Abstract
Acute respiratory infections (ARIs) are the leading cause of infectious disease–related morbidity, hospitalization, and morbidity among children worldwide. This study aimed to assess the viral and bacterial causes of ARI morbidity and mortality in children under 5 years in Senegal. Nasopharyngeal samples were collected from children under 5 years who had ARI. Viruses and bacteria were identified using multiplex real-time reverse transcription-polymerase chain reaction and conventional biochemical techniques, respectively. Adenovirus was the most prevalent virus (50%; n = 81), followed by influenza virus (45.68%, n = 74), rhinovirus (40.12%; n = 65), enterovirus (25.31%; n = 41), and respiratory syncytial virus (16.05%; n = 26), whereas Streptococcus pneumoniae (17%; n = 29), Moraxella catarrhalis (15.43%; n = 25), and Haemophilus influenzae (8.02%; n = 13) were the most commonly isolated bacteria. Virus pathogens seem more likely to be more prevalent in our settings and were often associated with bacteria and S. pneumoniae (6%; 16) coinfection.
Introduction
Respiratory tract infections (RTIs) such as acute otitis media, sinusitis, bronchitis, and community-acquired pneumonia are a leading cause of infectious disease–related morbidity, hospitalization, and mortality among children worldwide, particularly in low-income countries. 1
According to World Health Organization (WHO), the prevalence of hospitalized children under 5 years with acute respiratory infections (ARIs) is estimated to be 20% and 90% of those were due to pneumonia. 2 In addition, the number of childhood deaths annually related to ARIs is very important and is estimated between 1.9 and 2.2 million, and 70% of the deaths took place in Africa and Southeast Asia which are the most.1,3
Bacteria and viruses have been reported as the main causes of ARIs. In children under 5 years, ARIs are caused mainly due to viruses; respiratory syncytial viruses (RSVs), parainfluenza viruses, influenza virus A and B, and human metapneumovirus (hMPV) are the most common viruses isolated.4,5 However, primary infections with viral pathogens can predispose to secondary bacterial infections, and the most frequently isolated bacteria in ARIs include Streptococcus pneumonia and Haemophilus influenzae. 6
These bacteria were increasingly resistant to the most commonly used antibiotics for ARI treatment, 7 leading to increase in mortality rates, hospital durations, and health care–associated costs. 8
In resource-limited countries, bacteria have been the main cause of ARIs. This could be explained by the inaccessibility of molecular diagnostic tools thus leading to inadequate antibiotics prescription and consequently contributed to a rapid increase in antimicrobial resistance among bacteria causing RTIs.9,10
In Senegal, few studies on viral and bacterial etiologies of RTIs are available in pediatric settings. Respiratory tract infections are mainly empirically treated with antibiotics on a simple suspicion of bacterial infection. Indeed, this could be one of the major causes of high morbidity and mortality rates.
The aim of this study was to investigate the viral and/or bacterial infections associated with ARIs in children under 5 years.
Materials and Methods
Recruitment and samples
This study was conducted between January 2015 and January 2016 at the Albert Royer Paediatric hospital, the Roi Baudouin Hospital, and the Abass Ndao Hospital, all located in Dakar, Senegal. Inclusion criteria were as follows: children aged under 5 years attending with an upper or lower airway infection.
Bronchoalveolar lavage, sinus fluids, and throat swab samples were collected and referred to the biotechnology unit of the laboratory of bacteriology and virology of Aristide Le Dantec and to the virology unit of Pasteur Institute.
This study has been approved by the Ethics Committee for Research of the Cheikh Anta DIOP University of Dakar. Samples and information for questionnaires have been collected after patient’s informed consent.
Identification of bacterial isolates
Bacteria were isolated from appropriate culture media and incubation conditions prior identification following microbiological standard procedures based on morphological, cultural, and biochemical or antigenic characters. Strains were identified if the bacterial load was at least 105 CFUs/mL.
