Abstract
Background:
The emergence and spread of Extended-spectrum β-lactamases (ESBLs) in Enterobacteriaceae through the plasmid-mediated exchange have become a major threat to public health by complicating the treatment of severe infections in both animals and humans. Therefore, the current study focused on evaluating the manifestation of ESBLs production from the fecal isolates of E. coli, Shigella spp, Salmonella spp, and Klebsiella spps in commercial poultry production systems of Kiambu County, Kenya.
Materials and methods:
Out of 591 isolates identified as E. coli, Shigella spp, Salmonella spp, and Klebsiella spps from 437 fecal samples, only 78 were phenotypically suggestive to be ESBL producers. The possible ESBL producers were screened for the presence of blaTEM, blaCTX-M, blaOXA, and blaSHV using the PCR technique. These isolates were also screened for carriage of the QnrS gene that confers resistance to the fluoroquinolone class of drugs.
Results:
The most detected ESBL gene from the isolates was blaOXA (n = 20; 26%), followed by blaTEM (n = 16, 21%), with the majority of them detected in E. coli. The blaCTX-M was identified in all the 4 enteric’s bacteria-type isolates tested. Three E. coli and Salmonella spp respectively were found to harbor all the 5 antimicrobial resistance (AMR) gene types. The blaTEM, blaOXA, blaSHV, and QnrS genes were not detected from Klebsiella and Shigella spps. Additionally, most of the AMR gene co-carriage was detected in both E. coli and Salmonella spps as follows blaTEM + blaOXA (n = 4); blaTEM + QnrS (n = 3); blaTEM + blaOXA + QnrS (n = 3), concurrently.
Conclusion:
Our findings highlight the significance of commercial poultry production in disseminating transferable antibiotic resistance genes that act as potential sources of extensive drug resistance in livestock, humans, and the environment, leaving limited therapeutic options in infection management.
Introduction
The emergence and spread of Extended-spectrum β-lactamases (ESBLs)-producing Enterobacteriaceae from livestock and humans has become a significant global public health concern. 1 The widespread use of antibiotics in human, veterinary, and agricultural treatment has ominously led to the selection and global dissemination of resistant genes in the Enterobacteriaceae family over the past years. 2 It has been of great concern due to the irrational use of expanded-spectrum antibiotics in animal feeds for prophylaxis and treatment. The sub-therapeutic use of these drugs in livestock may lead to the transmission of potentially resistant strains of bacteria in the environment, which poses a serious hazard to human health. 3 These practices in EU member states’ countries have been regarded as illegal since they resulted in the transmission of ESBL-producers in modern times. This transmission has led to poor treatment outcomes of serious nosocomial infections culminating in extended hospital and at times mortality. The spread and extensive use of β-lactams in poultry and livestock production has continuously rendered this class of antibiotics less effective against both livestock and human infections. 4 These ESBLs producing superbugs have been isolated from poultry production systems and humans and domestic farm animals, which increases their ability to share these genes through genetic elements such as plasmids, transposons, and integrons. This condition may also lead to the transformation of non-pathogenic bacteria into resistant reservoirs in the natural bacterial ecosystem. According to Abrar et al, most Enterobacteriaceae acquire ESBL genes by mutation or horizontal transfer of plasmids, which results in oxyimino-cephalosporin resistance, with the most common ESBL-encoding genes being blaCTX-M, blaTEM, blaSHV, and blaOXA. 5 ESBLs are classified according to their primary sequences and substrate profiles into different families such as the TEM, the SHV, the OXA, and the CTX-M-family. 6 When this gene occurs in enteric bacteria this increases the propensity of these organisms for extended resistance to various beta-lactam drugs. It is upon this basis that the current study was carried out to molecular characterize the presence of blaTEM, bla CTX-M, bla OXA, blaSHV, and blaQnr from fecal isolates of E. coli, Shigella spp, Salmonella spp, and Klebsiella spps in commercial poultry production systems of Kiambu County, Kenya.
Materials and Methods
Study location
This cross-sectional laboratory-based study was carried out in purposively selected 6 sub-counties of Kiambu County—Kenya namely; Ruiru, Juja, Gatundu North, Gatundu South, Thika, and Kikuyu being the major commercial poultry production centers. A total of 437 samples that consisted of farmers’ fecal samples (n = 72) and cloacal swabs (Broiler [n = 80], Layer [n = 160], and Improved Kienyenji [n = 145]), in commercial poultry production systems were collected in Kiambu County, Kenya, between November 2020 to February 2021. From the 437 fecal samples analyzed, a total of 592 non-duplicate isolates (Cloacal isolates=544; human isolates= 47) of E. coli (n = 289), Klebsiella spp (n = 83), Salmonella spp (n = 108), and Shigella spp (n = 111) were isolated.
