Abstract
Widespread antimalarial drug resistance has prompted the need for new therapeutics and greater understanding of malaria parasite biology. To this end, the isoprenoid biosynthesis inhibitor fosmidomycin has been used to probe the metabolic regulation in the malaria parasite, Plasmodium falciparum. Genetic changes in the haloacid dehalogenase (HAD) superfamily member HAD2 conferred resistance to fosmidomycin, at the cost of decreased fitness. In the absence of fosmidomycin, parasites gained mutations to phosphofructokinase that restored growth and fosmidomycin sensitivity, thus revealing an intriguing example of plasticity in a core glycolytic process. Moreover, this study marks a second report of a HAD superfamily protein-modulating metabolic homeostasis in P falciparum parasites. Haloacid dehalogenase enzymes are distributed across all domains of life and have increasingly been found to influence central carbon metabolism and drug sensitivity in P falciparum. Investigating the mechanisms by which HAD superfamily members modulate metabolism may shed light on how metabolic networks are connected in apicomplexan parasites and other organisms and may guide future therapeutic endeavors.
Keywords
Guggisberg AM, Frasse PM, Jezewski AJ, et al. Suppression of drug resistance reveals a genetic mechanism of metabolic plasticity in malaria parasites. mBio 2018; 9(6): e01193-e01118. doi:10.1128/mBio.01193-18. PubMed PMID: 30425143; PubMed Central PMCID: PMC6234871.
Caused by infection with Plasmodium spp. parasites, malaria continues to be an enormous economic and global health burden. 1 Malaria control worldwide has been hampered by widespread parasite resistance to all commonly used antimalarial therapies, including delayed clearance by front-line artemisinin-based agents. Plasmodium spp. and other apicomplexan parasites possess an unusual plastid organelle, the apicoplast. As a result, malaria parasites deploy an essential plastidial pathway for isoprenoid biosynthesis, the methylerythritol phosphate (MEP) pathway, which is vital for parasite growth but not shared by the human host. The phosphonate antibiotic fosmidomycin (FSM) is a competitive inhibitor of the first dedicated step of the MEP pathway, catalyzed by deoxyxylulose reductoisomerase. Although poor drug-like characteristics of FSM have limited its development as a clinical antimalarial, its highly specific mechanism-of-action has been invaluable as a chemical tool to probe MEP pathway regulation and biology.
Guggisberg et al 2 used FSM resistance as a strategy to probe metabolic regulatory networks in asexual Plasmodium falciparum parasites. Wild-type parasites were subjected to FSM selective pressure, and whole-genome sequencing was performed on the drug-resistant parasites that emerged. The researchers focused on a particular nucleotide polymorphism in PF3D7_1226300, which encodes a haloacid dehalogenase (HAD) protein family member, termed HAD2. Fosmidomycin-resistant had2 parasites were found to have a dramatic dysregulation of central carbon metabolism and downstream MEP pathway metabolites, leading to poor asexual growth and FSM resistance. In the absence of FSM pressure, had2 parasites acquired additional suppressive mutations in the glycolytic enzyme phosphofructokinase (PFK9). Reduced PFK9 function restored growth and FSM sensitivity to parasites lacking HAD2. This work reveals an important and unexpected mechanism of plasticity in a core glycolytic process in P falciparum.
In addition, this study marks a second example linking a HAD protein family member to cellular metabolic homeostasis in malaria parasites. Loss of the HAD protein family member HAD1, a close homolog of HAD2, also results in FSM resistance in P falciparum, accompanied by remarkable changes in MEP pathway metabolites. 3 Together, these results suggest a common, conserved, and vital role for HAD superfamily members in metabolic regulation. Bolstering support for this model of HAD-mediated metabolic control, genetic disruption of a third HAD protein family member, the putative phosphoglycolate phosphatase (PGP; PF3D7_0715000) of P falciparum, also appears to lead to metabolic dysregulation. In this case, loss of HAD function results in increased sensitivity to FSM, rather than resistance. 4 Additional compelling studies suggest that this newly identified role of HAD proteins in regulating metabolism is not unique to malaria parasites, as Wang et al find that multiple phosphatases, including several HAD proteins, modulate production of isoprenoids in Escherichia coli. 5
HAD proteins are characterized by a Rossmannoid fold and 4 highly conserved sequence motifs in their catalytic core domain, in addition to the presence of a hinged cap domain. This cap domain provides substrate specificity through close interactions with substrates. The characteristic organization of this substrate-defining cap domain within the gene sequence is the basis for dividing HAD superfamily enzymes into 3 distinct subfamilies (subfamily I, II, and III, respectively; Figure 1). 6 Although the namesake of this large superfamily of proteins (>600 000 members of InterPro IPR023214) is a bacterial HAD, most HAD enzymes instead catalyze phosphate-transferring reactions. Most (nearly 70%) of the HAD proteins are phosphatases, with many more performing phosphomutase, phosphonatase, or phosphotransferase activities. 7 Biochemical analyses of HAD proteins in vitro have demonstrated that they dephosphorylate a wide range of phosphosubstrates, including nucleotide phosphates, sugar phosphates, phosphorylated cofactors, and occasionally phosphoproteins.8,9 Moreover, these studies reveal that many HAD proteins have wide and promiscuous substrate profiles, such that a number of HAD proteins are annotated as “non-specific” cellular phosphatases. By contrast, other HAD proteins have much more narrow substrate specificity profiles. Importantly, much of the substrate profiling of HAD enzymes relies on in vitro phosphatase assays, and few attempts have been made to validate these substrates in a cellular context, where substrate availability may determine which substrates are most commonly used. It is difficult, therefore, to distinguish between the broad in vitro substrate profiles of HAD proteins and their potentially narrower roles in a biological context.
