Abstract
Recently, Coronavirus has been given considerable attention from the biomedical community based on the emergence and isolation of a deadly coronavirus infecting human. To understand the behavior of the newly emerging MERS-CoV requires knowledge at different levels (epidemiologic, antigenic, and pathogenic), and this knowledge can be generated from the most related viruses. In this study, we aimed to compare between 3 species of Coronavirus, namely Middle East Respiratory Syndrome (MERS-CoV), Severe Acute Respiratory Syndrome (SARS-CoV), and NeoCoV regarding whole genomes and 6 similar proteins (E, M, N, S, ORF1a, and ORF1ab) using different bioinformatics tools to provide a better understanding of the relationship between the 3 viruses at the nucleotide and amino acids levels. All sequences have been retrieved from National Center for Biotechnology Information (NCBI). Regards to target genomes’ phylogenetic analysis showed that MERS and SARS-CoVs were closer to each other compared with NeoCoV, and the last has the longest relative time. We found that all phylogenetic methods in addition to all parameters (physical and chemical properties of amino acids such as the number of amino acid, molecular weight, atomic composition, theoretical pI, and structural formula) indicated that NeoCoV proteins were the most related to MERS-CoV one. All phylogenetic trees (by both maximum-likelihood and neighbor-joining methods) indicated that NeoCoV proteins have less evolutionary changes except for ORF1a by just maximum-likelihood method. Our results indicated high similarity between viral structural proteins which are responsible for viral infectivity; therefore, we expect that NeoCoV sooner may appear in human-related infection.
Introduction
Coronaviruses (CoVs) have a large-scale spreading worldwide and infect human and various animal hosts, causing diseases which range from mostly upper respiratory tract infections in humans to gastrointestinal tract infections, encephalitis, and demyelination in animals and can be lethal.
1
Bat CoVs have been given extraordinary consideration as 2 emerged CoVs have been linked to unpredicted human disease outbreaks in the 21st century resulted in high mortality rate and extremely economic disruption.
2
These 2 viruses are the Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) and the Middle East Respiratory Syndrome Coronavirus (MERS-CoV) which were suggested to be originated from bats.
2
Based on genotypic and serological characterization, the International Committee for Taxonomy of Viruses (ICTV) has reported 4 CoVs genera, namely
It has previously been identified that MERS-CoV is a beta CoV causing high morbidity and mortality in humans. 7 In Saudi Arabia, it has been detected in a patient clinically diagnosed with a severe respiratory infection. 8 In June 11, 2014, MERS-CoV infection has been diagnosed in 699 patients mainly from the Arabian Peninsula with a case fatality rate potentially exceeding the rate reported during the SARS-CoV pandemic. 9 A research has reported a high rate of neutralizing antibodies against MERS-CoV found in camels in the Arabian Peninsula showing high relationship at genetic level to those from human cases suggesting that these camels are constituting the source of human infections. 10 At molecular level, the CoVs have a high frequency of recombination due to their unique replication mechanism which increases the propensity to result in high rates of mutation allowing the viruses to acclimatize to new hosts and ecological niches. 11 De Benedictis et al have characterized small genomic sequence fragments of bat CoVs (BtCoVs) that were closely related to MERS-CoV and suggested that MERS-CoV ancestors may have evolved in bats. 12 In China, since 2002, SARS-CoV was implicated as a causative agent of SARS and caused atypical pneumonia that spread rapidly throughout parts of Asia, North America, and Europe during 2002 to 2003 with cases having been reported in 30 countries. 13 According to the World Health Organization report, the mortality rate of SARS-CoV was more than 10%. 5 Close person-to-person contact has been shown to be the major transmission way of SARS-CoV principally via contact with aerosolized droplets or other bodily fluids. 14 Shortly after SARS-CoV outbreak and the subsequent implication of bats as reservoir hosts of the causative agent, CoV drove numerous studies on bats and the viruses they harbor. A specimen from Neoromicia cf. zuluensis bat in 2011 yielded a novel betacoronavirus called NeoCoV. 15
According to the Ndapewa Ithete and his colleagues results (in 2013), NeoCoV differed from MERS-CoV by only one amino acid (a.a) exchange (0.3%) in the translated 816-nt
In this study, we have attempted to provide a better understanding of the relationship between MERS-CoV, SARS-CoV, and NeoCoV at the level of amino acids regarding 6 similar proteins, including E, M, N, S, ORF1a, and ORF1ab, using different bioinformatics tools. The leading force for this study was the previous studies which constructed phylogenetic tree between different species of Coronaviridae based on either structural protein and nonstructural protein or whole genome, and they have found that there was some relationship between MERS-CoV and SARS-CoV, while others studied the relationship between MERS-CoV and NeoCoV but there was no study included MERS-CoV, SARS-CoV, and NeoCoV in the same study to know whose is the most related to whom. Bioinformatics tools and Phylogenetic analysis enables us to understand relationships between ancestral sequences and its descendants.
Materials and Methods
Bioinformatics processing and data analysis
In this study, genome sequences of the 3 target species of CoV were retrieved from the National Center for Biotechnology Information (NCBI; genome and nucleotide databases; https://www.ncbi.nlm.nih.gov/genome, https://www.ncbi.nlm.nih.gov/nuccore), namely MERS-CoV (genome ID: 31360), SARS-CoV (genome ID: 10320), and NeoCoV (genome ID: KC869678). However, 4 structural proteins, E, S, N, and M, and 2 NS proteins, ORF1a and ORF1ab, of each species were obtained from the NCBI protein database (www.ncbi.nlm.nih.gov/Protein/). Table 1 presents general information about all retrieved both nucleotide and protein sequences. These Genome and protein sequences were then subjected for comparison using different bioinformatics prediction tools.
General information of retrieved genomes and protein sequences.
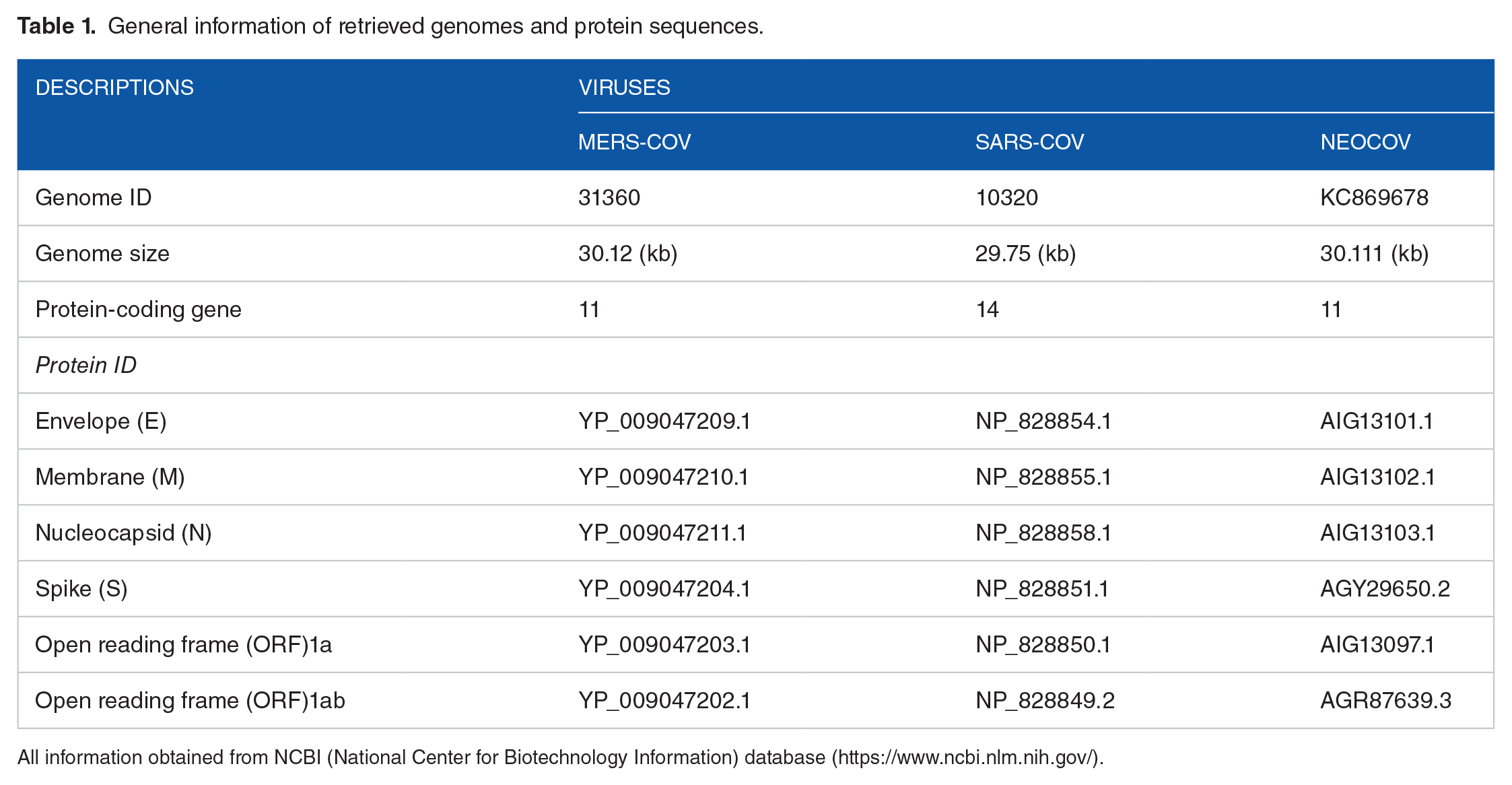
All information obtained from NCBI (National Center for Biotechnology Information) database (https://www.ncbi.nlm.nih.gov/).
Compute nucleotide composition and pair-wise alignments
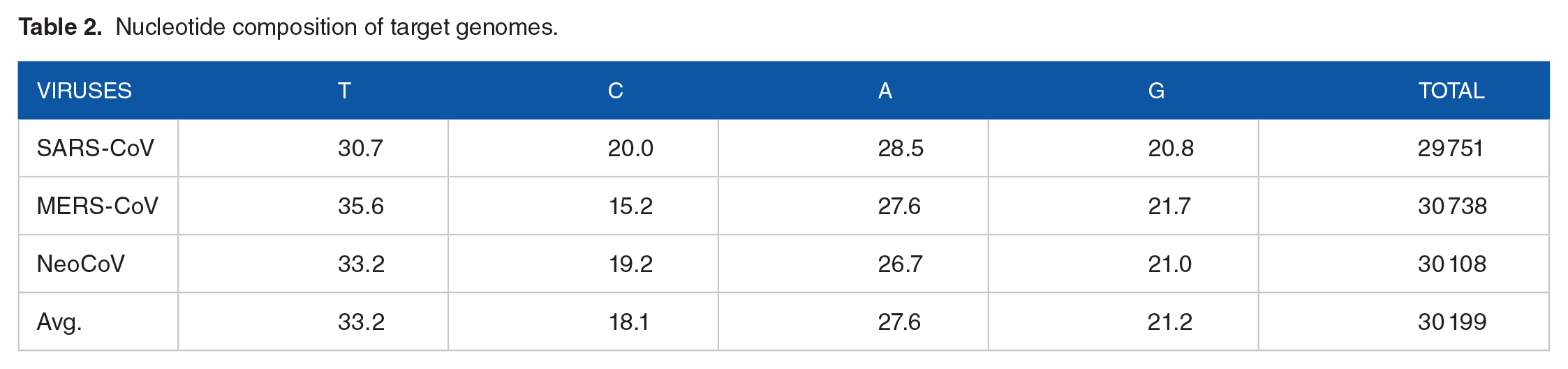
Nucleotide composition of the target genomes (MERS-CoV, SARS-CoV, and NeoCoV) was calculated as shown in Table 2 using Molecular Evolutionary Genetics Analysis Software Version 7.0 (MEGA7; https://www.megasoftware.net/home). Furthermore, pairwise alignment was done for each pair of target genomes using BLAST Needleman-Wunsch Global Align Nucleotide Sequences (https://blast.ncbi.nlm.nih.gov/Blast.cgi) as it is presented in Figure 1.
Nucleotide composition of target genomes.
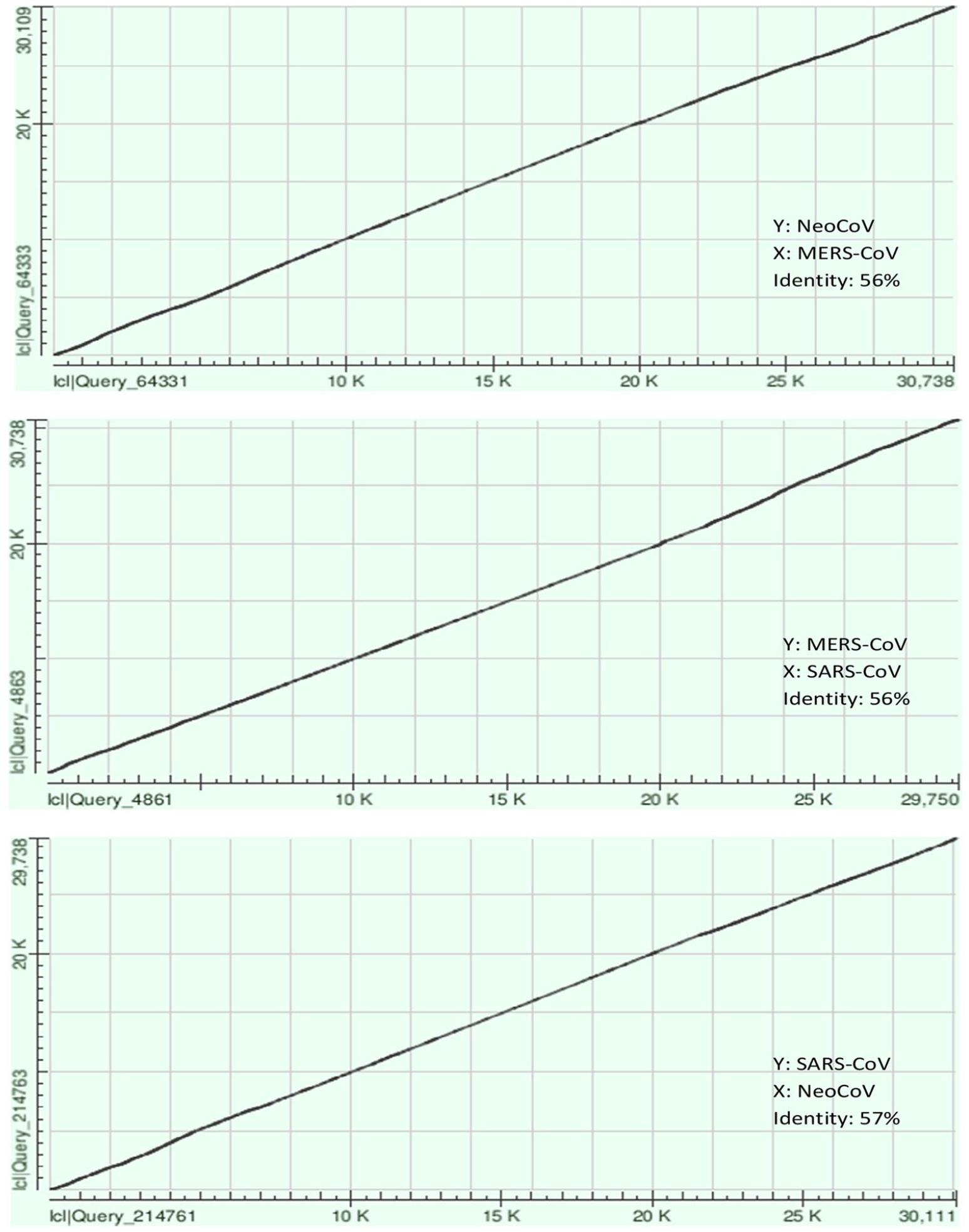
Global pairwise alignment results by Needleman-Wunsch method.
Multiple sequence alignment and phylogenetic tree
For the purpose of protein sequences comparison, first, the Multiple Sequence Alignments (MSA) was done using the Clustal method implemented in Clustal Omega tool (http://www.ebi.ac.uk/Tools/msa/clustalo/). Following the alignment, phylogenetic relationships were depicted in phylogram using distance matrix methods (Neighbor-Joining [NJ] and Unweighted Pair Group Method with Arithmetic mean [UPGMA]) in Phylogeny server (http://www.ebi.ac.uk/Tools/phylogeny/clustalw2_phylogeny/).17,18 Once trees were constructed, they were viewed by TreeDyn viewer tool (http://www.treedyn.org/) as shown in boxes B and C in Figures 4, 6, 8, 10, 12, and 14. By the same token, second scenario was as follows, and MSA was done by Multiple Sequence Comparison by Log-Expectation (MUSCLE) method using Muscle online tool (https://www.ebi.ac.uk/Tools/msa/muscle/). After that, alignment results in Phylip or Clustal format were subjected to Gblocks program version 0.91b (alignment curation tool). Furthermore, PhyML 3.0 (using maximum-likelihood method) and Protpars (using Parsimony method) were used to generate Newick format tree files which have been viewed by TreeDyn viewer tool 19 as shown in boxes A and D in Figures 4, 6, 8, 10, 12, and 14. Previous tools are available at Gblocks (http://molevol.cmima.csic.es/castresana/Gblocks_server.html), PhyML 3.0 (http://www.atgc-montpellier.fr/phyml/), and Protpars (http://www.trex.uqam.ca/). The third scenario was revolved around constructing ultrametric phylogenetic trees using Muscle method for multiple sequence alignments and RelTime method to generate the tree. Then, Time-trees were generated by the Molecular Evolutionary Genetics Analysis software Version 7.0 (MEGA7; https://www.megasoftware.net/home). Figures 2 and 3 show the constructed phylogenetic trees of this scenario that have been done by MEGA7.