Nucleic acid extraction and viral detection
Total viral nucleic acid (DNA and RNA) was extracted from 140 μL of each clinical specimen using the PureLink Viral RNA/DNA Mini Kit (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s recommendations. DNA/RNA was eluted with 60 μL nuclease-free water and stored at −80°C until use. A 2-step multiplex real-time reverse transcription-polymerase chain reaction was performed with a Bio-Rad CFX-96 thermo cycler (Bio-Rad Laboratories, Singapore) and the Anyplex II RV16 Detection Kit (Seegene, Seoul, Korea) for a simultaneous testing of influenza viruses (fluA and fluB), human respiratory syncytial virus (RSVA and RSVB), human adenoviruses (HAdV), HMPV, human coronavirus (229E, NL63, OC43), human parainfluenza virus (PIV1, 2, 3, and 4), human rhinovirus (HRV), human enterovirus (HEV), and human bocavirus (HBoV).
Statistical analysis
Data were analyzed using R statistical software version 3.3.1. 11
Results
Over a period 1 year, 162 children with ARIs were included in this study including 97 (59.9%) men. Overall, 30 children were under 6 months of age, 26 were 6 to 12 months old, 31 were 12 to 24 months old, 53 were 24 to 60 months old, and 23 were 60 to 112 months old.
Table 1 shows the distribution of major viruses isolated during this study period, the frequency of single or coinfection, and the number of infections stratified by age group.
Distribution of virus and number of coinfections stratified by age group.
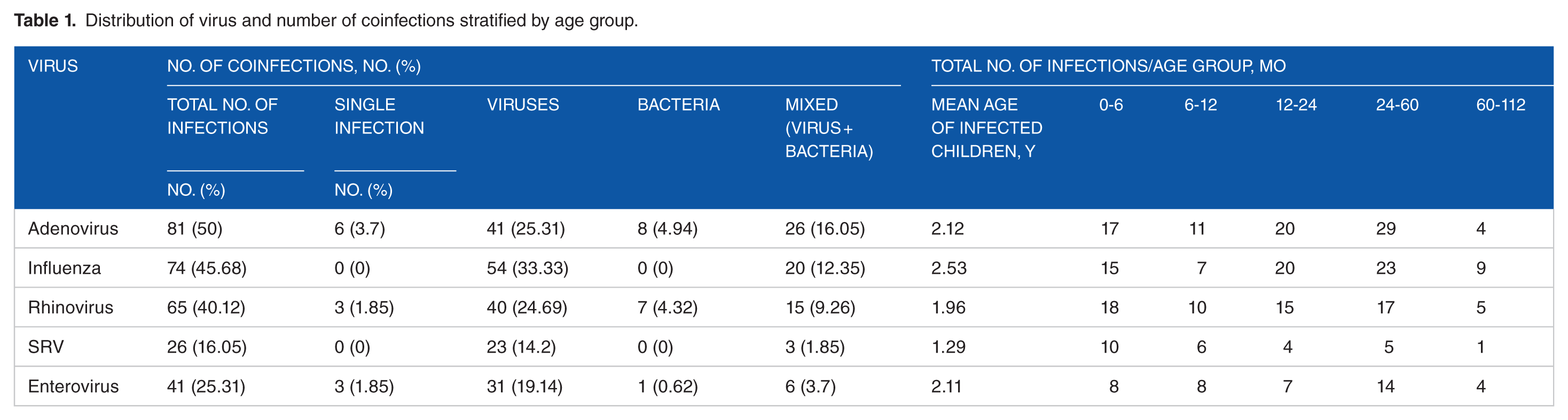
Among selected virus species, adenovirus was the most prevalent (50%, n = 81), followed by influenza virus (45.68%, n = 74), rhinovirus (40.12%, n = 65), enterovirus (25.31%, n = 41), and RSV (16.05%, n = 26).
Single infection with adenovirus was very rare (3.7%, n = 6). However, adenovirus was associated with other viruses and bacteria in 25.31% (n = 41) and 4.94% (n = 8), respectively.
In addition, adenovirus/other viruses/bacteria coinfections have been detected in 16.05% (n = 26) of children.
Influenza virus infections were associated with single-virus coinfections (33.3%, n = 54) or virus and bacteria coinfections (12.35%, n = 20). However, influenza virus single-infected children were not detected.
