Abstract
Background:
The study aimed to assess the prevalence of
Methods:
Two hundred samples were collected from surgical wound infections and 100 blood cultures from patients with suggested bacteremia to identify
Results:
Conclusion:
MDR in all isolates (100%) and high-level resistance to gentamicin, erythromycin, and vancomycin were reported in
Keywords
Introduction
Enterococci, which were initially considered to be harmless flora of the gastrointestinal tract, have emerged in the last two decades as a major cause of hospital-acquired infections (HAIs),
1
including surgical site infections, urinary tract infections, and bacteremia.1-3 In the past, the source of infection by enterococci was mainly endogenous
4
; after that, the transmission of enterococci among hospitalized patients was reported.
5
Patients and Methods
Patient population
This is a cross-sectional study including 300 patients who developed clinical signs of surgical wound infection and bacteremia at least 48 hours after hospital admission as identified by the Centers for Disease Control and Prevention National Healthcare Safety Network (CDC/NHSN), 19 from surgery departments at Minia University hospital (a teaching hospital provides care to adult and pediatric patients in 35 wards including 800 beds), between June 2017 and January 2018. Samples were collected as the following: 200 wound swaps from patients with clinical signs of septic wounds and 100 blood cultures from patients with suggested bacteremia. The study protocol was approved by the local institutional review board at the authors’ affiliated institution (Registration number: MUH15329) and consents were obtained from all participants.
