Abstract
The use of antibiotics on a regular and excessive basis is a major factor in the spread of antibiotic-resistant bacteria. Patients discharge un-metabolized or relatively low doses of non-metabolized antibiotics through urine and stool, which might enter into the environment through sewage disposal and promote the emergence of antibiotic resistant bacteria. This study is designed to investigate how excessive use of antibiotics in the hospital sector and their release into hospital wastes contribute to the spread of antibiotic-resistant bacteria in different environmental settings. In this study, liquid hospital waste was collected from the sewage of Chittagong Medical College Hospital (CMCH), Bangladesh as well as from its distribution position in Chittagong city, Bangladesh. A total of 5 samples were collected from different positions in Chittagong city, including CMCH liquid waste. After collection, total bacteria and total cefixime resistant bacteria were counted by the total viable count (TVC) method. The result of bacteriological enumeration showed that a high magnitude of cefixime-resistant bacteria were available in all the hospital’s associated waste samples. The highest proportion of cefixime resistant bacteria (23.35%) was found in sample 2, whereas 17.4%, 7.6%, 5%, and 1.32% were found in samples 1, 3, 4, and 5, respectively. The total number of cefixime-resistant bacteria decreased with the increase in distance between the sample collection site and the hospital drain. This means that resistant bacteria developed in the hospital effluent are transferred to the environmental distribution sites.
Introduction
Antibiotic resistance is a public health concern that promotes illness, death rates, and healthcare-associated costs. 1 Moreover, in developing countries like Bangladesh, the impact of antimicrobial drug resistance on illness is more severe because of the high prevalence of bacterial infections.1-3 Antimicrobial drug resistance in developing countries is exacerbated by several factors, including irrational antimicrobial drug usage and poor sanitation.1,2,4,5
Antibiotics are usually prescribed to take care of bacterial infections in human and animal health. Once the antibiotics are prescribed, various human activities promote the contamination of the environment by antibiotics. Antibiotics often get into the environment through excretion by humans and disposal as waste by pharmaceutical plants.6,7 Most antibiotics are not metabolized after ingestion through reactions such as hydroxylation, cleavage, or glucuronation; instead, a significant amount of antibiotics are excreted in the environment via urine and feces, which mix with receiving waters. 8 The presence of antibiotics in different environmental niches like municipal sewage, sewage treatment plants, surface and river water, groundwater, drinking water, sludge, manure, agricultural soil, coastal waters and sediments, etc., has been previously reported by several investigators.9-13 Antibiotics available in the municipal wastewater has the potential to contaminate receiving surface and ground water. 14
One of the major risks associated with hospital-associated bacterial antibiotic resistance is the management of health care-associated infections, also referred to as nosocomial infections. Nosocomial infections generally occur in hospitalized patients during the period of receiving treatment. Bacteria are the most frequent causative agents for nosocomial infection. 15 Bacteria responsible for nosocomial infection usually develop antibiotic resistance by several mechanisms, including hydrolysis of antibiotics by producing extended-spectrum beta lactamase, which is able to hydrolyze a broad spectrum of beta lactam antibiotics. By secreting carbapenemases that can resist different kinds of antibiotics and also through the mutation of the target of antibiotics. 16
Chittagong Medical College Hospital (CMCH) is the biggest hospital in Chittagong Division, Bangladesh. Every day, numerous patients come to these hospitals for treatment of various diseases. In this hospital, different types of antibiotics are used for the treatment and prevention of infectious diseases. Most of the compounds used in medicine are only metabolized partially by patients and are discharged into the hospital sewage system through the feces and urine of patients. Exposure of un-metabolized or partially metabolized antibiotics in hospital waste leads to the selection of resistant bacteria. Thus, this hospital contributes a great deal to the development of antibiotic-resistant bacteria.
Antibiotic resistance is probably the most threatening difficulty for treatment now. The resistance phenomenon has contributed to scientists’ finding newer antibiotics. Most dangerously, the phenomenon has been increasing with the extensive use of antibiotics in hospitals. Anti-microbial resistance is a serious and growing problem in both developed and developing countries around the world. This study is therefore designed to evaluate the occurrence of antibiotic-resistant bacteria in hospital effluents and their receiving environments.