Selection of bacterial isolates for ESBL screening
The identification of the recovered enteric bacterial isolates was performed through traditional bacteriological methods and biochemical tests as guided in the Clinical and Laboratory Standards Institute/NCCLS 7 guidelines with an API 32 E system (bioMerieux SA, Marcy l’Etoile, France) according to Wei and Charles. 8 The isolates were stored at −80°C in MicroBank cryovials containing 20% glycerol (Pro-Lab Diagnostics, Round Rock, TX, USA). Control strains used in this study included K. Pneumoniae ATCC 700603, and E. coli ATCC 25922. The carriage of ESBL and QnrS gene was screened on 78 ESBL-positive isolates which included 42 strains of E. coli, 7 strains of Klebsiella spp, 24 strains of Salmonella spp, and 5 strains of Shigella spp respectively. These bacteria genera were chosen on their phenotypic resistance profiles toward β-lactams and fluoroquinolone antimicrobial tested as described in previous related research according to Gundran et al. 9
DNA extraction through boiling methods
A single pure colony of each revived target bacterial isolate was suspended in 0.5 ml of extraction buffer (100 µl of 1 ml buffer Tris Borate and 2 µl of 0.5 EDTA). Thereafter, 400 µl buffer suspension known as reaction mixer in Eppendorf tube was boiled for 10 minutes at 100°C. Post-boiling process centrifugation was done at 14 000 rpm for 5 minutes at 4°C. This was followed with DNA-supernatant stored at −20°C for later use as a DNA template for PCR amplification according to Solberg et al. 10
Molecular characterization of ESBLs resistant genes
Out of 591 bacterial isolates from the Poultry and human fecal samples obtained in Juja, Ruiru, Kikuyu, Thika, Gatundu North, and Gatundu South sub-counties of Kiambu County, only 78 isolates showed the presence of ESBL genes and hence were screened for the presence of antimicrobial resistance genes using PCR technique. The Genes targeted were blaTEM, blaCTX-M, blaOXA, blaSHV, and QnrS. The DNA extract of each sample has used a template for the detection of blaTEM, blaCTX-M, blaOXA, blaSHV, and QnrS as described in the methods of Brody and Kern. 11 In each sample, a total volume of 26 µl of the reaction mixture per gene was mixed in the Eppendorf tube as follows; 12 µl Qiagen master mix, 12 µl DNA’s free PCR water, 1 µl forward primer, 1 µl reverse primer, and finally 2 µl DNA. The PCR amplification conditions were as follows; Initial denaturation at 95°C for 5 minutes, denaturation at 94°C for 1 minute, Annealing at 60°C for 1 minute, initial extension at 72°C for 30 seconds, and final extension at 72°C for 5 minutes for 35 cycles using a GeneAmp® PCR system 9700 thermocyclers. However, the amplification condition varied slightly depending on the primer type and manufacturers’ recommended annealing temperature. The PCR primer as indicated in Table 1 were used as per manufacturers’ guidelines. Separation of PCR amplicons was done using 1.5 % agarose gel (Agarose Hi-Res standard) stained with Sybr green (Sigma-Aldrich) in 1X TBE buffer at 100 volts for 1 hour. Gel viewing was done use a UV Gelmax® imager and extended productive size compared against a 1 kb plus DNA ladder (Invitrogen).
Primers sequences and annealing temperatures for the ESBL gene used in the study.
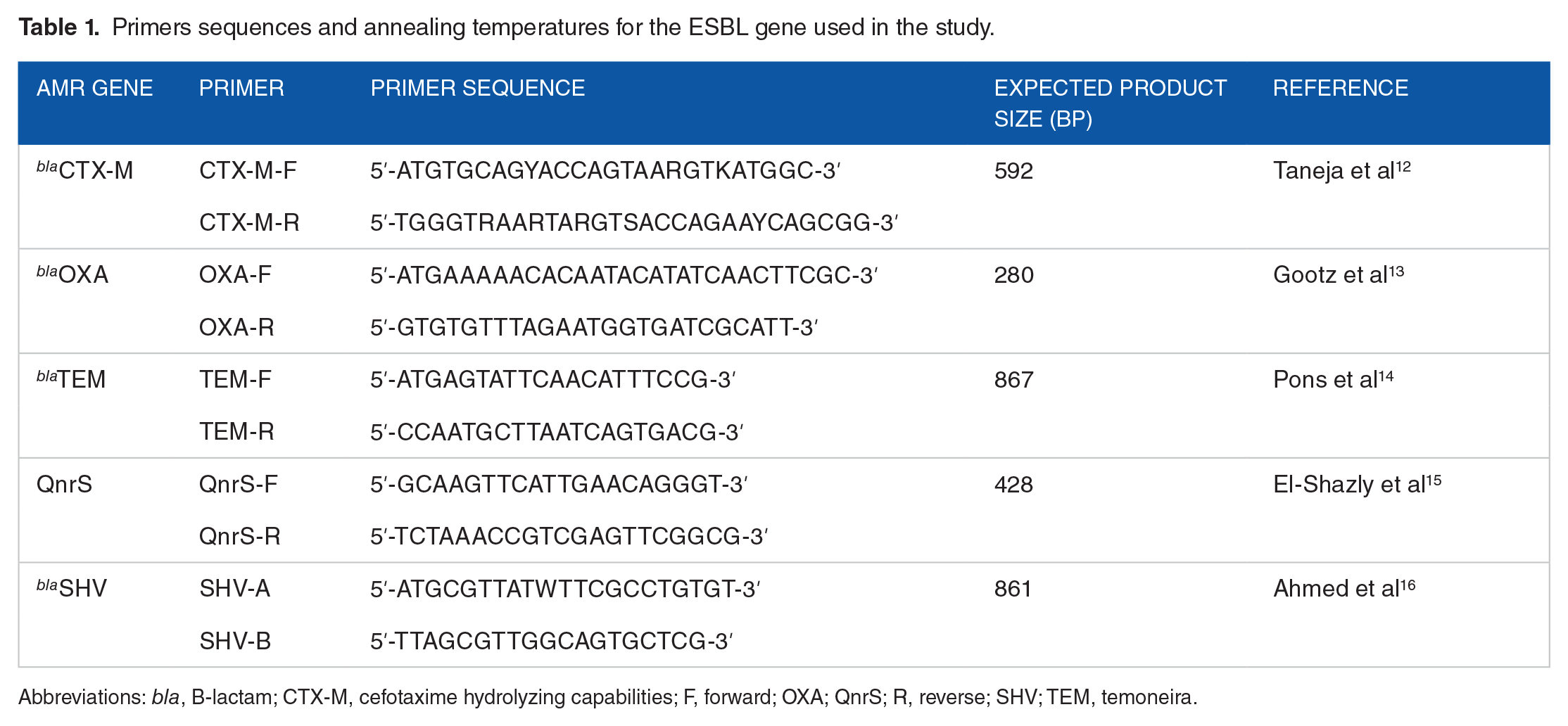
Abbreviations: bla, B-lactam; CTX-M, cefotaxime hydrolyzing capabilities; F, forward; OXA; QnrS; R, reverse; SHV; TEM, temoneira.
