.189) compared to Neural Networks (R2 = .952, RMSE = 0.332), Linear Regression (R2 = .949, RMSE = 0.343), and SVM (R2 = .931, RMSE = 0.398). This suggests its superior ability to capture the underlying patterns in the microRNA expression data. Given its robust performance, Random Forest holds promise for identifying novel biomarkers, developing more accurate diagnostic tools, and potentially guiding the stratification of patients for targeted therapeutic interventions in periodontal disease.
Conclusion:
The current study utilizes deep learning analysis of microRNA expression data to identify novel biomarkers associated with inflammasome activation and anti-apoptotic pathways. These findings hold promise for guiding the development of novel therapeutic strategies for periodontal disease. However, future studies are warranted to validate these biomarkers using independent datasets and experimental methods.
Introduction
Periodontitis leads to the destruction of tooth-supporting structures. The damage is initiated by Lipopolysaccharide, a key component of the cell wall of gram-negative organisms. 1 The healing of damaged periodontium occurs with the help of periodontal ligament stem cells. LPS, especially from P. gingivalis, increases the production of inflammatory factors, exacerbates periodontal inflammatory tissue damage, and delays the self-repairing of periodontal tissue, all of which restrict the proliferation of HPDLSCs either directly or indirectly through LPS. 2 Periodontal ligament stem cells exosomes release micro RNAs, altering regeneration and repair induced by bacterial LPS.3,4
Deep learning can integrate miRNA omics data with other data, such as gene expression or methylation. Identifying complex patterns in microRNA transcriptomics can reveal novel biomarkers in periodontitis. Multipotent PDLSCs can differentiate into fibroblasts, osteoblasts, cementoblasts, and adipocytes. Extracellular vesicles (EVs), 5 30 to 150 nm-sized membranous vesicles released by cells, are important cell-to-cell communication molecules. They govern injury repair and regeneration by influencing recipient cell proliferation, migration, differentiation, and immunological environment. Exosomal MicroRNAs6,7 can adversely affect posttranscriptional gene expression by interacting with the target mRNAs’ 3′ or 5′ untranslated region (UTR) or 5′ end region. Deep learning models are difficult to comprehend, especially in miRNA omics data, where biological relevance is critical. Thus, this study uses deep learning to uncover novel micro-RNA biomarkers in bacterial LPS-exposed PDLSC stem cells to elucidate the activation process.
Methods
When comparing healthy PDLSC cells to LPS-induced PDLSCs, using NCBI GEO DATA SET GSE163489, 8 researchers observed that the most differentially expressed micro RNAs were distinct. The data was cleaned by handling missing values, normalizing or standardizing features, and executing any necessary data transformations. Novel biomarkers can be predicted by artificial neural networks, SVM, random forest, and linear regression models, which were used with an orange tool to analyze the expression data. Data were split into 80 % training and 20 test data. The orange data mining toolkit was utilized for deep learning analysis of microRNA expression data, offering a user-friendly environment for machine learning tasks like classification, regression, and clustering. During the training phase, monitoring the model’s performance on the validation data is important to prevent overfitting. Based on the enrichr predictions, the Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment studies were carried out (Figure 1).

The Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses revealed the corticotrophin and apoptotic pathways.
Results and Discussion
Exosomes1,4,5 can regulate epigenetic processes by delivering miRNAs to recipient cells and regulate the biological function of recipient cells in bone regeneration. It has been indicated that the coculture of PDLSCs and BMSCs 7 can promote the osteogenic differentiation potential and capability of ECM formation of BMSCs, and the exosomes derived from PDLSCs were speculated to be important mediators of intercellular communication. Few studies found an altered miRNA pattern in gingival tissue, which is important for developing new periodontal disease biomarkers like miR-191-3p, miR-221-3p, miR-224-5p, and miR-1228-3p. The most differentially expressed micro RNAs were hsa-miR-7977, 7975, 3648, 4732-5p, 6794-3p, and 4526, and other study found differential expression of 108 exosomal miRNAs 4 (72 upregulated and 35 downregulated) from osteogenically differentiated PDLSCs, many of which are osteogenic-related. miR-122-5p, miR-142-5p, miR-25-3p, miR-192-5p, etc., which promote osteogenesis, were increased in exosomes from osteogenic differentiated PDLSCs. Functionally enriched pathway analysis of the NCBI dataset GSE163489 revealed the corticotrophin and apoptotic pathways. MiR-16 can modulate the cell cycle, inhibit cell proliferation, and promote cell apoptosis. These effects can be explained by several targets of miR-16, including the anti-apoptotic gene Bcl-2, numerous genes involved in the G1-S transition, such as cyclin D1, cyclin D3, cyclin E1, and CDK6 (cyclin-dependent kinase 6), and genes involved in the Wnt signaling pathway, such as (wingless-type MMTV integration site family, member 3A; Table 1).
R2 measures the goodness of fit of a Deep Learning Model.