Materials and Methods
Antibiotic selection
To envisage the correlation between waste disposal and resistance development, the treatment histories of antibiotics used in the CMCH were noted down. Analysis of prescribed antibiotics among 100 patients from the Surgery and Gyniatrics ward of CMCH revealed that Cefixime was the most frequently prescribed antibiotic (44%) among those patients at that particular period of time (Figure 1), and therefore, cefixime was selected as the target antibiotic.
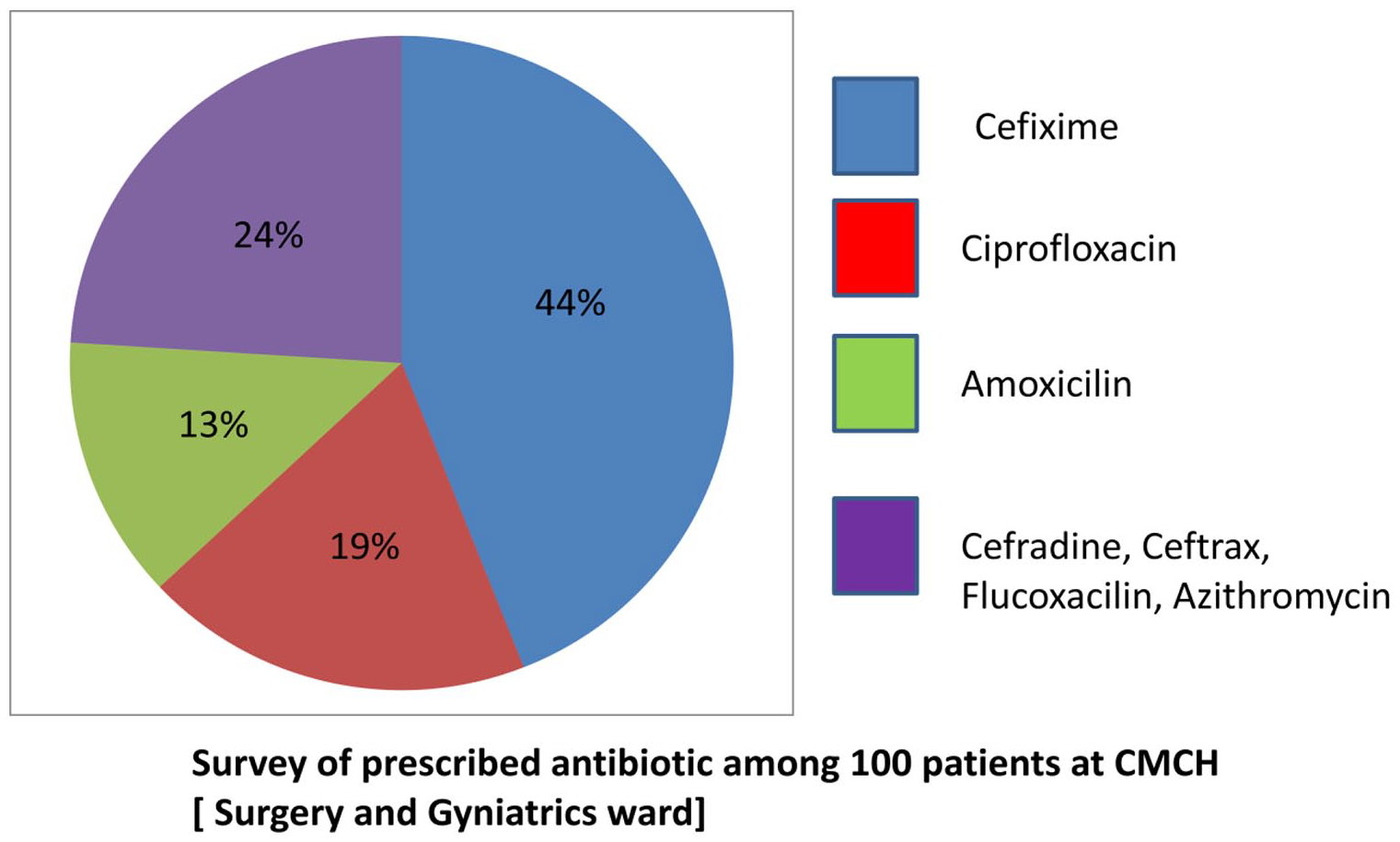
Survey of antibiotics prescribed to 100 patients at the CMCH Surgery and Gynecology ward: Prescriptions of patients admitted to surgery and gynecology wards were recorded and analyzed. Cefixime was prescribed to 44% of the total patient population.
