Abstract
Nasopharyngeal carcinoma (NPC) is one of the most common head and neck malignancies, and the primary treatment methods are radiotherapy and chemotherapy. Radiotherapy alone, concurrent chemoradiotherapy, and induction chemotherapy combined with concurrent chemoradiotherapy can be used according to different grades. Treatment options and prognoses vary greatly depending on the grade of disease in the patients. Accurate grading and risk assessment are required. Recently, radiomics has combined a large amount of invisible high-dimensional information extracted from computed tomography, magnetic resonance imaging, or positron emission tomography with powerful computing capabilities of machine-learning algorithms, providing the possibility to achieve an accurate diagnosis and individualized treatment for cancer patients. As an effective tumor biomarker of NPC, the radiomic signature has been widely used in grading, differential diagnosis, prediction of prognosis, evaluation of treatment response, and early identification of therapeutic complications. The process of radiomic research includes image segmentation, feature extraction, feature selection, model establishment, and evaluation. Many open-source or commercial tools can be used to achieve these procedures. The development of machine-learning algorithms provides more possibilities for radiomics research. This review aimed to summarize the application of radiomics in NPC and introduce the basic process of radiomics research.
Introduction
Nasopharyngeal carcinoma (NPC) is the most common type of head and neck squamous cell carcinoma. According to the 2018 Global Cancer Statistics, 129 100 new cases and 73 000 NPC deaths were expected to occur globally in 2018. 1 The incidence of NPC has apparent regional aggregation. Globally, NPC is more common in southern China, southeastern Asia, and northern Africa; in China, it is common in the southern and southwestern provinces.2,3 Medical imaging examinations, such as computed tomography (CT), magnetic resonance imaging (MRI), or positron emission tomography (PET), play an important role in the diagnosis and treatment of NPC and are widely used in the early detection, diagnosis, staging, and evaluation of treatment response. However, emerging radiomics studies have shown that much invisible high-dimensional information remains to be discovered. Recently, radiomics has provided information on tumor heterogeneity, such as tumor cellularity, degenerative changes, and neovascularization, which are difficult to assess visually. 4 As an effective tumor biomarker of NPC, the radiomic signature has been widely used in grading, differential diagnosis, prediction of prognosis, evaluation of treatment response, and early identification of therapeutic complications. The development of machine-learning algorithms provides powerful tools for processing and analyzing high-dimensional image data. Radiomic features combined with machine learning can achieve precise stratification of tumor patients, providing more evidence for individualized diagnosis and treatment. 5 The purpose of this review was to summarize the application of radiomics in NPC and to introduce the basic process of radiomics research.
Grading and stratification of risk
For patients with NPC, treatment options vary greatly among patients at different stages of disease. It is important to identify effective biomarkers for stratifying patients into different risk groups. Currently, the American Joint Committee on Cancer (AJCC) 8th tumor, nodes, and metastases (TNM) staging system, widely used in the staging of patients with NPC, has guidelines for the selection of treatment plans and the evaluation of prognosis. However, studies have shown that the currently used AJCC 8th TNM staging system has limitations regarding risk stratification; the survival curves for T2 and T3 patients almost overlap, without significant differences in locoregional recurrence-free survival (LRFS) (P = .606) and disease-free survival (DFS) (P = .735); similarly, the overall survival (OS) and DFS curves for stage II and III patients were overlapping. 6 Radiomic nomograms can be developed for the staging and stratification of patients with NPC, and studies have shown superiority over the current clinical TNM staging system. Kim et al established survival models based on MRI radiomics for early risk assessment by predicting the progression-free survival (PFS) of patients with NPC. The prognostic performances of clinical + stage + radiomics survival models (area under the receiver operating characteristics [ROC] curve [AUC], 0.80; 95% confidence interval [CI]: 0.80-0.81) was better than that of the clinical stage (AUC, 0.70, 95% CI, 0.69-0.71). 7 Moreover, Zhuo et al analyzed the feasibility of MRI-based radiomic features of 658 non-metastatic patients with NPC to stratify them into distinct survival subgroups. The obtained model achieved a better classification effect (C-index = 0.827, P < .004, and C-index = 0.814, P < .002) than the T-stage (C-index = 0.815, P = .002 and C-index = 0.803, P = .024). It showed a competitive but more stable classification effect than the TNM staging system (C-index = 0.842, P = .003 and C-index = 0.765, P = .050). 8 In addition, Feng et al used PET/MR image characteristics to establish models for grading patients with NPC. In the PET model, the AUC, accuracy, sensitivity, and specificity of the training cohort were 0.84, 0.75, 0.90, and 0.69, respectively. For the MR model, these were 0.85, 0.83, 0.75, and 0.86, respectively (Table 1). 9
NPC radiomics studies of grading and stratification of risk.