Ethical approval
The Ethical approval was obtained from the Department of Public Health, Pharmacology and Toxicology Research review board (UoN), faculty of veterinary medicine board of postgraduate studies (UoN), NACOSTI (NACOSTI/P/21/8761), and from County Government of Kiambu Livestock, Fisheries, and Veterinary services (KCG/ALF/ RESEARCH/VOL.1/49).
Results
The analysis found a majority of the ESBLs producing isolates from improved Kienyeji (32%; n = 25) poultry samples while humans/farmers fecal samples isolates had the least ESBL genes carriage at . . .% (n = 11). The ESBL genes detections were common in isolates from the Juja sub-county (n = 22), followed by Kikuyu (n = 17), Gatundu south (n = 16) with the least from Ruiru (n = 5), and Gatundu North (n = 3), respectively. We found that majority of the ESBLs genes were identified from both broiler (n = 24) and improved Kienyeji (n = 23) poultry systems with blaOXA (n = 20), and blaTEM (n = 16) genes being most present. Out of the 78 isolates with potential ESBLs production, only 66 were positive for the tested genes as shown in Table 2 below.
Distribution of the isolates and ESBLs genes in the commercial poultry production systems of Kiambu County.
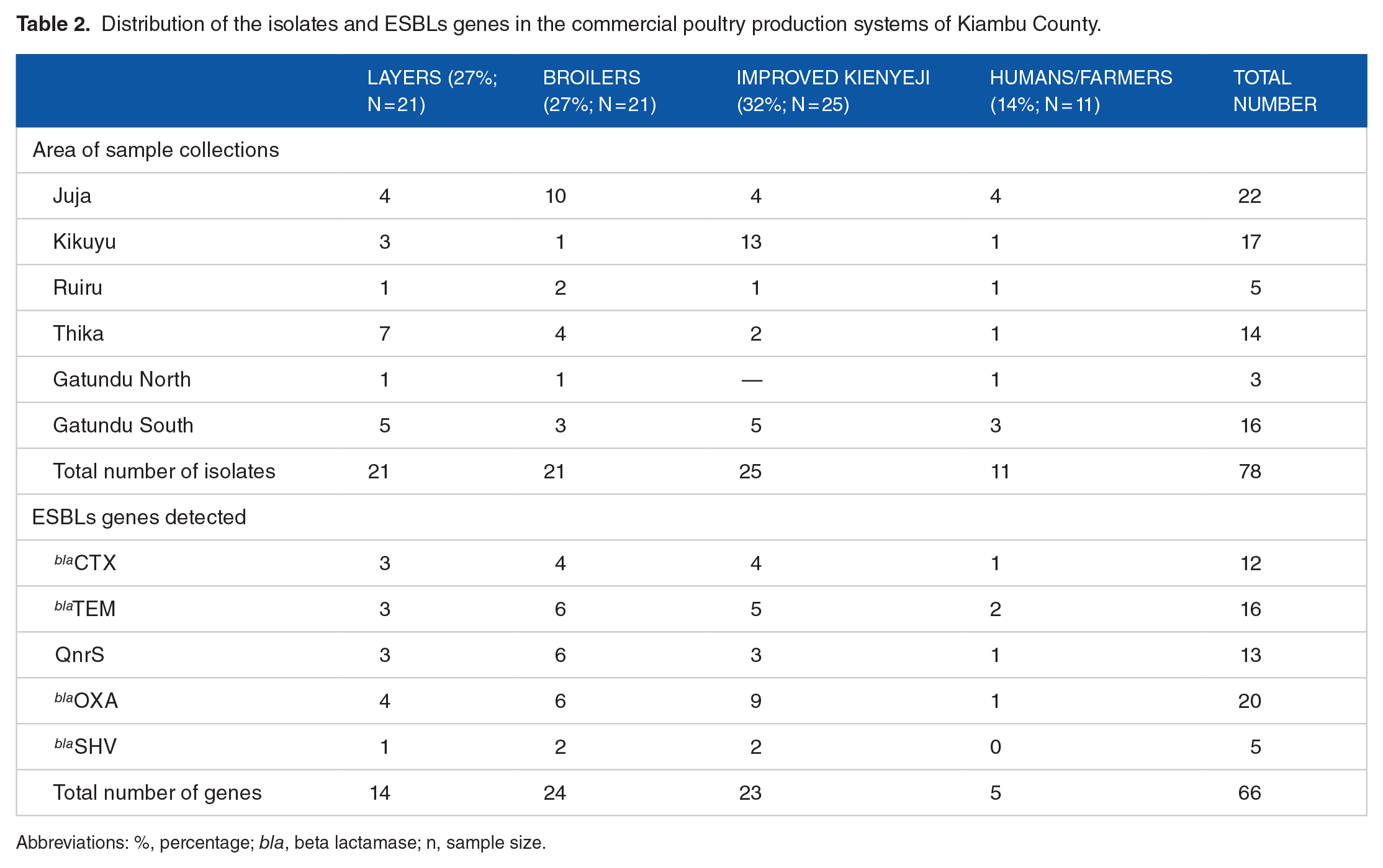
Abbreviations: %, percentage; bla, beta lactamase; n, sample size.