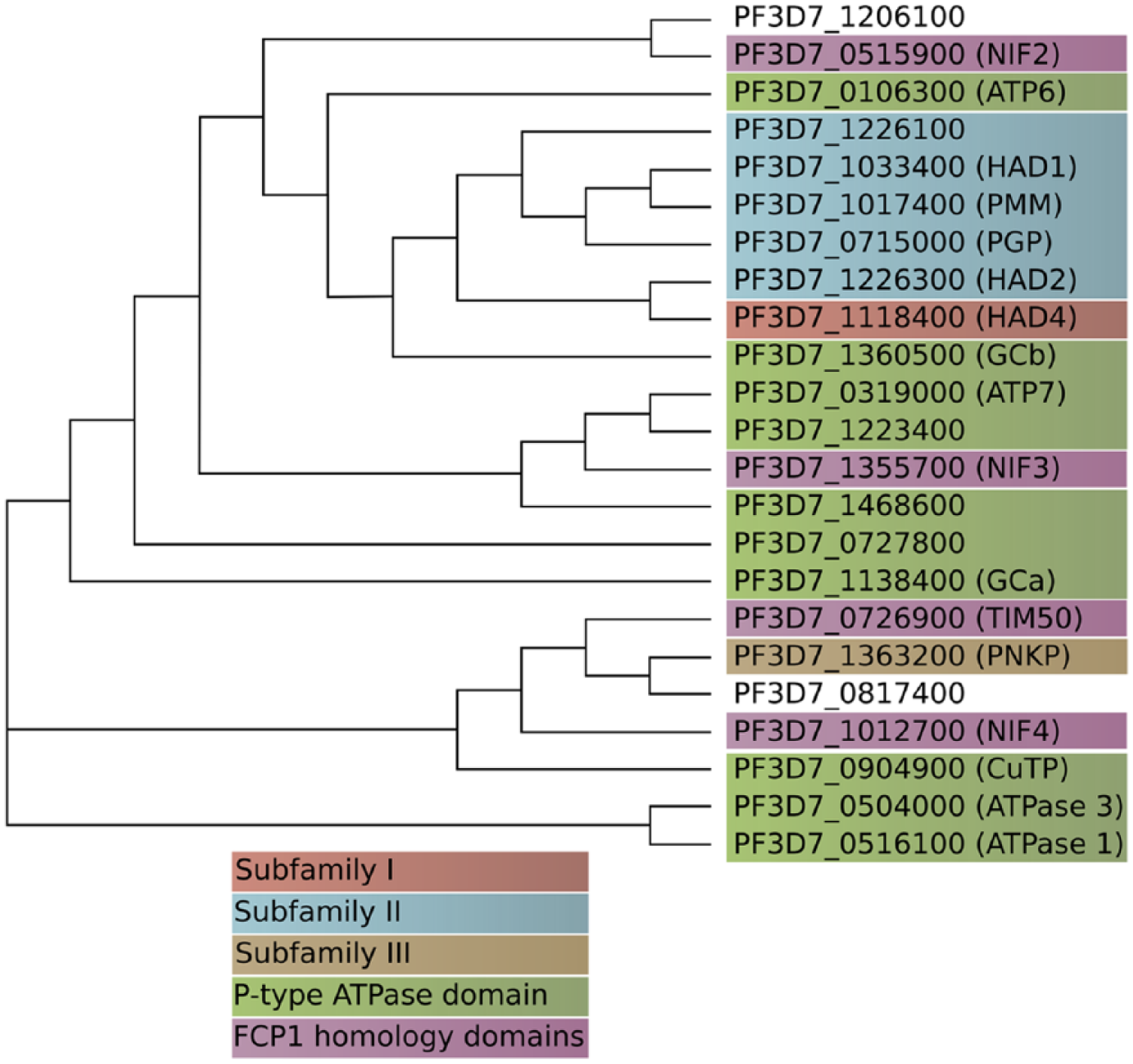
Phylogenetic tree of HAD superfamily (IPR023214) proteins from P falciparum. Established protein names are given in parentheses, where applicable. HAD1, HAD2, and PGP all fall within subfamily II of HAD proteins, and cluster near each other phylogenetically. Proteins are color-coordinated by the presence of InterPro domains as follows: HAD subfamily I (IPR006439), HAD subfamily II (IPR006379 and IPR006357), HAD subfamily III (IPR006549), P-type ATPase (IPR006539 and IPR008250), and FCP1 homology domain (IPR004274). Phylogenetic analysis was performed by ClustalW2 from EMBL-EBI, using the neighbor-joining method with distance correction. HAD indicates haloacid dehalogenase; PGP, phosphoglycolate phosphatase.
