Abstract
Antimicrobial resistance (AMR) is a major global health concern, with resistant bacteria and genes spreading between humans, animals, and the environment. However, waterborne AMR in Enterobacteriaceae remains underexplored in developing countries such as Ethiopia. This study assessed the AMR profiles of Enterobacteriaceae isolated from 66 surface water samples collected from waste-receiving streams and the Gulf of Lake Tana. Bacteria were identified using standard microbiological methods, and antimicrobial susceptibility was tested by the Kirby–Bauer disk diffusion method. The most frequent isolates were Providencia alcalifaciens, Citrobacter spp., and Enterobacter spp. Resistance to first-generation cephalosporins ranged from 10.9% to 45.8% in stream isolates compared with 1.7% to 25.6% in lake isolates, while ceftriaxone resistance was detected in 13.5% and 4.5% of isolates, respectively. Among species, 30% of Escherichia coli, 22% of Enterobacter cloacae, and 12% of P. alcalifaciens were resistant to ceftriaxone, whereas resistance to cefazolin reached 40%, 67%, and 44%, respectively. Stream isolates also showed higher resistance to ampicillin (63%) and amoxicillin–clavulanic acid (29%) compared with lake isolates (50% and 12%). Overall, 44.8% of Enterobacteriaceae were multidrug-resistant. Significant differences in resistance were observed between stream and lake isolates for ciprofloxacin, ceftriaxone, trimethoprim–sulfamethoxazole, piperacillin–tazobactam, and cefazolin. The high prevalence of resistance to commonly used antibiotics, including third-generation cephalosporins, highlights the environmental dimension of AMR and underscores the urgent need for antimicrobial stewardship and surveillance in Ethiopia and similar settings.
Introduction
Antibiotic resistance (AMR) is a major global health threat, causing high morbidity, mortality, prolonged hospital stays, and increased healthcare costs worldwide. Its impact is disproportionately severe in low-and middle-income countries, where infectious diseases, poverty, and malnutrition are prevalent. In countries such as Ethiopia, AMR significantly undermines progress toward Sustainable Development Goal 3 (good health and well-being).1,2 Infections caused by antimicrobial -resistant bacteria are associated with higher mortality and morbidity, longer hospital stays, and increased healthcare costs compared with infections by susceptible strains. 3
Enterobacteriaceae are a large family of Gram-negative bacteria, including pathogens and opportunistic pathogens, found in the gut of humans, animals, the environment, and aquatic organisms. Enterobacteriaceae members can potentially cause infections in both community and healthcare settings. 4 The Enterobacteriaceae members of bacteria currently of high concern include K. pneumoniae, Enterobacter spp., E. coli, Citrobacter freundii, and Proteus spp,. These bacteria are also categorized by the World Health Organization (WHO) as a priority Enterobacteriaceae related to AMR infections. 5
Surface waters are continuously polluted by human activities such as swimming, fishing, and irrigation. They also receive waste from humans and animals, including wastewater from treatment plants, septic systems, and industrial facilities. Consequently, Enterobacteriaceae from the gastrointestinal tracts of humans and warm-blooded animals are released into these environmental waters.4,6 Environmental surface waters are crucial in transmitting AMR bacteria and their resistance genes among humans and animals. 7 In Ethiopia, the presence of AMR in environmental surface water is primarily due to inadequate waste management and poorly regulated antimicrobial use across the human, animal, and agricultural sectors.8,9 Although antimicrobial resistance may occur naturally, it is aggravated by the excessive use and misuse of antimicrobials in humans and animals, as well as by residues in soil, crops, and water. 10 As a result, swimming, drinking water, bathing, aquatic sports, occupational exposure during agricultural irrigation, and eating food produced with reclaimed water can all expose people to antimicrobial-resistant bacteria. 11
Antimicrobial resistance will have a bleak future unless we can organize a more robust response to its growing threat by pooling existing data on surface water AMR. 12 As a result, Ethiopia has increased its efforts to raise public awareness about AMR and has developed a national strategy and action plan to prevent and control it. 13 Data on AMR in surface waters, including lakes and streams, are essential to understand the environmental spread of resistant bacteria and to inform strategies to combat AMR. In Ethiopia, however, such data are limited, with few studies focusing primarily on drinking water sources. 14 The main objective of the study was to determine the antimicrobial resistance profile of Enterobacteriaceae in surface water from streams receiving waste and the Gulf of Lake Tana in Bahir Dar City, Ethiopia.