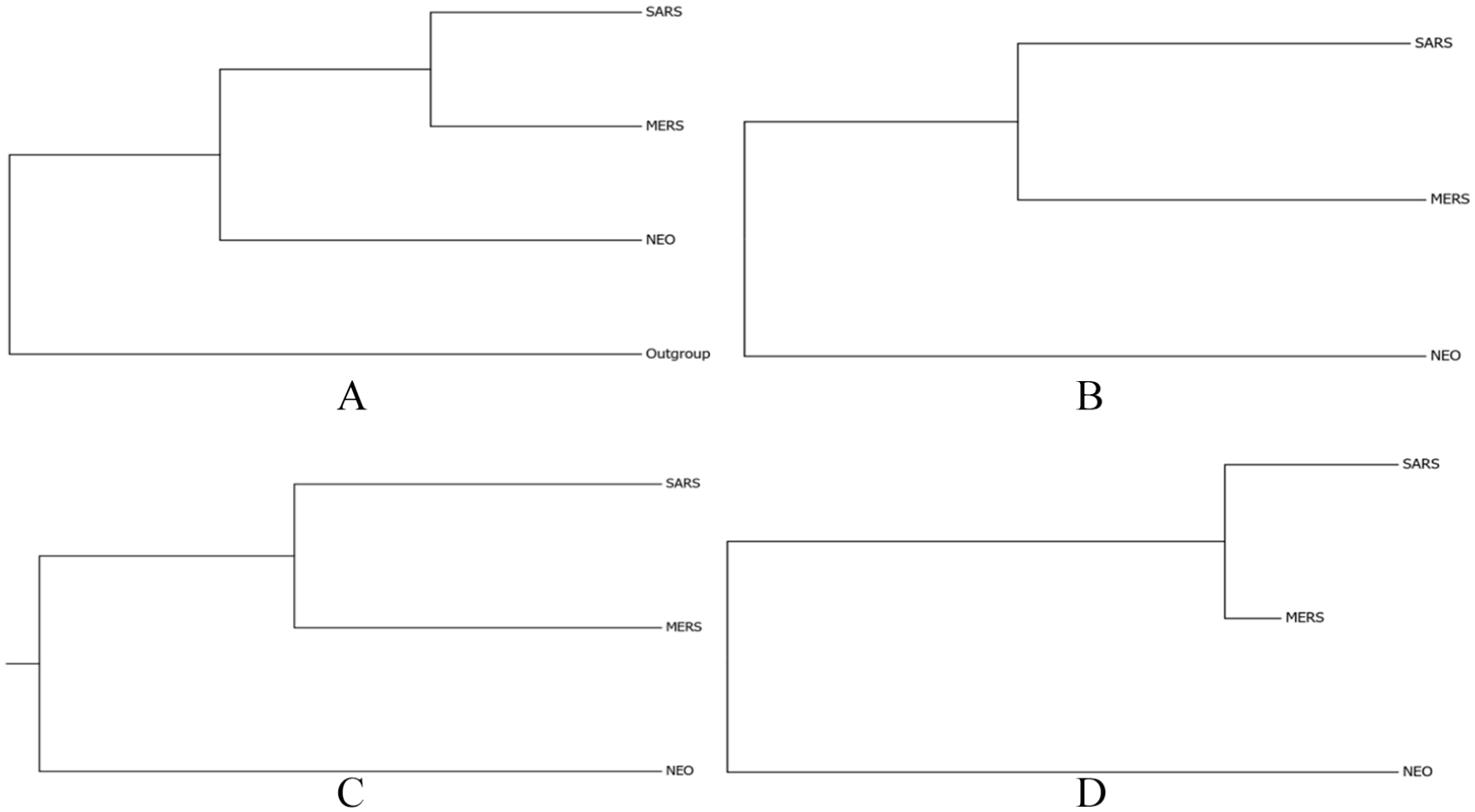
Phylogenetic trees comparing whole genomes of coronavirus species MERS-CoV, SARS-CoV, and NeoCoV, and trees “A-D” were built using different methods. (A) The evolutionary history was inferred using the Maximum Parsimony (MP) method. The most parsimonious tree with length = 42 663 is shown. The consistency index is 0.980475 (0.613278), the retention index is 0.369417 (0.369417), and the composite index is 0.362204 (0.226555) for all sites and parsimony-informative sites (in parentheses). The MP tree was obtained using the Subtree-Pruning-Regrafting (SPR) algorithm20(p126) with search level 0 in which the initial trees were obtained by the random addition of sequences (10 replicates). The analysis involved 4 nucleotide sequences. There were a total of 29 693 positions in the final dataset. (B) The evolutionary history was inferred using the Neighbor-Joining method. 21 The optimal tree with the sum of branch length = 18.91227594 is shown. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Maximum Composite Likelihood method 22 and are in the units of the number of base substitutions per site. The analysis involved three nucleotide sequences. All positions containing gaps and missing data were eliminated. There were a total of 29 690 positions in the final dataset. (C) The evolutionary history was inferred using the UPGMA method. 23 The optimal tree with the sum of branch length = 18.91227594 is shown. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Maximum Composite Likelihood method 22 and are in the units of the number of base substitutions per site. The analysis involved 3 nucleotide sequences. All positions containing gaps and missing data were eliminated. There were a total of 29 690 positions in the final dataset. (D) The evolutionary history was inferred by using the Maximum-Likelihood method based on the Tamura-Nei model. 24 The tree with the highest log likelihood (−121 024.68) is shown. Initial tree(s) for the heuristic search were obtained automatically by applying Neighbor-Join and BioNJ algorithms to a matrix of pairwise distances estimated using the Maximum Composite Likelihood (MCL) approach and then selecting the topology with superior log likelihood value. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. The analysis involved three nucleotide sequences. There were a total of 29 693 positions in the final dataset. All MSA of used sequences was curated by using Gblocks, and evolutionary analyses were conducted in MEGA7. 25 MSA indicates Multiple Sequence Alignments, UPGMA, Unweighted Pair Group Method with Arithmetic Mean.
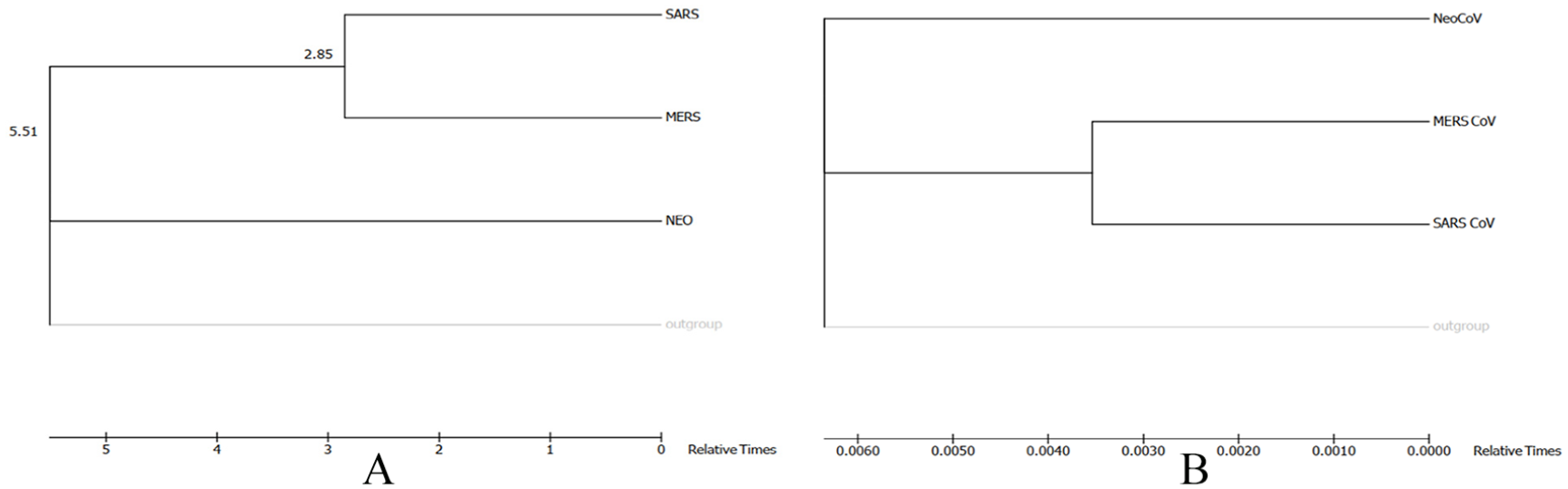
Molecular phylogenetic analysis of MERS-CoV, SARS-CoV, and NeoCoV genomes. The timetree shown was generated using the RelTime method. 26 Divergence times for all branching points in the topology were calculated using the Maximum-Likelihood method based on the Tamura-Nei model. 24 (A) The estimated log likelihood value of the topology shown is 135 729.24. The tree is drawn to scale, with branch lengths measured in the relative number of substitutions per site. The analysis involved 4 nucleotide sequences. There were a total of 30 738 positions in the final dataset. (B) The estimated log likelihood value of the topology shown is 79 370.24. The tree is drawn to scale, with branch lengths measured in the relative number of substitutions per site. The analysis involved 4 nucleotide sequences. There were a total of 22 620 positions in the final dataset. Evolutionary analyses were conducted in MEGA7. 25 (A) Without MSA curation. (B) With MSA curation. MSA indicates Multiple Sequence Alignments.
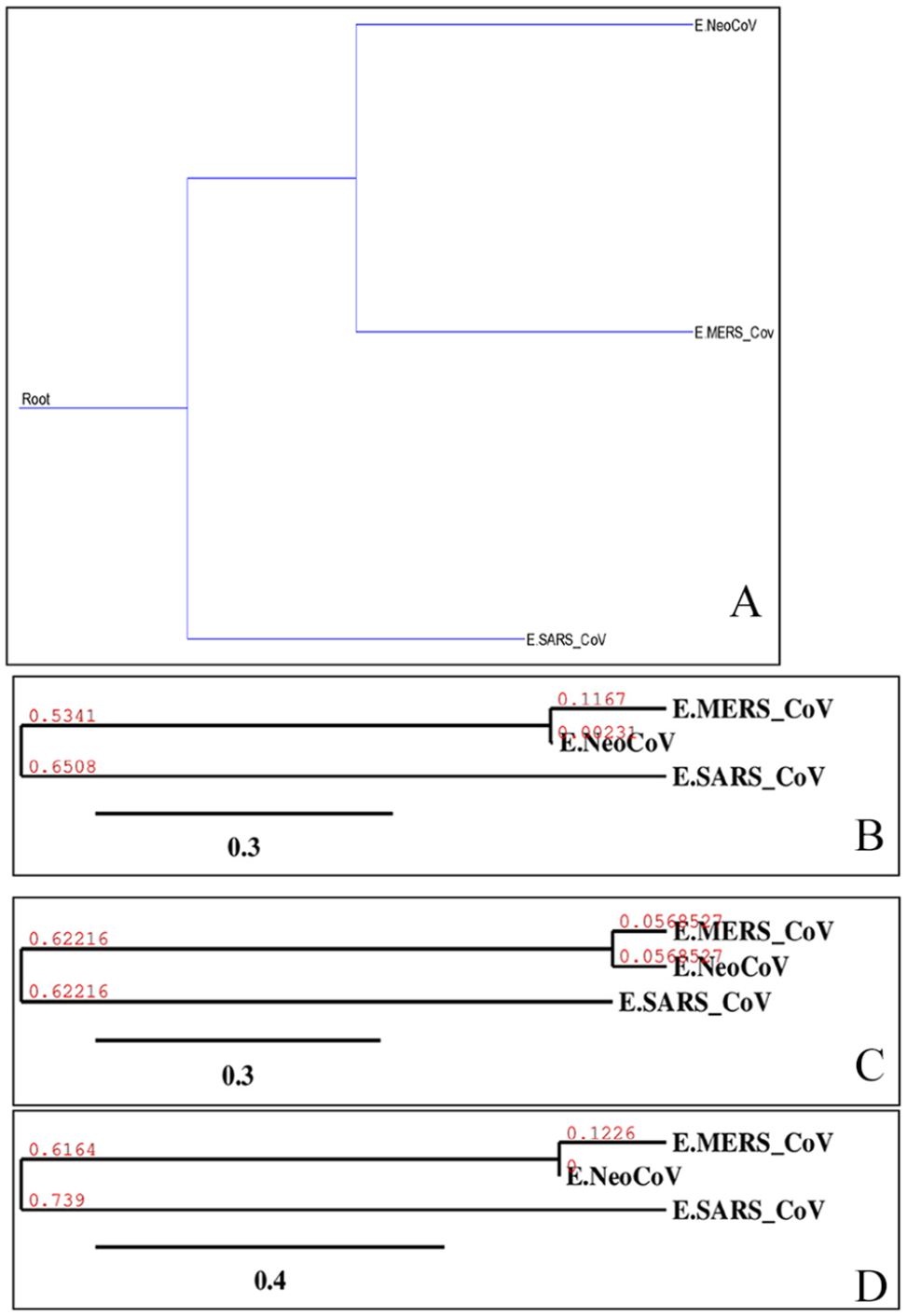
Phylogenetic of “E” proteins of target coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV). Trees (A)-(D) were built using different methods, and they are, respectively, Maximum Parsimony, Neighbor-Joining, UPGMA, and Maximum-Likelihood. UPGMA indicates Unweighted Pair Group Method with Arithmetic Mean.
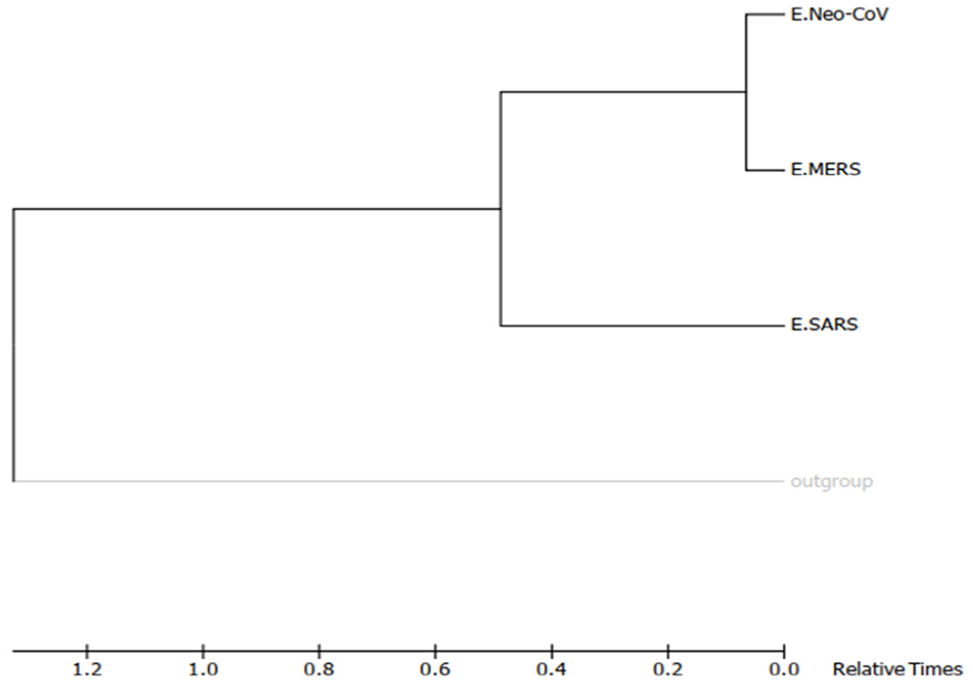
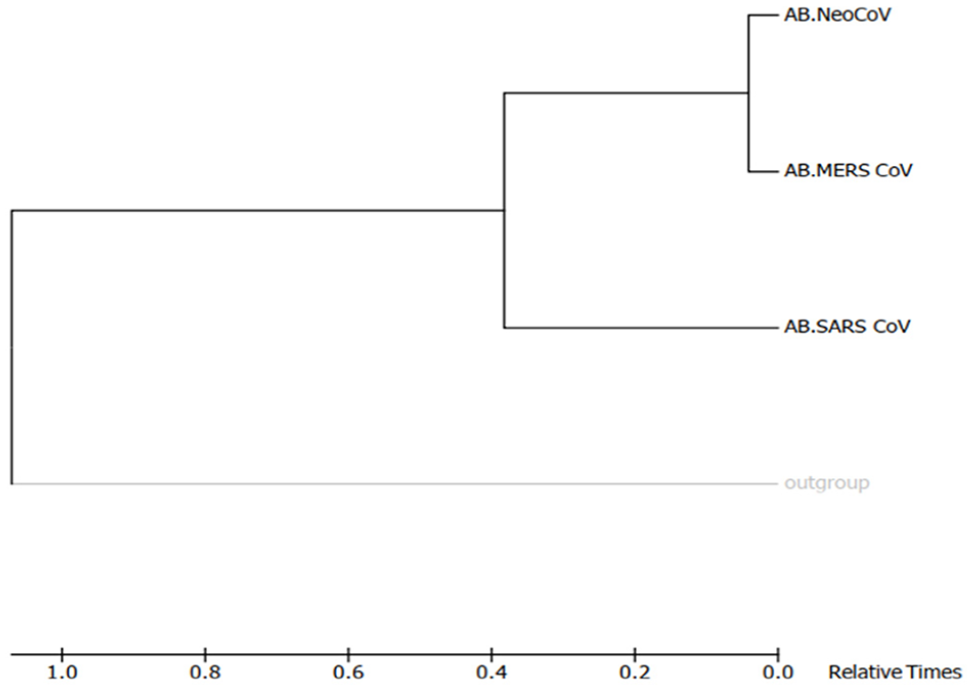
Molecular phylogenetic tree of target coronaviruses “E” proteins by Maximum-Likelihood method (timetree). The timetree shown was generated using the RelTime method. 26 Divergence times for all branching points in the topology were calculated using the Maximum-Likelihood method based on the Equal Input model. 27 The estimated log likelihood value of the topology shown is −703.32. The tree is drawn to scale, with branch lengths measured in the relative number of substitutions per site. The analysis involved 4 amino acid sequences. There were a total of 86 positions in the final dataset. Evolutionary analyses were conducted in MEGA7. 25
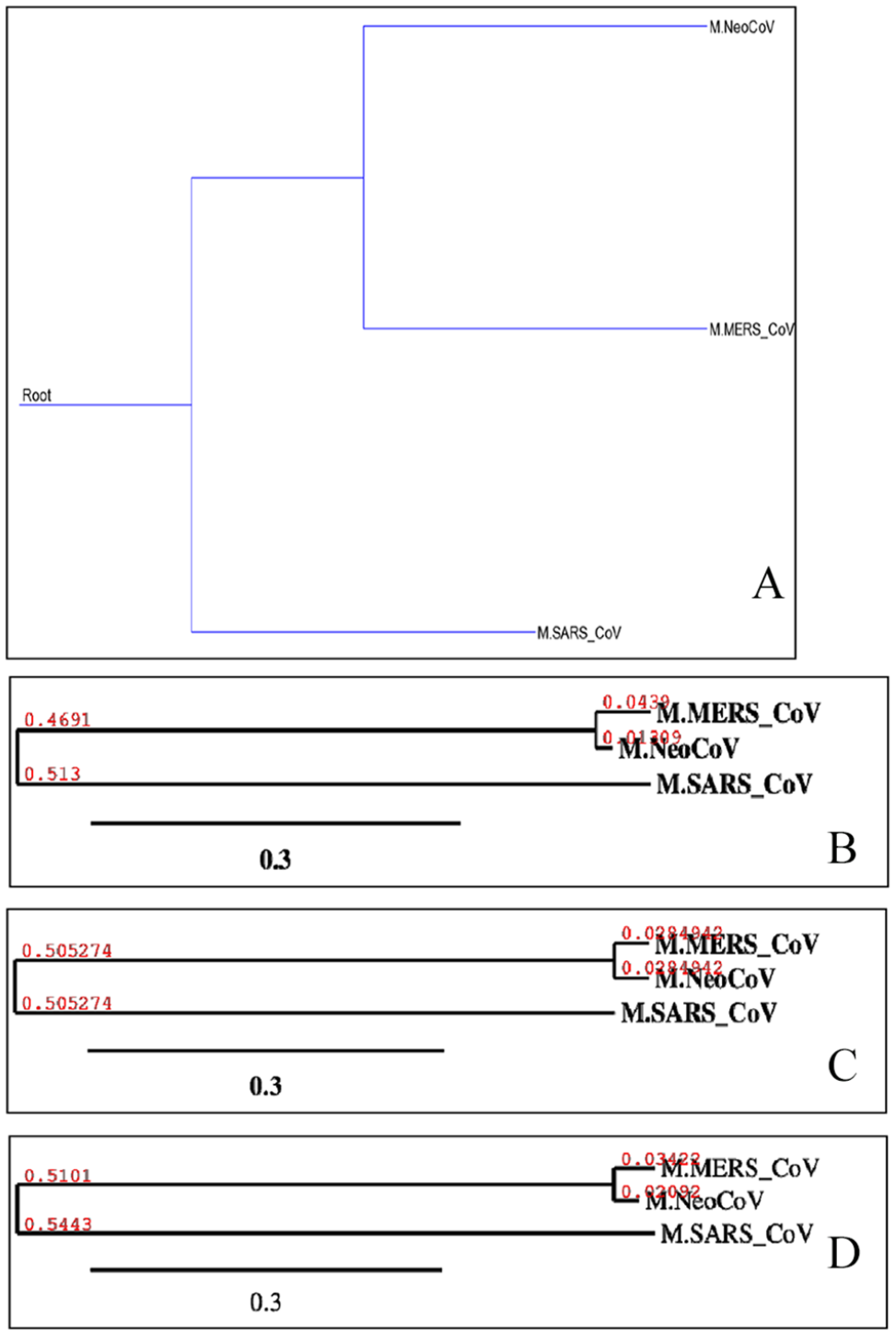
Shows phylogenetics of “M” proteins of target coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV). Trees “A-D” were built using different methods, and they are, respectively, Maximum Parsimony, Neighbor-Joining, UPGMA, and Maximum Likelihood. UPGMA, Unweighted Pair Group Method with Arithmetic Mean.
