Abstract
Background:
In recent years, abnormal expression of collagen triple helix repeat containing 1 (CTHRC1) has been found in some tumors, closely related to the poor prognosis of cancer patients. However, the clinical significance of CTHRC1 in gliomas is not completely understood.
Methods:
We investigated the expression, prognostic value, and potential biological function of CTHRC1 in different types of gliomas through bioinformatics analysis and experimental verification.
Results:
Bioinformatics analysis revealed several key findings regarding the expression and clinical significance of CTHRC1 in gliomas. First, the analysis demonstrated a positive correlation between CTHRC1 expression and the World Health Organization (WHO) grading of gliomas, a relationship that was validated through immunohistochemistry experiments. In addition, a trend was observed in which CTHRC1 expression increased with the extent of glioma invasion, as supported by Western blot experiments. Subsequent bioinformatics analysis identified the mesenchymal subtype of gliomas as having the highest levels of CTHRC1 expression, a finding reinforced by immunohistochemical staining. Moreover, high CTHRC1 expression was associated with poor prognosis in gliomas and emerged as an independent prognostic factor, with varying impacts on prognosis between low-grade gliomas (LGGs) and glioblastoma (GBM) subgroups. Notably, comparative analysis unveiled distinct patterns of immune infiltration of CTHRC1 in LGG and GBM. Furthermore, alterations in copy number variations and DNA methylation were identified as potential mechanisms underlying elevated CTHRC1 levels in gliomas. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis indicated that CTHRC1 and its associated genes mainly function in the extracellular matrix and participate in tumor-related signaling pathways.
Conclusions:
The CTHRC1 has shown significant clinical utility as a prognostic marker and mesenchymal subtype marker of glioma.
Introduction
Gliomas are the most common primary tumors of the central nervous system. The 2021 World Health Organization (WHO) classification categorizes gliomas into 4grades: grades I and II are low-grade gliomas (LGGs), which have low invasiveness and a better prognosis, whereas grades III and IV are high-grade gliomas (HGGs), which have high invasiveness and a poor prognosis, especially glioblastoma (GBM).1,2 The treatment approach for gliomas typically involves a combination of surgery, radiation therapy, and chemotherapy. Based on the gene detection results of gliomas, targeted therapy, immunotherapy, and molecular biology therapy have been explored as potential treatments to improve postoperative survival time. 3 Molecular targeted therapy, specifically targeting genes associated with glioma invasion and metastasis, is considered a promising approach to significantly improve the prognosis of gliomas.4-6 Studies have found a correlation between glioma progression and epithelial-mesenchymal transition (EMT), which promotes tumor proliferation, invasion, and treatment resistance.7-9 Therefore, the identification of EMT-related genes associated with glioma invasion is of significant importance for the clinical treatment of gliomas.
Collagen triple helix repeat containing 1 (CTHRC1) is a protein closely associated with tumor invasion and metastasis. 10 It has recently gained significant attention in cancer research.11,12 The CTHRC1 is a highly conserved secreted glycoprotein that was identified as a differentially expressed gene during arterial injury. 13 It shows transient high expression in the outer fibrous cells and inner smooth muscle cells involved in the repair of injured blood vessels, whereas its expression is not detected in normal arteries. The CTHRC1 is known to reduce collagen deposition and promote cell migration. 13 In recent years, abnormal expression of CTHRC1 has been observed in some tumors and is closely related to the survival prognosis of cancer patients.14-16 Studies have shown that CTHRC1 is associated with the invasiveness of glioma cell lines, and knocking down CTHRC1 can reduce glioma cell invasion.17,18 However, research on the role of CTHRC1 in gliomas is limited, and its exact function, particularly its clinical significance, is not yet fully understood.
To comprehensively assess the potential clinical significance of CTHRC1 in gliomas, this study integrates bioinformatics analysis and experimental validation to investigate the expression, prognostic value, and biological functions of CTHRC1 in different glioma subtypes. Initially, we examine CTHRC1 expression and its prognostic implications across various glioma types (grading, histology, subtype) through bioinformatics analysis. Subsequently, we validate these findings in vitro using glioma cell lines and in vivo using glioma tissue samples. Furthermore, we use bioinformatics tools to uncover additional insights into CTHRC1, such as tumor immune infiltration, mutations, copy number variations (CNVs), methylation patterns, associated genes, and Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis.
