Abstract
Background:
Biomarker testing is central to precision oncology, yet real-world implementation across cancer types and populations remains inconsistent. Social determinants of health (SDoH) may influence testing uptake and exacerbate disparities in access to targeted therapies.
Methods:
We conducted a retrospective cohort study using Version 7 of the NIH All of Us Research Program Curated Data Repository. Adults diagnosed with colorectal cancer (CRC), non–small cell lung cancer (NSCLC), or prostate cancer were identified using standardized condition codes. Biomarker testing was determined through Current Procedural Terminology (CPT) and Logical Observation Identifiers Names and Codes (LOINC) for panel-based and single-gene assays. Logistic regression assessed associations between sociodemographic factors and documented biomarker testing, using robust modeling for the combined cohort and stepwise regression for individual cancer types.
Results:
Among 11 415 eligible participants, only 2.4% (n = 277) had documented biomarker testing, with 71.1% receiving panel-based assays. In the combined model, unemployment was significantly associated with higher odds of testing (odds ratio [OR] = 1.68; 95% confidence interval [CI] = 1.06-2.66), while college education showed a marginal association (OR = 1.48; 95% CI = 0.95-2.30). In cancer-specific models, NSCLC testing was predicted by education alone (OR = 1.70), while CRC testing was associated with unemployment (OR = 2.44), higher income (OR = 1.90), and smoking history. No significant predictors were found for prostate cancer.
Conclusion:
Despite national guidelines, biomarker testing remains underutilized and unevenly distributed across sociodemographic groups. These findings should be interpreted as exploratory, reflecting the fidelity of structured electronic health record (EHR) documentation rather than definitive utilization. Leveraging the scale and diversity of All of Us highlights both equity gaps and documentation limitations, positioning the program as a valuable platform for hypothesis generation in precision oncology.
Keywords
Introduction
Molecular testing for identification of actionable genomic alterations has become a cornerstone of precision oncology, specifically in advanced non–small cell lung cancer (NSCLC), colorectal cancer (CRC), and prostate cancer. Accurate identification of these biomarkers informs targeted treatment selection, minimizes toxicity, and avoids ineffective therapies.1,2 In NSCLC, targeted therapies for mutations such as EGFR, ALK, or KRAS G12C have transformed outcomes for many patients, allowing for orally targeted therapeutic agents.1,2 In CRC, detecting KRAS, NRAS, and BRAF mutations, along with microsatellite instability (MSI) status, can inform whether monoclonal antibodies, immunotherapy or targeted small molecules are appropriate. 2 For prostate cancer, alterations in DNA repair genes like BRCA1, BRCA2, and ATM may indicate eligibility for therapy with PARP inhibitors. 2 Recent reviews have emphasized the practical application of molecular pathology in prostate cancer, highlighting the integration of prognostic and predictive biomarkers into routine practice.3,4 As precision medicine informs and expands the oncology therapeutic landscape, ensuring equitable access and documentation of biomarker testing is critical to delivering high-quality, individualized cancer care across diverse populations.
The Centers for Medicare & Medicaid Services support the clinical utility of next-generation sequencing through national coverage determinations for advanced cancer patients, incentivizing adoption of precision testing in clinical workflows. 5 Despite these recommendations and increasing technological capabilities, disparities remain in the real-world uptake and documentation of molecular or biomarker testing. Ronquillo and Lester 6 demonstrated the feasibility of using the NIH All of Us database to assess the landscape of genomic testing in cancer populations through informatics-driven analyses. This work highlighted significant variation in testing uptake across cancer types and demographic subgroups.
This study leverages the All of Us Curated Data Repository to conduct a multi-cancer, national-scale evaluation of documented biomarker testing patterns and social determinants of health (SDoH)-associated disparities.7,8 We focus on NSCLC, CRC, and prostate cancer to identify sociodemographic factors associated with biomarker testing and to assess variations in molecular panel testing of somatic tumor testing to identify targeted therapy. Our findings have direct implications for healthcare systems, payers, and policymakers seeking to improve equitable precision oncology delivery, guide electronic health record (EHR) standardization efforts, and inform future implementation strategies.