Table 3 presents results of the Genomic distribution of the ESBL genes among 4 targets Enterobacteriaceae Spps isolated from commercial poultry production systems. A total of 78 bacterial isolates that indicated ESBLs production were subjected to PCR test for the presence of blaTEM; blaCTX-M; blaOXA; blaSHV; and QnrS genes. The blaCTX-M was identified in all the 4 enteric’s bacteria isolates tested. Twenty-one percent of isolates showed the presence of blaTEM gene with the majority detected in E. coli. In both E. coli and Salmonella Spps all 5 genes were detected from these isolates. The blaTEM and QnrS genes were not detected from Klebsiella and Shigella spps. Additionally, the blaOXA and blaSHV were not detected in Klebsiella spps and Shigella spps respectively.
Percentage genotypic distribution of the ESBL genes among 4 targets Enterobacteriaceae Spp. isolated from commercial poultry production systems.
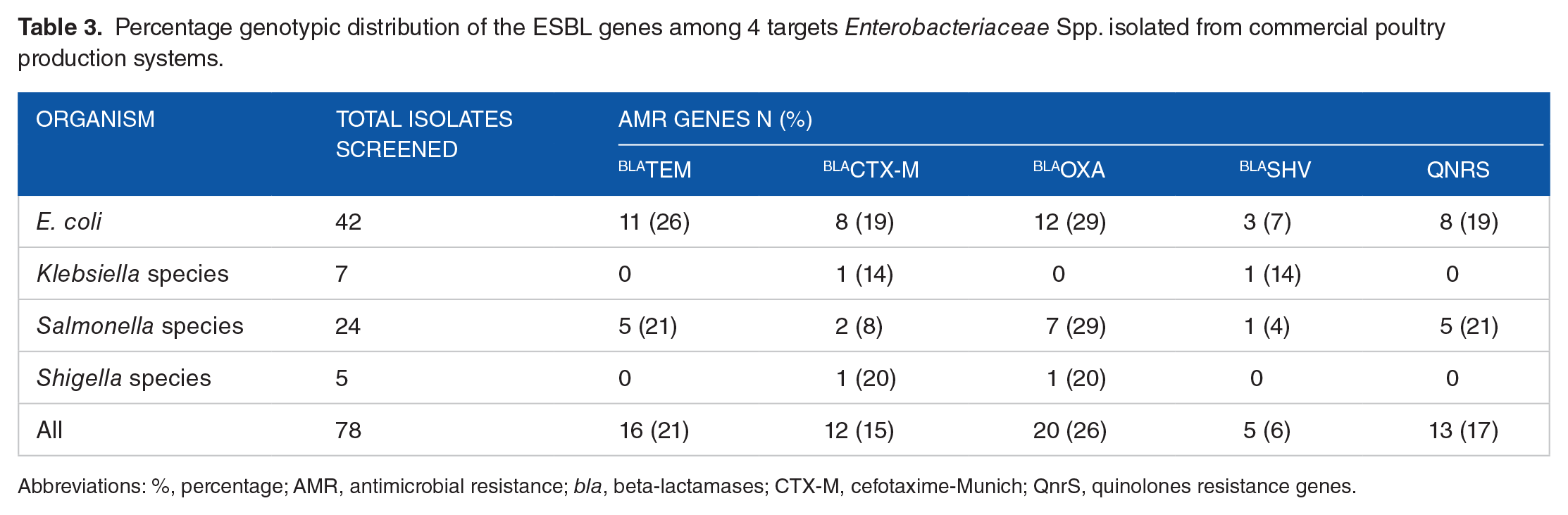
Abbreviations: %, percentage; AMR, antimicrobial resistance; bla, beta-lactamases; CTX-M, cefotaxime-Munich; QnrS, quinolones resistance genes.
Figure 1 presents the percentage distribution of ESBLs and QnrS genes among the 78 isolates. Twenty six percent of the isolates had blaOXA, 17% (QnrS), 16% (blaTEM), 12% (blaCTX-M), and 5% (blaSHV) respectively as shown in Figure 1.
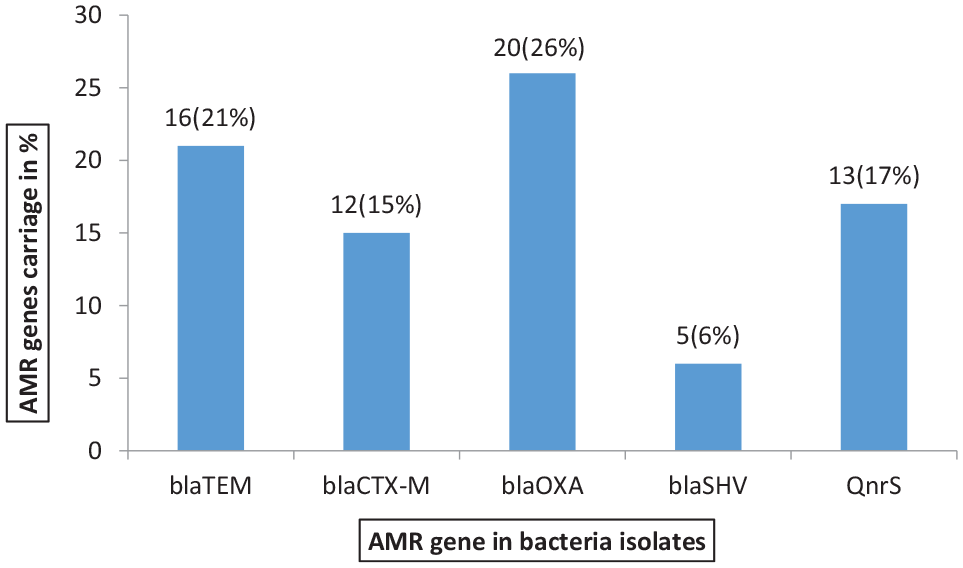
Distribution frequency of the AMR genes carriage among the isolates.