In the P falciparum genome, there are roughly 23 genes containing HAD superfamily domains (IPR023214). Based on sequence homology, many of these have been annotated as P-type ATPases. Several others contain FCP1-like domains, homologous to the essential protein serine phosphatase in yeast, FCP1. P falciparum possesses a single HAD subfamily I member, as well as a single subfamily III member. However, the HAD subfamily II is expanded in the P falciparum genome (Figure 1). Notably, the 3 HAD superfamily protein members that have been implicated in metabolic regulation and FSM sensitivity of P falciparum—HAD1, HAD2, and PGP—all fall within subfamily II of HAD superfamily proteins and cluster together phylogenetically (Figure 1). The other genes clustered within this group include 3 additional subfamily II members [annotated as a phosphomannomutase (PF3D7_1017400) and 2 hydrolases of unknown function (PF3D7_1226100 and PF3D7_1118400)], as well as guanylyl cyclase β (PF3D7_1360500). An important question for future investigation is whether the biological functions of HAD1, HAD2, and PGP in metabolic regulation and drug-sensitivity are more broadly applicable to other HAD proteins, or are limited to this subset of closely related HAD1-like homologs.
Although increasing evidence indicates that HAD protein family members, and specifically HAD subfamily II members, modulate metabolic homeostasis in P falciparum, the cellular mechanism by which HAD enzymes control metabolism remains yet undefined. We propose 2 hypothetical models of how HAD protein family members may regulate cellular metabolite levels (Figure 2). In the first model, HAD proteins regulate metabolism directly by enzymatically converting substrates that would otherwise directly feed into biosynthetic pathways (Figure 2A). Thus, loss of HAD activity causes a build-up of phosphosubstrate levels, leading to increased availability of these compounds for downstream metabolic reactions. To date, however, most of the genetic changes described in HADs not only reduce catalytic activity but also lead to substantially reduction of overall HAD protein levels. Therefore, the metabolic changes in had mutants may be due to the loss of HAD activity or, alternatively, due in part to some non-catalytic “moonlighting” function of HADs. In this alternative model of HAD-mediated metabolic regulation, enzymatic activity serves as a molecular switch between “active” and “inactive” conformational states (Figure 2B). Following the paradigm established by small GTPases (such as Ras and Rab homologs), 10 the conformational change in a given HAD might modulate localization or protein-protein interactions to mediate a regulatory effect. In support of this alternative model, structural studies of substrate-bound HAD1 by Park et al 11 find that HAD1 undergoes a large conformational change (from an “open” to “closed” conformation) during cap domain closure on substrate binding.