Low prevalence of both rhinovirus and enterovirus single infections was observed (1.85%, n = 3). These infections were most often associated with single virus or virus and bacteria coinfections.
Adenovirus, influenza virus, and enterovirus infections were more prevalent among children aged from 24 to 60 months.
The prevalence of RSV infections associated with other viruses or other viruses and bacteria was 19.14% and 1.85%, respectively; however, no RSV single infection was detected.
Children aged from 0 to 6 months were more likely to be more affected with rhinovirus and RSV infections.
The distribution of major bacterial clinical strains is shown in Table 2. Among isolated bacterial species, S. pneumoniae was the most prevalent bacteria (17%) followed by M. catarrhalis (15.43%) and H. influenzae (8.02%).
Distribution of bacteria and number of coinfections by stratified age group.
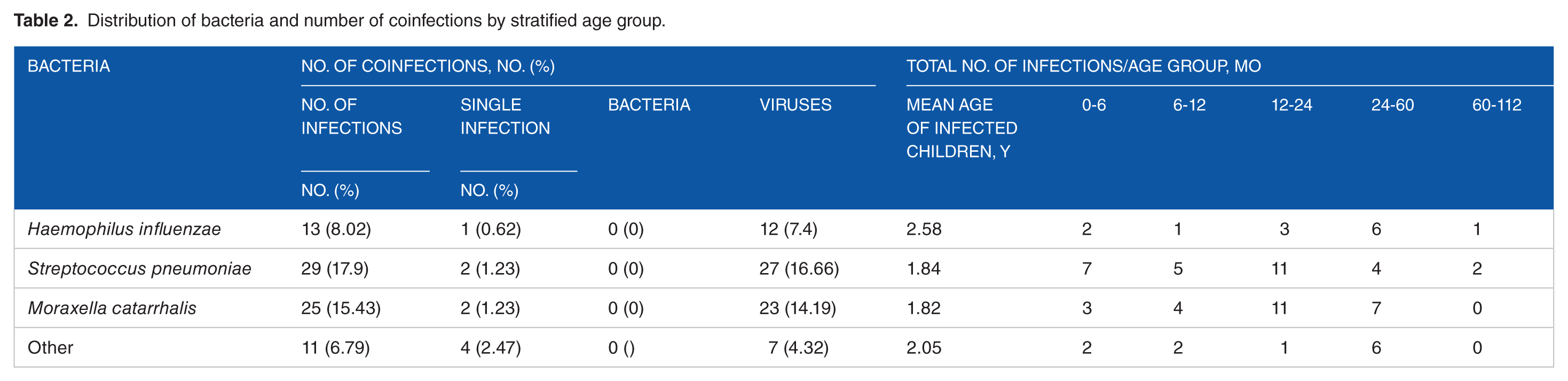
Bacterial single infections were also very rare: S. pneumoniae (2%), M. catarrhalis (2%), and H. influenzae (1%).
No bacteria/bacteria coinfections were detected. However, coinfections with bacteria and viruses had been observed in children having ARIs and coinfection with S. pneumoniae (16.66%) was most frequent followed by M. catarrhalis (14, 19%) and H. influenzae (4.32%).
The contribution of viral and bacterial pathogens to the clinical syndrome is depicted in Tables 3 and 4, respectively.
Contribution of viral pathogens to clinical syndrome.