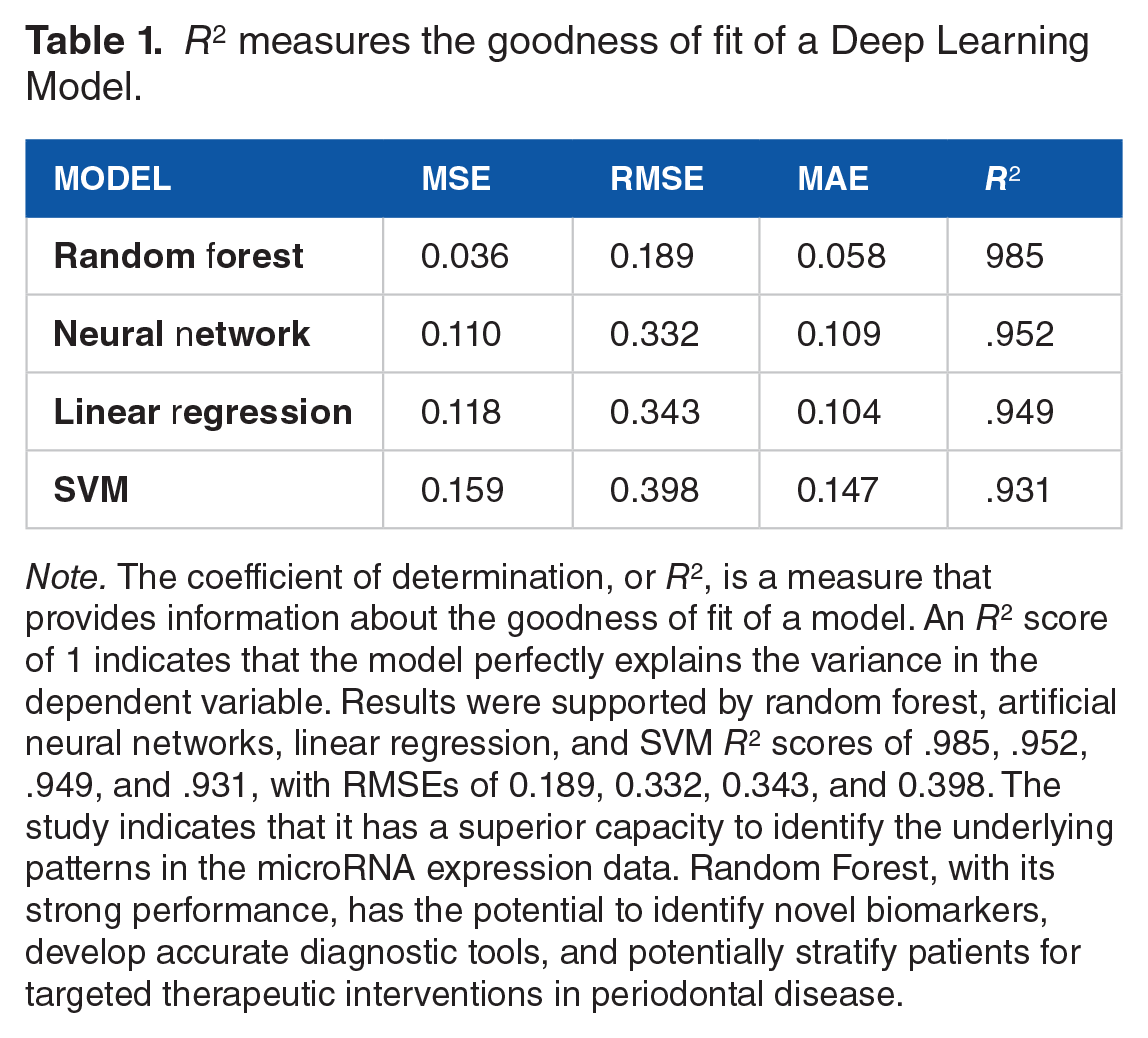
Note. The coefficient of determination, or R2, is a measure that provides information about the goodness of fit of a model. An R2 score of 1 indicates that the model perfectly explains the variance in the dependent variable. Results were supported by random forest, artificial neural networks, linear regression, and SVM R2 scores of .985, .952, .949, and .931, with RMSEs of 0.189, 0.332, 0.343, and 0.398. The study indicates that it has a superior capacity to identify the underlying patterns in the microRNA expression data. Random Forest, with its strong performance, has the potential to identify novel biomarkers, develop accurate diagnostic tools, and potentially stratify patients for targeted therapeutic interventions in periodontal disease.
Conclusion
The current study utilizes deep learning-based micro-RNA analysis to show novel biomarkers with inflammasome activation and anti-apoptotic pathways. This study highlights the therapeutic importance of exosomes and miRNAs in bone regeneration and periodontal disease. Exosomes regulate epigenetic processes and deliver miRNAs, suggesting the potential for enhancing bone regeneration. The study also identifies biomarkers for diagnosing and monitoring periodontal disease, highlighting potential therapeutic intervention targets.
Footnotes
Acknowledgements
None
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