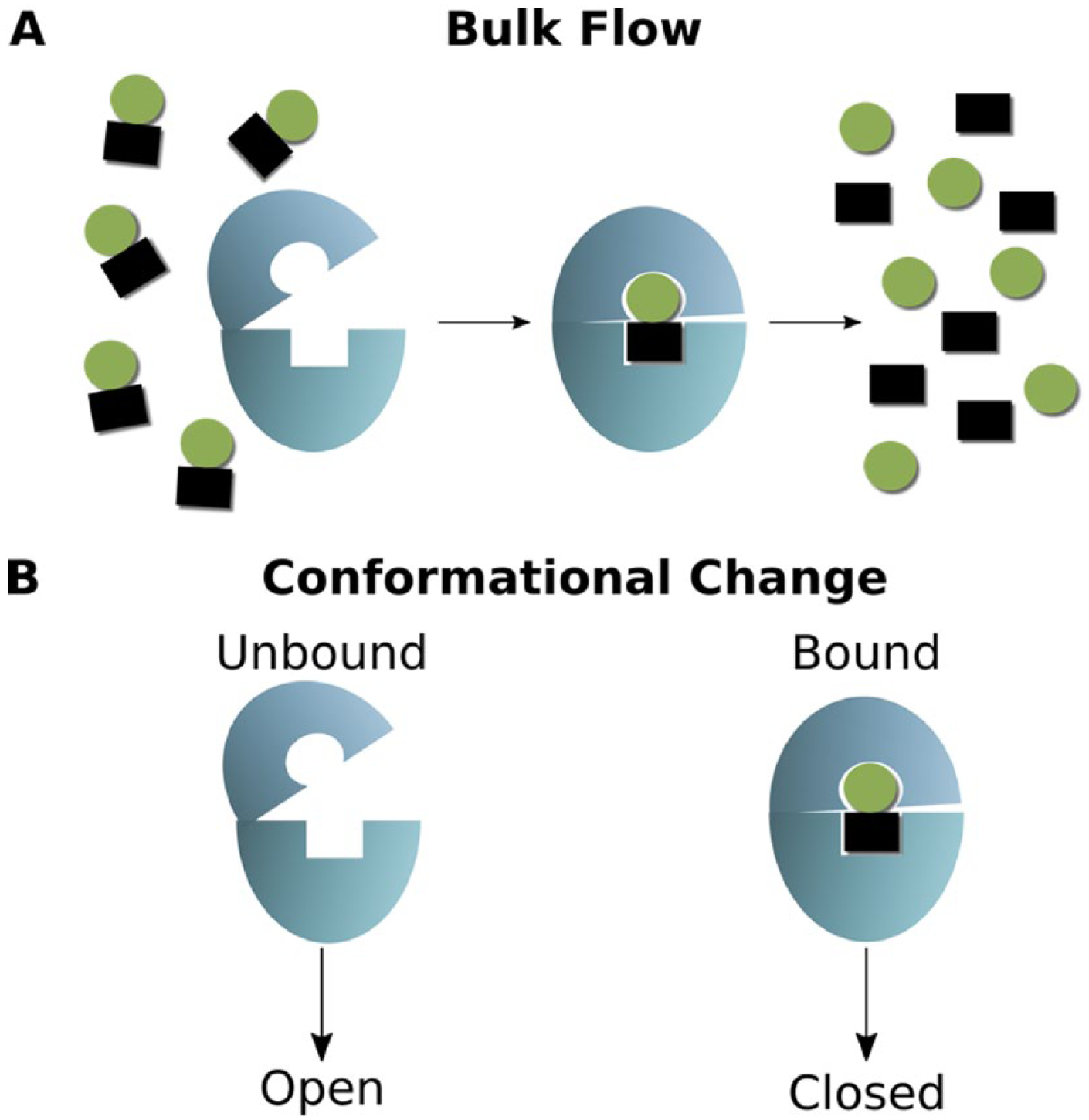
Mechanistic models of HAD-mediated metabolic regulation. (A) In Model 1, HAD proteins catalyze the bulk flow of metabolites from one state to another. (B) In Model 2, substrate binding and hydrolyses instead induce conformational changes that dictate the regulatory functions of HAD proteins, perhaps by mediating localization or protein-protein interactions. HAD indicates haloacid dehalogenase.
HAD proteins make up an enormous superfamily of enzymes that are involved in a wide range of metabolic pathways. In addition to the annotated biochemical activities of many HAD proteins across biology, a novel role for HAD proteins has emerged through studying malaria parasites: control of metabolic homeostasis and drug sensitivity. Parasites that lack certain HAD enzymes can be either sensitized to or resistant to FSM, at the expense of intra-erythrocytic growth rates, and exhibit global changes to their core metabolism. Understanding the underlying mechanisms by which malaria parasites control essential metabolism may guide development of future therapeutic strategies to combat malaria. In addition, studies of Plasmodium HADs are likely to reveal paradigms that may apply to the many thousands of HAD proteins across the domains of life, whose cellular functions are currently unknown.
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: P.M.F. was supported by NIH T32 GM007067. A.R.O.J. was supported by NIH/NIAID R01AI103280, R01AI123433, R21AI144472, and the Burroughs Welcome Fund (PATH).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
PMF and AROJ contributed equally to the design, writing, and editing of this manuscript.