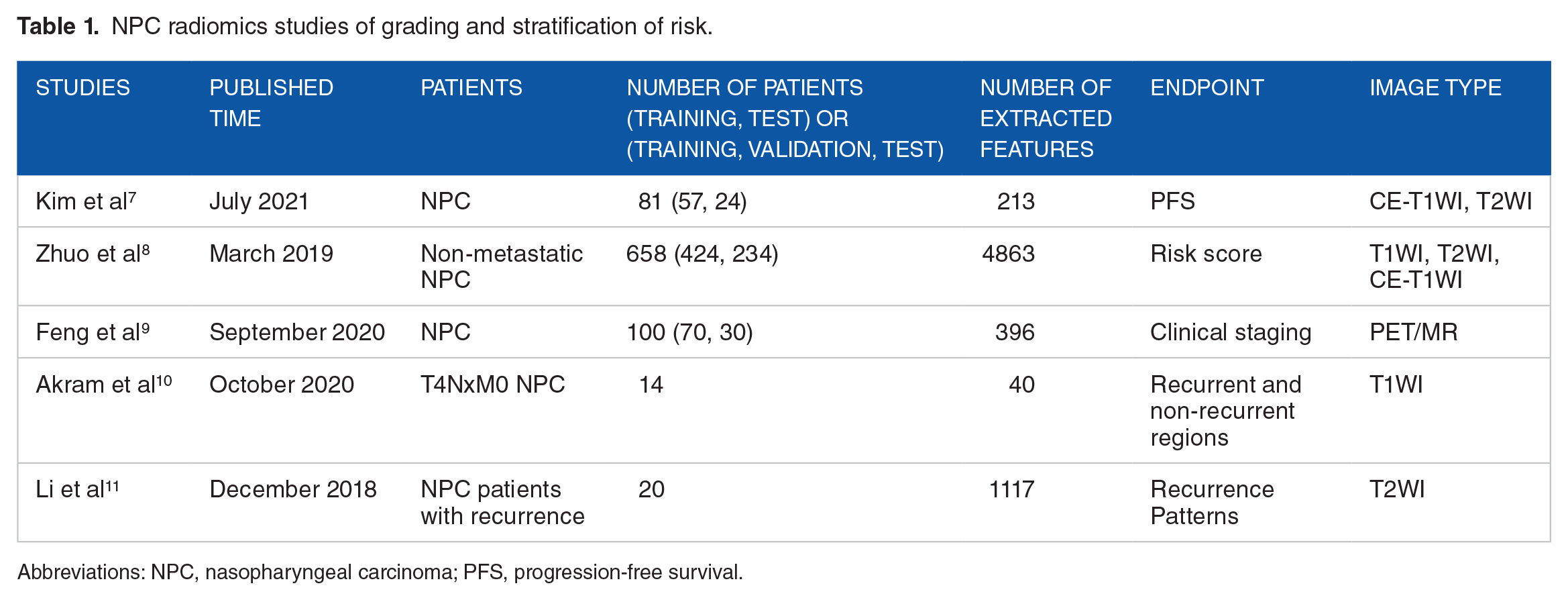
Abbreviations: NPC, nasopharyngeal carcinoma; PFS, progression-free survival.