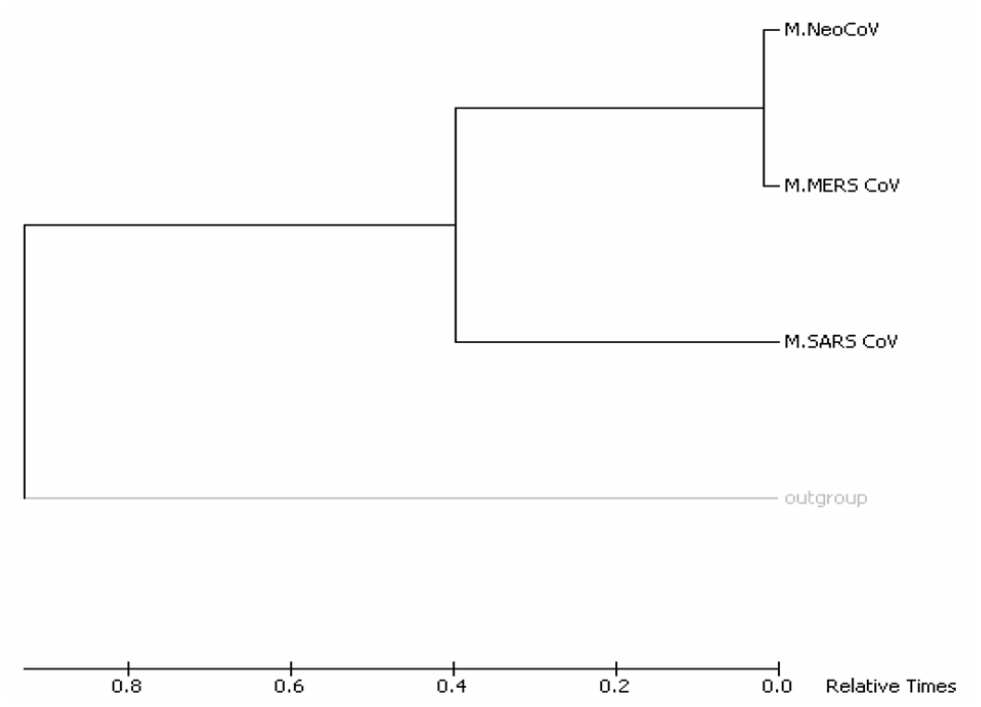
Molecular phylogenetic analysis by Maximum-Likelihood method (timetree). The timetree shown was generated using the RelTime method. 26 Divergence times for all branching points in the topology were calculated using the Maximum-Likelihood method based on the Equal Input model. 27 The estimated log likelihood value of the topology shown is −1802.98. The tree is drawn to scale, with branch lengths measured in the relative number of substitutions per site. The analysis involved 4 amino acid sequences. There were a total of 244 positions in the final dataset. Evolutionary analyses were conducted in MEGA7. 25
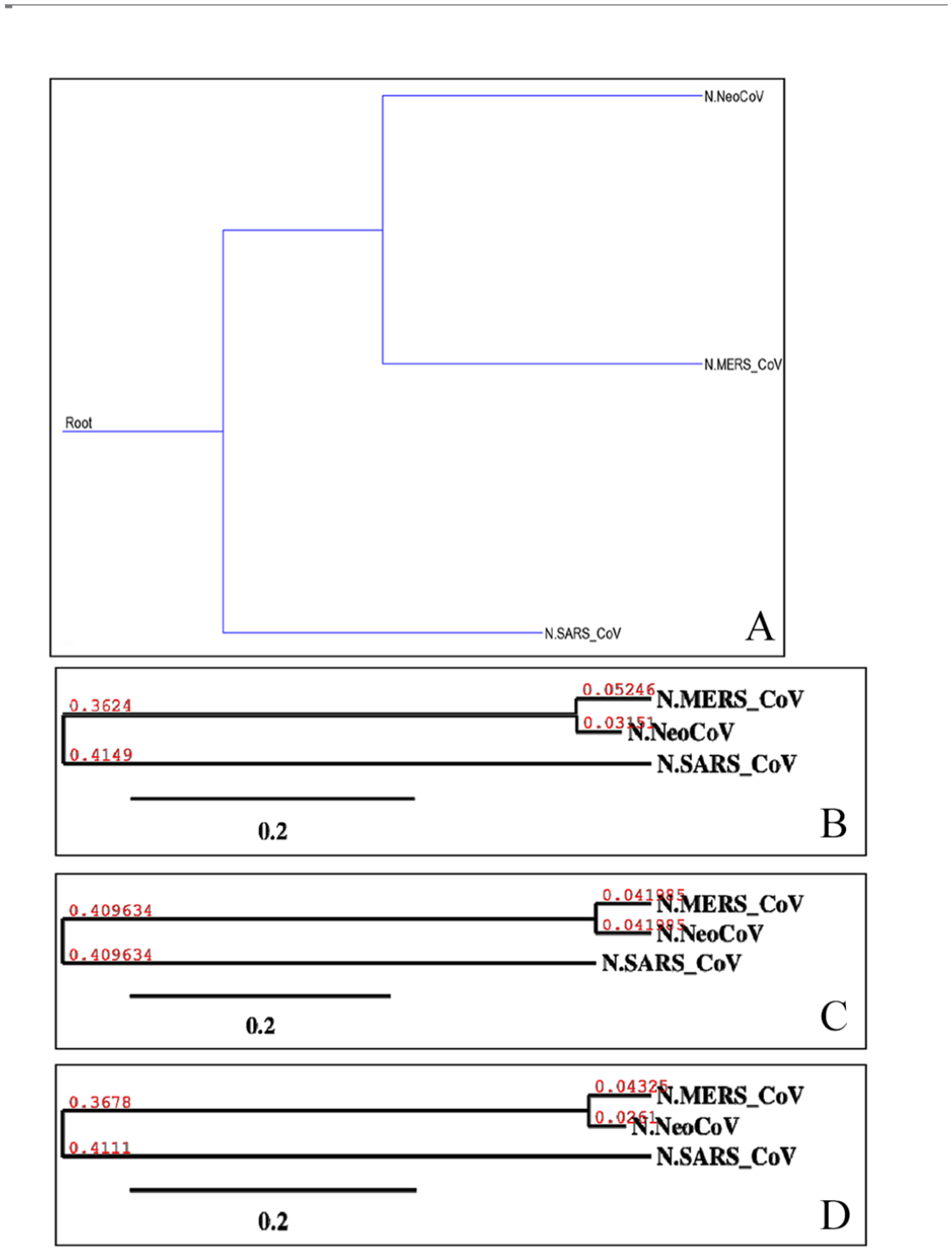
Phylogenetic of “N” proteins of target coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV). Trees (A)-(D) were built using different methods, and they are, respectively, Maximum Parsimony, Neighbor-Joining, UPGMA, and Maximum Likelihood. UPGMA indicates Unweighted Pair Group Method with Arithmetic Mean.
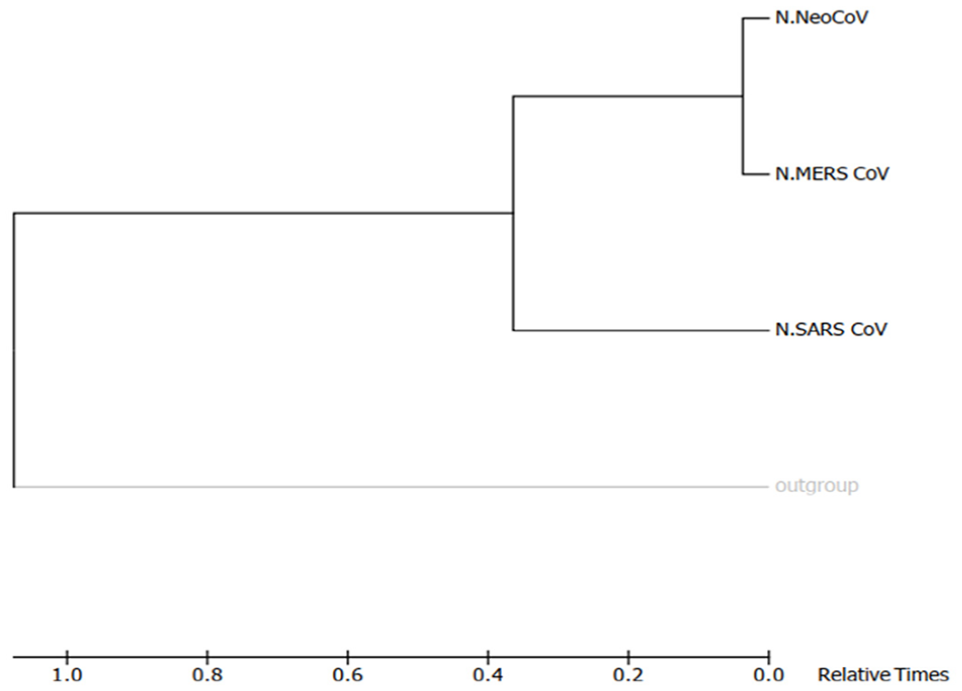
Molecular phylogenetic analysis by Maximum-Likelihood method (timetree). The timetree shown was generated using the RelTime method. 26 Divergence times for all branching points in the topology were calculated using the Maximum-Likelihood method based on the Equal Input model. 27 The estimated log likelihood value of the topology shown is −3425.67. The tree is drawn to scale, with branch lengths measured in the relative number of substitutions per site. The analysis involved 4 amino acid sequences. There were a total of 460 positions in the final dataset. Evolutionary analyses were conducted in MEGA7. 25
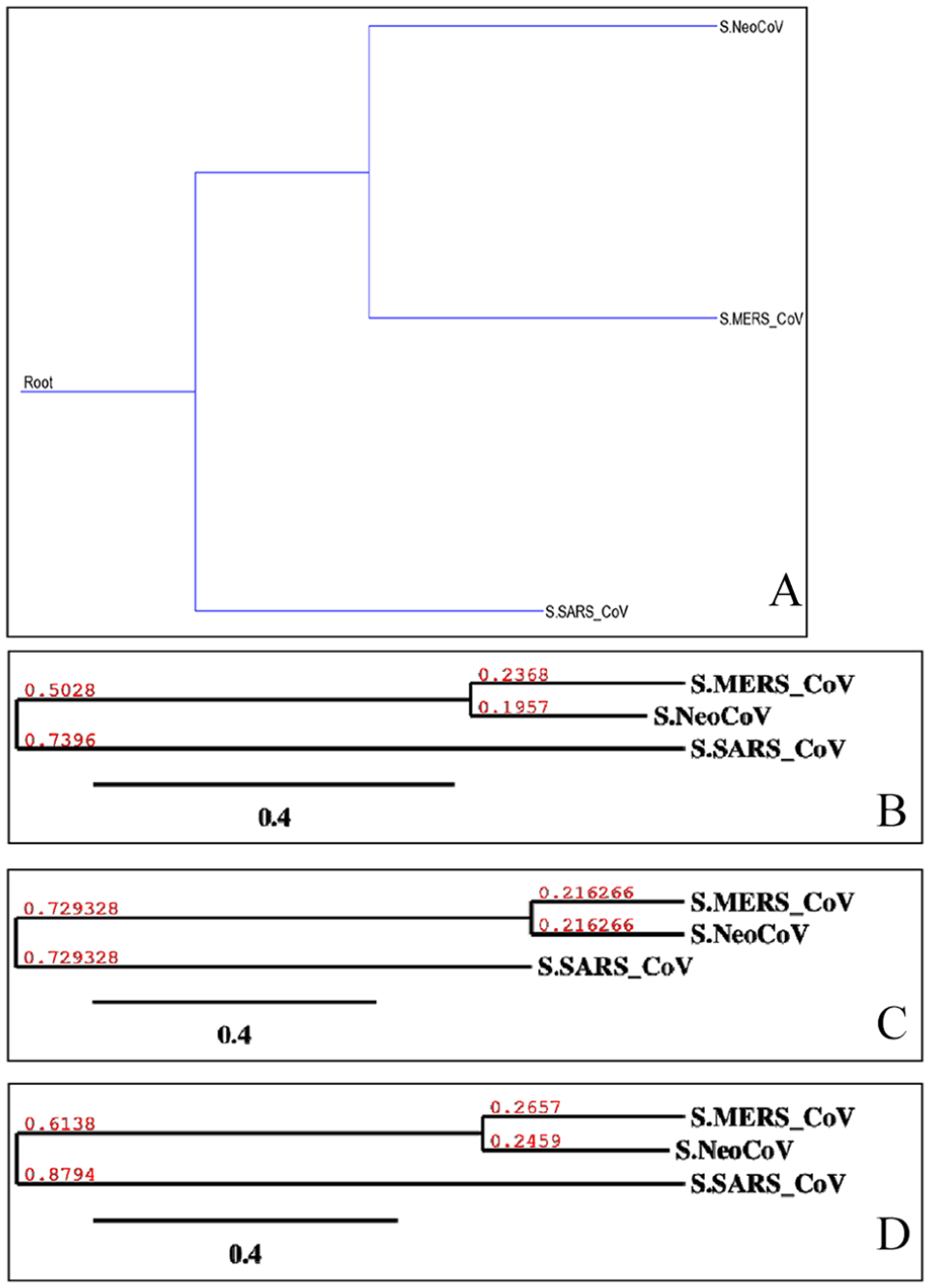
Shows phylogenetic of “S” proteins of target coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV). Trees (A)-(D) were built using different methods, and they are, respectively, Maximum Parsimony, Neighbor-Joining, UPGMA, and Maximum Likelihood. UPGMA indicates Unweighted Pair Group Method with Arithmetic Mean.
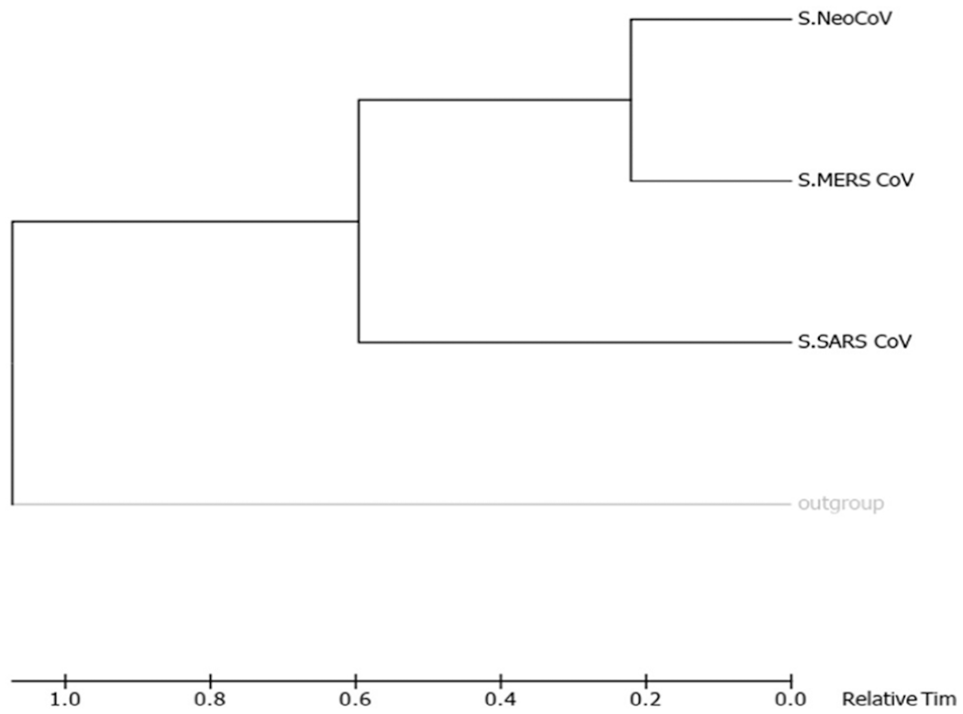
Molecular phylogenetic analysis by Maximum-Likelihood method (timetree). The timetree shown was generated using the RelTime method. 26 Divergence times for all branching points in the topology were calculated using the Maximum-Likelihood method based on the Equal Input model. 27 The estimated log likelihood value of the topology shown is −13 052.98. The tree is drawn to scale, with branch lengths measured in the relative number of substitutions per site. The analysis involved 4 amino acid sequences. There were a total of 1544 positions in the final dataset. Evolutionary analyses were conducted in MEGA7. 25
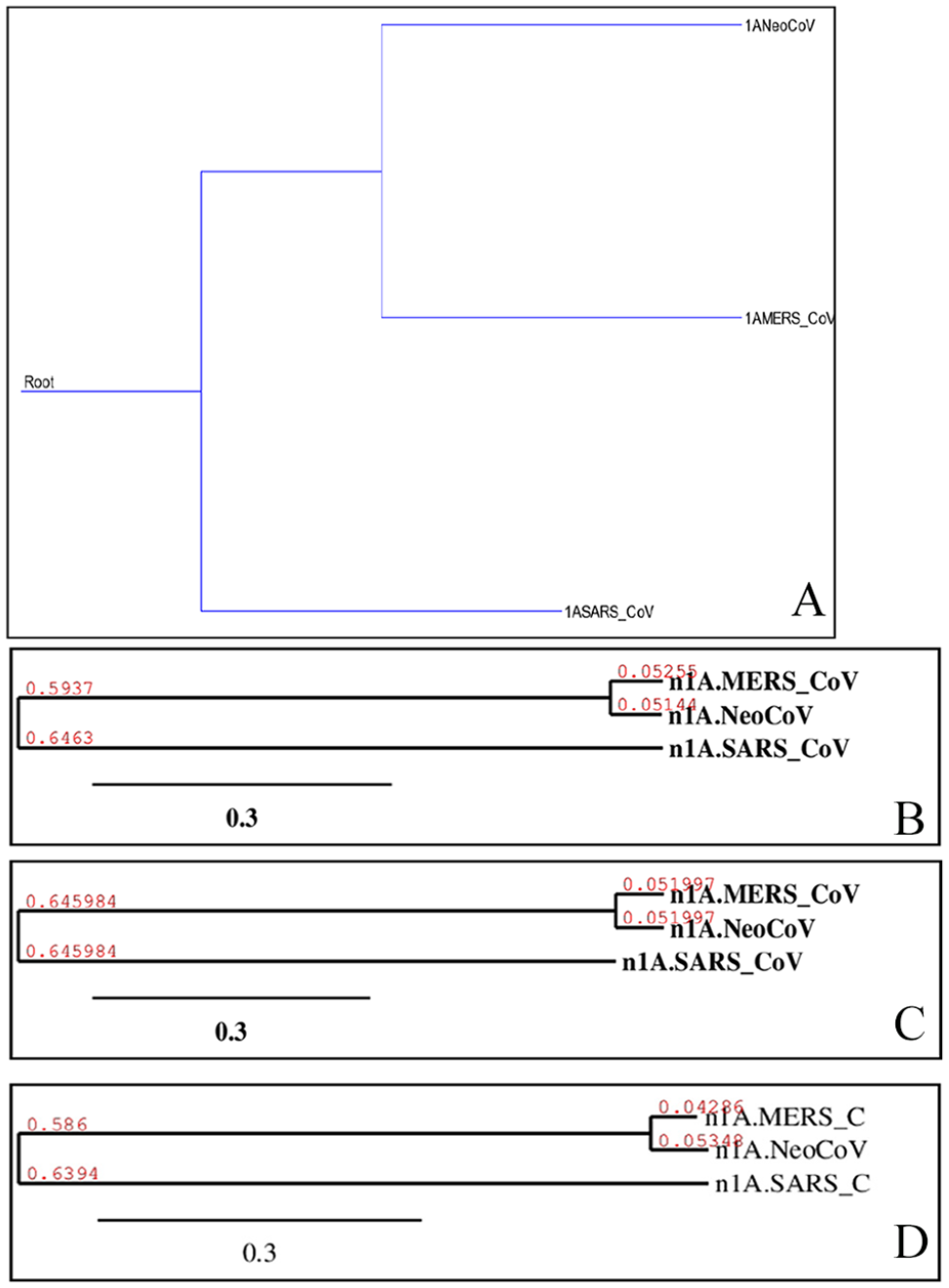
Shows phylogenetic of “1A” proteins of target coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV). Trees (A)-(D) were built using different methods, and they are, respectively, Maximum Parsimony, Neighbor-Joining, UPGMA, and Maximum Likelihood. UPGMA indicates Unweighted Pair Group Method with Arithmetic Mean.