Description of samples
Chittagong Medical College (CMC) Hospital has a direct connection with the municipal sewage system, which travels through the whole city and falls into a canal and finally into the Karnaphuli River (Figure 2). Thus, there is a very favorable condition for dispersing cefixime-resistant bacteria throughout the city and the areas around the city. The septic tank of the CMCH contains the excreta and medical waste of patients from the different wards like Gyniatrics, Surgery, Orthopedics, Medicine, Heart, and General Wards. These wastes are discharged from the septic tank through an underground pipe into a drain on the hospital’s back side. Wastes flow through this drain and into a second drain located outside the hospital. Thus, patients’ wastes pass to the different drains, canals, agricultural land, and ponds and are finally released into the Karnaphuli River (Figure 2). The objective of this study was to evaluate the effect of patients’ excreta on the spread of microbial resistance, so samples were collected from a hospital septic tank and its associated drains and also from an agricultural land near a major river located in Chittagong city, Bangladesh (Figure 2). A total of 5 samples were collected from 5 different locations within and outside the premises of the CMCH in Chittagong city (Figure 2).
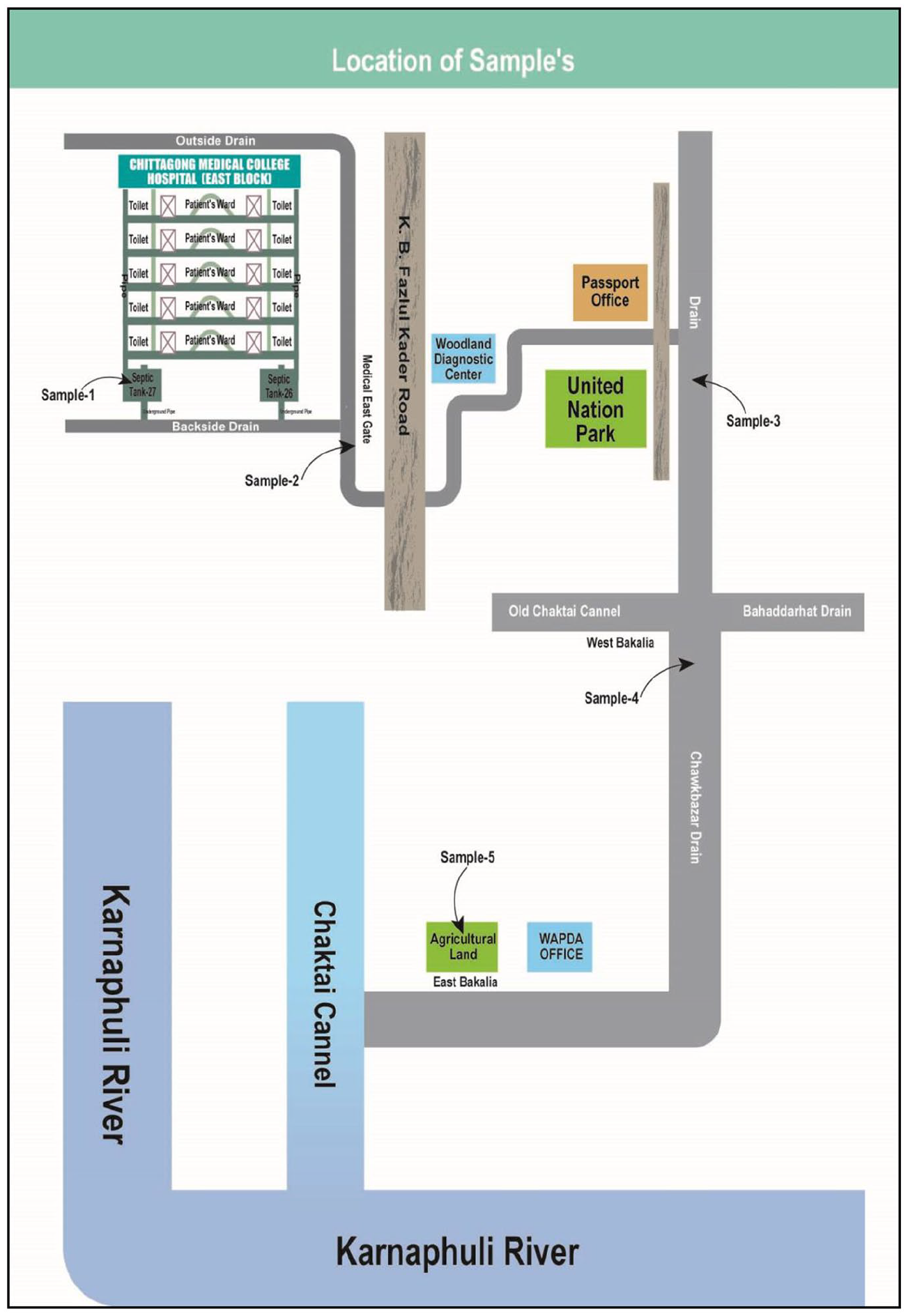
Location of the samples around Chittagong city, Bangladesh Sample 1: Sample 1 was collected from the 27-no. septic tank of CMCH in which surgery patients’ discharges (stools, urine, etc.) were deposited. The sample was a liquid portion of stool and other discharges. Sample 2: Sample 2 was collected from the position where hospital drains mixed with the outside drain (the drain by which hospital waste is distributed into different drains). The sample-2 was mainly drain water, consisting of all septic tank effluents of CMCH. Sample 2 was more diluted than sample 1. Sample 3: Sample 3 was collected from the large drain, which was situated in front of the Passport Office and the backside of the United Nations Park in Chittagong, Bangladesh. This drain was connected with the hospital’s outside drain through various small drains. Sample 3 was more diluted than sample 2. Sample 4: Sample 4 was collected from the large drain (Chawkbazar drain, Chittagong city) where hospital waste-containing drain was mixed with the Baharddarhat drain and the old Cakthai canal. It was situated in Fulthala, in western Bakolia. The sample-4 was diluted hospital waste, including most of the waste of the city. This drain ultimately falls into the Karnaphuli River through the Cakthai canal. Sample 5: Sample 5 was collected from an agricultural land that was situated beside the WAPDA Office in Eastern Bakolia to find resistance bacteria in agricultural soil. Since the land uses wastewater from the Chawkbazar drain situated beside it for irrigation purposes, there is a possibility of disseminating resistant bacteria in agricultural soil from the Chawkbazar drain that contains hospital effluents. Thus Waste water with soil from agricultural land was taken as sample-5.