Radiomic features can also be used to assess intratumor radio resistance so that patients with a high risk of recurrence can be identified early. Akram et al compared the components extracted from recurrent and non-recurrent tumor regions at pretreatment MRI of 14 patients with recurrent NPC. They found a difference in the radiomic pattern between the recurrent and non-recurrent regions within the tumor. 10 Li et al analyzed pretreatment MRI features to distinguish patients with in-field recurrence or non-progression after intensity-modulated radiation therapy (IMRT). The AUC values of the classification models ranged from 0.727 to 0.835. 11 The results showed that radiomic features could serve as imaging biomarkers to identify patients at risk of in-field recurrence.
Differential diagnosis
Several previous studies have shown that radiomics can be applied for the differential diagnosis of tumors.12,13 In NPC, PET-based radiomic features have achieved good performance that are superior to traditional PET parameters in differential diagnosis.14,15 Traditional parameters commonly used to reflect metabolic information in PET include standardized uptake value (SUV)max, SUVmean, metabolic tumor volume (MTV), and total lesion glycolysis (TLG), among others. These parameters are associated with the patients’ prognosis.16,17 Lv et al reported that some PET/CT-based radiomic features, such as SumEntropy, short-zone low gray-level emphasis (SZLGE), and low gray-level zone emphasis (LGZE), performed better than conventional metrics for differentiating NPC from chronic nasopharyngitis. 14 Du et al 15 showed that a PET/CT-based radiomic signature achieved a higher AUC value than conventional metrics in the differentiation between local recurrence and inflammation (Table 2).
NPC radiomics studies of differential diagnosis.

Abbreviations: CN, chronic nasopharyngitis; CT, computed tomography; NPC, nasopharyngeal carcinoma; PET, positron emission tomography.
Prediction of prognosis
In NPC, prognosis varies greatly among patients at different stages of disease. In addition, the prognosis may vary significantly after the standard treatment in patients at the same TNM stage of disease because the TNM stage is mainly based on the traditional imaging data of the anatomical structure. Therefore, a large amount of structural and functional information may be missed. 18 Radiomic signatures obtained from CT, MRI, and PET can be used as an effective tool for predicting local recurrence, distant metastasis, OS, and PFS in patients with NPC. A nomogram developed by the CT-based Radscore and potential clinical characteristics was used by Zhu et al to predict the risk of local recurrence in patients with NPC who received IMRT. The C-index, specificity, and sensitivity of the nomogram were 0.931 (95% CI, 0.8765-0.9856), 91.2%, and 82.8% in the training cohort, and 0.799 (95% CI, 0.6458-0.9515), 79.4%, and 69.2% in the validation cohort, respectively. 19 Yan et al developed a CT-based radiomic nomogram to predict PFS in patients with locoregionally advanced NPC. They found that the nomogram integrating the radiomic signature and clinical characteristics achieved better predictive performance (C-index, 0.873; 95% CI, 0.803-0.943) than that of the clinical nomogram (C-index, 0.729; 95% CI, 0.620-0.838) and the TNM staging system (C-index, 0.689; 95% CI, 0.592-0.787) (Table 3). 20
NPC radiomics studies on prediction of prognosis.

Abbreviations: AC, adjuvant chemotherapy; CCRT, concurrent chemotherapy radiation treatment; DFS, disease-free survival; D-FFS, distant failure-free survival; DM, distant metastasis; DMFS, distant metastasis-free survival; FFS, failure-free survival; IC, induction chemotherapy; IMRT, intensity-modulated radiation therapy; LR, local recurrence; LRFS, locoregional recurrence-free survival; LR-FFS, locoregional failure-free survival; OS, overall survival; WSI, whole slide images.