Materials and Methods
Data sets and bioinformatics analysis
This study conducted bioinformatics analysis from June to September 2023, followed by the validation of the results through immunohistochemistry (IHC) and Western blot experiments from October to December 2023. This study used the TCGA-GBMLGG data set, CGGA data set, TCGA-LGG data set, and TCGA-GBM data set for the bioinformatics analysis of gliomas. First, we performed bioinformatics analysis and mutual validation of CTHRC1 using the readily available TCGA-GBMLGG data set and CGGA data set from the GlioVis online database (http://gliovis.bioinfo.cnio.es/). 19 The primary objective was to investigate the relationship between CTHRC1 mRNA expression and grade, histology, subtype, and prognosis. Next, the authors downloaded and curated RNA-seq data and clinical data from TCGA-LGG and TCGA-GBM from the TCGA database (https://portal.gdc.cancer.gov) for subsequent prognostic subgroup analysis of CTHRC1 and univariate and multivariate Cox regression analysis. 20
Glioma tissue samples
To validate the results obtained from the bioinformatics analysis, the authors conducted IHC experiments to detect CTHRC1 in glioma tissue samples. Ten surgically resected specimens from different WHO grades (WHO II = 4, WHO III = 3, WHO IV = 3) of glioma were randomly selected (Supplementary material 1), adhering to the following inclusion and exclusion criteria. Inclusion criteria include (1) complete tumor resection and (2) pathological diagnosis confirmed as glioma (WHO grades II-IV). Exclusion criteria include (1) patients who received chemotherapy or radiotherapy before surgery and (2) patients with other malignant tumors. Furthermore, to investigate CTHRC1 expression in distinct molecular subtypes of GBM (classical, mesenchymal, proneural), 9 corresponding tissue samples were selected for immunohistochemical staining, consisting of 3 cases each for every molecular subtype (Supplementary material 2), in adherence to the aforementioned criteria. These glioma specimens were obtained from tumor tissues of patients undergoing surgery at the Affiliated Hospital of Guizhou Medical University and were subsequently paraffin-embedded and stored long-term in the pathology department. This study was conducted in accordance with the principles of the Helsinki Declaration, approved by the Ethics Committee of Affiliated Hospital of Guizhou Medical University (Ethics Approval No: 2022090), and written informed consent was obtained from patients or their guardians.
Cell lines
The GBM cell lines with different invasive capacities (U87, U251, LN229) and human astrocytes (NHA) were purchased from the Shanghai Institute of Cell Biology, Chinese Academy of Sciences, and authenticated using short tandem repeat (STR) profiling. Retrieve the frozen cells from the liquid nitrogen tank and immediately place them in a 37°C water bath for thawing. Gently agitate to accelerate thawing until only a small piece of ice remains in the vial, with a duration of approximately 1 minute. After disinfection with 75% ethanol, the cells were transferred to a laminar flow cabinet that had been irradiated with UV light for 30 minutes. Using a sterile pipette tip, the cell suspension was aspirated into a sterile 15 mL centrifuge tube and supplemented with 10% fetal bovine serum (FBS) (44.5 mL Dulbecco’s Modified Eagle Medium [DMEM] from Gibco, Waltham, MA, USA, 5 mL FBS from VivaCell, China, and 0.5 mL penicillin-streptomycin from VivaCell). The tube was centrifuged at 1000 r/min for 5 minutes, the supernatant was aspirated, and the cell pellet was resuspended in 10% FBS. The cell suspension was transferred to a vented culture flask and incubated at 37°C with 5% carbon dioxide in a humidified incubator. When the cell confluence reached 80%, the cells were digested with trypsin for 5 minutes and then subcultured at a 1:3 ratio.
Immunohistochemistry
Immunohistochemistry was performed to detect the protein expression of CTHRC1 in glioma tissues. First, tissue sections were placed in a 65°C oven for 20 minutes to dissolve the paraffin. Then, the tissue sections were deparaffinized in xylene and subsequently hydrated in gradually increasing concentrations of ethanol. Subsequently, the hydrated tissue sections were immersed in a citrate buffer solution for antigen retrieval under high temperature and high pressure. Nonspecific staining was removed by treating the sections with 3% hydrogen peroxide, followed by blocking with goat serum. Rabbit Anti-CTHRC1 antibody (Proteintech, Wuhan, China, diluted at 1:100) was then applied as the primary antibody, and the sections were incubated overnight at 4°C. Next, the sections were incubated with goat anti-rabbit IgG (HUABIO, Hangzhou, China, diluted at 1:100) as the secondary antibody and subjected to consistent DAB color development. Subsequently, the sections were counterstained with hematoxylin, rinsed with ammonia water, dehydrated, and coverslipped. Finally, the immunohistochemical staining results were observed under a microscope, and quantitative analysis was performed using FIJI software. Three repeated immunohistochemical tests were carried out for each patient’s pathological sections to evaluate the expression level of CTHRC1 protein.
Western blot
Western blot analysis was performed to detect the protein expression differences of CTHRC1 in NHA, LN229, U87, and U251 cell lines using β-actin as the internal reference. First, cells were lysed using radio-immunoprecipitation assay (RIPA) cell lysis buffer to extract total cellular proteins. Then, the total protein concentration in the samples was measured using the bicinchoninic acid assay (BCA) method. Subsequently, the Western blot experiment was carried out. The specific details are as follows: the proteins from the samples were separated by sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS-PAGE) gel electrophoresis. The separated proteins were then transferred to a polyvinylidene fluoride (PVDF) membrane through wet transfer. The membrane was blocked with a 5% skim milk solution to prevent nonspecific binding. Rabbit Anti-CTHRC1 antibody (Proteintech, Wuhan, China, diluted at 1:1500) was added for overnight incubation at 4°C. HRP-conjugated goat anti-rabbit secondary antibody (HUABIO, Hangzhou, China, diluted at 1:50000) was added for 2 hours of incubation at room temperature. The ECL chemiluminescence reagent was added to the membrane, and the protein band images were captured using an imaging system. ImageJ software was used for grayscale measurement and quantitative analysis of the protein bands to detect the differential expression of CTHRC1 in different cell lines (Supplementary material 3).