Materials and Methods
Study Area
The study was conducted in the Gulf of Lake Tana and the waste-receiving stream in Bahir Dar City (Figure 1). Bahir Dar City is the capital of the Amhara National Regional State, located 565 km northwest of the Ethiopian capital, Addis Ababa, and it is one of Ethiopia’s largest and fastest-growing cities. Bahir Dar city is located at 11.35′ 37.10' N and 37.23′ E. The city is estimated to cover 62.22 km2 and has an average altitude of 1995 m above sea level. The average annual rainfall is approximately 1360 mm, with 70% to 90% occurring between June and September. 15 The region is characterized by mild nocturnal temperatures and hot daytime conditions. The mean annual temperature is approximately 16°C, with average monthly maximum temperatures reaching up to 26°C. 16
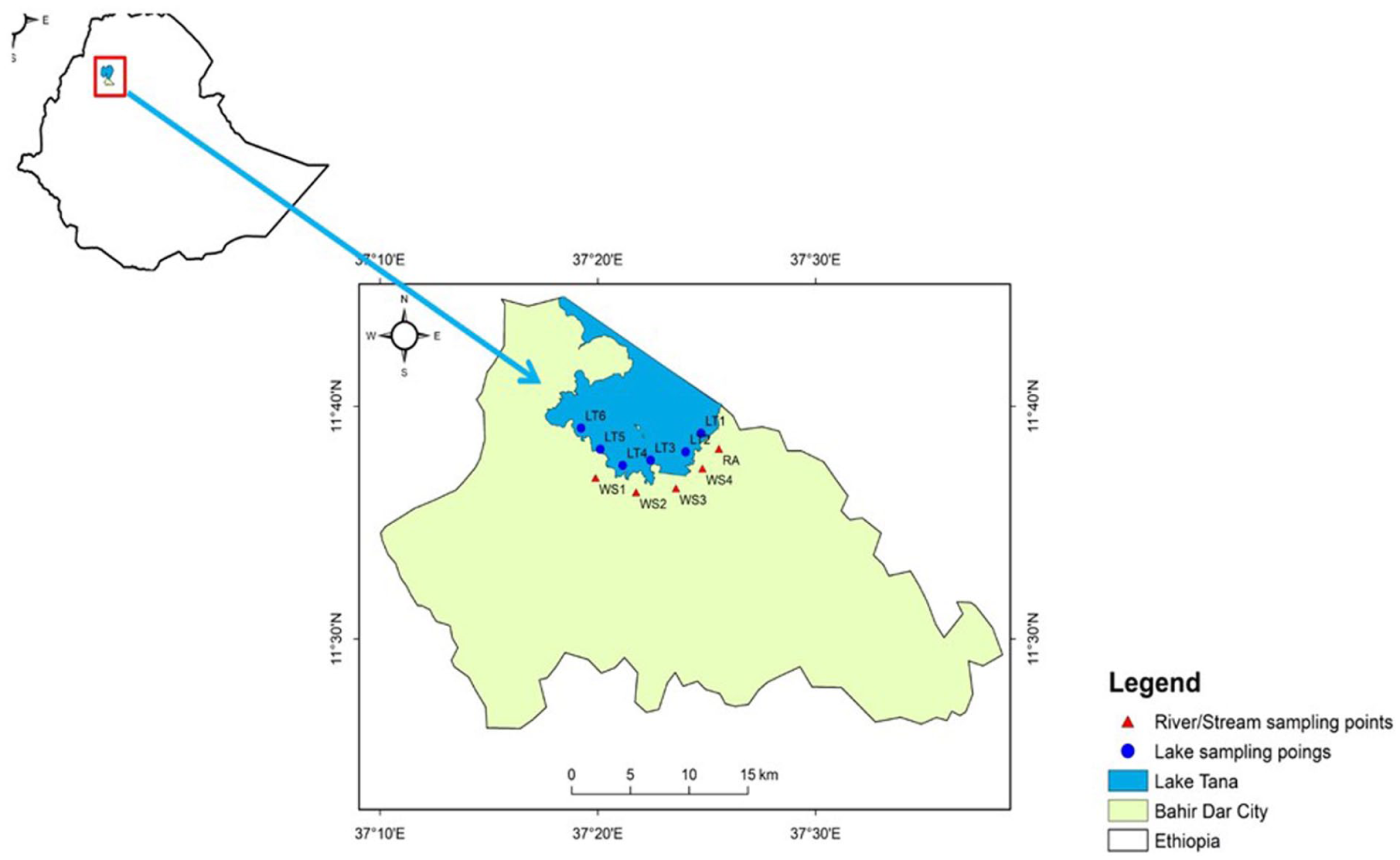
Maps of the study area, Ethiopia (Left top) and sampling areas (Right), southern gulf of Lake Tana water (blue circles) and stream waters (red filled triangles).