Moreover, recent evidence suggests that MRI-based radiomics, combined with clinical information, can be used as an effective tool for predicting local recurrence, distant metastasis, OS, and PFS in patients with NPC. Zhang et al built classifiers for the prediction of locoregional recurrences or distant metastases in patients with advanced NPC. Results showed that the combination of feature selection method random forest (RF) and classification method RF achieved the best performance (AUC, 0.8464 ± 0.0069; test error, 0.3135 ± 0.0088). 21 In another study, Zhang et al 22 showed that the radiomic classifiers based on the combined contrast-enhanced T1-weighted imaging (CET1WI) and T2-weighted imaging (T2WI) (AUC: 0.886, 95% CI, 0.815-0.956) achieved better predictive performance to identify patients with progression than classifiers based on CET1WI (AUC: 0.793, 95% CI, 0.698-0.889) or T2WI alone (AUC: 0.813, 95% CI, 0.721-0.904). Zhang et al also built a nomogram based on the radiomics signature and TNM staging system to predict PFS, which achieved better performance than the TNM staging system only (C-index, 0.761 vs 0.514; P < 2.68 × 10–9). In addition, a radiomics nomogram established by the radiomics signature and clinical characteristics achieved better performance than the nomogram based on clinical characteristics alone (C-index, 0.776 vs 0.649; P < 1.60 × 10–7). 23 Furthermore, Zhang et al built an MRI-based model with 7 features to classify patients into high- or low-risk distant metastasis groups. Results showed a significant difference in OS between the high- and low-risk groups (P < .001). In addition, a nomogram based on combined radiomic and clinical features achieved a significant predictive performance for distant metastasis (AUC, 0.827; 95% CI, 0.754-0.900 in the training cohort and AUC, 0.792; 95% CI, 0.633-0.952 in the validation cohort). 24 In another Zhang et al 25 study, a nomogram combining radiomic features with clinical features achieved good predictive performance of LRFS (C-index: 0.74 95% CI, 0.58-0.85) in the validation cohort, and successfully categorized the patients into low- or high-risk groups with significant differences in LRFS (P < .05). Bologna et al developed MRI-based radiomic signatures to predict the OS, DFS, LRFS, and distant metastasis-free survival (DMFS) of non-endemic Epstein–Barr virus (EBV)-related patients with NPC. The results showed that radiomic signature achieved good predictive performance for OS and LRFS, with C-index values of 0.68 and 0.72, respectively. They also found that the combination of radiomic and clinical features improved prognostic performance. 26 In another study, Ming et al also developed nomograms to predict the OS, DFS, LRFS, and DMFS of patients with NPC. They found that nomograms combining radiomic features with clinical characteristics could predict DFS with a C-index of 0.751 (0.639, 0.863) and OS with a C-index of 0.845 (0.752, 0.939), which improved the prediction accuracy for clinical characteristics only. 27 In addition, Ouyang et al established predictive models to stratify stage III to IVB patients with NPC into low- or high-risk groups for PFS. The results revealed radiomic signature as an independent predictor of PFS (hazard ratio [HR] = 5.14 in the training set, HR = 7.28 in the validation set). 28 Yang et al 29 developed a nomogram based on radiomic signatures of nasopharynx gross tumor volume (GTVnx) and GTV of cervical lymph node (GTVnd), planning score (PS), and clinical characteristics to predict the PFS of locoregionally advanced patients with NPC, with a C-index of 0.811 (95% CI, 0.74-0.882), and outperformed that of the TNM stage alone (C-index, 0.613, 95% CI, 0.532-0.694) in the validation cohort. In another study, 5 models based on clinical data, overall stage, radiomics, radiomics + overall stage, radiomics + overall stage + EBV-DNA, respectively, were established by Shen et al to predict the PFS of non-metastatic patients with NPC. Among them, the model incorporating radiomics, overall stage, and EBV DNA yielded the highest C-index (training cohort: 0.805, validation cohort: 0.874). 30
Moreover, patients have been stratified according to T-stage in some research, and predictive models were developed based on patients with a specific T-stage. Zhang et al established a radiomics signature to predict LRFS in multicenter patients with non-metastatic T4 NPC. A nomogram that combined the Radscore with clinical characteristics showed good prediction performance (C-index: 0.810 in the training cohort, 0.807 in the internal validation cohort, and 0.753 in the external validation cohort). 31 Wu et al 32 stratified patients with NPC into 2 groups (T1-2 and T3-4 stages) to explore the performance of a model for predicting PFS of MRI-based radiomics signatures and the radiomic models based on pre-stratified tumor stages had better predictive performance.