Methods
Research patient data repository
This retrospective cohort study utilized data from the All of Us Curated Data Repository Version 7, focuses on EHR and participant survey data. As of February 2025, the Curated Data Repository has over 400 000 participants with more than 280 000 individuals with electronic health care data and more than 240 000 biosamples, as described in prior studies.7,8 The health care provider organization network includes regional medical centers, community health centers, and medical centers run by the US Department of Veterans Affairs.
The reporting of this study adheres to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines, and the completed STROBE checklist is provided as a Supplementary File (Supplement). 9
All participants in the All of Us Research Program provided written informed consent at the time of enrollment for use of their de-identified data in research. This secondary analysis used only de-identified data and was determined by the Purdue University Institutional Review Board to be exempt from additional review under Category 4 for secondary research (IRB #2024-389).
Defining and characterizing population
We identified adults aged >18 years with a documented diagnosis of advanced NSCLC, CRC, or prostate cancer from the NIH All of Us Research Program database. Cancer diagnoses were defined using standardized condition codes from the OMOP vocabulary.
Non–small cell lung cancer-related codes included
Primary malignant neoplasm of right lung, left lung, or lung (unspecified), NSCLC, metastatic NSCLC, malignant tumor of lung, and squamous NSCLC.
Colorectal cancer–related codes included
Carcinoma of colon (stage IV), malignant neoplasm of colon and/or rectum, and malignant tumors of specific colon subsites including ascending, descending, transverse, sigmoid colon, and rectosigmoid junction.
Prostate cancer–related codes included
Hormone-refractory prostate cancer, hormone-sensitive prostate cancer, malignant tumor of prostate, metastasis from malignant tumor of prostate, metastatic castration-resistant prostate cancer, and prostate cancer metastatic to bone.
The NSCLC, CRC, and prostate cancer were selected a priori to represent high-prevalence solid tumors at different stages of precision oncology adoption, NSCLC as established, CRC as intermediate, and prostate cancer as emerging. This design enables comparison across cancers with varying levels of guideline maturity and molecular testing uptake. Breast cancer, although common, was intentionally excluded because its biomarker testing paradigm centers on hormone receptor and HER2 expression rather than somatic genomic sequencing, which was the focus of this analysis.
To identify biomarker testing, we used both Current Procedural Terminology (CPT) procedure codes and Logical Observation Identifiers Names and Codes (LOINC) laboratory codes from the OMOP data model. The CPT codes captured evidence of panel-based next-generation sequencing (NGS) (eg, 0037U, 81445, 81450, 81455), single-gene molecular pathology assays (81400-81408, including gene-specific tests such as EGFR, ALK, KRAS, BRAF, and BRCA1/2), and unlisted procedure codes (81479, 81599) to account for laboratory-developed or emerging tests not yet assigned specific designations (Supplement Table S1). The LOINC codes captured results of individual biomarker assays (eg, EGFR, KRAS, BRAF, ALK, NRAS, RET, microsatellite instability; Supplement Table S2). Germline BRCA1/2 codes were included to reflect prostate cancer guidelines. Only codes relevant to somatic (tumor-based) testing in NSCLC, CRC, and prostate cancer were retained; codes used exclusively for germline testing or unrelated molecular assays were excluded. Code selection was guided by the National Comprehensive Cancer Network (NCCN) guidelines, Medicare National Coverage Determinations, and expert clinical review to ensure consistency with current standards of care. All codes were reviewed by oncology and pharmacy experts for transparency and reproducibility. Patients were classified as having documented biomarker testing if at least 1 relevant CPT or LOINC code was observed in their record at any time during the study period.
As a quality-control measure, we validated the CPT and LOINC code lists against NCCN and Centers for Medicare & Medicaid Services (CMS) guidance to ensure inclusion of relevant biomarkers. We de-duplicated encounter-level codes per participant, harmonized OMOP concept IDs across sites, and performed logic checks to flag implausible or inconsistent entries (eg, single-gene and panel codes on the same encounter date). Two reviewers adjudicated discrepancies before final dataset lock.