Table 4 presents the results of the co-carriage of the AMR genes among the 4 selected enteric’s in this study. Most of the AMR gene co-carriage was witnessed in both E. coli and Salmonella spps as follows blaTEM + blaOXA (n = 4); blaTEM + QnrS; blaTEM + blaOXA + QnrS . Four percent (4%) showed the presence of blaTEM + OXA co-carriage among E. coli and Salmonella spps. blaTEM + CTX-M + OXA and blaOXA + QnrS AMR gene co-carriage were observed in Salmonella spps. Also, blaTEM + CTX-M, CTX-M + SHV, and TEM + SHV + QnrS AMR gene carriage were only observed in E. coli isolates. Both Shigella and Klebsiella spps showed no AMR gene co-carriage as indicated in Table 4 below.
AMR gene co-carriage among the isolates.
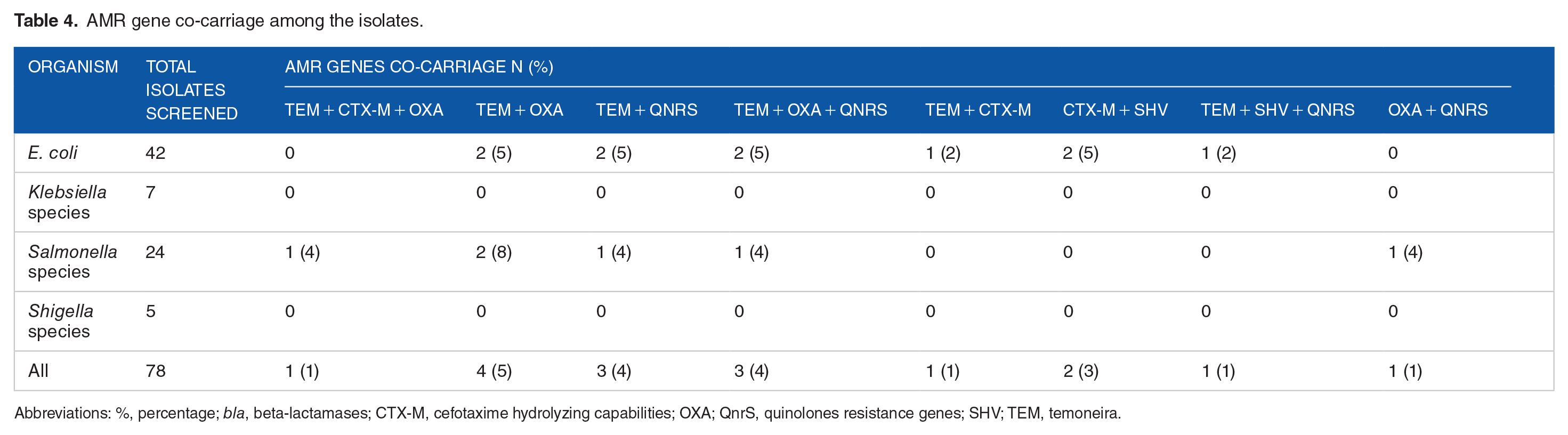
Abbreviations: %, percentage; bla, beta-lactamases; CTX-M, cefotaxime hydrolyzing capabilities; OXA; QnrS, quinolones resistance genes; SHV; TEM, temoneira.
Figure 2 presents the results of the electrophoretic reactions for the positive isolates with blaTEM genes among the isolates. With 11 E. coli and 5 Salmonella spps isolates testing positive for the blaTEM gene.
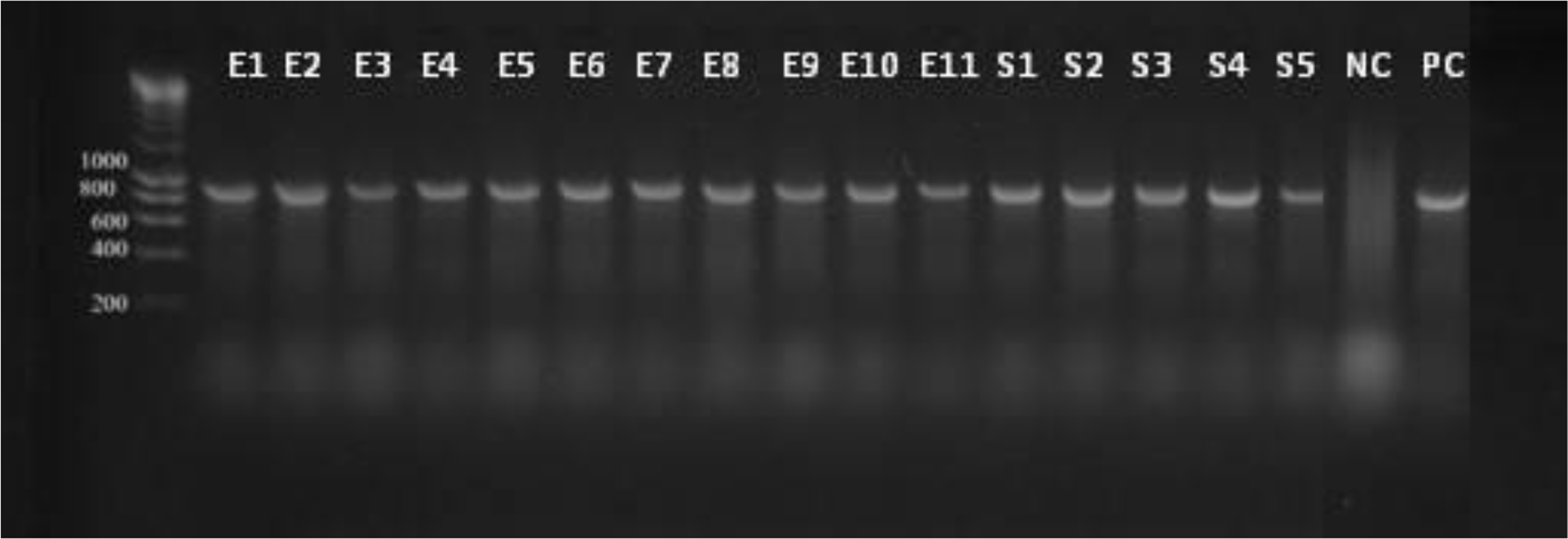
LM-DNA ladder for electrophoretic reaction with positive isolates for Peer Review blaTEM genes.
