Abstract
Whole genome sequencing is an important milestone of the genomics-assisted molecular biology studies. Sequencing the whole genome of plants unravels the complex molecular structure of these complex organisms. In this respect, whole genome sequencing of important small millet that is finger millet was subsequently released by 2 independent groups. This commentary outlines the major applications of these reports, which also provided some opinion about this.
Keywords
Whole genome and transcriptome sequences of finger millet provide novel perspectives to develop drought tolerance and nutraceutical properties in millets.
1
The modern sequencing technologies and the transcriptome analysis offer more possibilities to analyze complex genomes.2,3 To date, only little attention has been paid on genome sequencing and crop improvement programs, and as a result, this millet has been categorized as an orphan crop for modern genetics studies.
4
The presence of duplicated genes derived from genome duplication, size, and complex nature slowed down the progress in genome sequencing of finger millet.
1
The genome size of wild species varied from 580 Mb (
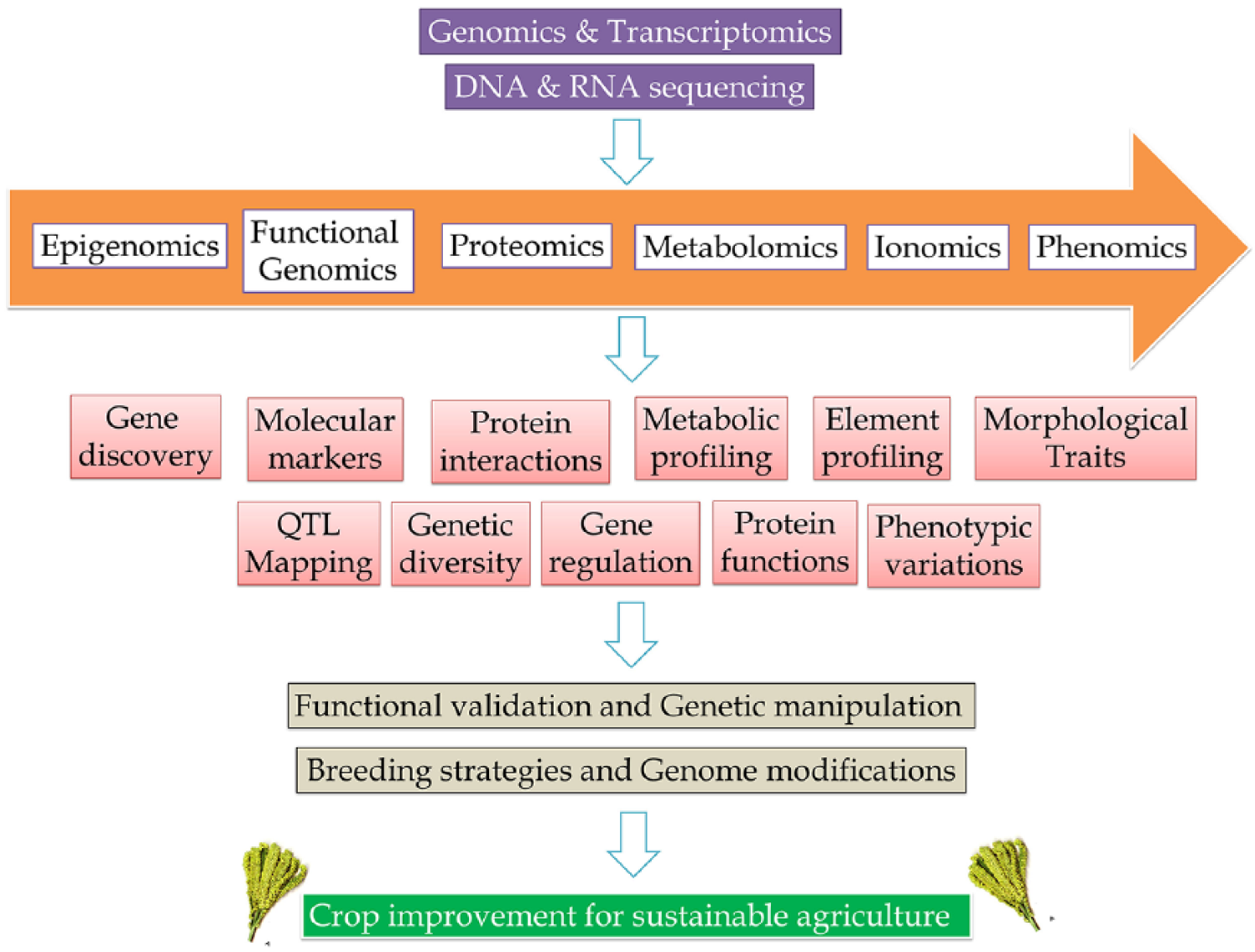
Importance of other omics technologies in crop improvement.
The comparative genomics and transgenic research will help to transfer key genes of drought tolerance and nutrient fortification into other millets and non-millet cereals. The number of genes identified from this study 1 is also higher compared with those identified by Hatakeyama et al. 5 Pathway prediction analysis using KEGG Automatic Annotation Server revealed that the finger millet genes are also predicted to be involved in carbohydrate and amino acid metabolism. This drought-hardy crop can grow well even in low moisture and hot environmental conditions, because it has an effective carbon assimilating mechanism through C4 pathway. 7 High-resolution pathway analysis coupled with computational biology on finger millet genome sequence will help to understand key traits like drought tolerance and grain nutrient filling, which can be transferred to other cereals.
Genome collinearity analysis revealed that there was a high synteny between genomes of finger millet and rice followed by foxtail millet, and the least synteny was witnessed between finger millet and maize.
1
Comparative analyses between genomes of grasses belonging to the subfamilies such as Ehrhartoideae, Panicoideae, and Pooideae have shown high conservation of collinearity despite some 60 million years of independent evolution of the major grass lineages.7,8 Genomes of finger millet, rice, and foxtail millet have found to show high synteny, which might be due to the fact that these three species shared some genes that are common among these crops.
9
Comparative maps of finger millet and rice demonstrated that other than the rearrangements that are necessary to account for the difference in the chromosome number between these 2 genomes, the finger millet genome has remained highly conserved as its divergence from a common ancestor with rice some 60 million years ago.
10
Changes in photosynthetic pathway are clustered in similar time period, suggesting that global climatic forces in addition to temperature may have facilitated transitions between C3 and C4. Assembling the sequence data (
Genes that code proteins of several important functions, namely, plant transcription factors, drought responsive genes, resistance genes for various diseases, calcium transport, and accumulation-related genes, were identified by homology-based analysis.
1
Drought tolerance is regulated by various genes, including transcription factors that allow plants to withstand harsh conditions.
14
Moreover, the identification of genes and its function is necessary for dissecting and engineering the regulatory network for enhanced productivity, selection of quantitative trait loci (QTL), and crop breeding. In the recent past, only a limited number of studies were performed on the characterization of finger millet genes involved in important agronomical traits. For example, heterologous expression of finger millet
Footnotes
Acknowledgements
The authors acknowledge Dr Stanislaus Antony Ceasar for his valuable suggestions to improve this commentary.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The author Mr. S. Pandian acknowledges the University Grants Commission, New Delhi, India, for providing Research Fellowship in the form of UGC BSR-SRF (UGC order no: F.25-1/2014-15(BSR)/7-326/2011/BSR).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
All authors listed have made substantial, direct, and intellectual contribution to the work and approved it for publication.