Baseline demographics and social determinants of health
Variables included race, ethnicity, smoking status, disability, education level, employment status, marital status, insurance type, and income. Patient race was categorized into white vs not white. Ethnicity was classified as Hispanic and non-Hispanic. All survey data were self-reported. Disability included at least one of the following categories: deaf, blind, difficulty with errands alone, difficulty concentrating, difficulty dressing or bathing, or difficulty walking or climbing stairs. Education level was categorized as any college education or no college education, and employment status was categorized as employed, unemployed, or retired. Sex was documented according to sex at birth. Marital status was self-reported and classified as married or not married. Self-reported annual income was categorized as at least $75 000 or less than $75 000.
Statistical analysis
In accordance with All of Us guidelines, missing values were incorporated into other categories when the missing cell size was fewer than 20 cases, ensuring data privacy and compliance with obscuring rules. This approach preserves confidentiality while retaining the full sample size for analysis. All statistical analyses were performed in R (version 4.4.0).
Multivariate models were restricted to participants with complete data on all covariates; no imputation was performed. Missing data for regression analyses were handled using listwise deletion, whereby any participant with missing values in the covariates of interest was excluded to ensure complete-case analysis and consistent model estimation. We used logistic regression to evaluate associations between sociodemographic factors and the presence of documented biomarker testing (by CPT or LOINC code). For the combined cohort (NSCLC, CRC, and prostate), we implemented a robust logistic regression model. For each cancer type individually, we used stepwise logistic regression to determine the most parsimonious model, using the Akaike Information Criterion (AIC) for model selection.
Multicollinearity was assessed using Generalized Variance Inflation Factor (GVIF), with no values exceeding thresholds of concern. Sensitivity analyses were conducted for the combined cohort model by re-running it after excluding influential observations using Cook’s distance and by refitting it using robust logistic regression techniques. Model stability for the combined model was further assessed via 10-fold cross-validation. Sensitivity analyses were not performed for the individual cancer-type models (NSCLC, CRC, prostate) due to limited sample sizes and the exploratory nature of those stratified analyses.
Results
Current state of biomarker testing in colorectal cancer, non–small cell lung cancer, and prostate cancer
A total of 11 415 patients with CRC, NSCLC, or prostate cancer were identified, of whom 277 had documented biomarker testing (Table 1). Patients with biomarker testing were more frequently male compared with those without documented testing (54.5% vs 70.8% overall). The racial and ethnic distribution was predominantly white across both groups (69.1% no testing vs 67.9% tested), with approximately 10% identifying as Hispanic or Latino. A slightly higher proportion of Black or African American patients was noted among those with biomarker testing (15.5%) compared with those without (14.6%). Most patients were between 60 and 80 years of age, with patients aged 70-80 making up the largest age group overall (40.2%), and those receiving biomarker testing skewing slightly younger (21.7% aged 60-70 vs 26.4% without testing). Across all 3 cancer types, only 2.4% of patients (n = 277 of 11 415) had documented biomarker testing within structured EHR data. When stratified by cancer type, testing documentation was most frequent in NSCLC (6.2%), followed by CRC (2.1%) and prostate cancer (0.8%) (Table 1 and Figure 1). These rates reflect capture in structured CPT and LOINC fields of All of Us, which may underestimate true utilization.
Baseline demographic characteristics by cancer type and biomarker testing status.