Figure 3 presents the results of the electrophoretic reactions for the positive isolates with blaCTX-M genes among the isolates. With 8 E. coli, 2 Salmonella spps, 1 Shigella, and 1 Klebsiella isolates testing positive for the blaCTX-M gene.
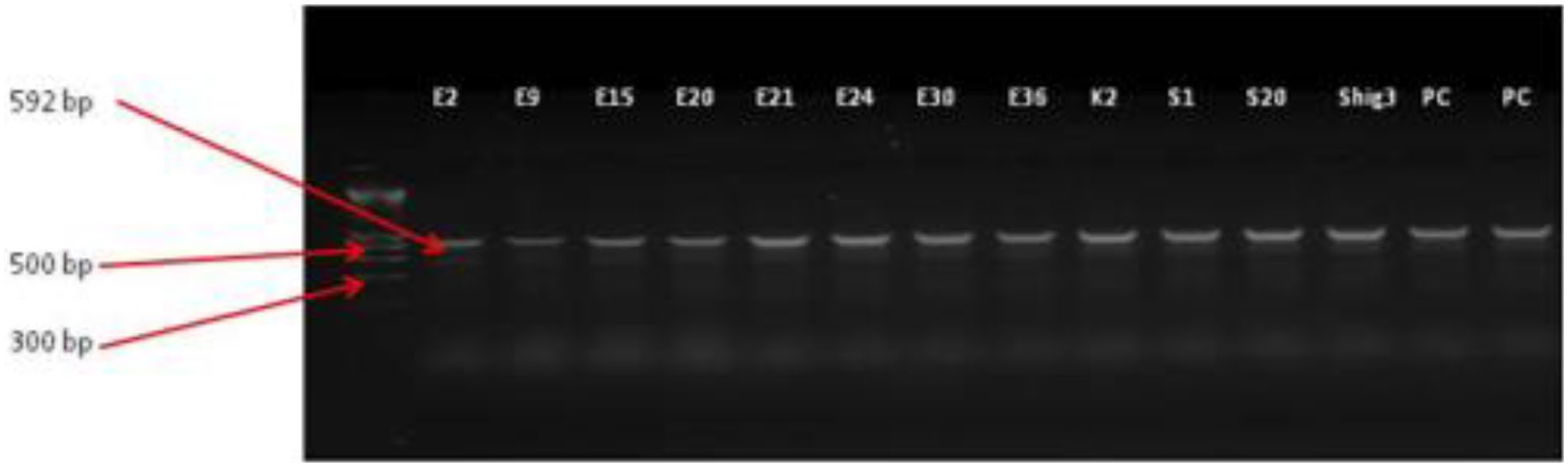
LM-DNA ladder (100 bp for gene size determination), for electrophoretic reaction with positive isolates for blaCTX-M genes among the isolates.
Figure 4 presents the results of the electrophoretic reactions for the positive isolates with QnrS genes. We found that 8 E. coli and 5 Salmonella spps isolates testing positive for the QnrS gene.
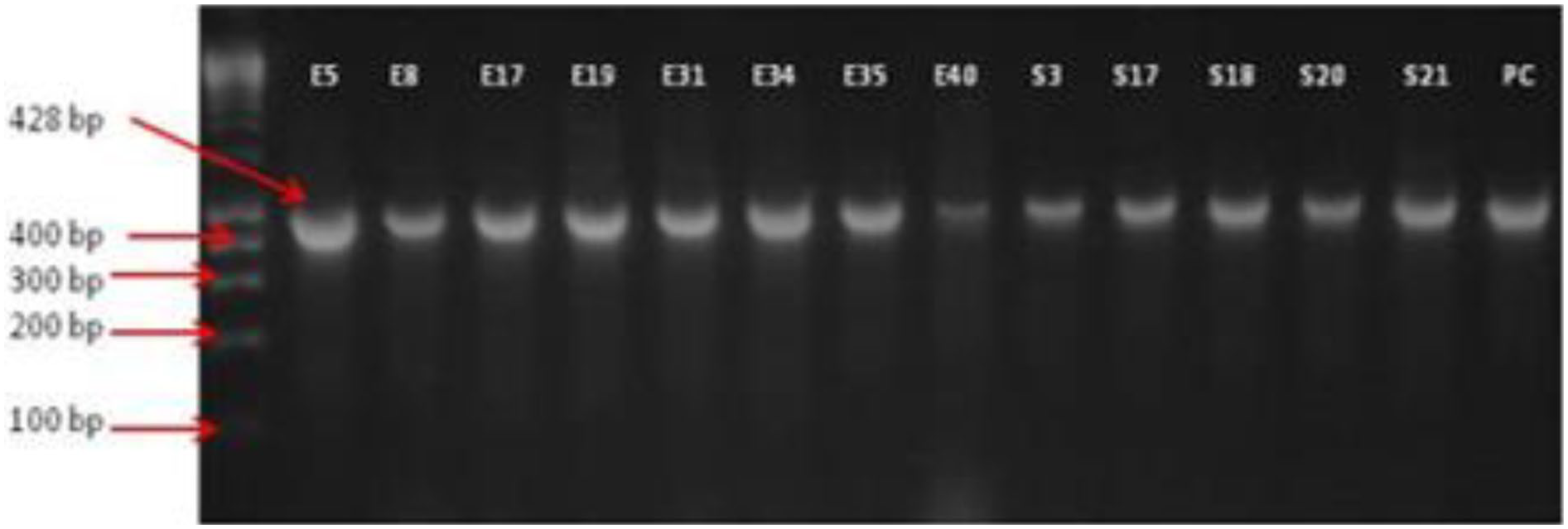
LM-DNA ladder, for electrophoretic reaction with positive isolates for QnrS genes among the isolates.