Sample collection, transportation, and storage
All the samples were collected by maintaining an aseptic procedure in sterilized screw cap test tubes. After collection, the samples were kept in ice bags and transported to the Molecular Biology Lab of the Genetic Engineering and Biotechnology Department at Chittagong University. The samples were stored at 4°C in the laboratory refrigerator and analyzed within 24 hours.
Serial dilution of the sample
In 9 ml of sterile water, 1 ml of the sample was taken. The mixture was shaken well. This is dilution number 10−1. From a 10−1 dilution, 1 ml was added to 9 ml of sterile water and mixed well. This is dilution number 10−2. 1 ml of the 10−2 tube was transferred to 9 ml of sterile water and thoroughly mixed. This is dilution number 10−3. By repeating the same procedure, the dilution was made up to 10−8. The procedures were done in laminar air flow.
Preparation of antibiotic solution
Raw powder (active ingredients of pharmaceuticals-API) of Cefixime Antibiotic was taken in sterile distilled water in such an amount that the final concentration would be 50 µg/ml (such as 0.005 g in 100 ml). The solution was mixed thoroughly. After mixing, the tubes were then stored at a 4°C temperature in the refrigerator.
Total viable count from samples 1, 2, 3, 4, and 5 without antibiotic
Total viable count was performed by following pour plate method. 17 The media was cooled to 45°C and kept liquid until it was poured into the plates. About 1 ml of the samples from 10−6, 10−7, and 10−8 dilutions was poured on to sterile petri-plates with sterile pipettes. Then, media at about 45°C was poured on to the petri-plates containing samples so that the thickness of the media would be about 3 to 3.50 mm. Then the plates were stirred by hand gently clockwise and anti-clockwise for about 20 times so that the samples were mixed thoroughly with the media. Then the plates were allowed to stand steady for the solidification of the media. The plates were placed on such a surface that an equal thickness was maintained throughout the media. After the solidification of the media, the plates were placed in an incubator in an inverted position at a 37°C temperature for 24 hours. The procedures were done in laminar air flow. After 24 hours of growth, plates with 30 to 300 bacterial colonies were counted. The total number of bacteria will be the multiplication of the colony number and the dilution factor. Plates for all the 3 dilutions were counted like this. Then an average of the numbers of the 3 dilutions was done, and this was the total number of bacteria in the sample. The total bacteria of all the 5 samples was counted by this procedure.