Correlation analysis of immune infiltration
Comparative analysis of CTHRC1 in LGG and GBM regarding immune infiltration differences was conducted using the TIMER online database (https://cistrome.shinyapps.io/timer/). 21 The analysis included correlation analysis between CTHRC1 levels and immune cell infiltration levels, detection of the prognostic value of CTHRC1 levels and tumor-infiltrating immune cells using Cox proportional hazards models, and correlation analysis between CTHRC1 and immune cell type markers. In addition, the authors further investigated the relationship between CTHRC1 levels in LGG and GBM and 28 tumor immune-infiltrating cell subtypes using the TISIDB online database (http://cis.hku.hk/TISIDB/). 22
Mutation, copy number variation, and methylation analysis of collagen triple helix repeat containing 1
Aberrant gene expression is closely associated with gene mutations, CNV, and DNA methylation. We further investigated the molecular mechanisms underlying the CTHRC1 abnormalities in gliomas using the UCSC Xena database (http://xena.ucsc.edu/), including gene mutations, CNV, and DNA methylation. 23 We also validated our findings using the cBioPortal database (https://www.cbioportal.org/). 24
Gene Ontology and Kyoto Encyclopedia of Genes and Genomes enrichment analysis
We obtained CTHRC1-related genes using the STRING online tool (https://cn.string-db.org/). 25 We performed GO and KEGG enrichment analysis on CTHRC1 and its related genes. In R software (v.4.2.1), we used the clusterProfiler package (v.4.4.4) for GO and KEGG enrichment analysis. To facilitate the presentation and highlight the results of the GO and KEGG enrichment analysis of CTHRC1-related genes, we used the ggplot2 package (v.3.3.6) to visualize the enrichment analysis results, showing only the top 3 results.
Statistical analysis
The statistical analysis results for the online databases (GlioVis, TIMER, TISIDB) were generated automatically by the authors using the specified parameters in the databases. The following statistical analyses required the authors to perform the data analysis for nononline databases. Differences in CTHRC1 protein expression among different glioma WHO grades in the IHC experiment and among different cell lines in the Western blot experiment were determined using Student’s t-test. Data visualization was performed using GraphPad Prism software (v.8.0.2). The prognostic value of CTHRC1 expression in the TCGA-LGG and TCGA-GBM data sets was assessed using the Kaplan-Meier survival curves. Specifically, in R software (v.4.2.1), the survival package (v.3.3.1) was used for proportional hazards assumption testing, fitting survival regression, and visualization of the results using the survminer and ggplot2 packages (v.3.3.6). Univariate and multivariate Cox regression models were used to investigate whether CTHRC1 expression is an independent prognostic indicator in TCGA-LGG and TCGA-GBM patients. Specifically, in R software (v.4.2.1), the survival package (v.3.3.1) was used for proportional hazards assumption testing and Cox regression analysis. All bioinformatics analysis and experimental results mentioned above were considered statistically significant with a P < .05.
Results
The expression of collagen triple helix repeat containing 1 is positively correlated with tumor grade
According to the analysis results from the GlioVis database, the expression of CTHRC1 mRNA is positively correlated with glioma WHO grade in the TCGA-GBMLGG dataset (Figure 1A). These findings were validated in the CGGA data set (Figure 1B). Furthermore, IHC staining results showed consistent CTHRC1 protein expression patterns with the aforementioned mRNA expression results (Figure 1C and D). Specifically, the expression of CTHRC1 protein was the highest in WHO grade IV gliomas, followed by WHO grade III gliomas, and the lowest in WHO grade II gliomas, with statistically significant differences observed between different grades of gliomas (P < .05). These results indicate a positive correlation between the expression of CTHRC1 and tumor grade, suggesting that CTHRC1 expression may increase with the malignancy of the tumor.

The expression of CTHRC1 is positively correlated with glioma WHO grade. (A) TCGA-GBMLGG dataset; (B) CGGA data set; (C) IHC staining of CTHRC1 in different grades of glioma (200×); (D) Statistical analysis of CTHRC1 expression detected by IHC experiment, with a sample size of 4 for WHO II, 3 for WHO III, and 3 for WHO IV, each with 3 repetitions per sample, was conducted using Student’s t-test.