Furthermore, deep-learning-based radiomics has also been widely used as a tumor biomarker for predicting the prognosis of patients with NPC. It is usually combined with other clinical characteristics and achieves good predictive performance. In a study by Zhang et al, radiomic and deep-learning-based histopathologic signatures were established to predict the failure-free survival (FFS) of patients with NPC, both of which have presented good predictive performance for treatment failure (C-index: 0.689-0.779, all P < .050). The multiscale nomogram showed better performance than the clinical model in the external test cohorts (C-index: 0.834 vs 0.679, P < .050). 33 Zhong et al developed 3 deep-learning-based radiomic signatures based on T1WI, T2WI, and CET1WI to predict DFS of patients with T3N1M0 NPC, all of which were significantly correlated with DFS (C-index: 0.695-0.731, all P < .001 in the training cohort and C-index: 0.706-0.755, all P < .001 in the validation cohort). The combined model of radiomic signatures and clinical features improved the predictive effect compared with the clinical model (C-index: 0.771 vs 0.640, P < .001 in the training cohort and C-index: 0.788 vs 0.625, P = .001 in the validation cohort). 34 In a multicenter study, Zhong et al revealed that deep-learning-based radiomic nomograms for predicting the DFS of advanced patients with NPC who received concurrent chemoradiotherapy (CCRT) or induction chemotherapy (ICT) + CCRT showed excellent prognostic ability in all groups. In the ICT-preferred group, patients who received ICT + CCRT showed better survival than those who received only CCRT (HR: 0.17, P < .001 in the internal cohort and 0.24, P = .02 in the external test cohort), whereas the opposite was observed in the CCRT-preferred group (HR: 6.24, P < .001 in the internal cohort and 12.08, P < .001 in the external test cohort). 35
In addition, recent studies have suggested that PET/CT-based radiomics can be used to predict PFS in patients with NPC.36,38 Local recurrence and distant metastasis of locally advanced NPC can be predicted using the radiomic signature of pretreatment PET/CT. 37
Prediction of treatment response and identification of therapeutic complications
For patients with NPC, the treatment response of patients in the same stage may differ because of different levels of sensitivities to the treatment. The selection of radiotherapy dose and chemotherapy regimen often requires careful consideration of disease control and prevention of therapeutic side effects. Research has shown that when CCRT is performed, patients may experience increased toxicity while achieving good local control. 39 Radiomics has been used to evaluate therapeutic responses after radiotherapy or CCRT in some studies.40,41 In addition, Yu et al analyzed radiomic features extracted from pretreatment MRI to predict tumor shrinkage. The results showed that the model obtained by combining elements of T1 and T2 images had the best prediction effect, and the AUCs in the training and testing sets were 0.984 (95% CI, 0.983-0.984) and 0.930 (95% CI, 0.928-0.933), respectively. For patients with obvious tumor shrinkage, adaptive radiotherapy can be applied so that treatment plans can be optimized in a timely manner. 41 Furthermore, the efficacy of ICT in patients with locally advanced NPC remains controversial, and there is uncertainty about whether a patient can benefit from ICT.42,43 Therefore, whether ICT is needed for patients requires more precise evaluation criteria. Recent studies have shown that CT, 44 MRI,45 -47 and PET 48 radiomics could be used to predict ICT or neoadjuvant chemotherapy response, and it is expected to be an effective biomarker to evaluate whether a patient could benefit from ICT (Table 4).
NPC radiomics studies on prediction of treatment response and recognition of therapeutic complication.

Abbreviations: DFS, disease-free survival; IC, induction chemotherapy; NACT, neoadjuvant chemotherapy; NPC, nasopharyngeal carcinoma; RTLI, radiotherapy-induced temporal lobe injury.