Lake Tana represents more than 50% of the country’s surface freshwater and provides several ecosystem services for a population of more than three million people. Lake Tana supports multiple activities, including providing drinking water, recreation, agriculture, and fisheries for local communities. However, growing amounts of solid and liquid waste from human activities around Bahir Dar City are directly discharged into open fields and ditches. The area’s inadequate sanitation infrastructure, including pit latrines and septic tanks with poorly sealed interfaces, is frequently contaminated by these wastes. 16 As a result, the level of fecal contamination in Lake Tana exceeds the WHO drinking and recreational water standards. 17
Sampling locations were selected by purposive sampling methods considering potential sources of pollution (eg, swimming sites, waste-receiving streams), accessibility, and pristine. A total of 11 sites, which include 4 waste-receiving streams, 6 swimming areas of Lake Tana, and 1 pelagic area of Lake Tana, were studied (Figure 1 and Table 1).
Description of Water Sampling Areas for Isolation of Enterobacteriaceae.

Waste receiving streams refer to the flowing water bodies that receive various locally produced solid and liquid wastes.
Gulf of Laka Tana: Littoral zones or the shoreline of Lake Tana near Bahir Dar City (Figure 1), Ethiopia
Water Sample Collection
Water samples were collected from July 25 to October 29, 2024, following EPA surface water sampling guidelines. Therefore, the catch sampling technique 18 was applied to obtain samples from waste-receiving streams. Water samples from wastewater intake streams were collected between 10:00 and 14:30, coinciding with peak discharge times, to capture variations in waste release. A boat was used to collect water samples from the pelagic zone of Lake Tana. At each sampling site, 250 ml of water was collected directly from the surface of the water (0 m) using sterilized glass bottles with 300 ml. The neck of the bottle was cleaned with 70% alcohol and stored in a refrigerator box during transport. Overall, we collected 66 samples from Lake Tana (n = 36), the waste-receiving streams (n = 24), and the Blue Nile River (n = 6). For Lake water, surface water sampling techniques have been used to capture water that swimmers come into contact with during swimming activities.
Identification of Bacteria
The bacteriological analysis of water samples was performed in the accredited microbiological laboratory at the Amhara Regional State Institute of Public Health (APHI) in Ethiopia (www.aphi.gov.et), following established standard operating procedures. Therefore, 2 separate 100 ml of water were passed through 2 sterile membrane filters with a 0.45 μm diameter to keep the Enterobacteriaceae on the surface. The membrane filters were then placed on a broth saturated with pre-lactic acid sulfate and 2 separate sterile pads on separate metal plates. Samples were incubated at 37°C for 24 hours for other Enterobacteriaceae, followed by an additional 24-hour incubation at 44°C for E. coli, in accordance with American Public Health Association (APHA, 1998). 19 Enterobacteriaceae at the genus level were identified based on colony morphology on MacConkey agar, followed by biochemical assays including oxidase, triple sugar iron, sulfur reduction and motility, citrate utilization, and decarboxylase and urease tests. 20
Antibiotic Susceptibility Testing (AST)
The Kirby-Bauer diffusion test on Mueller-Hinton agar was used to test for the antimicrobial susceptibility of Enterobacteriaceae. The antimicrobials tested are: amoxicillin (10 µg), gentamicin (10 µg), cefazolin (30 µg), piperacillin (10 µg), ceftriaxone (30 µg), tetracycline (30 µg), cephalothin (30 µg), and kanamycin (5 µg). The inhibitory zones have been interpreted following CLSI. 21 To standardize the inoculum density of bacterial suspension for the antimicrobial susceptibility testing, 0.5 MacFarland standard was used.
Data Analysis
The Chi-squared tests (X2) were computed to compare the differences levels of antimicrobial resistance between Lake Tana water and waste-receiving flowing water (WS) using IBM SPSS Statistics version 26. The P-values < .05 were considered statistically significant.
Results
Distribution of Enterobacteriaceae Isolates
In total, 107 Enterobacteriaceae isolates were identified from the surface water of Lake Tana and waste-receiving streams. The bacteria most commonly detected were P. alcalifaciens followed by Citrobacter spp., Enterobacter spp., and E. coli (see Figure 2). In particular, of the Enterobacteriaceae isolates, 63.6% (68/107) were recovered from waste-receiving streams and 36.4% (39/107) from Lake Tana (Figure 3) . In waste-receiving streams, P. alcalifaciens and Enterobacter spp., were the most frequently identified bacteria, while in lake water samples, Citrobacter spp. and Enterobacter spp. were the predominant isolates Figure 3a.