The expression trend of collagen triple helix repeat containing 1 increases with the degree of glioma invasion
According to the analysis results from the GlioVis database, the mRNA expression of CTHRC1 in GBM is significantly higher than in oligodendroglioma, oligoastrocytoma, and astrocytoma in the TCGA-GBMLGG dataset (P < .05) (Figure 2A). Similarly, in the CGGA data set, the mRNA expression of CTHRC1 in GBM is significantly higher than in oligoastrocytoma, anaplastic oligoastrocytoma, oligodendroglioma, astrocytoma, anaplastic oligodendroglioma, and anaplastic astrocytoma (P < .05) (Figure 2B). These database analysis results indicate that the expression of CTHRC1 is significantly higher in GBM compared with other types of gliomas. It is well known that GBM exhibits the highest invasive potential, suggesting that CTHRC1 expression is higher in tumor types with greater invasiveness. This hypothesis has been verified by cell experiment in vitro. According to Western blot analysis results, the expression of CTHRC1 in all GBM cell lines was significantly higher than in normal human astrocytes (NHA) (P < .05) (Figure 2C). The CTHRC1 exhibited a trend of increasing expression with increased invasion levels of GBM cell lines. Specifically, the experimental results showed a lower expression level of CTHRC1 in LN229, a slightly higher expression level in U87 compared with LN229, and a slightly higher expression level in U251 compared with U87. Previous studies have shown that LN229 exhibits lower invasiveness than U87, 26 and CTHRC1 expression is lower in U87 compared with U251. 18 Overall, CTHRC1 demonstrates a trend of increasing expression with increased invasion in GBM.

The expression of CTHRC1 increasing with the degree of tumor invasiveness. (A) TCGA-GBMLGG data set; (B) CGGA data set; (C) Western blot analysis to examine the expression of CTHRC1 in different malignant glioma cell lines (LN229, U251, U87) and normal human astrocyte cells (NHA); (D) quantitative analysis of CTHRC1 protein expression in different cell lines was performed by Western blot experiment with at least 3 repetitions, and statistical analysis was conducted using Student’s t-test.
Collagen triple helix repeat containing 1 is a potential biomarker for mesenchymal subtype of glioma
We further evaluated the expression pattern of CTHRC1 in different molecular subtypes of glioma, including the classical, mesenchymal, and proneural subtypes. Bioinformatics analysis revealed that CTHRC1 expression was significantly higher in the mesenchymal subtype compared with other subtypes in the CGGA data set (P < .05) (Figure 3A). Subsequently, we performed experimental validation. Glioblastoma tissue sections corresponding to the classical, mesenchymal, and proneural subtypes, as defined by the expression of epidermal growth factor receptor (EGFR), neurofibromin 1 (NF1), and platelet-derived growth factor receptor alpha (PDGFRA)/isocitrate dehydrogenase 1 (IDH1), respectively, were selected for CTHRC1 immunohistochemical staining. 27 The results showed that CTHRC1 expression was significantly higher in the mesenchymal subtype of glioma compared with the classical and proneural subtypes (P < .05) (Figure 3B and C). These findings suggest that CTHRC1 is a biomarker for the mesenchymal subtype of glioma.

Significant upregulation of CTHRC1 expression in the mesenchymal subtype of glioma. (A) CGGA dataset; (B) IHC staining of CTHRC1 in different molecular subtypes (200×); (C) statistical analysis of CTHRC1 expression detected by IHC experiment, the number of samples (Classical = 3, Mesenchymal = 3, Proneural = 3), repetitions per sample (n = 3), Student’s t-test was used.
Overexpression of collagen triple helix repeat containing 1 is associated with poor prognosis of glioma
Using the median expression level of CTHRC1 mRNA as the cutoff, we evaluated the relationship between CTHRC1 mRNA expression level and survival time in different data sets. The results showed that patients with high CTHRC1 expression in the TCGA-GBMLGG data set had a lower overall survival (OS) in all grades of glioma (P < .05) (Figure 4A). Similar results were obtained in the CGGA data set (P < .05) (Figure 4B). These results suggest a poor prognosis associated with CTHRC1 overexpression in glioma. To further investigate whether the prognostic value of CTHRC1 differs across subgroups, we performed survival analysis using the TCGA-LGG data set and TCGA-GBM data set separately. Our analysis revealed a significant association between high CTHRC1 expression and poor prognosis in LGG (P < .05) (Figure 4C). In the survival analysis of GBM patients, although high CTHRC1 expression was associated with shorter OS compared with low expression, no statistical difference was observed (Figure 4D). This indicates a differential effect of CTHRC1 overexpression on the prognosis of LGG and GBM.

KM curves demonstrate poorer prognosis and subgroup differences in glioma patients with higher CTHRC1 expression. (A) TCGA-GBMLGG data set; (B) CGGA data set; (C) LGG subgroup; (D) GBM subgroup.
Collagen triple helix repeat containing 1 is an independent prognostic factor for glioma
To further investigate whether CTHRC1 is an independent prognostic factor in glioma, we performed univariate and multivariate Cox regression analyses for CTHRC1 expression and other clinical features (age, sex, race) in the TCGA-GBMLGG data set. The results showed that both univariate and multivariate Cox regression analyses demonstrated a significant association between age and CTHRC1 with OS (P < .05) (Table 1). These results suggest that CTHRC1 is an independent prognostic factor in glioma.
Univariate and multivariate Cox analysis in TCGA-GBMLGG RNA-seq set.