Figure 5 presents the results of the electrophoretic reactions for the positive isolates with blaOXA genes. We found that 12 E. coli, 7 Salmonella spps, and 1 Shigella spps isolates testing positive for the blaOXA gene.
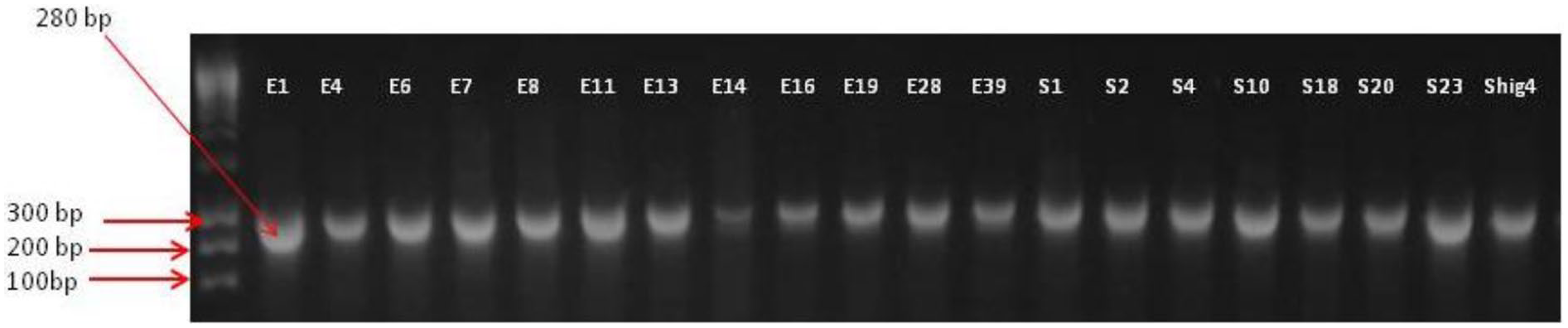
LM-DNA ladder, for electrophoretic reaction with positive isolates for blaOXA genes among the isolates.
Figure 6 presents the results of the electrophoretic reactions for the positive isolates with blaSHV genes. We found that 3 E. coli, 1 Salmonella spp, and 1 Klebsiella spp isolates tested positive for the blaSHV gene.
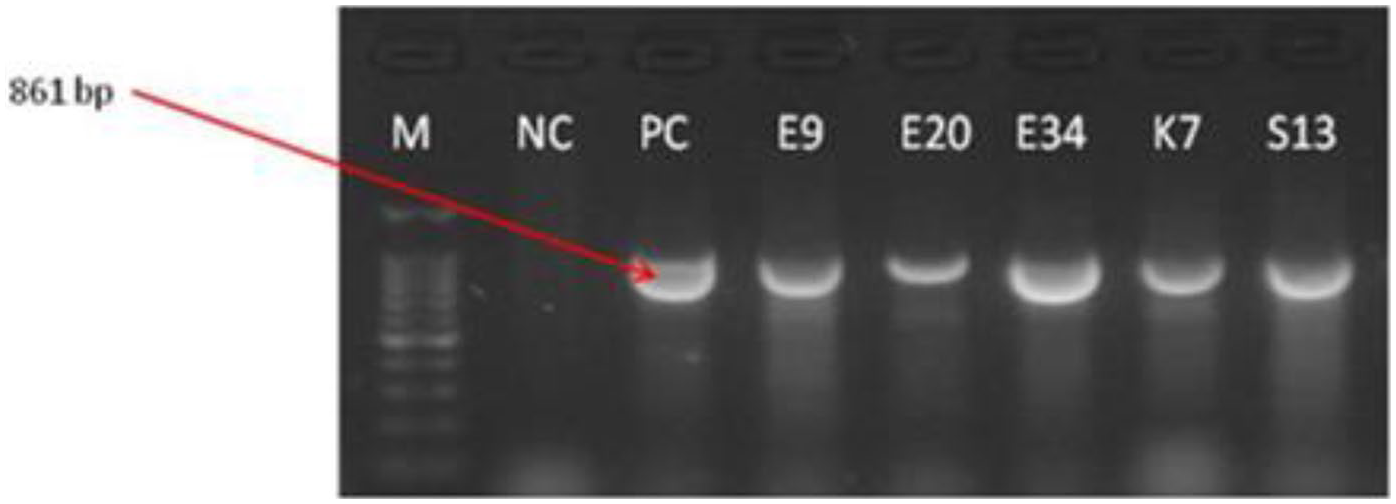
LM-DNA ladder, for electrophoretic reaction with positive isolates for blaSHV genes among the isolates.
Discussion
Research-based knowledge and understanding of antimicrobial resistance trends among bacterial isolates from livestock and humans are paramount for recommendations in the experimental antibiotic management of infections. The current study described the molecular detection of ESBLs producers among E. coli, Shigella spp, Salmonella spp, and K. pneumonia spp isolates from commercial poultry production systems of Kiambu County, Kenya. Data from our recent related work recorded high antimicrobial resistances among the present bacterial isolates especially toward Sulfamethoxazole (79%), Trimethoprim (71%), Tetracyclines (59%), Ampicillin (49%), and Amoxicillin/Clavunallic acid (39%). Our results were consistent with findings of previous studies that have also recorded high AMR profiles toward Sulfamethoxazole, and Trimethoprim among E. coli isolates from poultry samples as per studies carried out by Egypt, Sudan, and Kenya.17-21 These findings could be a reflection of AMR emergence due to the wide empirical use of Human drugs in the management of poultry diseases prophylaxis and as growth promoters.