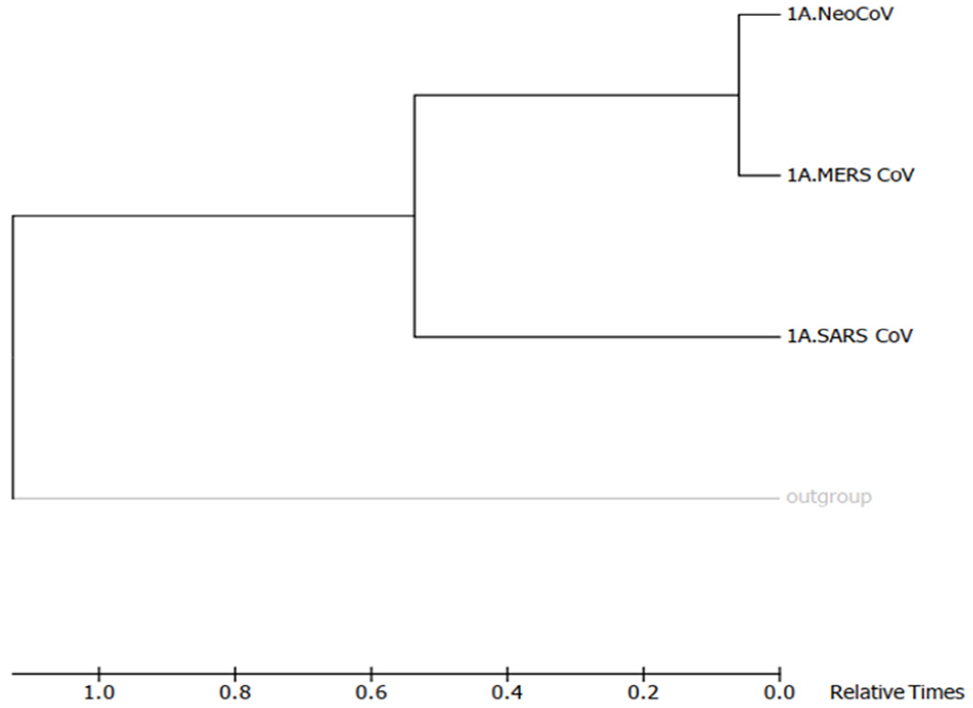
Molecular phylogenetic analysis by Maximum-Likelihood method (timetree). The timetree shown was generated using the RelTime method. 26 Divergence times for all branching points in the topology were calculated using the Maximum-Likelihood method based on the Equal Input model. 27 The estimated log likelihood value of the topology shown is −38 685.85. The tree is drawn to scale, with branch lengths measured in the relative number of substitutions per site. The analysis involved 4 amino acid sequences. There were a total of 4988 positions in the final dataset. Evolutionary analyses were conducted in MEGA7. 25
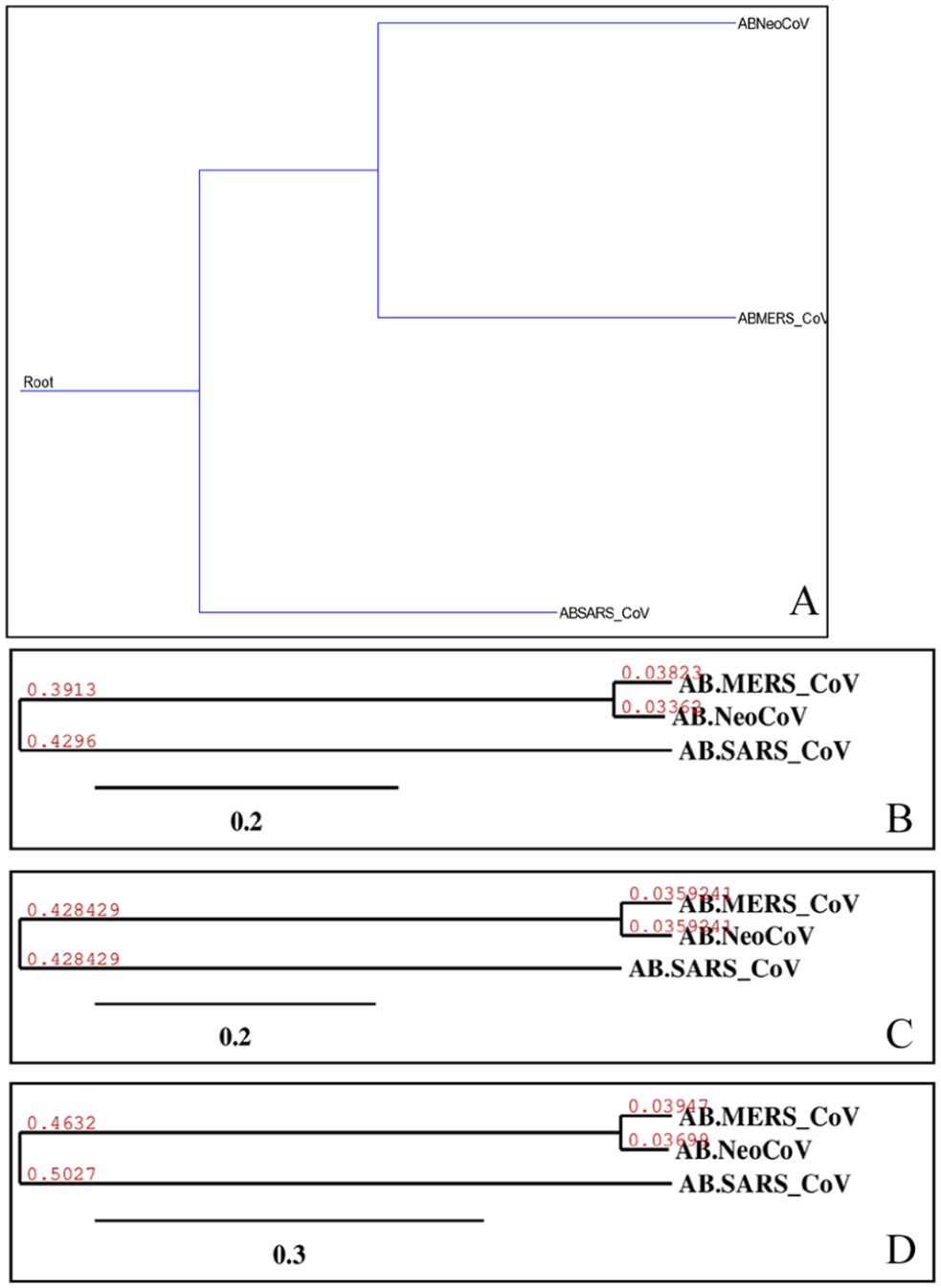
Phylogenetic of “1AB” proteins of target coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV). Trees (A)-(D) were built using different methods, and they are, respectively, Maximum Parsimony, Neighbor-Joining, UPGMA, and Maximum Likelihood. UPGMA indicates Unweighted Pair Group Method with Arithmetic Mean.
Molecular phylogenetic analysis by Maximum-Likelihood method (timetree). The timetree shown was generated using the RelTime method. 26 Divergence times for all branching points in the topology were calculated using the Maximum-Likelihood method based on the Equal Input model. 27 The estimated log likelihood value of the topology shown is −59 576.47. The tree is drawn to scale, with branch lengths measured in the relative number of substitutions per site. The analysis involved 4 amino acid sequences. There were a total of 8041 positions in the final dataset. Evolutionary analyses were conducted in MEGA7. 25
Calculate physical and chemical parameters
To determine physical and chemical properties of the protein sequence, ProtParam tool http://web.expasy.org/protparam/) has been used (which gives the computation of various physical and chemical parameters for a given protein stored in Swiss-Prot or TrEMBL databases or for a user entered sequence. The computed parameters are the molecular weight, theoretical pI, amino acid composition, atomic composition, extinction coefficient, estimated half-life, instability index, aliphatic index, and grand average of hydropathicity (GRAVY) as presented in Tables 3 to 14. 28
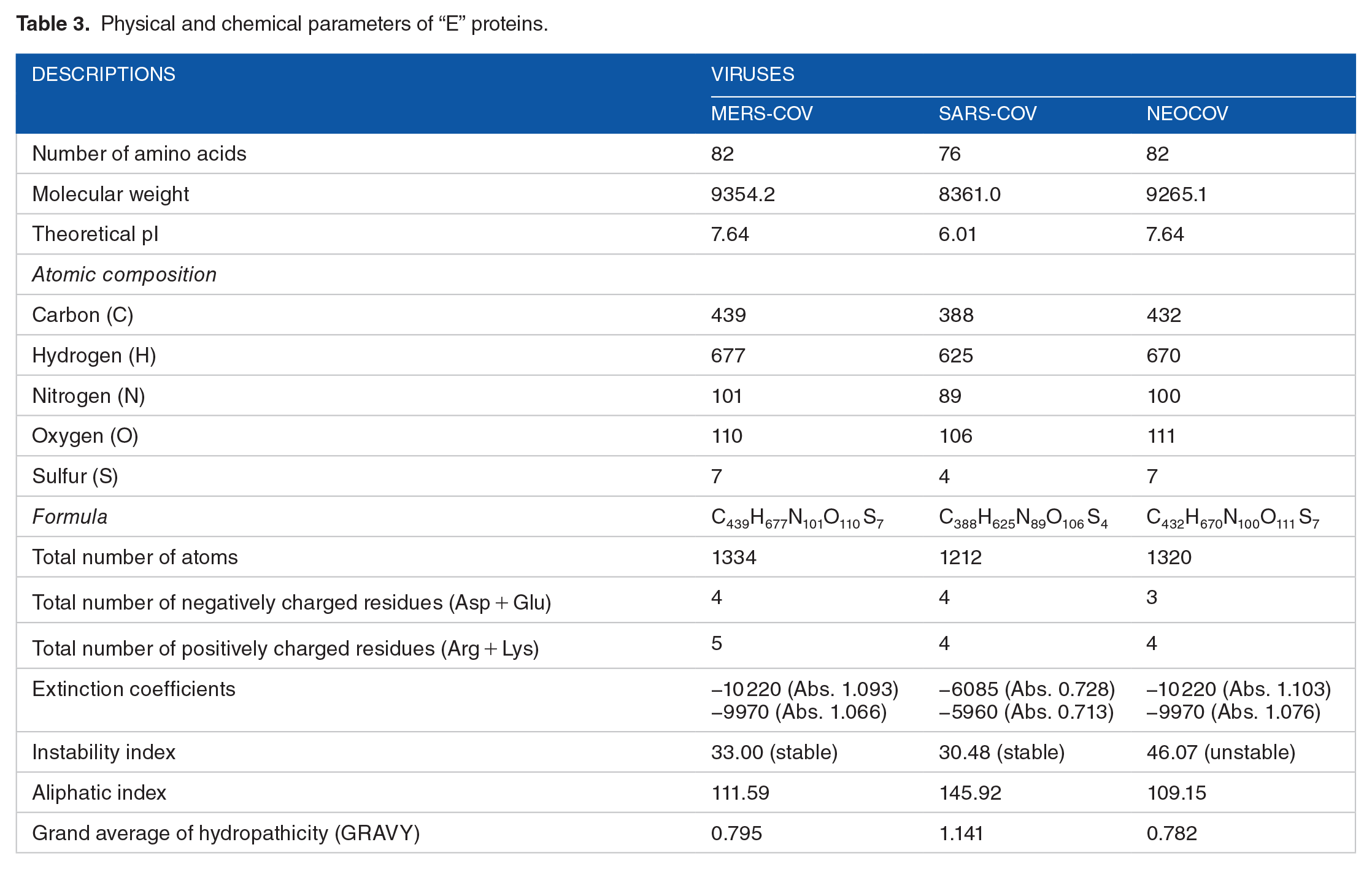
Physical and chemical parameters of “E” proteins.
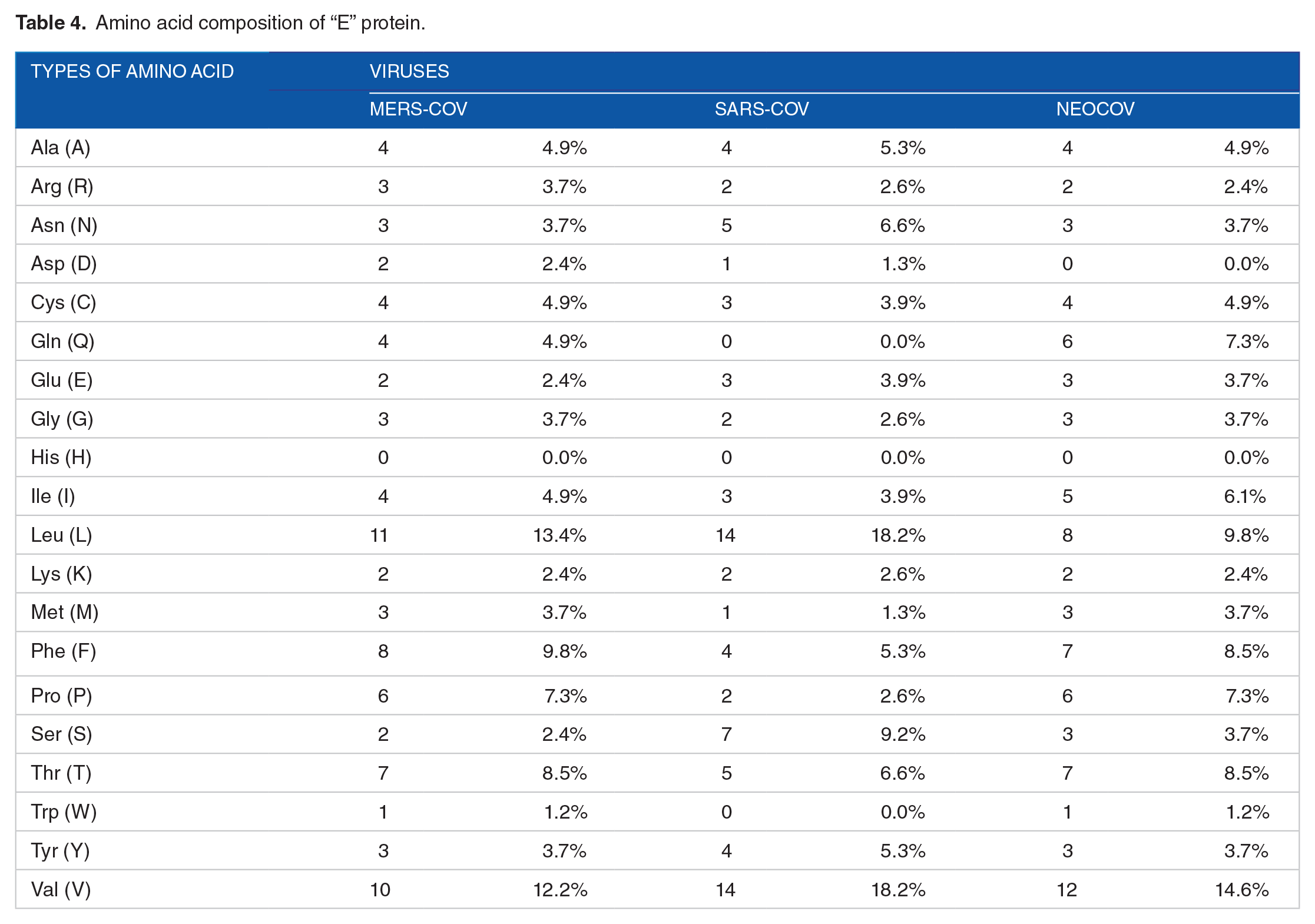
Amino acid composition of “E” protein.
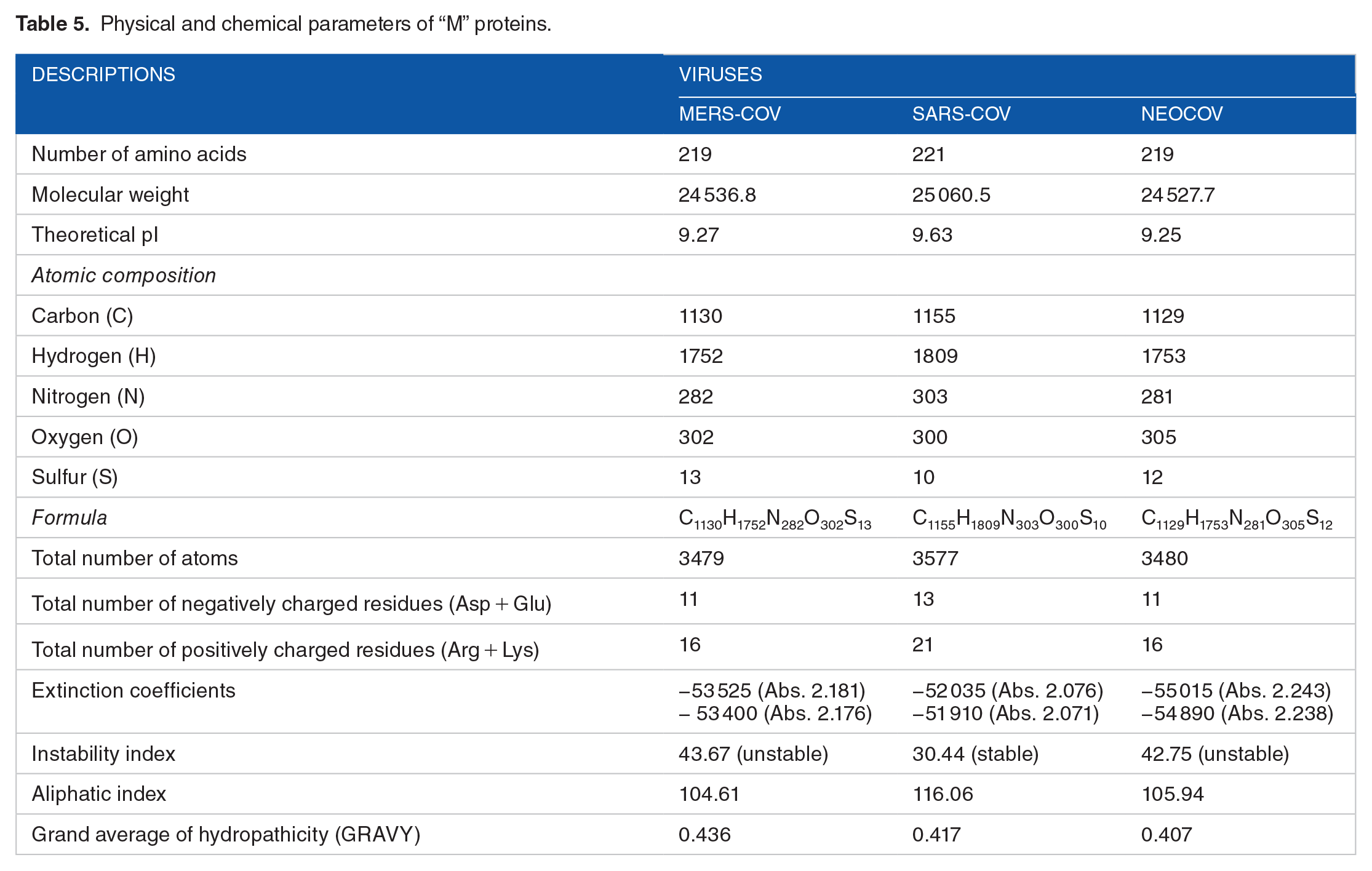
Physical and chemical parameters of “M” proteins.