Correlation analysis of CTHRC1 and tumor immune cells
According to the analysis results of the TCGA LGG and GBM cohorts in the TIMER database, the expression of CTHRC1 in LGG tissues is positively correlated with CD8+ T cells (r = 0.36, P = 4.15e−16), neutrophils (r = 0.175, P = 1.23e−04), and dendritic cells (r = 0.159, P = 4.92e−04). However, it is not associated with purity (r = 0.014, P = 7.54e−01), B cells (r = 0.047, P = 3.03e−01), CD4+ T cells (r = −0.005, P = 9.15e−01), or macrophages (r = 0.089, P = 5.28e−02). On the contrary, in GBM tissues, the expression of CTHRC1 is negatively correlated with B cells (r = −0.255, P = 1.12e−02), CD4+ T cells (r = −0.281, P = 1.09e−03), and neutrophils (r = −0.269, P = 2.01e−03), and positively correlated with dendritic cells (r = 0.187, P = 3.15e−02). However, there is no correlation with purity (r = −0.149, P = 8.05e−02), CD8+ T cells (r = −0.074, P = 4.00e−01), or macrophages (r = −0.123, P = 1.62e−01) (Figure 5). These results indicate differential immune cell infiltration associated with CTHRC1 expression in LGG and GBM.
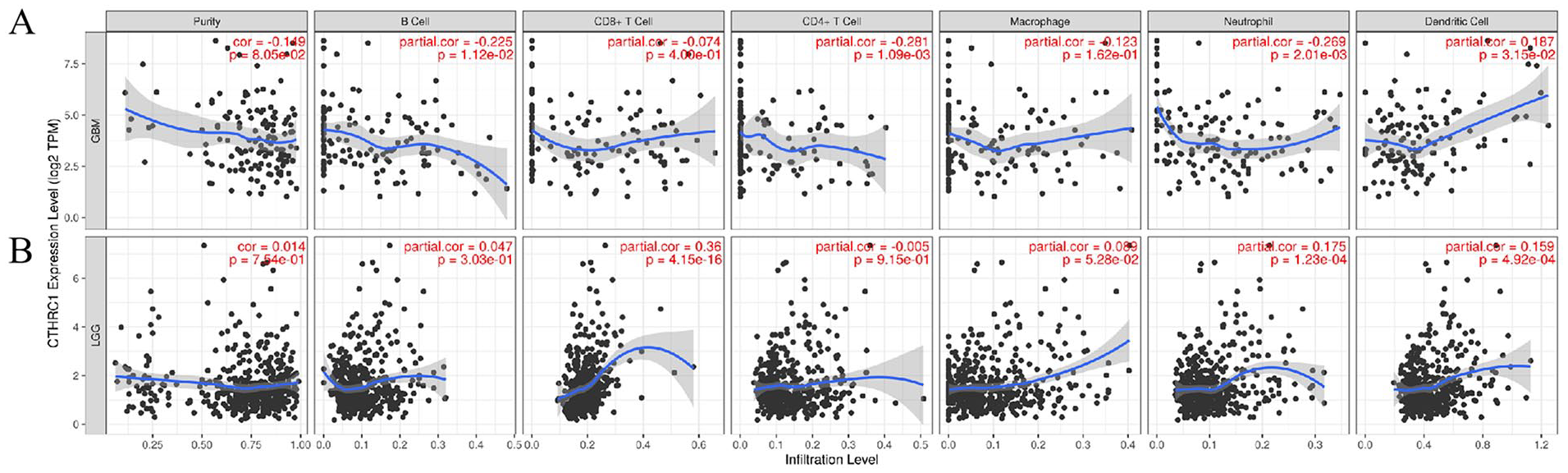
Correlation analysis of CTHRC1 with immune cell infiltration levels. (A) GBM; (B) LGG.
Cox proportional hazards models of collagen triple helix repeat containing 1 and tumor-infiltrating immune cells in low-grade glioma and glioblastoma
In the TIMER database, we used Cox proportional hazards models to assess the prognostic value of CTHRC1 levels and tumor-infiltrating immune cells in LGG and GBM. Low-grade glioma and GBM showed significant differences. The analysis results indicated a significant correlation (P < .05) between macrophage, CTHRC1, and clinical prognosis in LGG and between CD4+ T cells, dendritic cells, and clinical prognosis in GBM (P < .05). However, there was no significant correlation (P > .05) between CTHRC1 and clinical prognosis in GBM (Table 2). It is worth noting that this analysis revealed a correlation between CTHRC1 and clinical prognosis in LGG but not in GBM, further confirming the differential prognostic significance of CTHRC1 in LGG and GBM (Figure 5).
Cox proportional hazards models of CTHRC1 and 6 tumor-infiltrating immune cells in GBM and LGG.