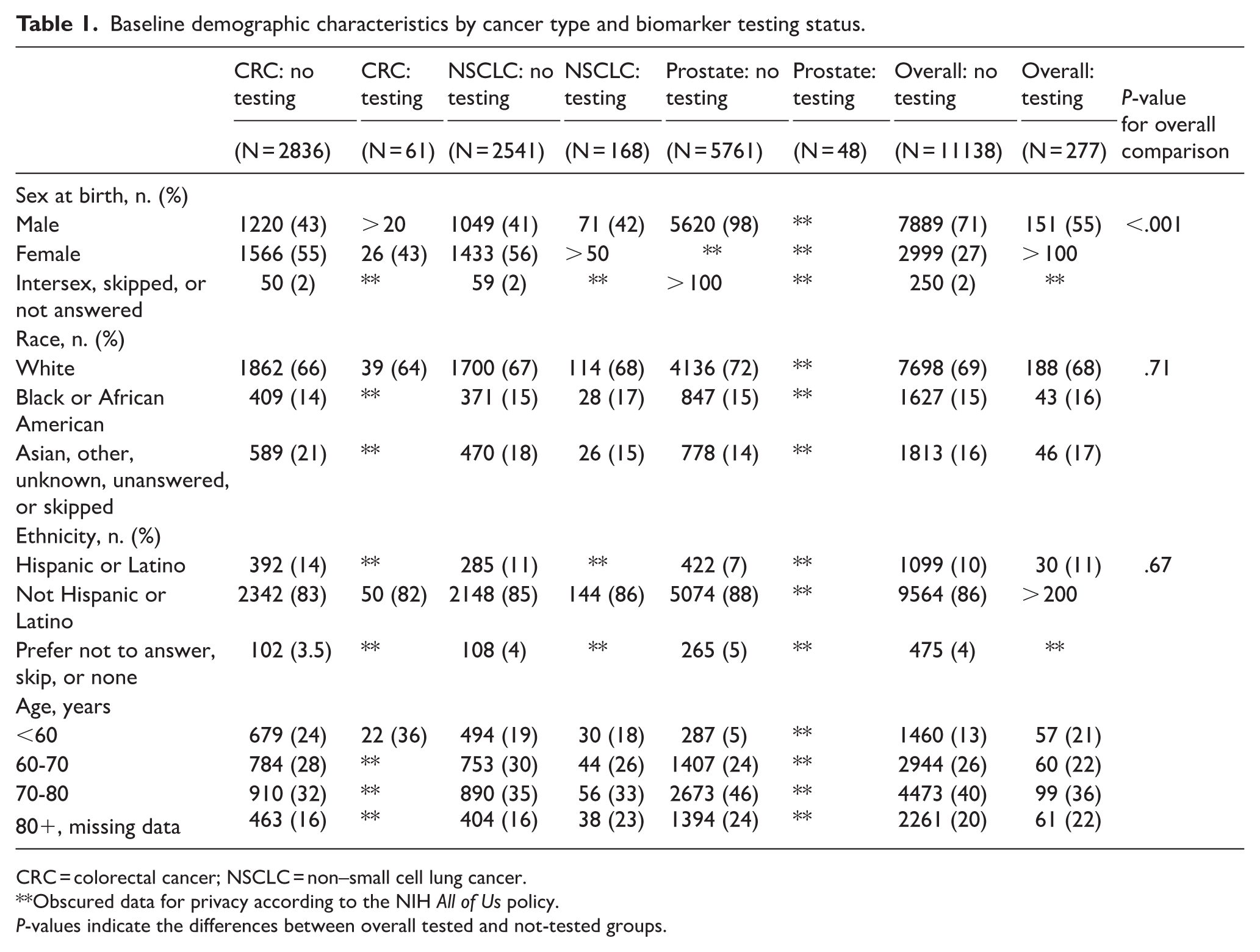
CRC = colorectal cancer; NSCLC = non–small cell lung cancer.
Obscured data for privacy according to the NIH All of Us policy.
P-values indicate the differences between overall tested and not-tested groups.

Documented biomarker testing.
Across all cancer types, a total of 277 patients received documented biomarker testing. Of these, 197 (71.1%) underwent panel-based testing, while 80 (28.9%) received single-gene or other forms of testing (P = NS). Among patients who underwent biomarker testing, the distribution of testing modality varied by cancer type (Figure 1). In the CRC cohort, 38 patients received targeted panel testing, while 23 received single-gene testing. In contrast, single-gene testing was less common among patients with NSCLC, with 36 individuals receiving this modality compared with 132 who underwent panel testing. For prostate cancer, 21 patients received single-gene testing, and 27 received panel testing.