Moreover, the early identification of therapeutic complications using radiomics has also achieved excellent results. Liu et al conducted a study to analyze changes in CT-based radiomics in the parotid glands during radiotherapy to predict the incidence of acute xerostomia. They obtained a prediction model with a precision of 0.9220 and sensitivity of 100%. 49 Zhang et al obtained 3 prediction models based on MRI-based radiomic features extracted from the medial temporal lobe, gray matter, and white matter to predict radioactive brain injury. The mean AUCs were 0.830 (95% CI, 0.823-0.837), 0.773 (95% CI, 0.763-0.782), and 0.716 (95% CI, 0.699-0.733), respectively. 50 Hou et al selected 14 T2WI-based features to establish a radiomic signature to identify radiotherapy-induced temporal lobe injury (RTLI) in patients with NPC. A radiomics nomogram combining the radiomic signature with clinical characteristics achieved better classification performance (AUC, 0.87; 95% CI, 0.82-0.91) than the radiomic (AUC, 0.71; 95% CI, 0.65-0.78) and clinical (AUC, 0.73; 95% CI, 0.67-0.79) models. 51
Workflow
Image segmentation
At present, many open-source or commercial software programs can be used for 2-dimensional (2D) and 3-dimensional (3D) image segmentation. The commonly used open-source software includes LIFEx (https://www.lifexsoft.org), itk-snap (http://www.itk-snap.org), and 3Dslicer (https://www.slicer.org). Due to the complex surrounding anatomical structures, irregular tumor shapes, and indistinct boundaries between the tumor and surrounding tissue, regions of interest (ROIs), which are often manually delineated by experienced radiologists or radiation oncologists, are usually affected by the subjective judgment of different observers. Therefore, interobserver and intra-observer intraclass correlation coefficients (ICCs) can be calculated to verify the repeatability of ROI delineation.27,30 Unenhanced and enhanced CT are commonly used in CT-based radiomic studies. Previous studies, which revealed that enhanced and unenhanced CT-based radiomic features can reflect different aspects of tumor heterogeneity, showed inconsistent results.52,53 For NPC, compared with unenhanced images, enhanced CT can better display tumor boundaries and enlarged lymph nodes, which is more helpful for image segmentation. The most used MR images are T1WI, T2WI, and enhanced T1WI. Enhanced T1WI can better show tumor boundaries, but may cause an overestimation of tumor size due to surrounding edematous areas. 26 In differential diagnosis, prediction of prognosis, and treatment response studies, the ROIs are usually based on the main tumor and the largest lymph node. Cluster analysis results showed that the main tumor characteristics correlated with T-stage and overall stage but not with N-stage, and lymph node characteristics correlated with N-stage. 27 In the prediction of treatment side effects, ROIs were selected according to different research objectives. To extract stable features, image preprocessing is essential to exclude the influence of different machines, scanning parameters, reconstruction algorithms, and noise.54,55 The procedures of image preprocessing usually include resampling, standardization, normalization, smoothing, and filtering (Figure 1).

Workflow of radiomics.
Feature extraction
Feature extraction is usually achieved using open-source software such as Pyradiomics (https://pyradiomics.readthedocs.io), LIFEx, and 3D slicer. The number of features extracted by the different tools vary slightly. Standard image features include the following categories:
Intensity histogram-based features
Shape-based features
First-order texture features
Second-order texture features
Higher-order texture features
Previous studies have shown that feature extraction is affected by different extraction tools and has poor repeatability and stability.56,57 Zwanenburg et al 58 standardized the extraction of 169 standard features that can be used to verify and calibrate other software and provide a direction for subsequent research. In addition, the Image Biomarker Standardization Initiative (IBSI) Reference Manual provides guidelines for feature extraction.58,59
Feature selection
The purpose of feature selection is to reserve meaningful features from the many extracted features, for further analysis. It is important to select the appropriate features for the models. Too many features will cause overfitting, while too few will result in unsatisfactory prediction performance of the models. Feature selection methods commonly used mainly consist of filtering, wrapping, and embedding methods. 60 Filtering methods is used to filter the features according to a certain threshold, such as by the correlation coefficient method, t test, chi-square test, and variance selection method. Wrapping methods perform multiple iterations and retain several characteristics according to the objective function, such as recursive feature elimination (RFE), RF, support vector machine (SVM), and K-nearest neighbor (KNN). Embedding methods such as least shrinkage and selection operator (LASSO), ridge regression, and gradient boosting decision tree (GBTD) are combined with machine-learning algorithms to obtain the weight coefficients of each feature so that the features can be sorted and filtered. 60 These 3 methods can be used in combination. Currently, LASSO is the most used feature selection algorithm in NPC radiomic research and has achieved high selection efficiency. However, LASSO often ignores the pairwise correlations of selected features. To address this problem, Pearson correlation coefficient (PCC) can be computed to exclude features with pairwise correlations.10,41 In addition, dimensionality reduction methods, such as principal components analysis (PCA) and linear discriminant analysis (LDA), are also often used for variable selection. Dimensionality reduction methods mainly combine several variables to obtain new variables based on the relationship between variables. It changes the original features and is different from the feature selection methods.