Correlation analysis of collagen triple helix repeat containing 1 and markers of immune cell type
We evaluated the correlation between CTHRC1 expression and gene marker levels of tumor-infiltrating immune cells in LGG and GBM tissues by searching the TIMER database. Our overall findings revealed significant differences between LGG and GBM, with a significantly higher number of correlations between CTHRC1 and tumor-infiltrating immune cell markers in LGG tissues compared with GBM (Table 3). Specifically, in LGG tissues, CTHRC1 levels were closely associated with immune markers of CD8+ T cells, dendritic cells, natural killer cells, Treg cells, general T cells, tumor-associated macrophages (TAMs), Th1 cells, and Th17 cells. In contrast, in GBM tissues, except for immune markers of M2 macrophages and TAMs, there was no significant correlation between CTHRC1 and other tumor-infiltrating immune cell markers. These findings suggest a higher involvement of CTHRC1 in regulating tumor immune infiltration in LGG compared with GBM. The results indicate differential correlations between CTHRC1 and tumor-infiltrating immune cell markers in LGG and GBM, which may contribute to the differential prognostic significance of CTHRC1 in these 2 types of tumors.
Correlation analysis of CTHRC1 and immune cell type markers in LGG and GBM.

P < .05; **P < .01; ***P < .001; ****P < .0001.
Relationship between collagen triple helix repeat containing 1 and 28 tumor immunoinfiltrating cell subtypes
Furthermore, we further explored the relationship between CTHRC1 levels and 28 subtypes of tumor-infiltrating immune cells using the TISIDB database (Table 4). These results indicated that in LGG, CTHRC1 was positively correlated with 21 immune cell subtypes, especially activated CD8 T cells (r = 0.459, P < 2.2e−16). In GBM, CTHRC1 was positively correlated with 23 immune cell subtypes, particularly Gamma delta T cells (r = 0.548, P < 2.2e−16), Type 1 T helper cells (r = 0.522, P < 2.2e−16), and Regulatory T cells (r = 0.506, P < 2.2e−16). In both LGG and GBM, CTHRC1 showed no significant correlation with activated B cells, immature B cells, memory B cells, CD56bright natural killer cells, and eosinophils, indicating consistency. However, in LGG, CTHRC1 was not associated with effector memory CD8 T cells and macrophages, whereas in GBM, it was correlated, reflecting differences between LGG and GBM. Based on these results, LGG and GBM showed overall consistency in their correlations with 28 subtypes of tumor-infiltrating immune cells, but subtle differences were observed between LGG and GBM subgroups.
Relationship between CTHRC1 expression and human tumor lymphocyte infiltration.

Mutation, copy number variation, and methylation of collagen triple helix repeat containing 1
Collagen triple helix repeat containing 1 expression is significantly upregulated in gliomas. Therefore, we investigated the reasons for the elevated levels of CTHRC1. We analyzed the DNA methylation, gene mutations, and CNV of CTHRC1 in TCGA-GBMLGG using the UCSC Xena database. The heatmap showed that CTHRC1 mRNA expression was associated with CNV and DNA methylation but not with somatic mutations (Figure 6A). The mutation plot from cBioPortal for Cancer Genomics further confirmed the lack of somatic mutations in CTHRC1 in the 812 included patients/samples (Figure 6B). Therefore, we speculate that CNV and DNA methylation may contribute to the increased levels of CTHRC1 in gliomas.

Mutation, CNV, and methylation analysis of CTHRC1 in glioma. (A) Heatmap showing the correlations between CTHRC1 mRNA and somatic mutations, CNV, and methylation in TCGA-GBMLGG; (B) the correlation between CTHRC1 mRNA and somatic mutation in TCGA-GBMLGG.
Enrichment analysis of Gene Ontology and Kyoto Encyclopedia of Genes and Genomes genes associated with collagen triple helix repeat containing 1
We obtained 10 CTHRC1-related genes including COL1A1, FZD3, FZD9, POSTN, COL1A2, MMP2, VANGL2, TGFB1, COL5A1, and SULF1 using the STRING online tool (Figure 7A). The GO and KEGG enrichment analysis of these 11 genes, including CTHRC1, revealed enrichment in terms such as extracellular matrix organization, extracellular structure organization, external encapsulating structure organization, collagen-containing extracellular matrix, fibrillar collagen trimer, banded collagen fibril, platelet-derived growth factor binding, extracellular matrix structural constituent, and Wnt-protein binding (Table 5 and Figure 7B). The KEGG pathway enrichment analysis identified pathways such as Proteoglycans in cancer, AGE-RAGE signaling pathway in diabetic complications, and Relaxin signaling pathway (Table 5 and Figure 7B). Based on the results of the GO and KEGG enrichment analysis, it suggests that CTHRC1 mainly functions in the extracellular matrix and participates in tumor-related signaling pathways. These findings further support that CTHRC1 is a gene associated with tumor metastasis.

GO and KEGG enrichment analysis of CTHRC1-related genes. (A) CTHRC1-related genes; (B) Top3 enrichment of GO and KEGG in CTHRC1-related genes.
Results of GO and KEGG enrichment analysis of CTHRC1-related genes.