Bacterial isolation
Identification of the isolated
Antimicrobial susceptibility testing
Antimicrobial susceptibility of the isolates was determined by disk diffusion method for the following antibacterial agents; erythromycin (15 µg), gentamicin (120 μg), tetracycline (30 μg), ampicillin (10 µg), amoxicillin-clavulanic (30 µg), Cefepime (30 µg), vancomycin (30 μg), teicoplanin (30 μg), linezolid (30 μg), ciprofloxacin (5 μg), Imipenem (10 µg), and rifampin (5 μg) (Bioanalyse, Turkey). Muller-Hinton agar plates were inoculated with 0.5 McFarland standard suspension of the strains, antimicrobial disks were placed into plates and then were incubated at 37°C for 24 hours. Minimum inhibitory concentrations (MICs) of vancomycin and erythromycin were determined by the agar dilution method. Zone diameters were assessed according to the Clinical Laboratory Standard Institute guidelines. 20
DNA extraction
DNA was extracted using genomic BYF DNA extraction Mini Kit (Intron Biotechnology, Korea) according to the manufacturer’s instructions.
Identification of E faecalis Gene and Resistance Genes Using Real-Time PCR
The
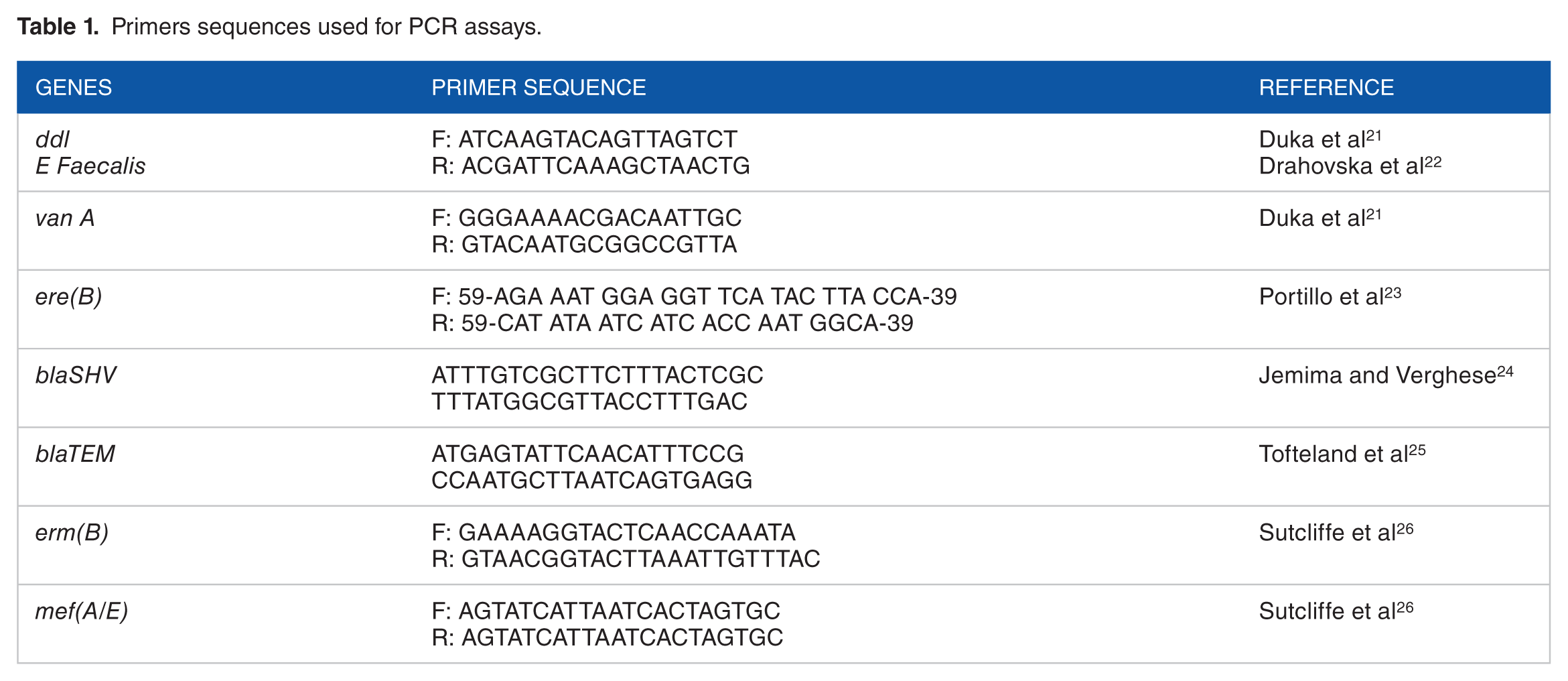
Primers sequences used for PCR assays.
Detection of erythromycin resistance genes, erm(B), and mef(A/E) by conventional PCR