Amino acid composition of “M” protein.
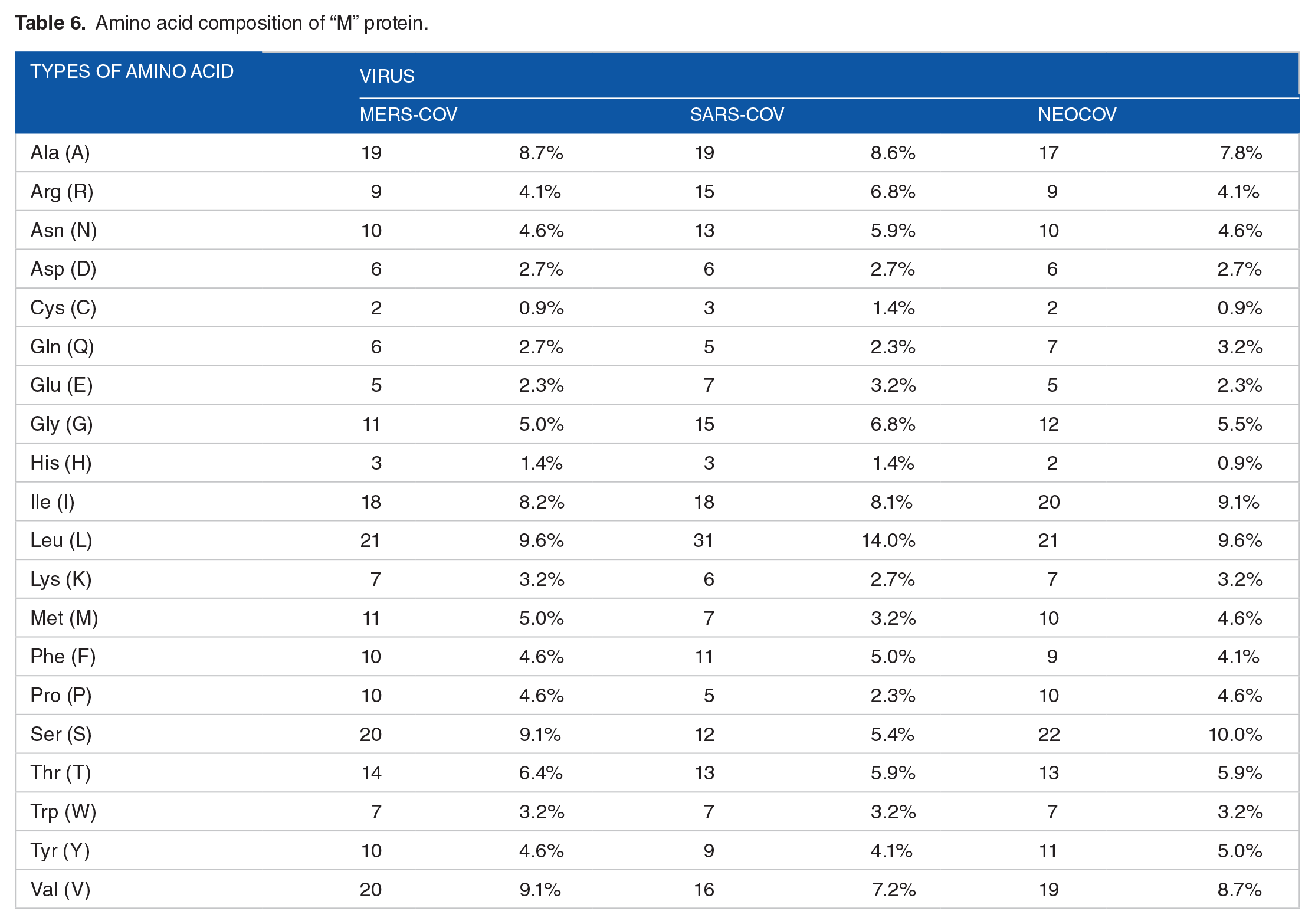
Physical and chemical parameters of “N” proteins.
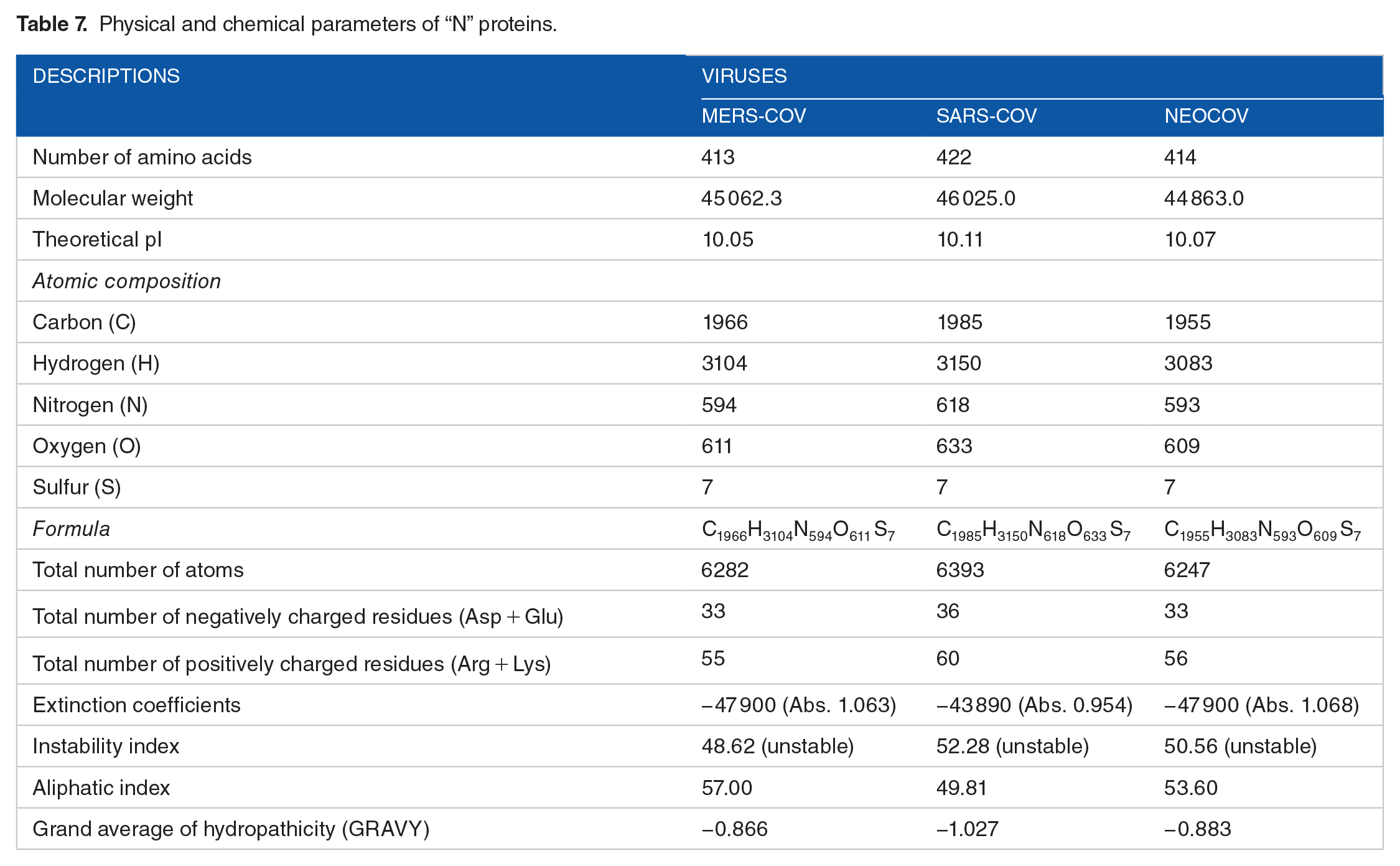
The amino acid composition of “N” protein.
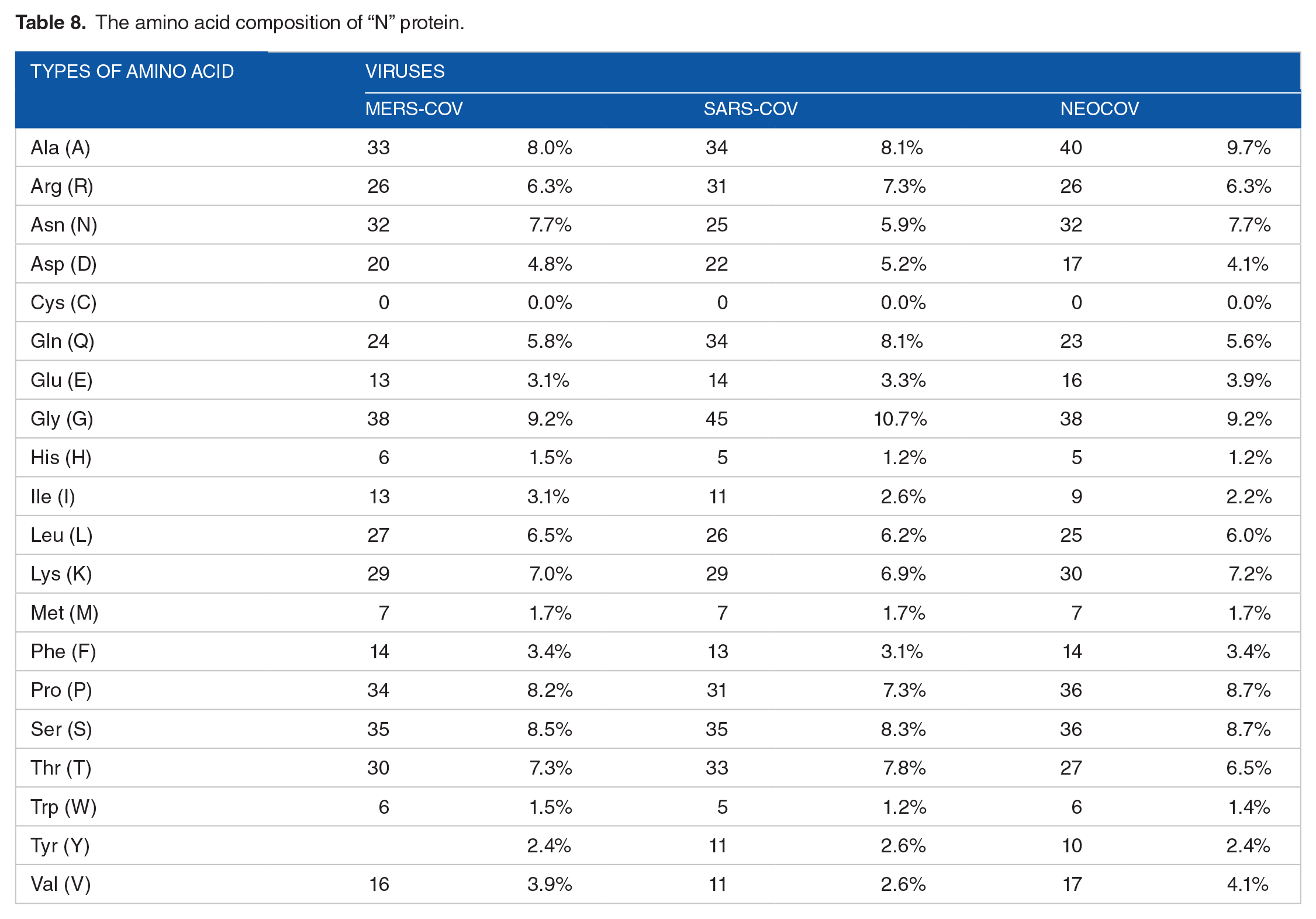
Physical and chemical parameters of “S” proteins.
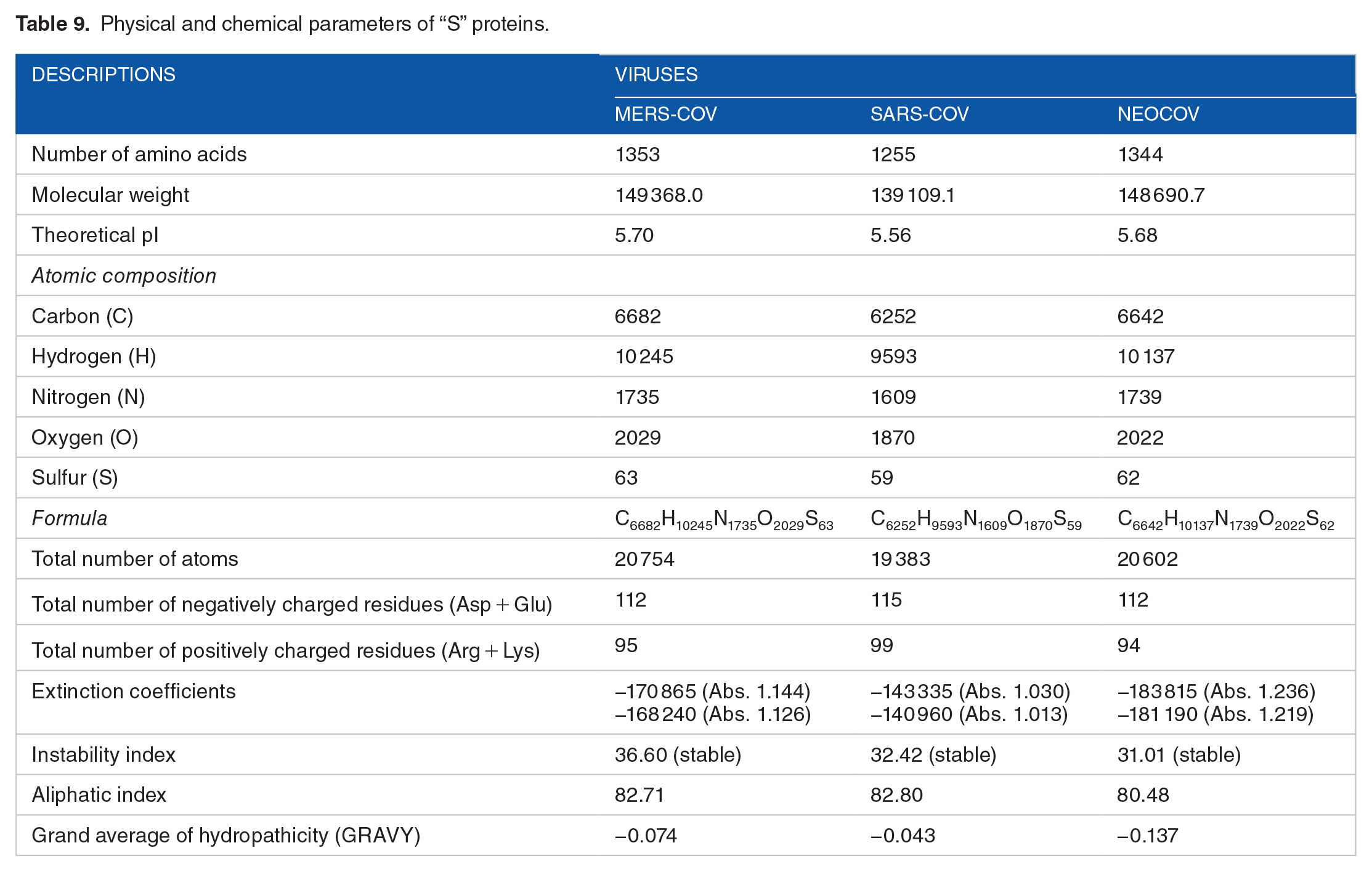
Amino acid composition of “S” protein.
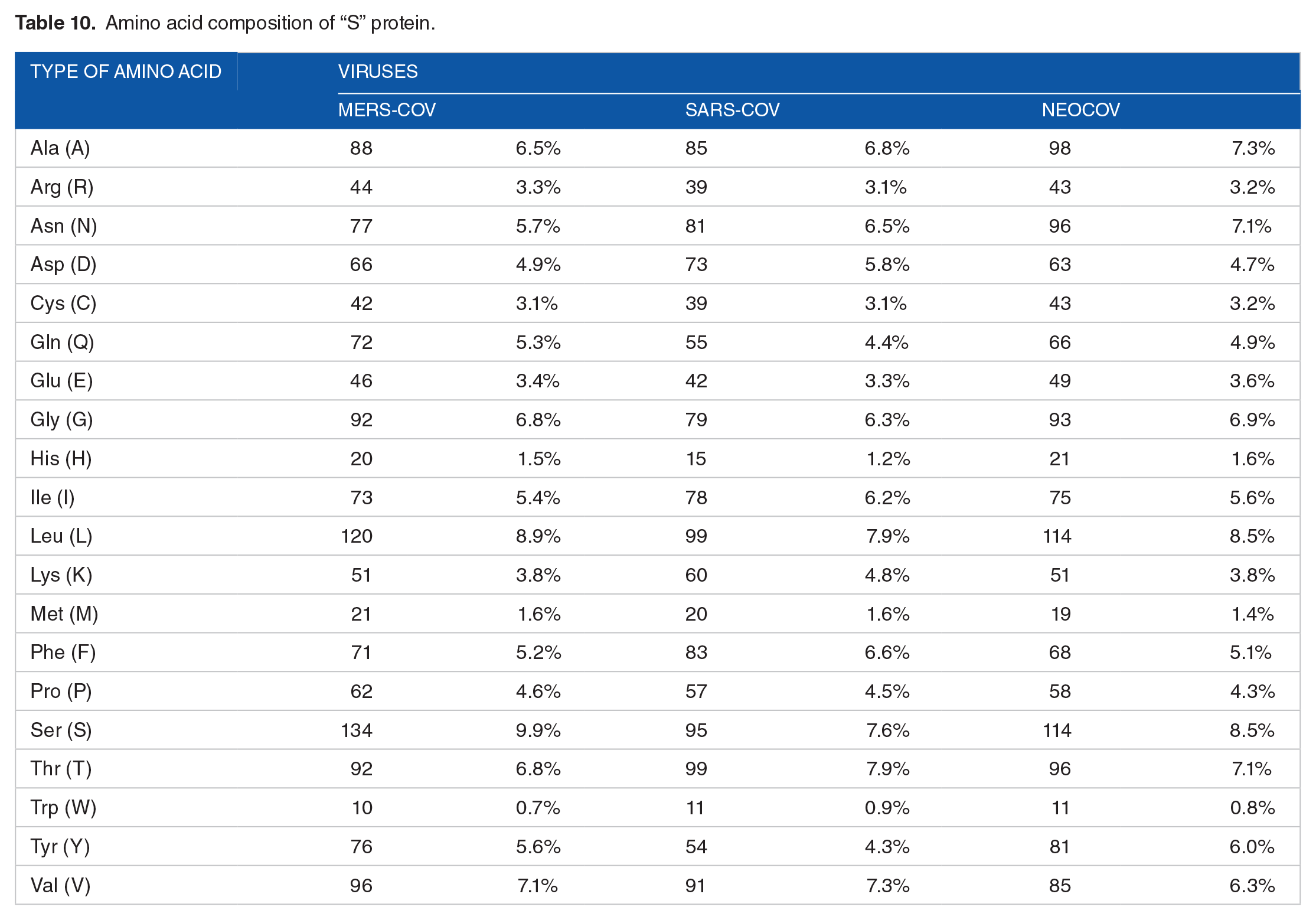
Physical and chemical parameters of “ORF1a” proteins.