Social Determinants of Health in Cancer Biomarker Testing
Among all cancer types (Table 2), the robust logistic regression model retained college education and employment status as predictors (Figure 2). College education was associated with higher odds of biomarker testing and should be interpreted as exploratory (odds ratio [OR] = 1.48; 95% confidence interval [CI] = 0.95-2.3; P = .085), and being unemployed was significantly associated with increased odds compared with being employed (OR = 1.68; 95% CI = 1.06-2.66; P = .029). Collinearity was minimal (GVIFs ~1.05), and sensitivity analysis without outliers yielded an unstable model, reinforcing the robustness of the original fit.
Social determinants of health.

CRC = colorectal cancer; NSCLC = non–small cell lung cancer.
Obscured data for privacy according to the NIH All of Us policy.
Missing, skipped, unknown, or unanswered data are not displayed.

Forest plot of robust logistic regression model on documented biomarker testing.
In the NSCLC cohort stepwise model, only college education remained a significant predictor of biomarker testing in the final model (OR = 1.70; 95% CI = 1.01-2.87). For CRC, the stepwise model retained employment, income, and smoking status. Unemployment (OR = 2.44; 95% CI = 1.11-5.36), income ⩾75k (OR = 1.90; 95% CI = 1.02-3.55), and a history of smoking (OR = 0.48; 95% CI = 0.25-0.91) were all associated with higher likelihood of biomarker testing. The prostate cancer cohort did not have any significant predictors of biomarker testing. The absence of significant predictors in prostate cancer may reflect both lower adoption of testing in practice and limitations in documentation within All of Us.
Discussion
The NIH All of Us database represents a significant advancement in precision medicine by integrating EHRs nationwide to enable large-scale analysis of RWD. This study leverages documented biomarker testing (as captured in structured CPT and LOINC codes) in participants with various cancers to examine current testing patterns and the influence of SDoH. Rather than providing a definitive estimate of utilization, our findings should be viewed as an exploratory assessment of documentation fidelity within All of Us, highlighting both opportunities and limitations in how biomarker testing is recorded. Notably, it provides insights not only into the adoption of biomarker testing but also into the retrievability of these test results as an essential component for guiding targeted therapies. The novelty of this work lies in applying a uniquely diverse, national-scale resource to identify equity-related patterns and documentation gaps, positioning All of Us as a platform for hypothesis generation and future validation. A novel aspect of this work is the use of survey data to assess the impact of SDoH on biomarker testing across multiple cancer types.
Molecular testing has become a central tenet in oncology due to the evolution of prognostic and actionable biomarkers in cancer. In NSCLC, CRC, and prostate cancer, biomarker testing via single-gene assays or, ideally, broad panel-based NGS approaches is often recommended or mandated by the US Food and Drug Administration (FDA) for identifying eligibility for targeted therapies. Among these, NSCLC has seen the most advanced implementation of precision medicine, with multiple genomic alterations, including ALK, EGFR, BRAF, MET exon 14 skipping (METex14), RET, KRAS G12C, ROS1, ERBB2, and NTRK1/2/3, influencing or guiding treatment decisions.10 -20 In CRC, recommended testing includes KRAS, NRAS, and BRAF mutations, as well as MSI or mismatch repair deficiency (dMMR), which inform the use of anti-EGFR therapies and immunotherapy, respectively.21 -27 In prostate cancer, genomic testing is increasingly important in advanced stages for identifying alterations in DNA damage repair genes such as BRCA1, BRCA2, and ATM, which can predict response to PARP inhibitors.3 -5,28,29 Other biomarkers like MSI-H/dMMR, PTEN, TP53, and those affecting androgen receptor signaling are also under investigation. 30 Beyond these established targets, an expanding array of emerging biomarkers from tumor mutational burden, homologous recombination repair deficiency, circulating tumor cells, to multiomics and AI further adds to the urgency of developing comprehensive, standardized molecular testing and documentation frameworks to ensure equitable implementation across malignancies. 31 This expanding biomarker landscape continues to evolve with new prognostic and predictive markers across tumor types, underscoring the need for harmonized integration into clinical pathology reporting and decision-making. 32 Despite the progress of biomarker-driven therapy in various cancers, our findings indicate that biomarker testing in prostate cancer remains underutilized compared to NSCLC and CRC. This lag may reflect both true underuse in clinical practice and incomplete documentation within All of Us, reinforcing the need for improved structured capture of prostate cancer biomarkers. 33
Adoption of biomarker testing in clinical practice is influenced by multiple implementation barriers and facilitators. Although NCCN guidelines provide clear recommendations for molecular profiling in advanced NSCLC and CRC, clinician awareness, institutional infrastructure, and reimbursement constraints contribute to inconsistent uptake across healthcare systems.2,19,32,33 Broader adoption of panel-based testing also depends on access to validated assays, EHR-integrated ordering pathways, and multidisciplinary support through molecular tumor boards. 6 Food and Drug Administration approvals of targeted therapies commonly require companion diagnostic testing, which can stimulate uptake, yet gaps between drug approvals, payer coverage, and laboratory capacity often delay real-world implementation.5,29 The regulatory and infrastructural challenges overlap with the SDoH observed in our analysis, highlighting that equitable precision oncology requires alignment among clinical guidelines, diagnostic infrastructure, and patient access.