Total viable count from samples 1, 2, 3, 4, and 5 with antibiotic (cefixime)
This experiment was done by following the above procedure. 17 In this case, different concentrations of cefixime were added to the agar media before being poured into the plates. The cefixime concentrations were 50 to 500 µg/ml.
Results
Bacterial counts of 5 samples with and without antibiotics (at a concentration of 50 µg/ml cefixime)
Bacterial counts of samples 1, 2, 3, 4, and 5 with and without cefixime are listed in Table 1. In this investigation, the total number of bacteria and the total number of cefixime-resistant bacteria were enumerated for all samples (Table 1 and Figure 3). TVC without antibiotics showed almost similar results for samples 1 to 4, with the highest number of bacteria being found in sample 1, and the lowest number of bacteria being found in sample 5. The occurrence of cefixime-resistant bacteria was comparatively higher in sample 2 than in the other 4 samples. In sample 2, 23.35% of bacteria were resistant to cefixime. This was followed by sample-1, which comprised about 17% (17.40% accurately) cefixime resistant bacteria. In sample 3, resistance to cefixime constituted 7.60%, slightly higher than in sample 4, which made up 5% resistance. Even so, 1.32% of bacteria showed resistance against cefixime in sample 5, which was very alarming as the sample source was an agricultural field. This high abundance of cefixime-resistant bacteria in hospital sewage might be the result of frequent cefixime consumption by the patients of Chittagong Medical College Hospital (CMCH).
Percentage of cefixime resistant bacteria in samples 1, 2, 3, 4, and 5.
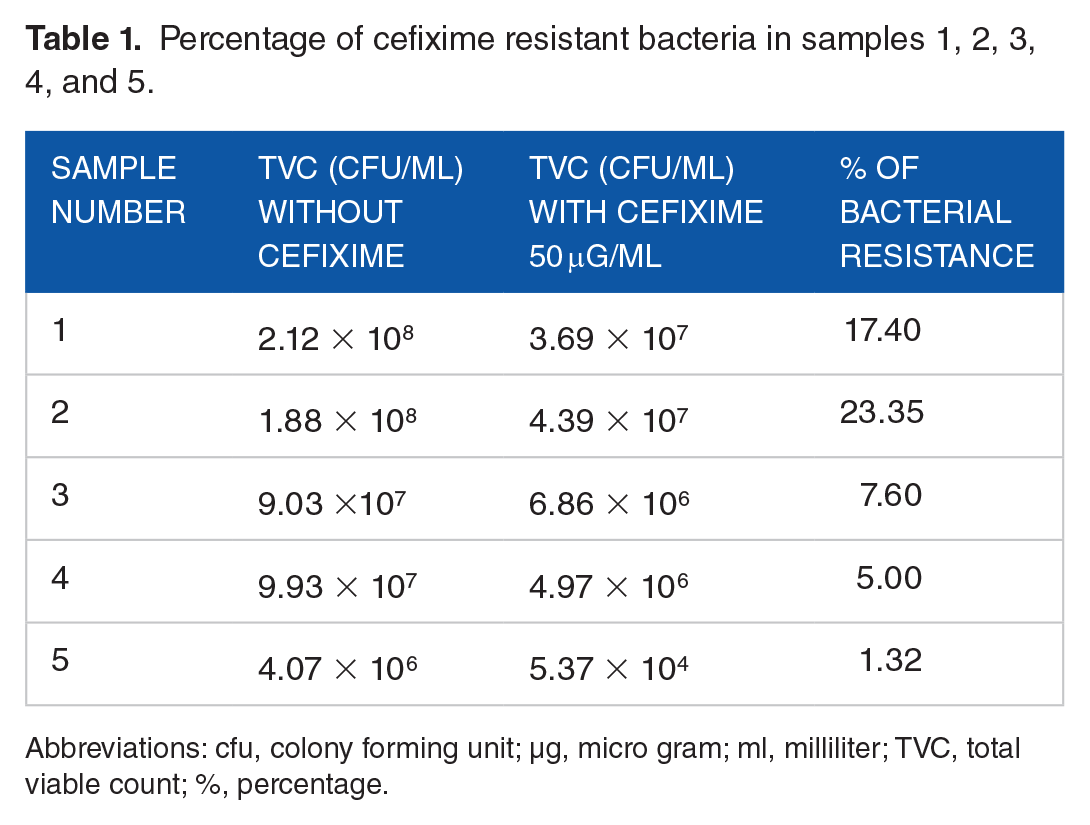
Abbreviations: cfu, colony forming unit; µg, micro gram; ml, milliliter; TVC, total viable count; %, percentage.