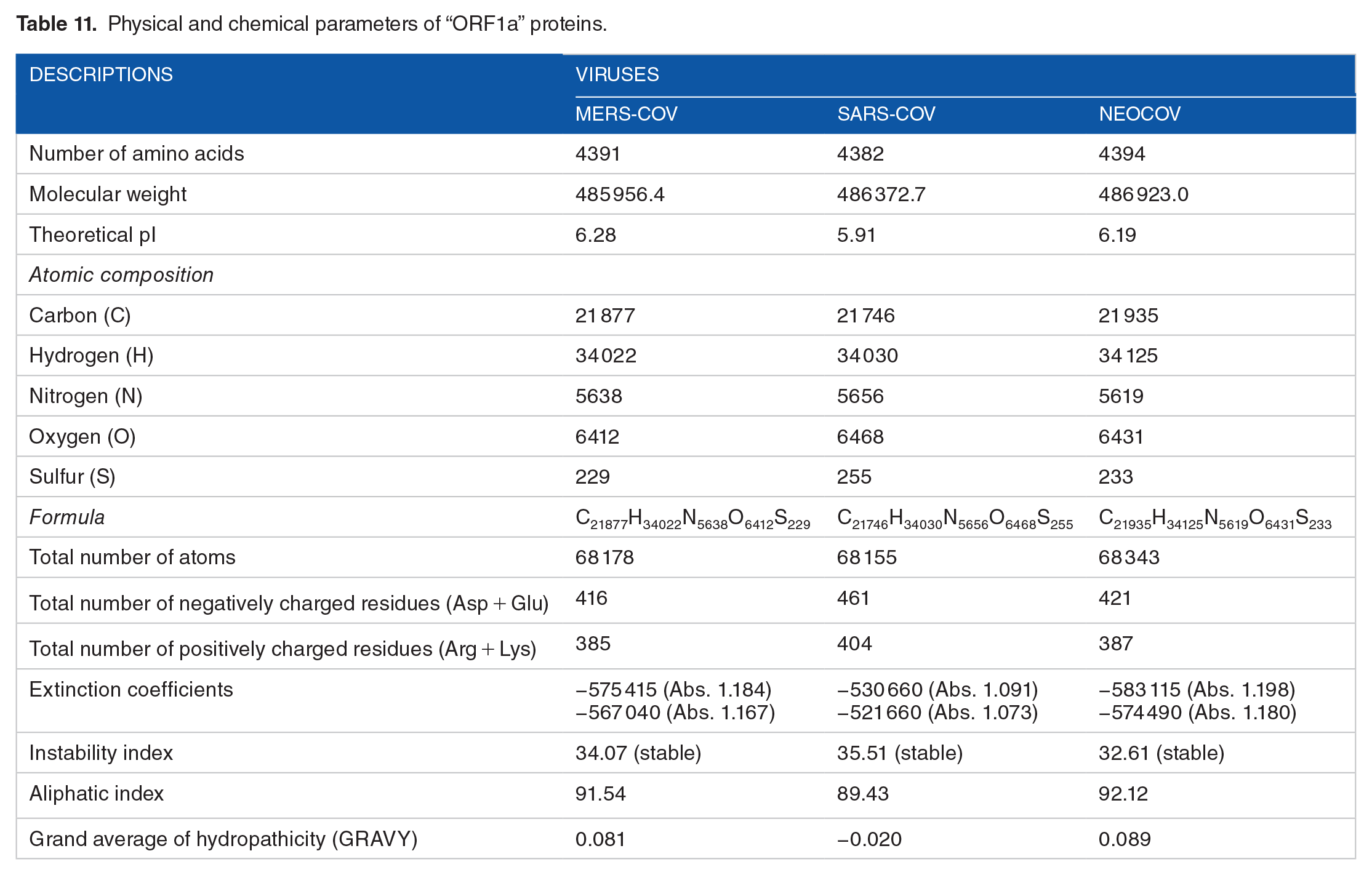
Amino acid composition of “ORF1a” proteins.
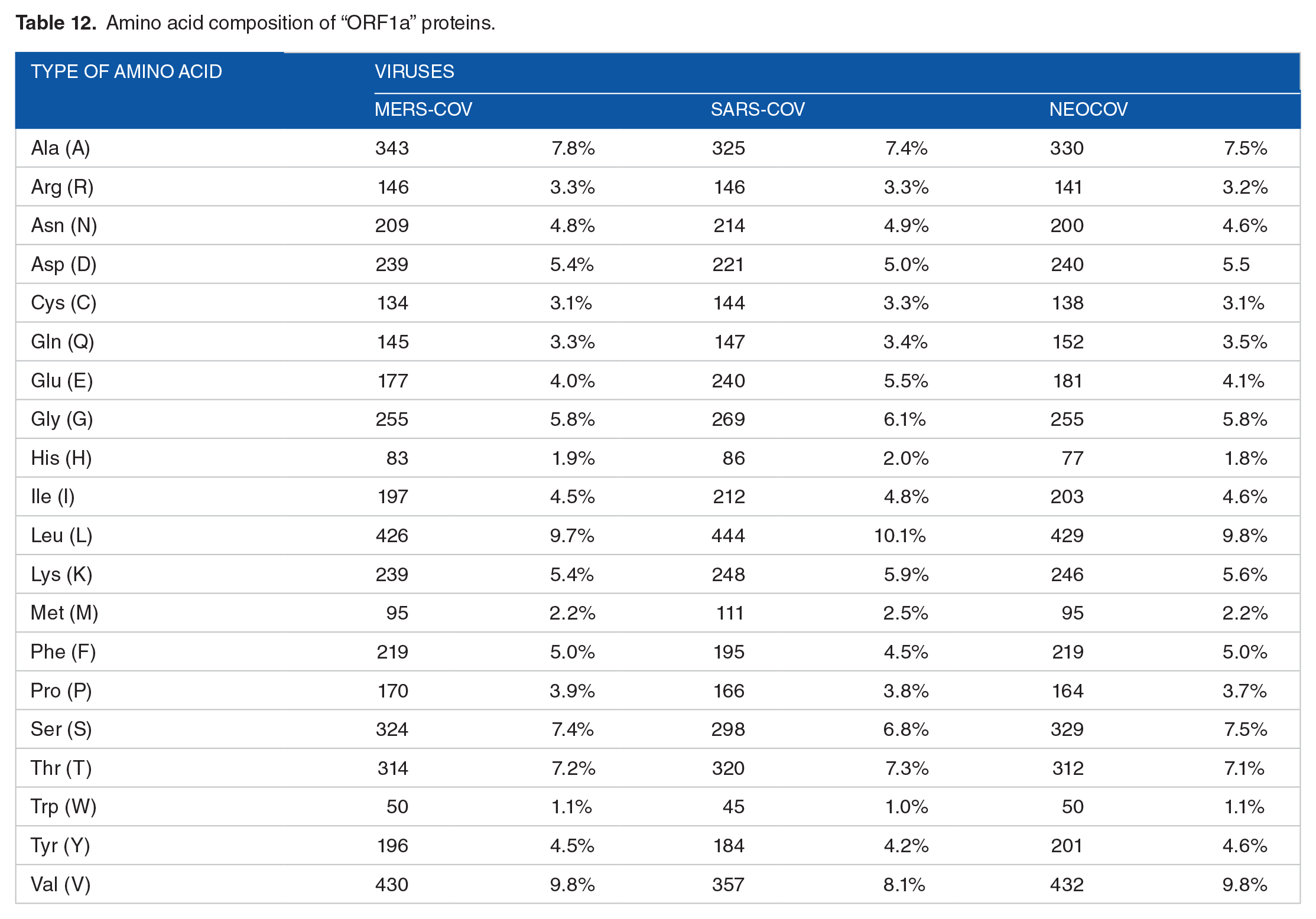
Physical and chemical parameters of “ORF1ab” proteins.
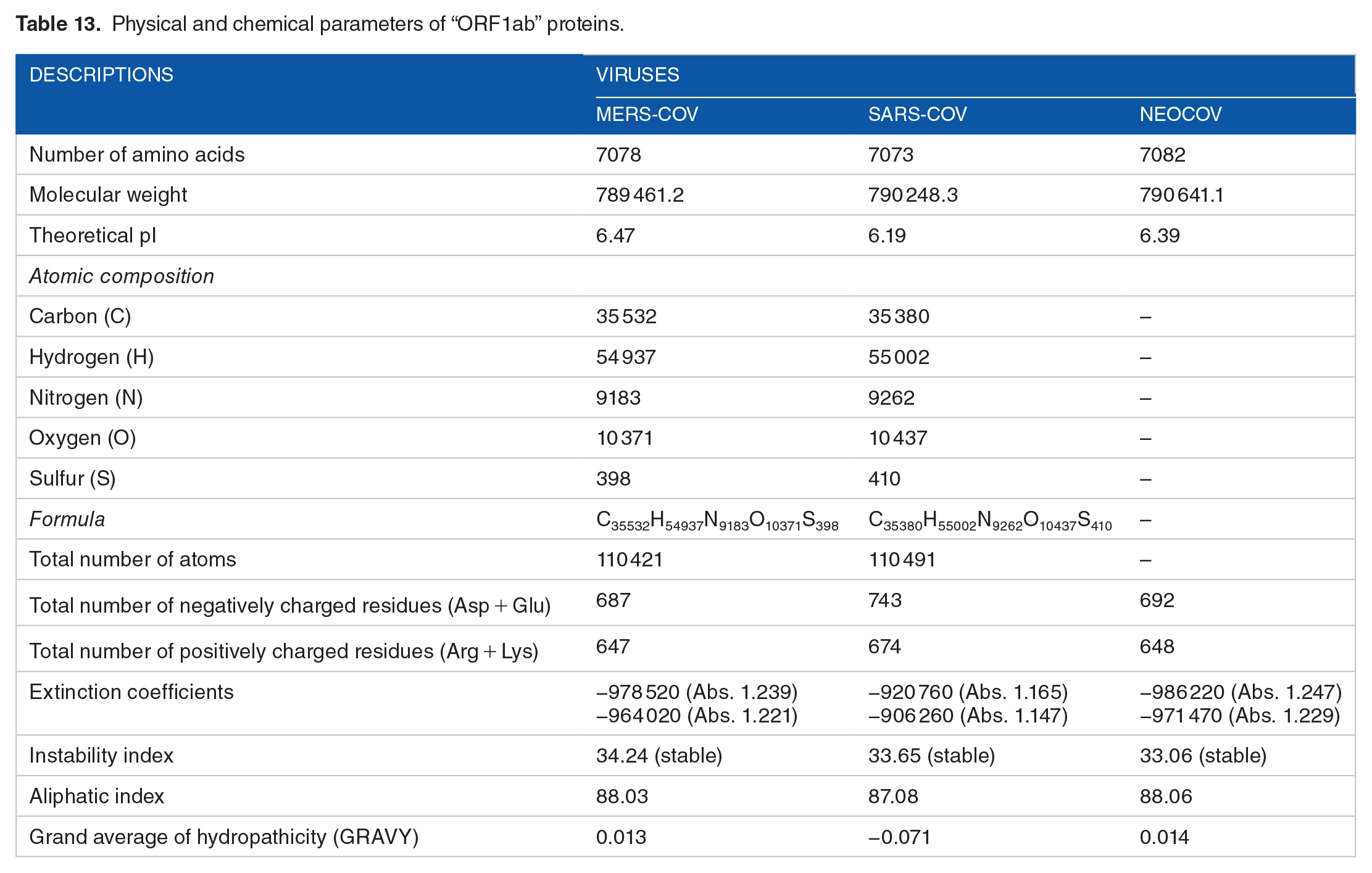
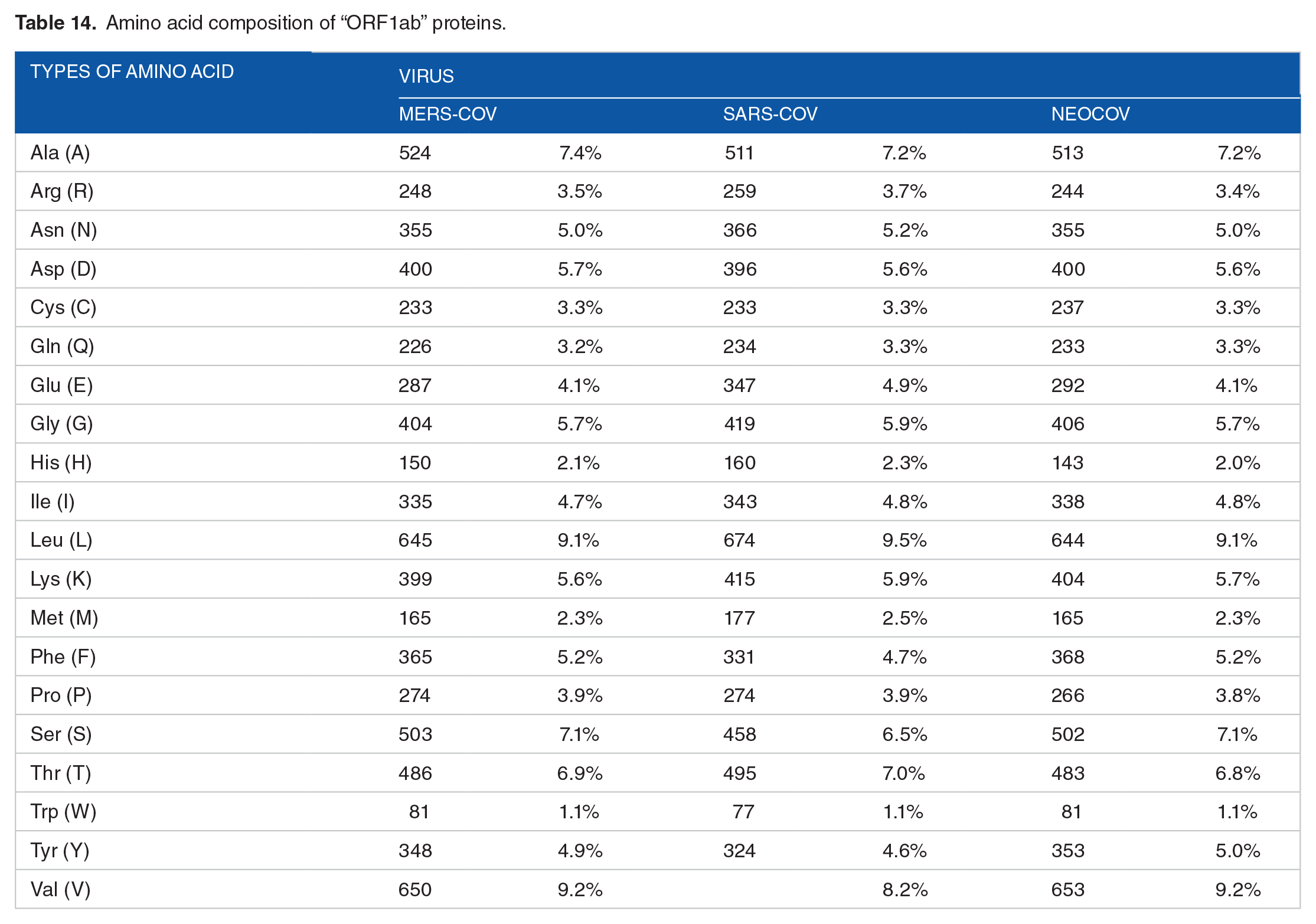
Amino acid composition of “ORF1ab” proteins.
Pairwise alignment for protein sequences (primary structure)
Same type of protein sequences of the CoV species of interest was compared using Basic Local Alignment Search Tool (BLAST; https://blast.ncbi.nlm.nih.gov/Blast.cgi). Pairwise alignment was done to determine the matched regions and the number of identical/similar amino acids as described in Table 15.
Pairwise alignment for protein sequences (primary structure) of coronavirus species.
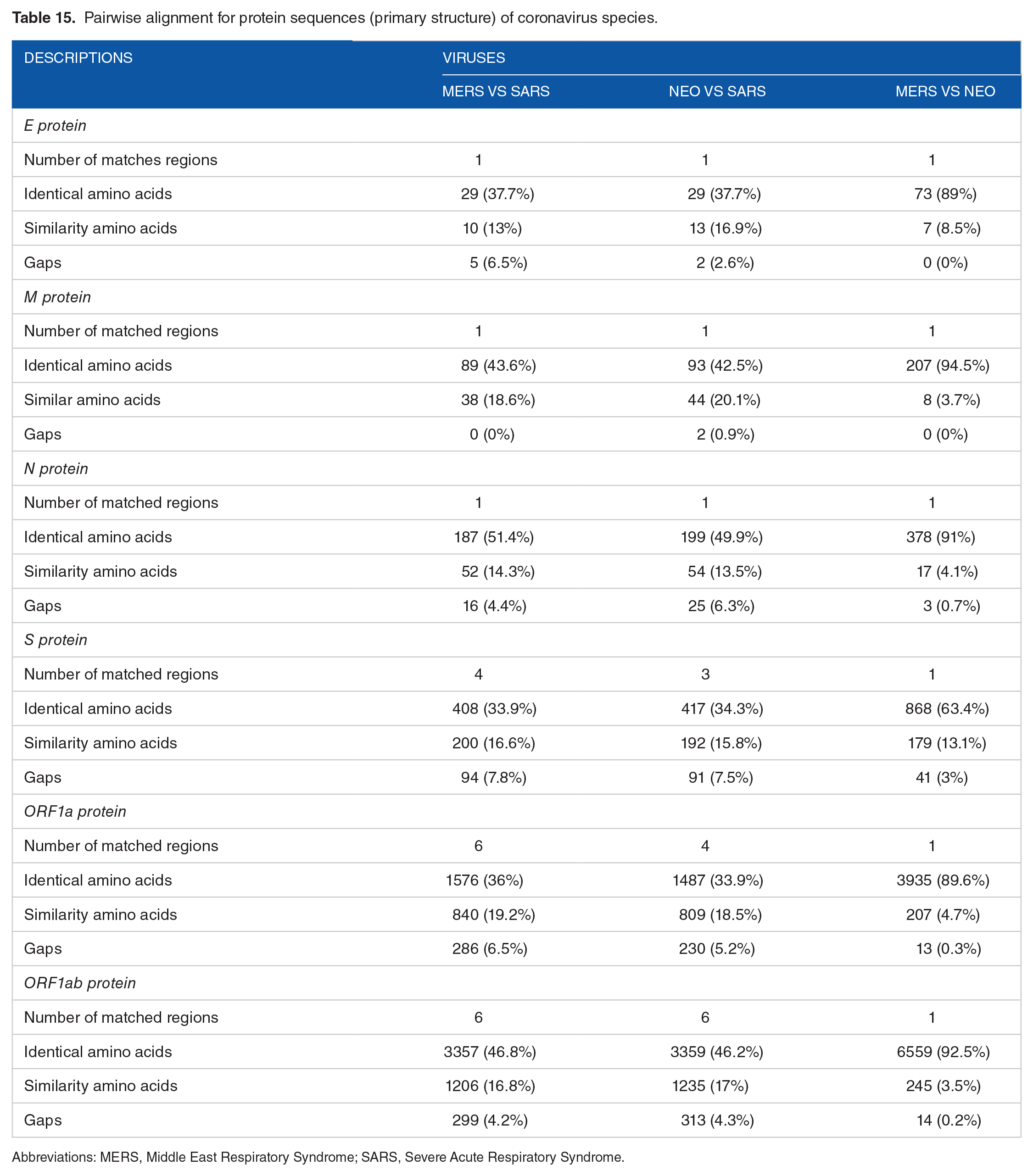
Abbreviations: MERS, Middle East Respiratory Syndrome; SARS, Severe Acute Respiratory Syndrome.