Interestingly, unemployment was associated with higher odds of receiving biomarker testing in both the combined cohort and the CRC subgroup. These findings contrast with expected observations in cancer care access by socioeconomic status. A possible, but unverifiable with our data set, explanation is that unemployed patients may more frequently receive care at academic or safety-net institutions, where adherence to guideline-based biomarker testing may be more consistent and embedded in clinical workflows.21,23 Another reason could be clinical severity bias, where unemployed patients may present with more advanced disease and thus be prioritized for molecular testing. Given the limitations of the dataset, these results should be interpreted as hypothesis-generating and highlight the value of All of Us in surfacing novel associations that warrant follow-up in cohorts with more complete clinical annotation. Additional research is required to understand how care setting, insurance coverage, and health status interact to influence access to precision oncology among socioeconomically disadvantaged groups.
This highlights a central challenge in precision oncology: translating genomic advances into equitable clinical practice. While the All of Us platform offers a robust resource for evaluating real-world trends, our findings underscore ongoing gaps in biomarker documentation and implementation. This study has several limitations. First, there is potential for selection bias especially given that the stage of disease could not be delineated, which may have included participants into the denominator who would not have needed biomarker testing. The most significant limitation is the reliance on structured CPT and LOINC fields to identify biomarker testing. Many test results are stored as scanned pathology reports or unstructured text within EHRs, leading to under-documentation in structured data sets. Consequently, our findings likely underestimate the true prevalence of biomarker testing and should be interpreted primarily as a reflection of data capture and documentation fidelity rather than clinical adoption. Furthermore, cancer stage at diagnosis could not be delineated within the All of Us data set. Inclusion of early-stage cases, which would not typically undergo molecular testing may have inflated denominators and lowered observed testing rates. This limitation represents a key confounder when interpreting testing prevalence. Future research could integrate unstructured EHR data using natural language processing approaches to extract biomarker results from pathology and molecular reports. Linking structured and unstructured data, as well as incorporating cancer stage and treatment variables, will be essential to validate and extend these findings. Documentation of biomarker or molecular reports varies across oncology clinics; results may be uploaded as structured EHR data, stored as PDF documents, or recorded in provider notes. Of these, scanned PDFs into a patient EHR represent the mostinaccessible format for data extraction and may not be captured in the All of Us database. Data completeness and consistency are additional concerns, as missing or inaccurate entries can undermine the robustness of findings. While the All of Us’s diverse participant pool supports generalizability, our sample may not fully represent the broader cancer patient population.
These findings emphasize the need for standardized, structured documentation of biomarker testing across EHR systems to ensure retrievability, guide treatment decisions, and support research. Practical strategies include embedding structured biomarker fields in EHRs, implementing automated prompts for guideline-concordant testing, expanding reimbursement for molecular diagnostics, and developing equity-focused quality metrics. Strengthening data infrastructure by integrating unstructured pathology data and advancing natural language processing pipelines to capture non-billable and scanned biomarker results will be essential to close current documentation gaps and fully realize the potential of initiatives such as All of Us to inform national cancer care and policy.