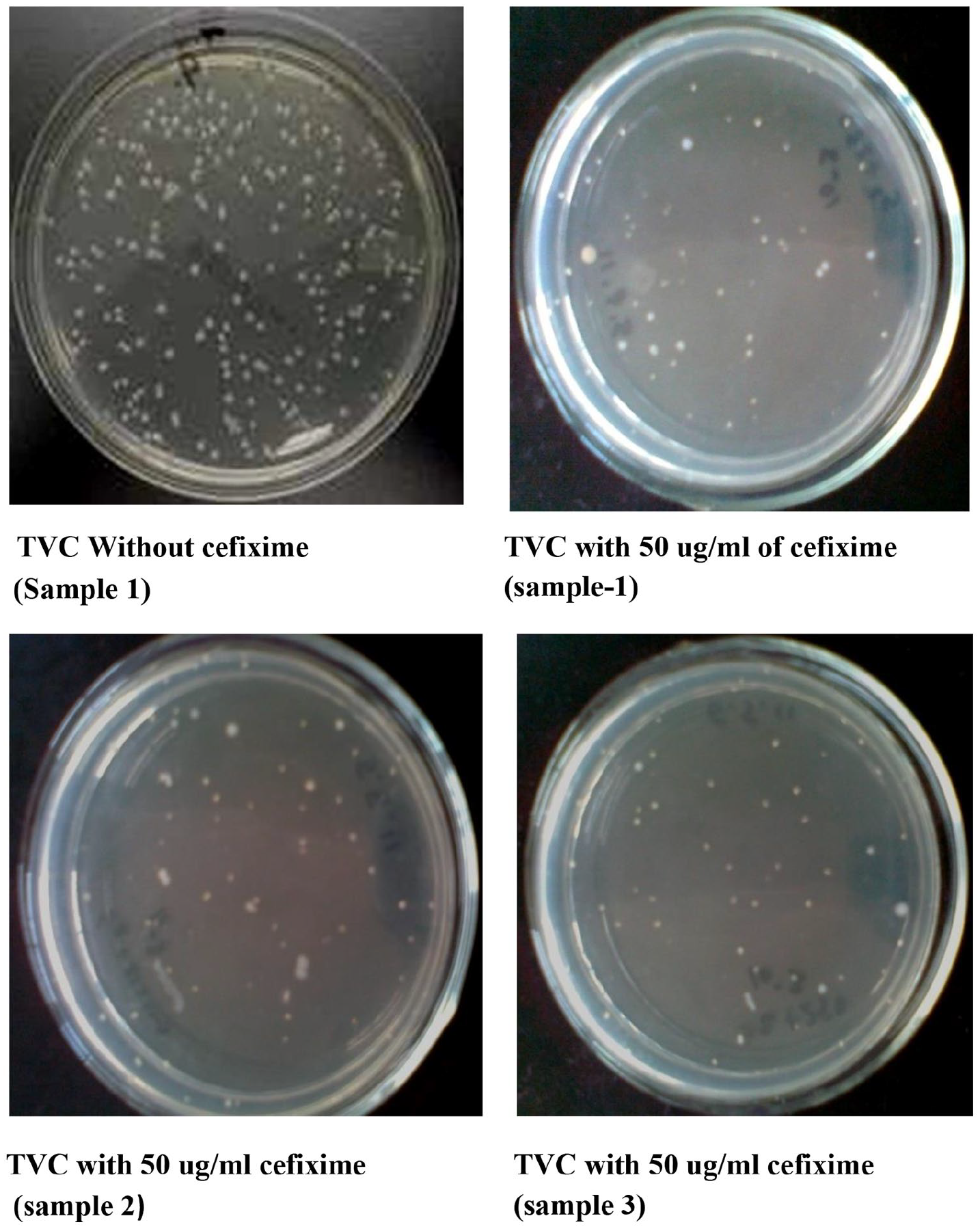
Total viable count (TVC) before and after antibiotic treatment (50 µg/ml cefixime concentration). Representative photographs of some selected plates.