PCR reactions were performed using thermal cycler (UNO II thermocycler, Biometra, Germany), 50 µL reaction: 25 µL DreamTaq Green PCR Master Mix (2X), 20 pmoles/µL for forward and reverse primer, 300 ng of DNA and completed to 50 µL with nuclease-free deionized water. Each gene was amplified using its specific primer 26 (Table 1) (Eurofins Genomic Co., Germany). Positive and negative controls from previous research were used. 27 PCR products were resolved on 2% agarose gel and visualized under a UV transilluminator (Biometra).
Statistical analysis
Categorical variables were analyzed using the chi-square test, using SPSS software (version 20).
Results
Characteristics of the study population
Out of 300 bacterial isolates, 26 (8.6%)
Characterization of
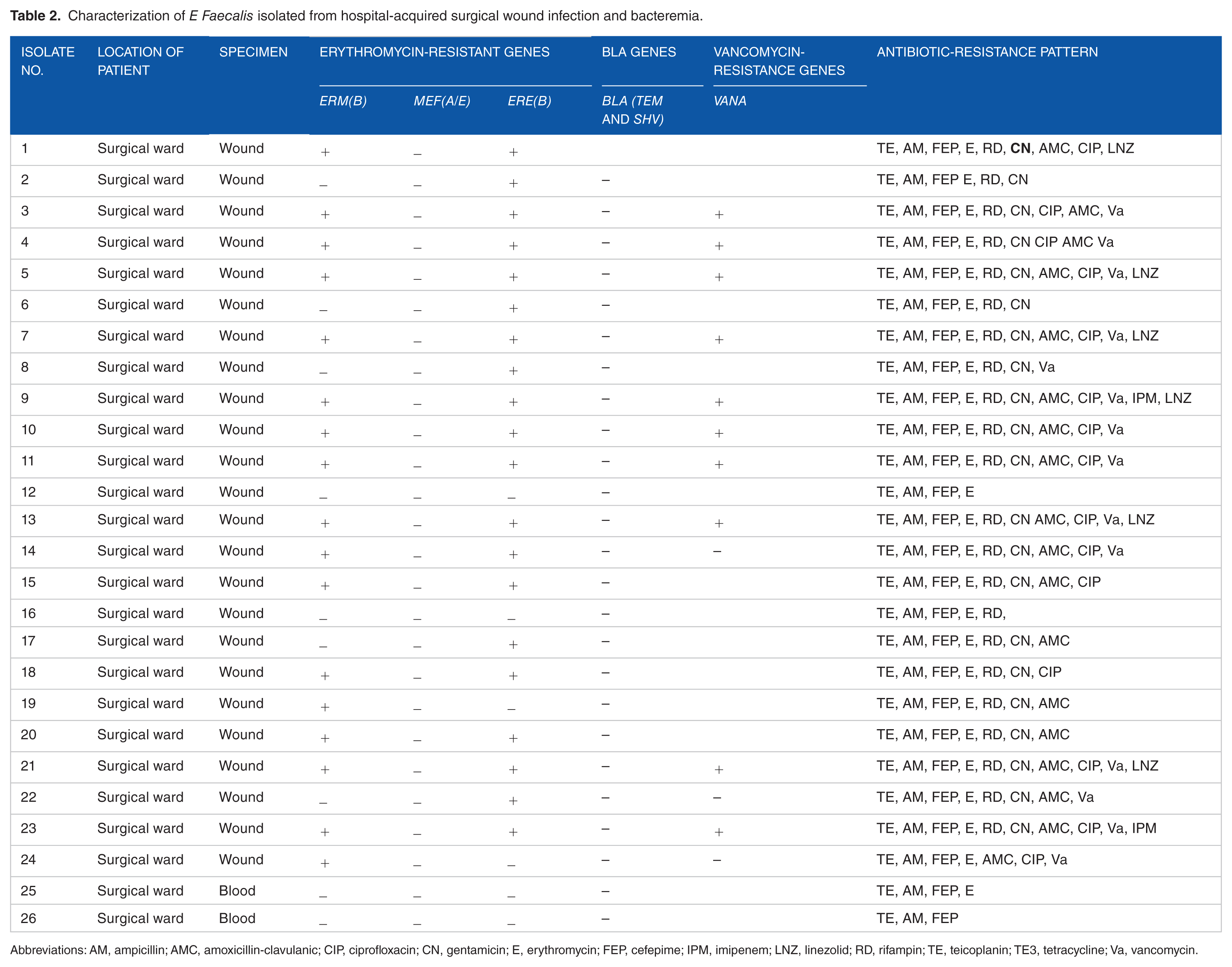
Abbreviations: AM, ampicillin; AMC, amoxicillin-clavulanic; CIP, ciprofloxacin; CN, gentamicin; E, erythromycin; FEP, cefepime; IPM, imipenem; LNZ, linezolid; RD, rifampin; TE, teicoplanin; TE3, tetracycline; Va, vancomycin.
Antimicrobial susceptibility
Among the 26 tested
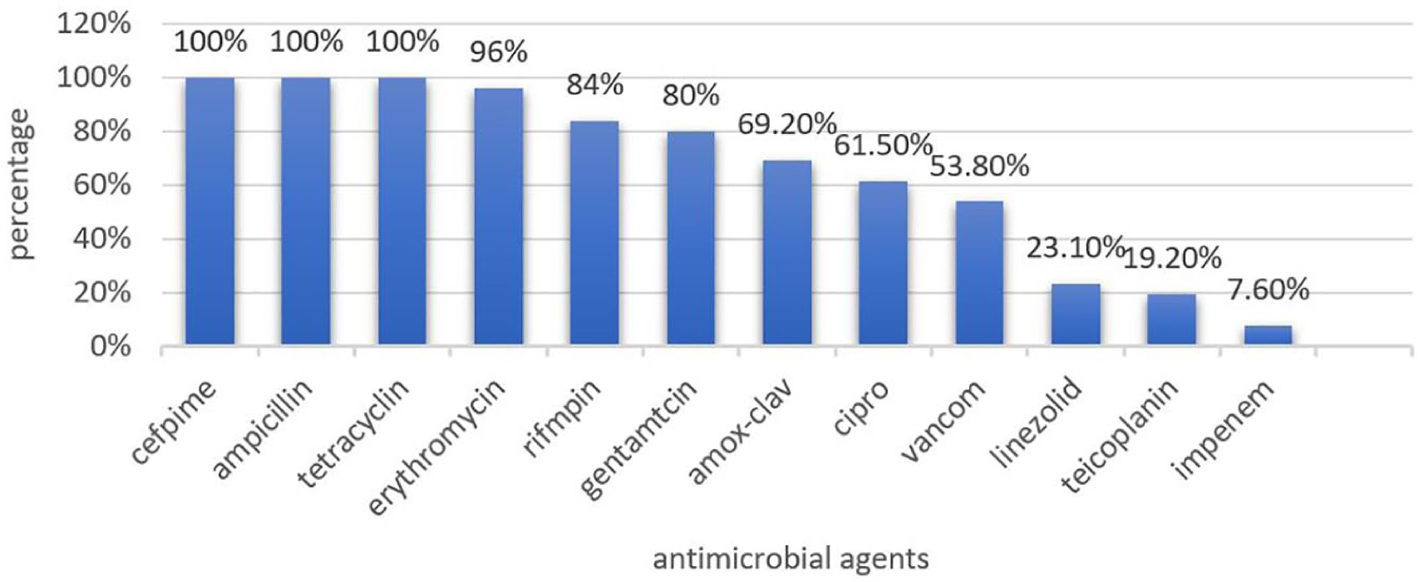
Antimicrobial resistance patterns of
Detection of resistance genes
Out of 25 erythromycin-resistant isolates, 20 (80%) were found to be positive for
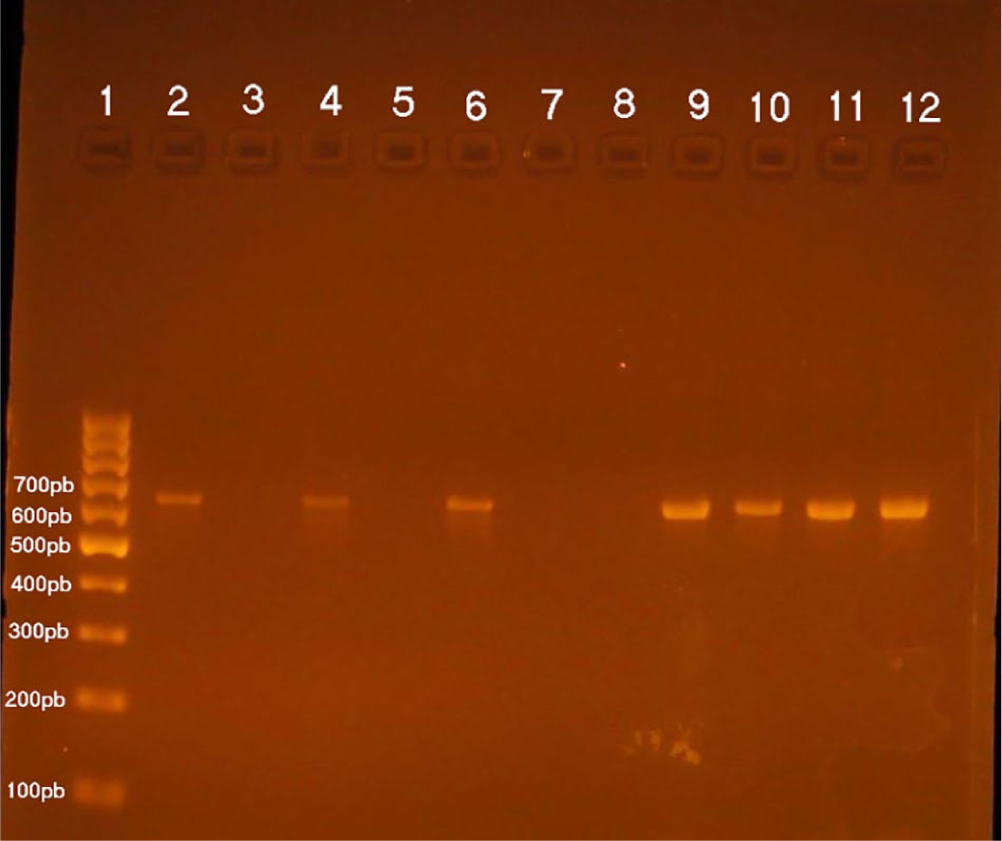
Gel electrophoresis for PCR products detecting erm B gene; lane 1: 100 bp molecular weight marker, lane 2: positive control, lane 3: negative control, lane 4, 6, and lanes 9-12: positive strains (639 bp), lanes 5, 7, and 8: negative strains.
Discussion
Enterococci are frequently isolated from health care settings. They reported as the third most common hospital-acquired pathogen.
28
They are increasingly isolated from traumatic and surgical wounds
29
and from bacteremia.
10
In the present study, the prevalence of E. faecalis isolation was 8.6% among 300 Egyptian patients with hospital-acquired infections. The isolation rate was higher in surgical wound samples; 24/200 (12%) than blood samples (2%). Our findings were higher than other reports, where
Multidrug resistance (MDR) was detected in all isolates (100%), defined by resistance to three or more Antimicrobials from different antimicrobial families
39
indicating a big challenge in treating infections by
Macrolides are still effective for treatment of important human infections.
40
Cross-resistance to macrolides is caused by mutations in
In summary, this study showed a prevalence of 8.6%
In conclusion, occurrence of cross-resistance to macrolides between gram-positive pathogens due to increasing rate of mutations in resistance genes and gene transfer between different species make the studying of such mechanisms between the
Limitations
There are two major limitations in this study that could be addressed in future research: first is that the study focused on
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
All authors conducted the the research, wrote the manuscript and analyzed the results. All authors reviewed the final manuscript.