Discussion
Currently, there is limited research on the relationship between CTHRC1 and gliomas, and our understanding of it is not comprehensive. According to the literature search results from the PubMed database, only 5 studies have described the relationship between gliomas and CTHRC1, with only 2 studies specifically focusing on the relationship between gliomas and CTHRC1, whereas the other 3 studies only mentioned CTHRC1.17,18,28- 30 In a 2017 in vitro study, it was found that CTHRC1 expression was elevated in GBM cell lines, and knocking down CTHRC1 could inhibit GBM cell EMT and migration. 17 Another study in 2017 investigated glioma tissues and cell lines and demonstrated that CTHRC1 in gliomas could regulate cell invasion, migration, and adhesion through multiple pathways. 18 However, these study lacked research on the prognostic value of CTHRC1 and did not conduct more detailed investigations based on tumor grade, resulting in an incomplete understanding of CTHRC1.17,18 In a 2021 pan-cancer analysis, CTHRC1 was detected using qPCR, and the results showed that CTHRC1 expression in HGG tissues and glioma cell lines was higher compared with LGG and normal cell lines, which is consistent with our study findings. 28 However, this study was based on a single database and was a pan-cancer analysis, and the prognostic value and molecular function of CTHRC1 in gliomas have not been fully demonstrated. 28 In a 2022 bioinformatics analysis, 32 characteristic genes of GBM were identified, and CTHRC1 was mentioned to be highly expressed in GBM, but there was limited description of its role, and there was a lack of experimental validation and prognostic value research. 29 Based on the research gap in existing studies, our study comprehensively investigated the expression, prognostic value, and biological function of CTHRC1 in different types of gliomas, and made some new discoveries.
First, our study showed that the expression of CTHRC1 increases with the malignancy and invasiveness of gliomas, especially in the mesenchymal subtype, which is the most malignant and invasive subtype. Further IHC experiment confirmed that CTHRC1 is highly expressed in the mesenchymal subtype of GBM, suggesting its potential as a biomarker for diagnosing mesenchymal molecular subtype gliomas. In addition, studies have shown that different subtypes of GBM have different responses to aggressive treatments, with the classical subtype benefiting the most, the proneural subtype showing no benefit, and the mesenchymal subtype having the shortest median survival and poorest response to radiotherapy and chemotherapy. 31 Therefore, these molecular subtyping strategies are considered promising for predicting GBM prognosis and guiding treatment decisions. 32 Considering the close relationship between CTHRC1 and the mesenchymal subtype found in our study, CTHRC1 may also serve as a potential biomarker for predicting adverse prognosis and evaluating treatment outcomes in gliomas.
What puzzles us is that our study found that high expression of CTHRC1 is associated with poor prognosis in all grades of gliomas, but there are differences between LGG and GBM. In LGG, high expression of CTHRC1 predicts poor prognosis, whereas in GBM, it loses its predictive function. Although there is a trend toward predicting poor prognosis in GBM, it did not reach statistical significance. This suggests that CTHRC1 has a higher predictive value for poor prognosis in LGG, indicating that the accuracy of predicting poor prognosis based on high expression of CTHRC1 is higher in LGG than in GBM. It suggests that strong overexpression of CTHRC1 detected in LGG, which is traditionally considered to have a better prognosis, might be worth paying attention to because high expression of CTHRC1 may predict poor prognosis in LGG. The prognostic value of CTHRC1 in LGG may be greater than in GBM.
This study found that high expression of CTHRC1 is associated with poor prognosis in gliomas, as determined by both univariate and multivariate Cox analyses, indicating that CTHRC1 is an independent prognostic factor for gliomas. Similar findings have been observed in other tumor types, including gastric cancer and pancreatic ductal adenocarcinoma, where CTHRC1 has been identified as an independent prognostic factor for these cancers.33,34 However, our study results suggest that there are prognostic differences of CTHRC1 between LGG and GBM, indicating that the role of CTHRC1 may vary in these 2 tumor types. To better understand the role of CTHRC1 in different grades of gliomas, we compared the immune infiltration differences of CTHRC1 in LGG and GBM. We found significant differences in immune infiltration between LGG and GBM. In these differences, we noted that CTHRC1 is positively correlated with many markers of TAMs, and the correlation values (Cor) between CTHRC1 and TAM markers are generally higher in GBM compared with LGG. Studies have shown that TAMs promote cancer metastasis.35,36 It is well known that GBM is more prone to metastasis than LGG. These findings suggest that CTHRC1 may regulate TAMs involvement in the metastasis of both LGG and GBM, particularly in GBM.
This study found that CNV and DNA methylation may contribute to the increased levels of CTHRC1 in gliomas. DNA methylation and CNV play crucial roles in the regulation of epigenetics.37,38 Studies have shown that epigenetic gene reprogramming could be a fundamental driving force in gliomas. 39 This suggests that the abnormal elevation of CTHRC1 may be regulated by epigenetic mechanisms, thereby enhancing the invasive ability of gliomas. In addition to this study, other research has also found a close association between the abnormal elevation of CTHRC1 and CNV and DNA methylation in kidney renal papillary cell carcinoma and kidney renal clear cell carcinoma. 40 This provides further evidence for the close relationship between the abnormal elevation of CTHRC1 and epigenetic regulation. These findings open up new avenues and directions for future targeted interventions against CTHRC1. Therefore, the results of these studies provide new insights and directions for future targeted interventions against CTHRC1.