Over the past decades, ESBL producing Gram-negative bacilli, especially E. coli, Salmonella, Shigella, and K. pneumonia have emerged as serious pathogens, both community, and hospital-acquired infections, globally. The occurrence of these ESBLs among livestock and human isolates greatly differs worldwide and geographically and is promptly varying over time. 22 In our current study, ESBL phenotypes were found to be positive in 78 isolates (13.2%) out of 593 isolates. We have demonstrated that the majority of the ESBLs producing isolates were gotten from an improved Kienyeji (n = 25) poultry production system with the least number of isolates gotten from humans/farmers fecal samples (n = 11). We further found that most of the isolates came from the Juja sub-county (n = 22). We further found that majority of the ESBLs genes were identified from both broiler (n = 24) and improved Kienyeji (n = 23) poultry systems with blaOXA (n = 20), and blaTEM (n = 16) genes being most present. According to Ambler et al 23 , one of the significant antibiotic resistance mechanisms in Enterobacteriaceae is founded on plasmid-mediated production of extended-spectrum β-lactamases (ESBLs) which inactivate β-lactam-antibiotics including Cephalosporins and monobactams by hydrolyzing their β-lactam ring.
In our study, out of 78 isolates which showed potential ESBLs production, only 66 were positive for the tested genes. We also found a high prevalence of ESBL production by E. coli (54%) and K. pneumonia (31%) isolates, in commercial poultry production systems of Kiambu County. Similarly, this has also been found in Kenya,24,25 Sudan, 26 Jordan, 27 India, 28 and china 29 where theyo found 31.4 % E. coli infestation. Moreover, in countries like Nigeria 20 and China 21 high prevalence of ESBLs producers has been reported. This high prevalence of ESBLs producers in our study could be associated with the extensive use of third-generation cephalosporins in poultry production systems in Kenya.
In this study, we also found that blaOXA (26%) and blaTEM (21%) we the most identified, this agrees with a study carried in brazil and disagrees with the previous studies by Naseer and Sundsfjord 30 who found blaCTX to be the most predominant. Further to this, it is until 2000, SHV and TEM remained the dominant variants of ESBL; however, CTX-M enzymes have taken their place over the last decades. Maybe, this could be because the ESBL genes are located on a plasmid that can be transferred from 1 organism to another, rather easily, and can incorporate genetic material coding for other resistance genes. We also noted that both E. coli and Salmonella isolates were resistant to fluoroquinolones as confirmed by the presence of the QnrS gene. This could be associated with increased irrational use of fluoroquinolones in commercial poultry production, as Zahraei and Farashi 31 witnessed in their study.
Furthermore, we found a high co-carriage of AMR resistance genes among E. coli and Salmonella isolates with blaTEM + OXA, blaTEM + QnrS, and blaTEM + OXA + QnrS being the most occurring. This can be explained by the fact that the genes are located on plasmids and can be transferred from 1 organism to another and hence coding resistance to other antimicrobial classes. Additional, enterobacterial found in animals and the human intestinal tract have been important reservoirs for resistance genes leading to the dissemination of ESBL-producers in the community especially if ESBL genes are coded by resistance pathogenic bacteria leading to hospital and community infections. In this study, our genotypic survey on 78 confirmed ESBL phenotype strains by PCR revealed that 85% (n = 66) of the isolates were positive for at least one of the studied genes. According to Xi et al 32 antibiotic-resistant bacteria and resistance genes are considered environmental pollutants and responsible for a tenacious public health crisis throughout the world. The health problems associated with antibiotic-resistant microorganisms are more about restricted therapeutic remedies in most developing countries that lack access to good quality treatment, thus, emphasizing infection as an important root of morbidity and mortality.
In conclusion, the results of this study advocate for the importance of ESBL-producing E. coli, Salmonella, Shigella, and K. pneumonia, as common causes of infectious diseases among the commercial poultry production systems of Kiambu County, Kenya. Furthermore, the high majority of multidrug-resistant organisms should be taken into account considered when choosing therapeutic agents. At the same time, the continuous local monitoring of resistance patterns is necessary to select an empirical antimicrobial therapy adequately. Further studies aimed at unraveling the molecular mechanisms of resistance will better understand the epidemiology associated with ESBL-producing species of Enterobacteriaceae.
Footnotes
Acknowledgements
The authors of this manuscript express their gratitude to everyone who contributed immensely to this research. Special thanks are due to John Maina and Sharon Chepkurui for their tireless contributions and commitment during Lab work research assistance. The authors also register special thanks to Prof. Samuel Kariuki, Acting Executive Director (KEMRI), for his guidance and material support in the conduct of this study.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research study was partially funded by the National research fund-Kenya (NRF/Ph.D./02/166).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
JGN, JKG, GOA, and JMM conceptualized and designed the study. JGN conducted the study, collected data, and performed data analysis. JGN, JKG, GOA, JMM, JM, and JM-2, interpreted the results and participated in the writing of the draft script, and affirmed that the manuscript was an accurate and transparent account of the study being reported. JGN accepts full responsibility for the work and the conduct of the study and organized the decision to publish. The corresponding author confirms that all listed authors meet authorship criteria and that no others have been omitted.
Ethical Approval
The Ethical approval was obtained from the Department of Public Health, Pharmacology and Toxicology Research review board (UoN), faculty of veterinary medicine board of postgraduate studies (UoN), NACOSTI (NACOSTI/P/21/8761), and from County Government of Kiambu Livestock, Fisheries, and Veterinary Services (KCG/ALF/ RESEARCH/VOL.1/49).