Deep-learning-based radiomics
In recent years, deep learning based on neural networks has provided a new approach for radiomic research. Deep learning can achieve image feature selection in a fully automated and end-to-end manner without the need for accurate ROI segmentation, so that the influence of image preprocessing, manual segmentation, and feature extraction can be reduced.34,61 However, a sufficient sample size is required for deep learning.
Establishment of nomogram or machine-learning classifiers
In previous studies of NPC radiomics, in addition to the selected radiomic features, some meaningful clinical indicators could also be included, such as age, stage, grade, complete blood count, and biochemical analysis. Other tumor biomarkers, such as C-reactive protein and EBV-DNA, can also be combined with radiomic features. After obtaining meaningful features, nomograms or machine-learning classifiers were established for further analysis. Nomograms are often based on multivariate Cox proportional hazards models. Risk scores for each patient and survival curves were obtained for prognosis prediction and comparison between different survival subgroups. Machine-learning classifiers are often established using machine-learning algorithms. Many machine-learning methods, such as SVM, LDA, KNN, and decision tree, have achieved good prediction accuracy. These machine-learning algorithms can be used directly in Python (https://www.python.org), R software (https://www.r-project.org), and MATLAB (https://www.mathworks.com).
Validation and evaluation
Research data are often divided into training and validation sets, which often included internal and external validation sets. Internal validation sets are used to obtain the optimal model parameters, while external validation sets are used to validate and evaluate the predictive performance of the models. The number of patients in the training and validation sets often needs to be divided according to the requirements and specific conditions. For studies with a small sample size, internal verification can be carried out by holdout and k-fold cross-validations. The holdout cross-validation method divides the training sets into 2 parts: (1) used for training and (2) used for parameter adjustment. The model parameters obtained using this method are easily affected by the division ratio. In k-fold cross-validation, the training sets are divided into k-folds without replacement, with k – 1 folds used for training and 1-fold for validation. This process is repeated k times to obtain k models and their performance evaluations. The results of k times were averaged to obtain the final predicted performance. When a model is obtained through the training sets, it is verified through the validation sets. An independent external validation set that is not involved in the training process is very important for the evaluation and verification of the model’s predictive performance. Some evaluation parameters are calculated to evaluate the obtained models. For classification models, commonly used evaluation parameters include but are not limited to accuracy, sensitivity, specificity, AUC, and F1-scores. For Cox proportional hazards models or nomograms, commonly used evaluation indicators include but are not limited to the C-index and AUC.
Disadvantages in Radiomics Studies of NPC
Although excellent results have been achieved in the application of radiomics in NPC, there are still many problems. First, image feature extraction is affected by various factors, and its stability and repeatability are still insufficient. Further research is needed to standardize the feature extraction. Second, most of the studies were single centered and lacked external validation. Finally, although the models can achieve high accuracy, the significance of many features cannot be explained. The biological meaning behind radiomic features is worth exploring, such as the relationship between radiomic features and computational pathology features or gene expression. Zhang et al explored the correlation between the main radiomic features and genetic alterations. The results showed that the 2 texture features were related to the genetic alterations of chromatin remodeling pathways. 33 Gao et al 62 also found a significant correlation between gene expression and Radscore, especially the mRNA expression of CDKL2, PLIN5, and SPAG1. Moreover, several studies have shown that radiomics features are correlated with human papilloma virus (HPV) expression status in head and neck squamous cell carcinoma. 63 In NPC, more research is needed to explore the interpretability of radiomics before it can be broadly adopted.