Bacterial counts of 5 samples at different concentrations of cefixime (concentration of 100 µg/ml to 500 µg/ml of cefixime)
Primary evaluation by 50 µg/ml of cefixime revealed that significant numbers of resistant bacteria are available in all the 4 samples directly associated with CMCH. So a question has arisen: what is the tolerance level of cefixime? To answer the question, we evaluated the TVC of bacteria among those samples at different concentrations of antibiotics, which ranged from 100 µg to 500 µg/ml of cefixime. We found, bacteria among samples 1, 2, 3, and 4 can resist up to 500 µg of cefixime/ml of media, and this result actually represents the devastating misuse of antibiotics in CMCH and its associated environment (Table 2). We have also repeated this experiment with sample 5, which was collected from agricultural land near a major river in Chittagong, Bangladesh, and subsequently found high tolerances of cefixime by bacteria on this agricultural land (Table 2).
Bacterial counts of 5 samples at various cefixime concentrations (100-500 g/ml).
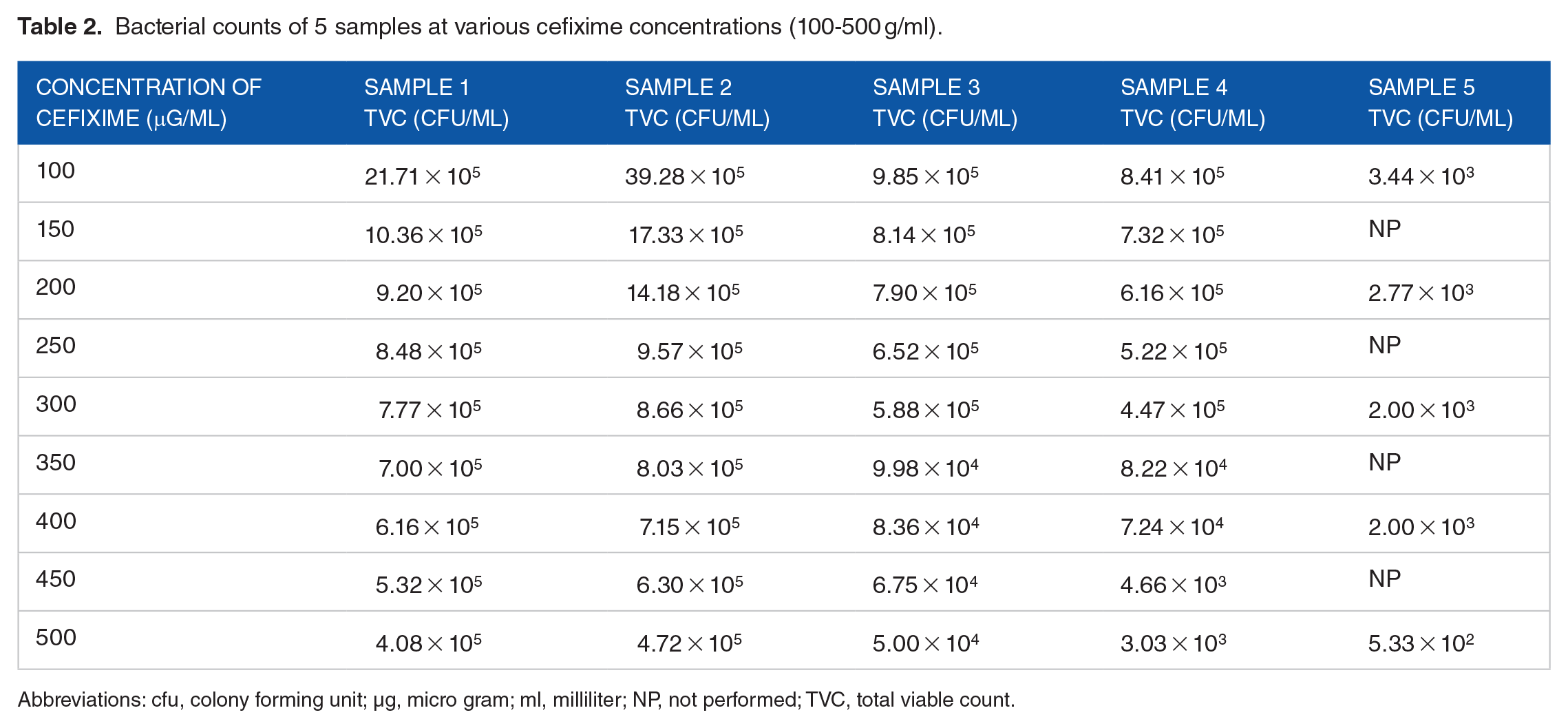
Abbreviations: cfu, colony forming unit; µg, micro gram; ml, milliliter; NP, not performed; TVC, total viable count.