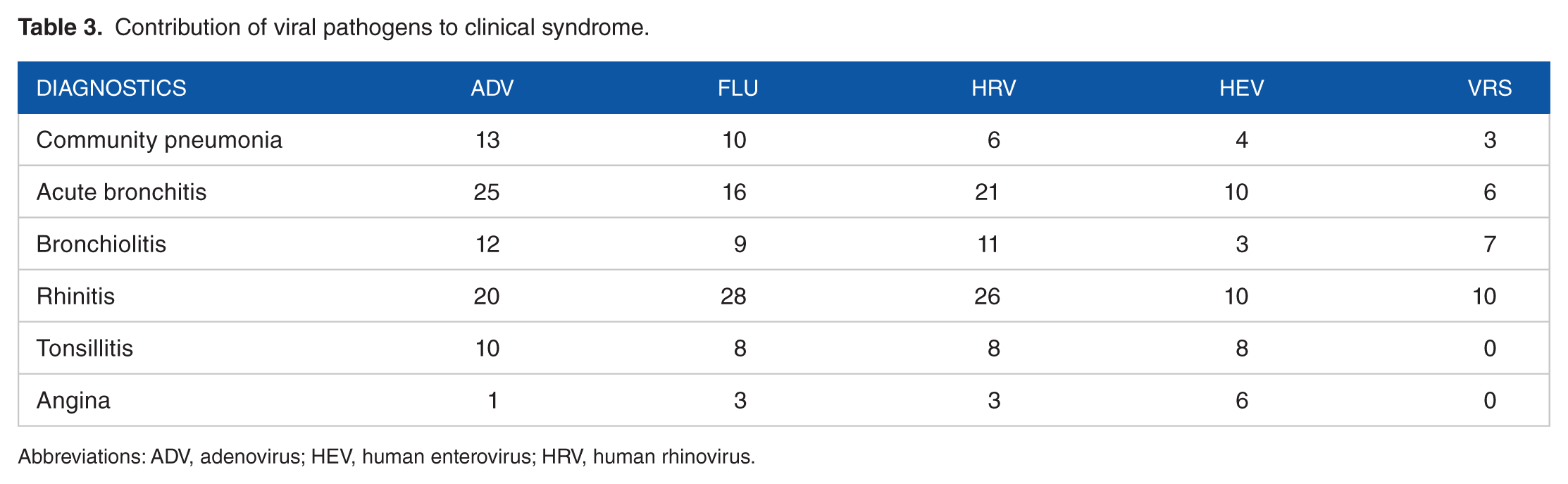
Abbreviations: ADV, adenovirus; HEV, human enterovirus; HRV, human rhinovirus.
Contribution of bacterial pathogens to clinical syndrome.
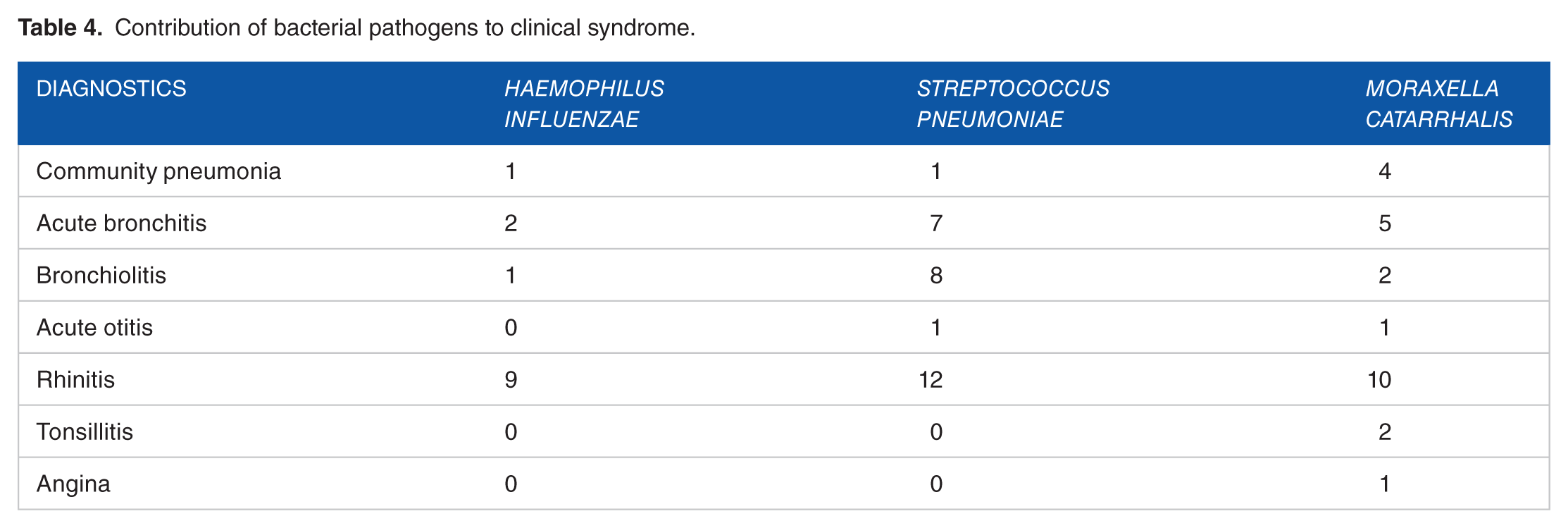
Discussion
Acute respiratory infections may be caused by viruses, bacteria, or both. Thus, determination of the causative agent is crucial to limit the abuse of antibiotics.