Proteins secondary structure prediction
For the purpose of converting the primary protein structure to secondary protein structure, GOR IV Tool has been used (version 4.0; https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=npsa_gor4.html), which was based on the information theory which gives the 2 outputs. The first output comprised the sequence and the predicted secondary structure in rows, H = helix, E = extended or beta strand, and C = coil. The second presents probability values for each secondary structure at each amino acid position. The program gives the predicted secondary structure with the highest probability compatible with a predicted helix segment of at least 4 residues and a predicted extended segment of at least 2 residues 29 as shown in Figures 16 to 21.
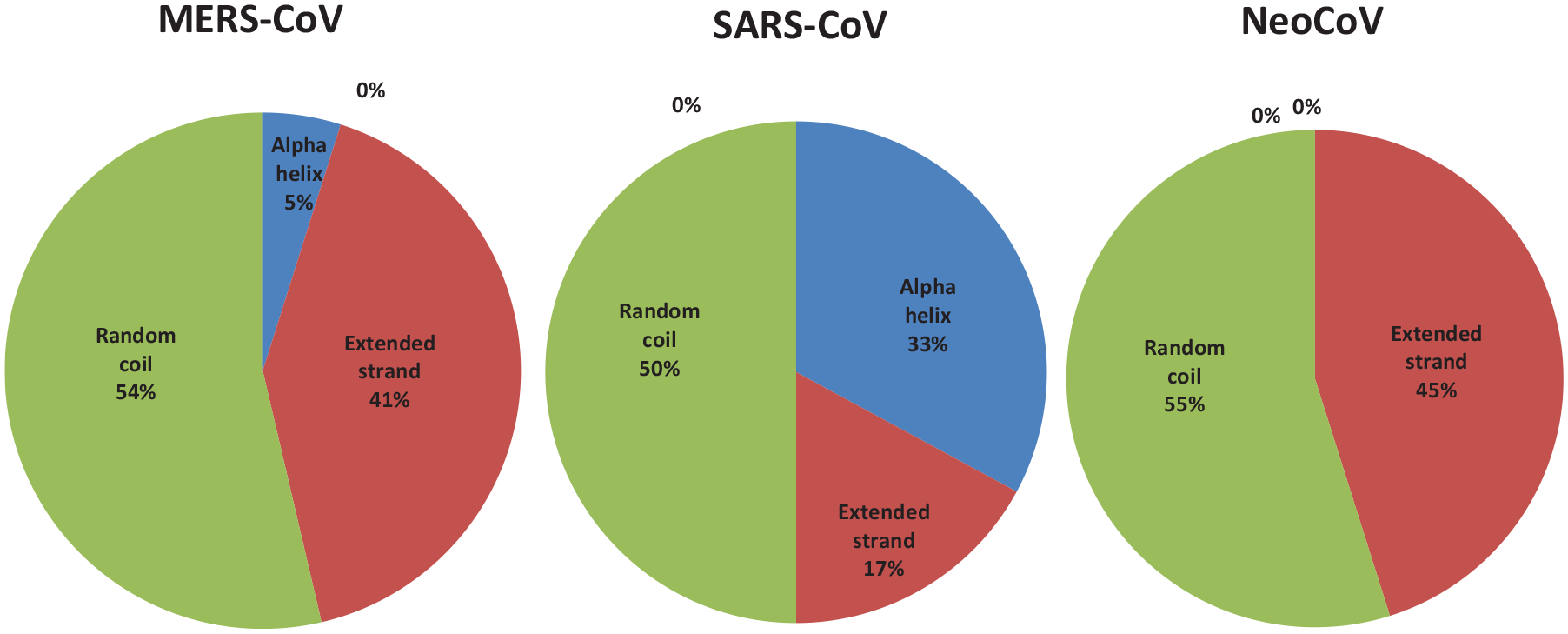
Percent of secondary structure component of E proteins. Blue color for alpha helix, brown for extended strand, and green color for the random coil.
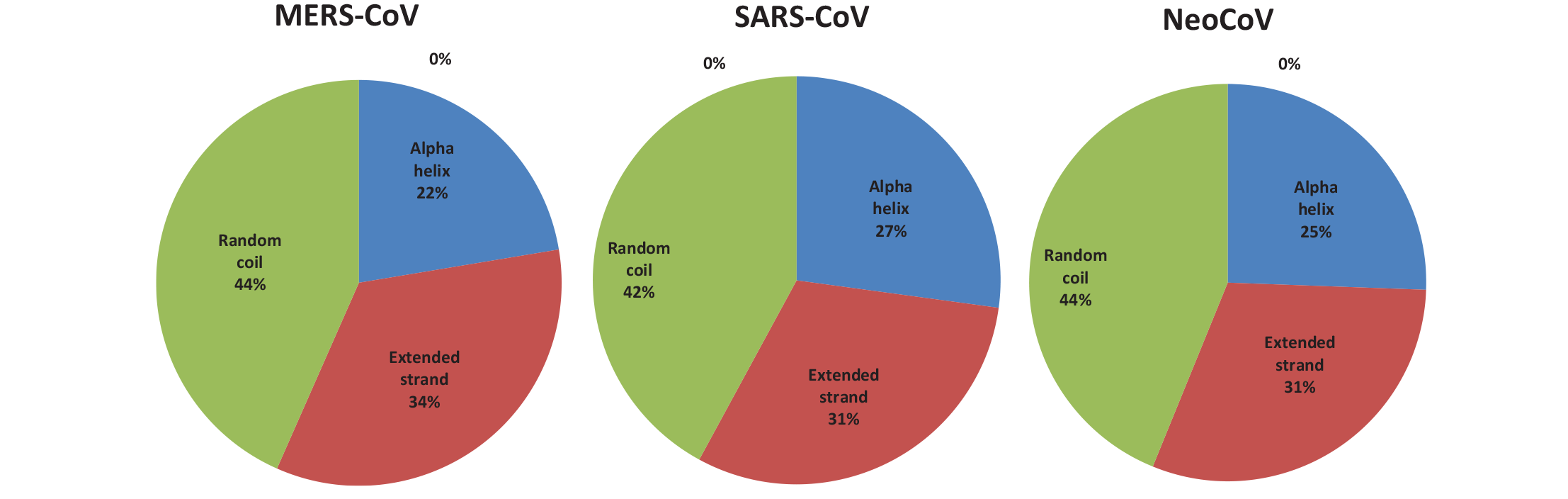
Percent of secondary structure component of M proteins. Blue color for alpha helix, brown for extended strand, and green color for the random coil.
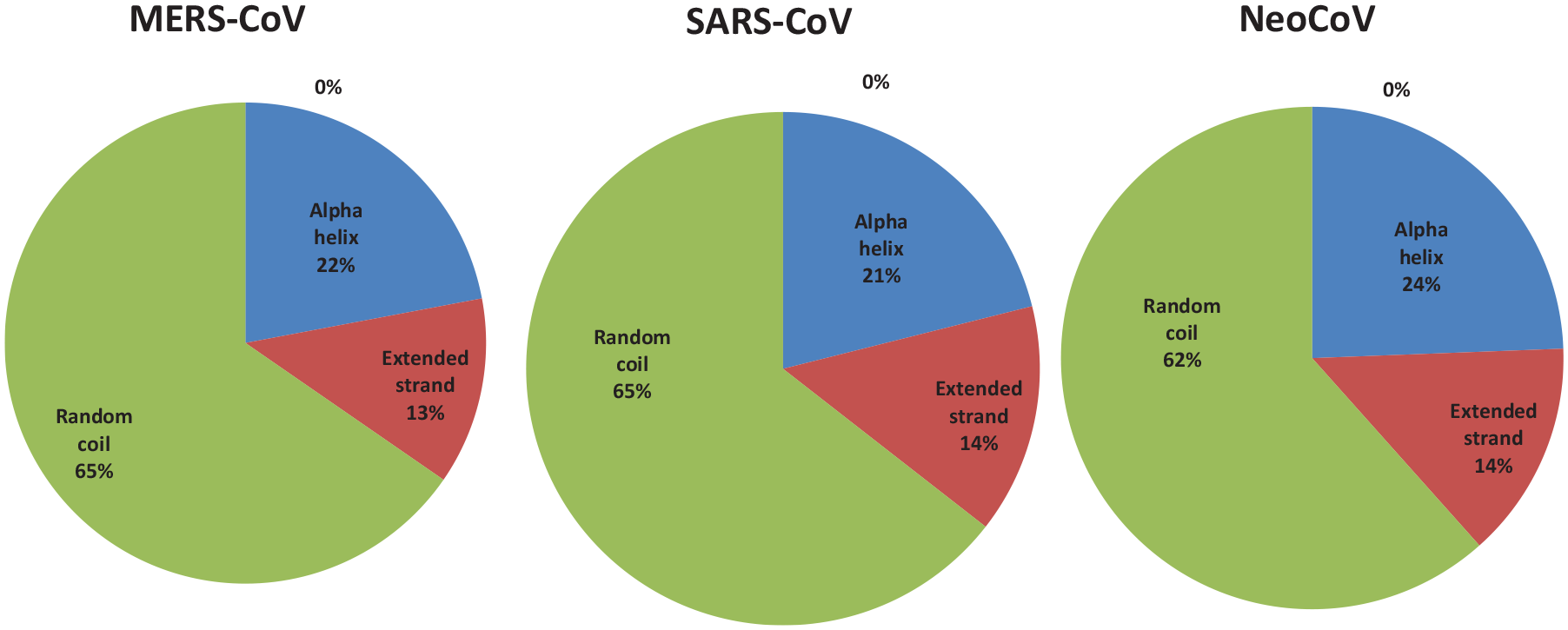
Percent of secondary structure component of N proteins. Blue color for alpha helix, brown for extended strand, and green color for the random coil.
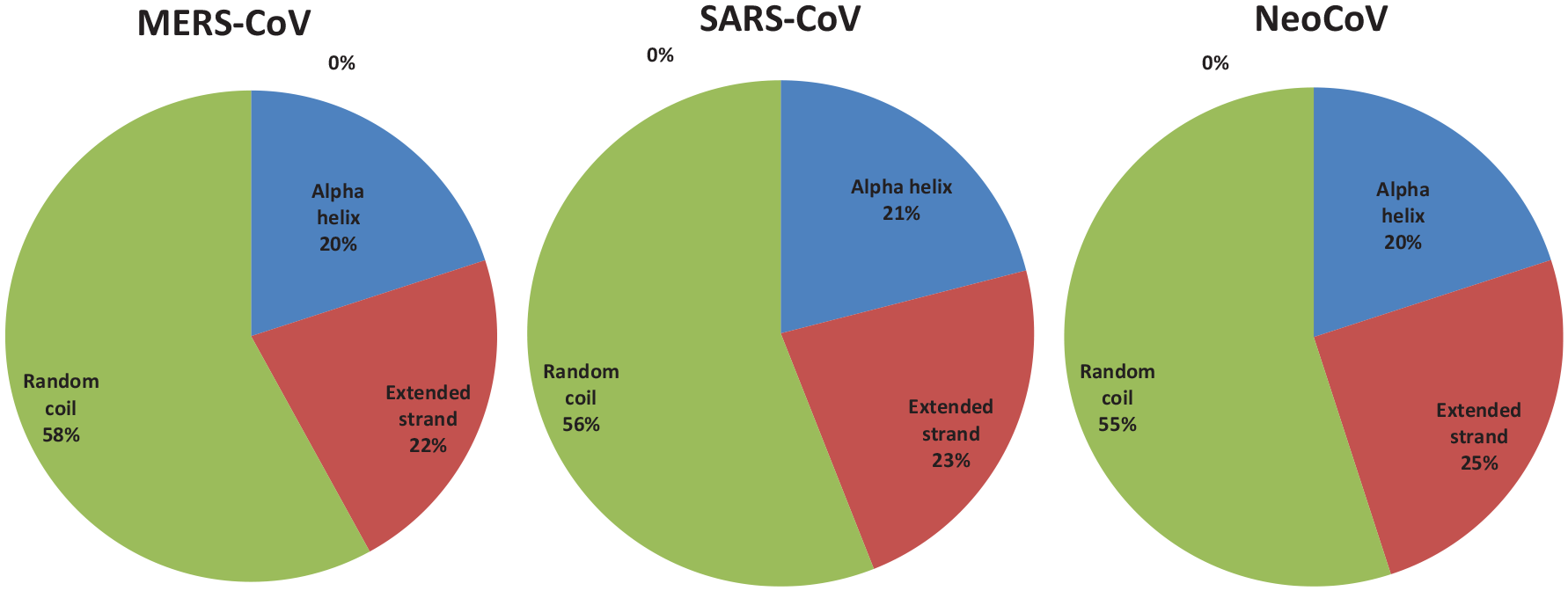
Percent of secondary structure component of S proteins. Blue color for alpha helix, brown for extended strand, and green color for the random coil.
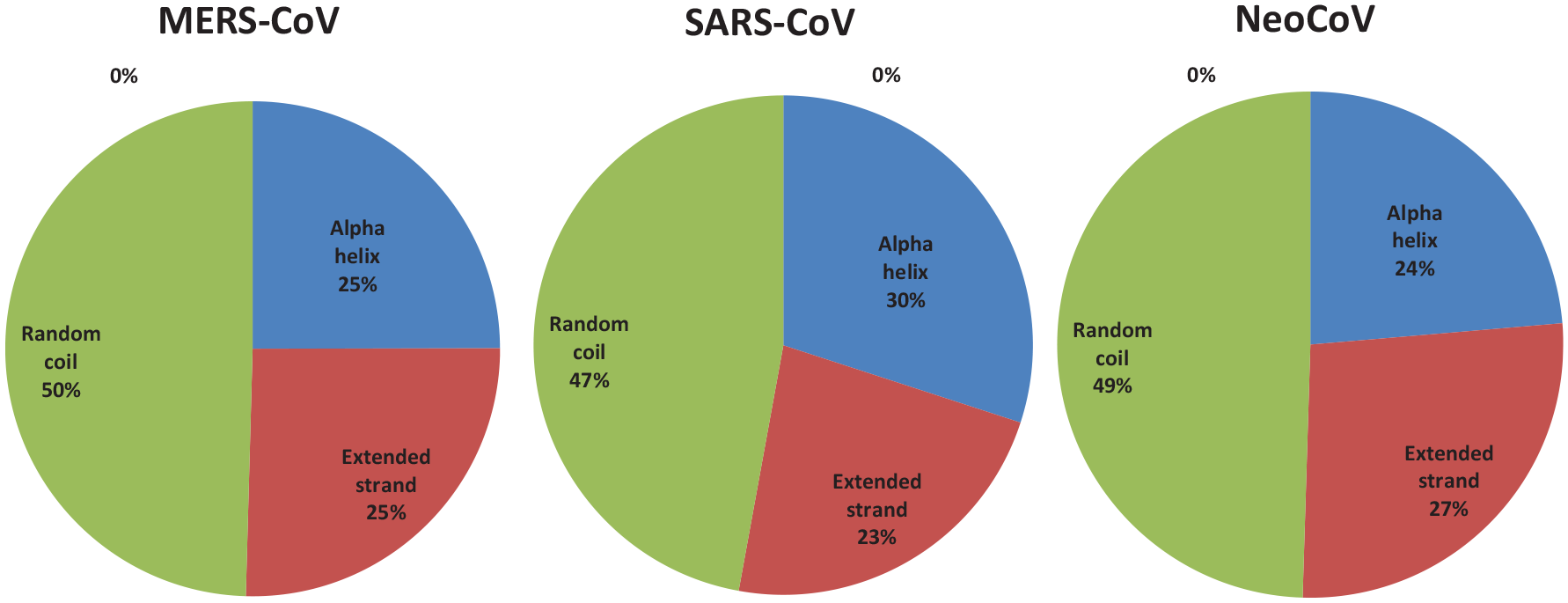
Percent of secondary structure component of ORF1a proteins. Blue color for alpha helix, brown for extended strand, and green color for the random coil.
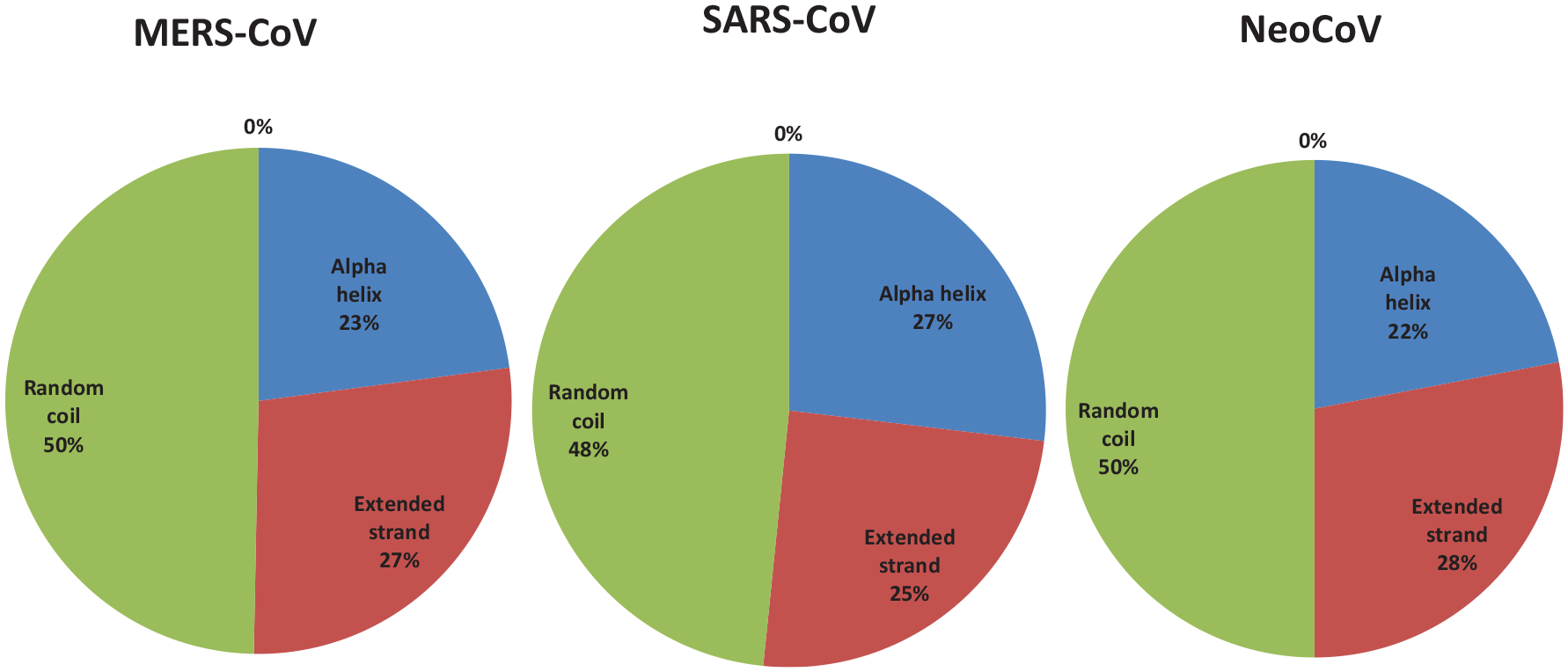
Percent of secondary structure component of ORF1ab proteins. Blue color for alpha helix, brown for extended strand, and green color for the random coil.
Proteins homology modeling prediction
The three-dimensional (3D) structure prediction of target structural proteins (E, M, N, and S) was obtained by using CPH models and RaptorX servers. In CPH server, the template recognition is based on profile-profile alignment guided by secondary structure and exposure predictions (http://www.cbs.dtu.dk/services/CPHmodels/).