Conclusion
This study provides one of the first national-scale evaluations of documented biomarker testing in structured EHR fields across colorectal, lung, and prostate cancers using the NIH All of Us Research Program. Documented testing was uncommon, particularly in prostate cancer, and exploratory associations with education, employment, and income highlight inequities in access to precision oncology. These findings reflect the fidelity of biomarker documentation in All of Us rather than definitive clinical utilization rates, underscoring both the promise and current limitations of real-world data. Improving structured capture of biomarker information, linking it to treatments and outcomes, and embedding equity-focused strategies in policy and practice will be essential to realize the potential of precision oncology for all patients. The novelty of this work lies in leveraging the scale and diversity of All of Us to surface exploratory associations and identify documentation gaps, positioning it as a uniquely powerful platform for hypothesis generation in precision oncology.
Supplemental Material
sj-docx-1-onc-10.1177_11795549261417371 – Supplemental material for National Patterns of Biomarker Testing in Colorectal, Lung, and Prostate Cancers: Insights From the NIH All of Us Research Program
Supplemental material, sj-docx-1-onc-10.1177_11795549261417371 for National Patterns of Biomarker Testing in Colorectal, Lung, and Prostate Cancers: Insights From the NIH All of Us Research Program by Patrick J Kiel, Mark W McGiffin and Michael A Preston in Clinical Medicine Insights: Oncology
Supplemental Material
sj-docx-2-onc-10.1177_11795549261417371 – Supplemental material for National Patterns of Biomarker Testing in Colorectal, Lung, and Prostate Cancers: Insights From the NIH All of Us Research Program
Supplemental material, sj-docx-2-onc-10.1177_11795549261417371 for National Patterns of Biomarker Testing in Colorectal, Lung, and Prostate Cancers: Insights From the NIH All of Us Research Program by Patrick J Kiel, Mark W McGiffin and Michael A Preston in Clinical Medicine Insights: Oncology
Footnotes
Acknowledgements
We gratefully acknowledge All of Us participants for their contributions, without whom this research would not have been possible. We also thank the National Institutes of Health’s All of Us Research Program for making available the participant data examined in this study. This study used data from the All of Us Research Program’s Controlled Tier Dataset (v8) available to authorized users on the Researcher Workbench. Special thanks to Todd C. Skaar, David R. Foster, and Karen Suchanek Hudmon for their technical review and critical feedback.
Ethical Considerations
This study was reviewed by the Purdue University Institutional Review Board and determined to be exempt from additional review under Category 4 for secondary research involving de-identified data (IRB #2024-389). The analysis utilized de-identified data from the NIH All of Us Research Program.
Consent to participate
All participants in the NIH All of Us Research Program provided written informed consent at the time of enrollment for the use of their de-identified data in research. No additional consent was required for this secondary analysis.
Consent for publication
Not applicable.
Author contributions
P.J.K. conceived and designed the study, performed data extraction and statistical analyses, interpreted the results, and drafted the manuscript. M.W.M. contributed to manuscript revision. M.A.P. contributed to study design, interpretation, and supervision. All authors approved the final version of the manuscript and agree to be accountable for all aspects of the work.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work of Dr. M.A.P. and the research team was supported in part by the Reaching the Underserved, Rural, and Low-Income (RURaL) Lab for D&I Research in Cancer Disparities and the Center for Health Equity and Innovation.
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: P.J.K. is an employee of Amgen Inc and holds stock in the company. These affiliations are disclosed in the interest of transparency and do not alter the author’s adherence to the journal’s publication ethics.
Data availability statements
The data underlying this study are not publicly available due to participant privacy and security requirements of the All of Us Research Program. Researchers may access the All of Us Controlled Tier Dataset (v8) through the All of Us Researcher Workbench (![]() ) after completing the required registration, training, and project approval.
) after completing the required registration, training, and project approval.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