In our study conducted over a period of 1 year, we showed that viruses were the most common pathogens detected in ARIs. Similar findings have also been reported in a study conducted in United States by Obasi et al 12 in 2013. Several studies conducted in other countries such as Burkina Faso, 10 Zambia, 9 and Niger 13 provided evidence of the role of viruses in upper and lower airway diseases.
In Senegal, adenovirus was the most common pathogen detected from ARIs in children. This study confirms previous results reported between 2013 and 2015 in Senegal by Niang et al. 14 The overall detection rate of adenovirus was 50% among the 162 selected children. This adenovirus infection prevalence seems very high compared with rates documented from other countries, namely, Ghana, 15 Burkina, 10 and Zambia. 9 However, our results are in agreement with data reported in Cameroon. Overall, this confirms the overall high prevalence of adenovirus ARI in Africa. 16
In addition to adenovirus, influenza virus, rhinovirus, and enterovirus are the most common viruses detected, as reported in other studies.10,17
Our study revealed viral codetection is frequent in ARIs. Similar results were found in other countries.9,10,15 Viral codetection in clinical settings is becoming more common since the introduction of molecular-based multiplex tests. Although the clinical significance of these findings remains unclear, this seems to have no impact in disease severity. 18
In this study, we also found that ARIs were associated with bacterial pathogens; among those, S. pneumoniae (17.9%), M. catarrhalis (15.43%), and H. influenzae (8%) were the most common. In contrast with our findings, high rate (56%) of S. pneumoniae detection was observed in Niger among children having ARIs. 13
In our study, prevalence of mono-infections with S. pneumoniae (2%), M. catarrhalis (2%), or H. influenzae (1%) was very low. Thus, this low prevalence of bacterial pathogens in ARIs proves that antibiotics should no longer be systematically used in the treatment of ARIs. However, among low-prevalence bacterial isolates, codetections with viruses were more frequent and S. pneumoniae (16, 6%) coinfection was the most frequent in our findings. Similar results were reported in other studies.19,20
There results highlight the pivotal role of viruses in ARIs because primary infection with viral pathogens can predispose children to subsequent bacterial infections. 21
Our study has limitations; among others, the sample size which does not really allow to determine the direct association between viral or bacterial infection and disease severity. Moreover, a case-control study would be more suitable to determine the exact role of viruses in ARIs pathogenesis.
In conclusion, this study reports the profile of viral and bacterial pathogens among children under 5 years hospitalized with ARIs in Senegal. The high prevalence rates of viral infections and its clinical impact highlight the need to implement a systematic surveillance program for a better management of ARIs in children, particularly in resource-limited settings.
Footnotes
Acknowledgements
The authors are grateful to CEA-SAMEF (Dakar, Senegal), Micro CSB System (Dakar, Senegal), and Institute Pasteur Dakar (Dakar, Senegal) who have contributed to the success of this study.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partly supported by Micro CSB System (Dakar, Senegal), CEA-SAMEF (Dakar, Senegal), GlaxoSmithKline (GSK), and Institute Pasteur (Dakar, Senegal).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
BCS and DA conceived and designed the experiments. LC and DA analyzed the data. DA and CM wrote the first draft of the manuscript and jointly developed the structure and arguments for the paper. DA, CM, DAb, FA, BD, DA, DJBN, DA, LC, DN, NM, and FL contributed to the writing of the manuscript. DA, CM, and BCS made critical revisions and approved final version. All authors agree with manuscript results and conclusions and reviewed and approved the final manuscript.