Discussion
Antimicrobial resistance has been noted as one of the major microbial threats of the 21st century. 18 Number of articles on the evolving multidrug resistance in microbes have been published in many international journals, reflecting that a disastrous effect may occur in the near future. Several reports have shown that antimicrobial-resistant strains of bacteria are present in most hospital effluent discharges that lead to the spread of antibiotic-resistant bacteria into the environment.19,20 Therefore, the aim of this study was to evaluate the prevalence of antibiotic-resistant microflora in hospital effluents and their receiving environments.
Analysis of treatment histories collected from the CMCH showed that they mainly used Cefixime (44%) for treatment purposes. The level of cefixime resistance was remarkably high among the bacterial counts of each sample (sample 1, 2, 3, 4, and 5) at different concentrations of cefixime. It has been clearly shown that bacteria available in hospitals and their environmental wastes can resist cefixime, even up to 500 g/ml of growth media (Table 2). This high resistance to cefixime by bacteria from samples 1, 2, and 3 might be the consequence of excessive and irrational use of cefixime antibiotics by physicians in order to treat various diseases of patients who visited this hospital for treatment. This observation is also supported by several previous reports.21,22 In addition, the bacterial count in the wastes of agricultural land, which is situated near a major river in Chittagong, Bangladesh, was also high (Table 2). It may be due to the fact that the agricultural land might be flooded by waste water from the large drain by excessive rainfall or irrigation practice by local people using drain water (Figure 2). A high quantity of bacterial resistance in the agricultural field might promote the transfer of antibiotic resistance marker genes to the plants grown on that land, which could cause a potential health hazard. In the future, genomic screening of plants should be done to detect the presence of antibiotic resistance marker genes. According to our observations, the reason for the high resistance to antibiotics may be a combination of several reasons: The first is the unnecessary use of antibiotics in any type of illness, including viral infection; and the second is the widespread purchase of antibiotics without a physician’s prescription.
Conclusion
The fear of antibiotic resistance is that the resistance is transferable. So, there is a possibility of transferring the phenotype to other bacteria. The majority of antibiotics used are only partially metabolized after administration and are released via patient excreta into the municipal sewage system. Previous reports suggest that resistant bacteria from the environment may be transmitted to humans, 23 in whom they cause disease that cannot be treated by conventional antibiotics. Questions have evolved about how to solve the problem of antibiotic resistance. General improvements in public health measures and improved sewage systems in developing nations can contribute to preventing infection. This study shows the high prevalence of resistant bacteria spread in Chittagong city by CMCH. Further studies on molecular screening and genomic analysis may be conducted to identify specific genes related to antimicrobial resistance.
Footnotes
Acknowledgements
We are thankful to the Department of Genetic Engineering and Biotechnology, University of Chittagong, Bangladesh. We are also thankful to the staffs of the Dean office of Faculty of Biological Sciences, University of Chittagong, Bangladesh.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Author Contributions
A M M A C designed this study. K N U performed the experiments; A M M A C analyzed the data; A M M A C, K N U wrote the original draft. A M M A C reviewed the several versions of the manuscript. Both authors read and approved the final manuscript.
Data Availability
All the data are included in this manuscript.