In addition, we further validated the function of CTHRC1 based on bioinformatics analysis. Previous studies have demonstrated that CTHRC1 promotes the migration of glioma cells, and its mechanism is closely associated with CTHRC1-induced tumor cell EMT. The results of GO and KEGG enrichment analysis in this study indicate that CTHRC1 primarily functions in the extracellular matrix and participates in tumor-related signaling pathways. This suggests a close correlation between CTHRC1 and tumor cell migration, which is consistent with most of the current research findings. Most studies have confirmed the strong association between CTHRC1 and tumor cell migration. Overall, these findings provide further evidence for the role of CTHRC1 in glioma and its involvement in tumor cell migration.14-16
There are several limitations to this study. First, the study was unable to further clarify the reasons behind the association between CTHRC1 overexpression and poor prognosis in gliomas, as well as the differences observed between LGG and GBM. It can only be speculated that the differences may be related to variances in immune infiltration, which warrants further investigation in the future. Second, this study merely superficially validated the previously reported biological functions of CTHRC1 in gliomas through bioinformatics analysis, without conducting in-depth experiment to explore its specific mechanisms of action. Although the study confirmed the role of CTHRC1 in promoting tumor cell migration through GO and KEGG enrichment analysis, the specific mechanisms are still unclear. In summary, this study mainly affirms the potential clinical value of CTHRC1, but further research is needed to elucidate the underlying mechanisms.
Conclusions
This study revealed previously unknown clinical value of CTHRC1 in gliomas. For example, it identified CTHRC1 as a potential biomarker for the mesenchymal molecular subtype of gliomas. It also found that high CTHRC1 expression is associated with poor prognosis in gliomas, with differences observed between LGG and GBM. Collagen triple helix repeat containing 1 was identified as an independent prognostic factor in gliomas. Furthermore, the study demonstrated significant differences in immune infiltration of CTHRC1 between LGG and GBM. The CNV and DNA methylation were identified as potential factors leading to elevated levels of CTHRC1 in gliomas. These findings provide novel directions and insights for the potential identification of therapeutic targets in the treatment of gliomas.
Supplemental Material
sj-docx-1-onc-10.1177_11795549241260576 – Supplemental material for Expression, Prognostic Value, and Biological Function of CTHRC1 in Different Types of Gliomas: A Bioinformatic Analysis and Experiment Validation
Supplemental material, sj-docx-1-onc-10.1177_11795549241260576 for Expression, Prognostic Value, and Biological Function of CTHRC1 in Different Types of Gliomas: A Bioinformatic Analysis and Experiment Validation by Xueping Shi, Xi Zeng, Rukai Jiao, Yushi Yang, Xiaolin Du, Jiacai Qian, Jian Liu and Guangtang Chen in Clinical Medicine Insights: Oncology
Supplemental Material
sj-docx-2-onc-10.1177_11795549241260576 – Supplemental material for Expression, Prognostic Value, and Biological Function of CTHRC1 in Different Types of Gliomas: A Bioinformatic Analysis and Experiment Validation
Supplemental material, sj-docx-2-onc-10.1177_11795549241260576 for Expression, Prognostic Value, and Biological Function of CTHRC1 in Different Types of Gliomas: A Bioinformatic Analysis and Experiment Validation by Xueping Shi, Xi Zeng, Rukai Jiao, Yushi Yang, Xiaolin Du, Jiacai Qian, Jian Liu and Guangtang Chen in Clinical Medicine Insights: Oncology
Supplemental Material
sj-docx-3-onc-10.1177_11795549241260576 – Supplemental material for Expression, Prognostic Value, and Biological Function of CTHRC1 in Different Types of Gliomas: A Bioinformatic Analysis and Experiment Validation
Supplemental material, sj-docx-3-onc-10.1177_11795549241260576 for Expression, Prognostic Value, and Biological Function of CTHRC1 in Different Types of Gliomas: A Bioinformatic Analysis and Experiment Validation by Xueping Shi, Xi Zeng, Rukai Jiao, Yushi Yang, Xiaolin Du, Jiacai Qian, Jian Liu and Guangtang Chen in Clinical Medicine Insights: Oncology
Footnotes
Acknowledgements
The authors extend their heartfelt appreciation to the patients who generously donated glioma tissue samples for this study.
Author Contributions
XS conducted the IHC and Western blot experiment. XZ performed the bioinformatic analysis. RJ wrote the original manuscript. YY assisted with IHC experiment. XD assisted with data analysis. JQ assisted with data analysis. JL provided the funding and guided the experiments. GC designed the project and revised the manuscript.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by Key Discipline Construction Funds for Neurosurgery (ZDPT-2023-01) and the Guizhou Surgical Talent Training Base (The fourth batch) (RCJD-02).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The original contributions presented in the study are included in the article. Further inquiries can be directed to the corresponding authors.
Ethics Approval and Consent to Participate
This study was approved by the Ethical Committee of Affiliated Hospital of Guizhou Medical University (Ethics Approval No. 2022090), and the written informed consent of the patient or guardian was obtained.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