30
Proteins that do not have close 3D structures were subjected to RaptorX server, which was developed by Xu group. It is excelling at predicting 3D structures for protein sequences without close homologs in the Protein Data Bank (PDB). Additionally, it predicts secondary and tertiary structures, contacts, solvent accessibility, disordered regions, and binding sites with many confidence scores to indicate the quality of the predicted 3D model including
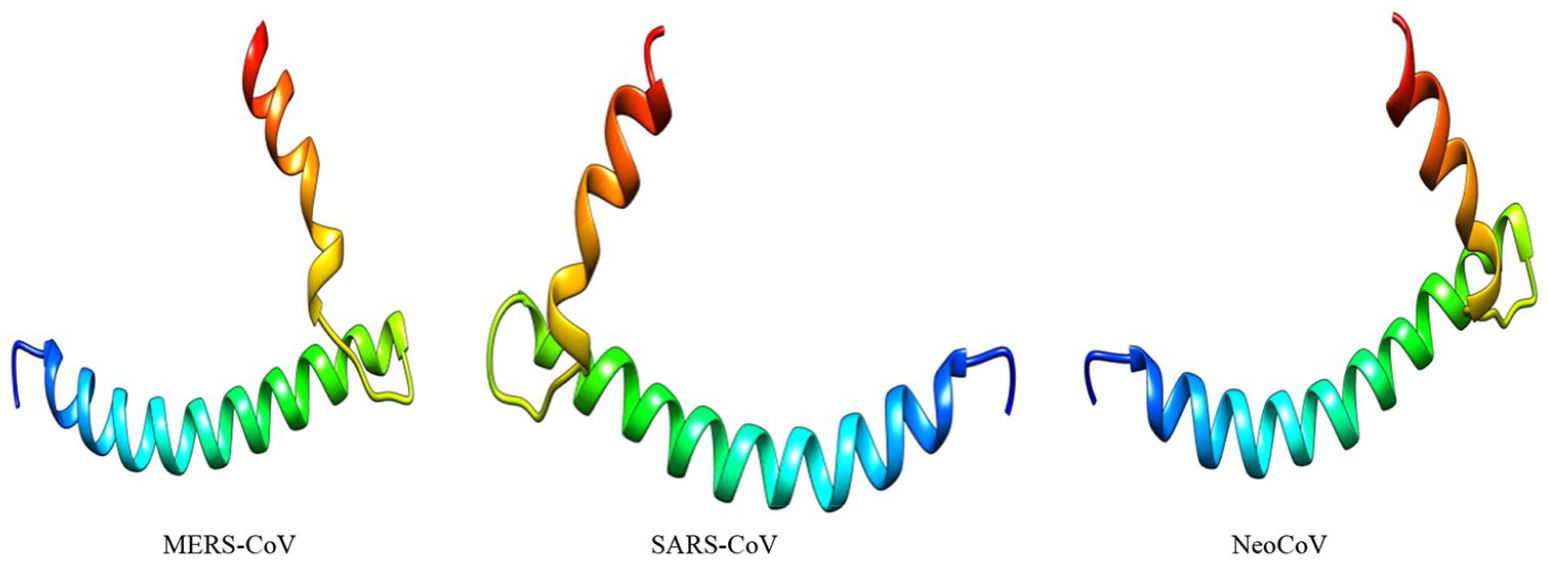
Three-dimensional (3D) structures of “E” proteins of 3 coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV).
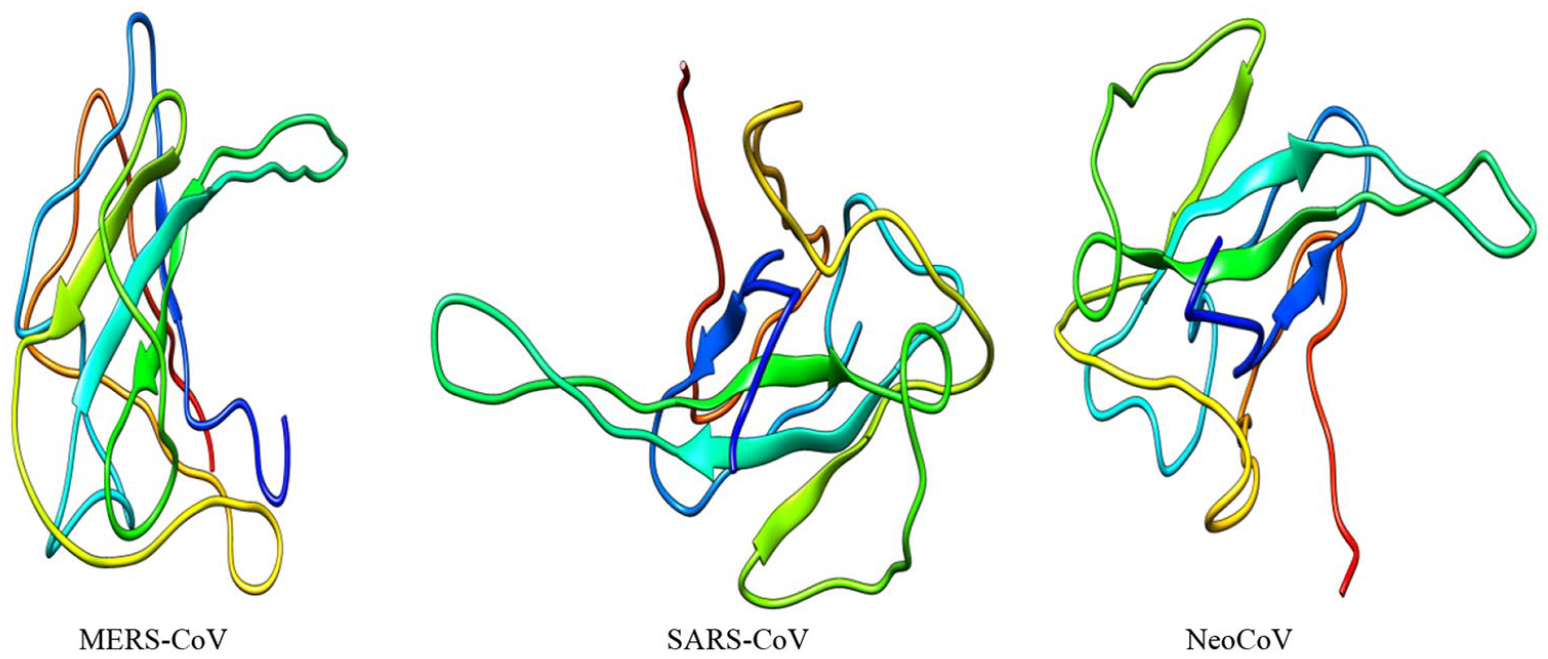
Three-dimensional (3D) structures of “M” proteins of 3 coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV).
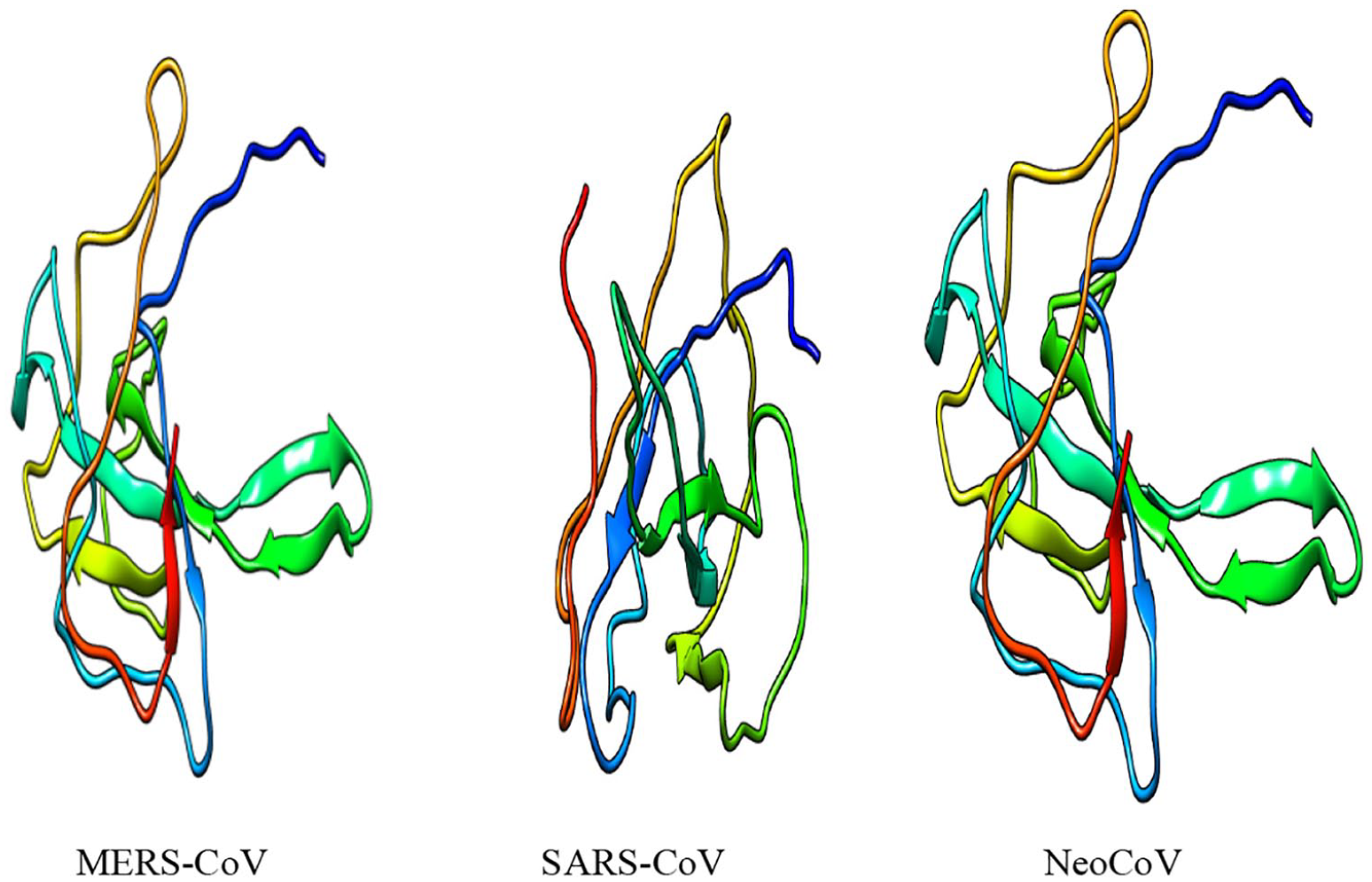
Three-dimensional (3D) structures of “N” proteins of 3 coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV).
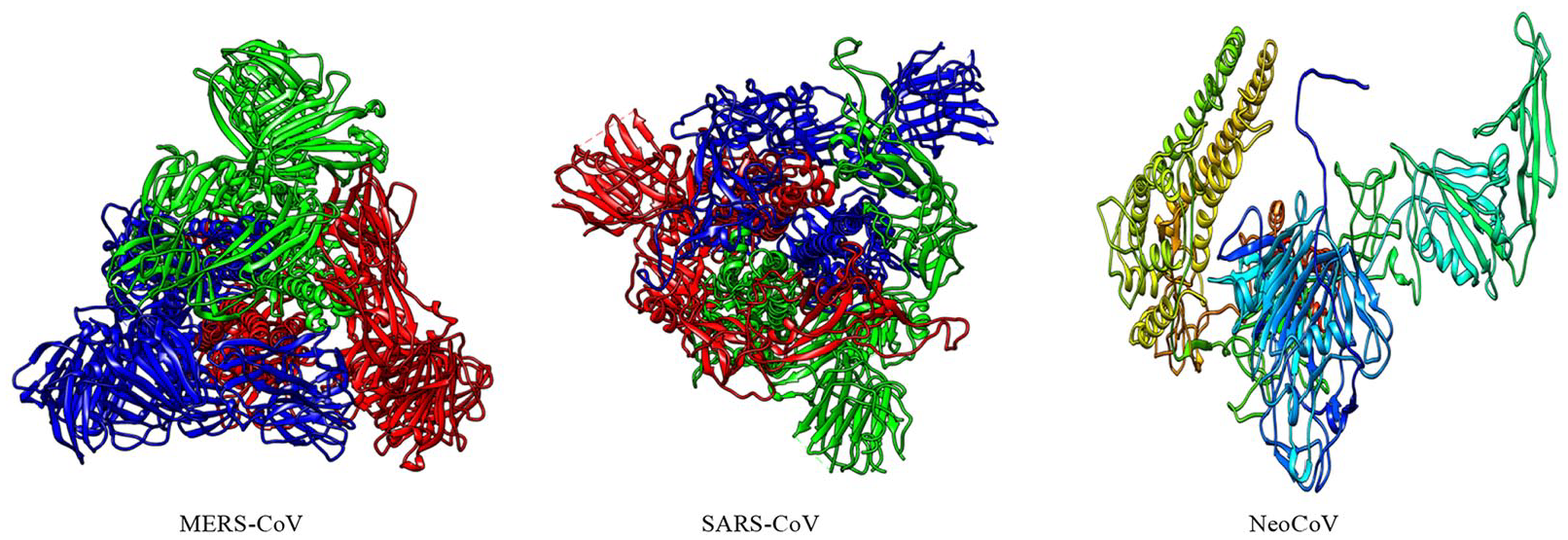
Three-dimensional (3D) structures of “S” proteins of 3 coronaviruses (MERS-CoV, SARS-CoV, and NeoCoV).
Measuring of template modeling score (TM-score) using Zhang-lab tool
The TM-score is defined to assess the topological similarity of 2 protein structures. 33 Zhang tool is designed to solve 2 major problems in the traditional metrics such as root mean square deviation (RMSD): (1) TM-score measures the global fold similarity and is less sensitive to the local structural variations and (2) magnitude of TM-score for random structure pairs is length-independent. TM-score has the value between 0 and 1, where 1 indicates a perfect match between 2 structures. Following strict statistics of structures in the PDB, scores below 0.17 correspond to randomly chosen unrelated proteins, whereas with a score higher than 0.5 assume generally the same fold in SCOP/CATH (https://zhanglab.ccmb.med.umich.edu/TM-score/) 34 (Table 16).
TM-score to the measure of similarity between couple 3D protein structures.
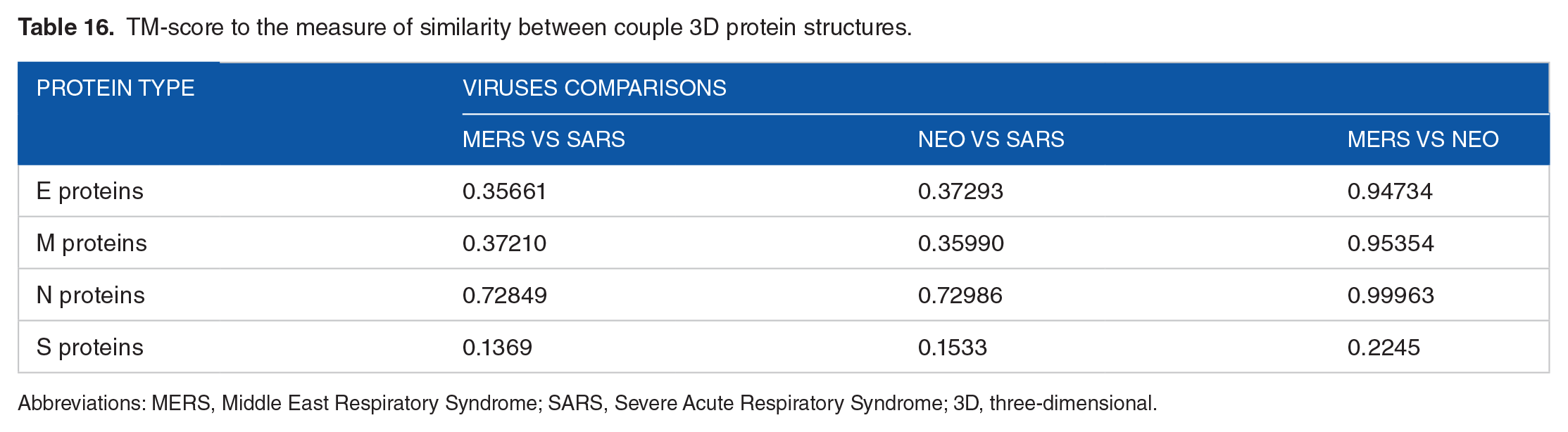
Abbreviations: MERS, Middle East Respiratory Syndrome; SARS, Severe Acute Respiratory Syndrome; 3D, three-dimensional.
Results
Discussion
In this study, we have endeavored to provide a deep understanding of the relationship between MERS-CoV, SARS-CoV, and NeoCoV at the amino acids level as the proteins are representing the functional unit of the genome and are directly involved in chemical processes essential for life. The proteins are species and organ-specific in which the proteins of one species or organs differ from those of another species or organs. However, proteins of similar function have similar amino acid composition and sequence. Despite the difficulties in explaining functions of protein from its amino acid sequence, understanding the correlations between structure and function is the key role of protein function.
With respect to the aim of determining the properties of amino acids that compose proteins of this study, Table 1 shows the physical and chemical properties of these proteins’ amino acids that are present in all CoV species of interest.
35
We have found that the number of amino acids of E, M, N, and S proteins in addition to all other parameters including molecular weight, atomic composition, theoretical pI, and structural formula of MERS-CoV and NeoCoV were close to each other if not identical, and this has supported the previous finding that NeoCoV was closely related to MERS-CoV and suggested that MERS-CoV’s ancestors may have evolved in bats.
12
This finding is in contrary to Victor Max Corman et al
10
results which have reported that NeoCoV and MERS-CoV belonged to one viral species and that the presence of a genetically divergent
In this study, we used 5 different methods of phylogenetic tree construction including Maximum Parsimony (MP), Neighbor-Joining (NJ), Unweighted Pair Group Method with Arithmetic Mean (UPGMA), Maximum Likelihood (ML), and RelTime (RT) to depict the relatedness, evolution change, and relative time between the viruses of interest (in the level of genome and protein). According to Phylogenetic results of the whole genomes which had relied on MUSCLE alignment, results have shown that joining of MERS-CoV and SARS-CoV with the nearest common ancestor and MERS-CoV has the lowest evolutionary change (Genetic distances). The RelTime method showed that NeoCoV was the oldest while MERS-CoV and SARS-CoV were belonged to the same time, based on the relative time. Furthermore, according to phylogenetic results of protein sequences which had relied on MUSCLE and CLUSTALW alignment methods, in general, trees have shown that joining NeoCoV and MERS-CoV proteins in same clades indicates that they are closest on the basis of all used methods. Furthermore, according to the horizontal branch length through used methods, most NeoCoV proteins have the shortest branch length comparing to others.
Regarding protein’s primary and secondary structures, most of the comparison results showed the most similarity between NeoCoV and MERS-CoV. Another comparison tool has template modeling score (TM-score), which is used to measure the topological similarity between the structure of proteins, and this method is insensitive to local structural variation. The TM results confirmed that NeoCoV was more close to MERS-CoV than SARS-CoV. Generally, phylogenetic analysis of the 6 proteins (E, S, M, N, ORF1a, and ORF1ab) revealed that there were high similarities between the 3 viruses although NeoCoV appeared close to MERS-CoV. This result indicated that they have the same common ancestor and NeoCoV may implicate in human-related infection sooner because of high similarity in portions involved in viral infectivity.
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
MMH conceived the idea and designed the methodology. AAE, ORG, SAA, RAA, and AMAM performed the initial draft analyses; in addition, MMH carried the final analysis. MMH and SBM interpreted the results. MAH, KS, and MMH wrote the manuscript and developed the final draft. All authors read and approved the final manuscript.
Availability of Data and Materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