Prospects of Radiomics Studies on NPC
For radiomics in NPC, many issues need to be studied further. First, more multicenter NPC radiomics studies should be conducted to improve the applicability and stability of radiomic features and prediction models. Second, deep learning based on neural networks can be applied. This approach can solve the problems of poor repeatability and stability of manual segmentation and feature extraction. Finally, radiogenomics, a combination of radiomics and genomics can be further studied in NPC. The relationship between image features and gene expression needs to be further explored, which is conducive to the interpretation of radiomics.
In addition, radiomics could be used as a tumor biomarker to predict whether patients will benefit from cetuximab, nimotuzumab, or immune checkpoint inhibitors, which can provide a new basis for the selection of individualized treatment plans for patients. To the best of our knowledge, most radiomic studies of NPC focus on the whole tumor, ignoring the heterogeneity of different regions within the tumor. A study conducted by Xu et al 36 showed that a PET/CT-based subregional radiomic signature achieved better prediction performance than that of the whole tumor. Subregional radiomics analysis can be applied to NPC. Moreover, radiotherapy causes a series of radiation side effects, such as oral mucositis, xerostomia, taste failure, mouth opening difficulty, and hearing damage. Radiomic features can be used to identify the side effects of radiotherapy. It is beneficial to take individual preventive measures in patients. For patients with NPC who undergo radiotherapy and chemotherapy, food intake may be affected, due to oral and laryngeal mucositis, taste failure, nausea, vomiting, and other adverse reactions, leading to weight loss.64,65 Studies have shown that nutritional status has a particular impact on the prognosis of patients with NPC. CT-based body composition information has been used as healthy and prognostic indicators in prostate cancer. 66 This method can be extended to patients with NPC. It can be used to identify patients at risk of malnutrition. In addition, radiomics can be used as a reference for the design of radiotherapy plan in the future. Studies by Akram et al 10 and Li et al 11 have shown that radiomic features can be used to assess intratumor radioresistance. These results can be used as a reference to accurately define areas with a high risk of recurrence, which may require a higher dose. In addition, cone beam computed tomography (CBCT) is one of the most commonly used imaging guidance methods in radiotherapy. It is used for patient positioning before radiotherapy and can detect the treatment target and external contour changes in a timely manner. The response of the tumor to the radiation dose can be studied by analyzing the CBCT images. The prediction of prognosis and treatment response based on the features extracted from CBCT images has been applied in lung cancer67,68 and prostate cancer 69 but rarely in head and neck squamous cell carcinoma.
Conclusions
The purpose of this review was to summarize the application of radiomics in NPC and to introduce the basic process of radiomics research. As mentioned in this article, radiomics has been widely used in grading and stratification of risk, differential diagnosis, prediction of prognosis, prediction of treatment response, and identification of therapeutic complications, and good results have been achieved. A mature process has been established for the research method, including image segmentation, feature extraction, feature selection, model establishment, and evaluation. However, there are still many problems, such as insufficient stability and repeatability of feature extraction, and lack of interpretation of features. Although radiomics has promising application prospects in patients with NPC, more research is needed to further promote the application of radiomics in clinical practice. Radiomics has good prospects for NPC as a rapidly developing field.
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The publication of this article was supported by People’s Liberation Army The General Hospital of Western Theater Command.
Declaration Of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
WD summarized the articles and wrote the manuscript. BX, TT, XZ, and ZH helped for article selection and gave valuable advice to the manuscript. LZ presented the idea and revised the manuscript. All authors have read and approved the final manuscript.