The overall proportion of Enterobacteriaceae isolates from waste-receiving streams and the southern gulf of Lake Tana.

The distribution of Enterobacteriaceae per site from waste-receiving streams water (a) and the southern gulf of Lake Tana (b).
Antimicrobial Resistance Profiles of Enterobacteriaceae
Overall, Enterobacteriaceae from waste-receiving streams and Lake Tana waters showed varying levels of antimicrobial resistance (Figure 4 ). Compared to surface water sources, AMR Enterobacteriaceae were generally higher in waste-receiving streams than in lake water (Figure 4a and b). Statistically significant differences in AMR were observed between isolates from Lake Tana and waste-receiving streams for ciprofloxacin (CIP; χ2 = 5.7, P = .01), ceftriaxone (CRO; χ2 = 10.3, P = .001), trimethoprim-sulfamethoxazole (SXT; χ2 = 4.2, P = .040), piperacillin-tazobactam (PTZ; χ2 = 5.8, P = .015), and cefazolin (CZ; χ2 = 4.33, P = .037). Considering bacteria-AMR combinations, 68% (243/357) of Enterobacteriaceae from waste-receiving streams and 32% (114/357) from Lake Tana showed resistance to the tested antimicrobials (Supplementary information Supplemental Table 1). Enterobacteriaceae isolates from waste-receiving streams (Figure 4a) showed resistance to first-generation cephalosporins (cephalothin and cefazolin) ranging from 10.9% to 45.8%, while isolates from the lake water (Figure 4b ) revealed 1.7% to 25.6% resistance levels. Similarly, resistance to third-generation cephalosporins (ceftriaxone) was observed in 13.5% and 4.5% of Enterobacteriaceae from waste-receiving streams and lake water, respectively. Stream-water Enterobacteriaceae revealed 63% resistance to ampicillin and 29% amoxicillin/clavulanic acid, whereas lake water isolates showed 50% resistance to ampicillin and 12% amoxicillin/clavulanic acid (Figure 4).

The overall antimicrobial resistance profiles of Enterobacteriaceae from surface water waste-receiving streams (a) and the southern gulf of Lake Tana (b).
Specifically, 30% of E. coli, 22% of E. cloacae, and 12% of P. alcalifaciens showed resistance to ceftriaxone, a third-generation cephalosporin. However, 40% of E. coli, 67% of E. cloacae, and 44% of P. alcalifaciens were resistant to first-generation cephalosporin (cefazolin; Table 2). In total, 44.8% of Enterobacteriaceae were multidrug-resistant (MDR) to 3 or more different classes of antimicrobials (R3 to R6; Table 3). Among these, the most prevalent MDR phenotypes observed were E. cloacae, P. retigeri, and P. vulgaris, followed by Citrobacter spp., E. aerogenes, and K. pneumoniae. However, only 4.6% of Enterobacteriaceae were 100% susceptible to the tested antibiotics. All Enterobacteriaceae isolates revealed sensitivity to gentamycin and Piperacillin/tazobactam (Figure 4).
Antimicrobial Susceptibility Results of Enterobacteriaceae Isolates from Surface Water.

Note. – indicates zero result.
Abbreviations: AMC, amoxicillin/clavulanic acid; AP, ampicillin; CIP, ciprofloxacin; CN, gentamycin; CRO, ceftriaxone; CZ, cefazolin; KA, kanamycin; KF, cephalothin; NA, not applicable; PTZ, piperacillin/ tazobactam; R, resistant; S, susceptible; SXT, trimethoprim/ sulfamethoxazole; TE, tetracycline.
Multidrug Resistance Profiles of the Enterobacteriaceae Isolates from Surface Water.

Abbreviations: R0 = sensitive for all classes of antibiotics; R1 = resistance for 1 class of antibiotics; R2 = resistance for 2 classes of antibiotics; R3 = resistance for 3 different classes of antibiotics; R4 = resistance for 4 different classes of antibiotics; R5 = resistance for 5 different classes of antibiotics; R6 = resistance for 6 different classes of antibiotics; - = sensitivity; MDR, multidrug-resistance.
Discussion
The development and dissemination of AMR are currently the primary challenges confronting the global healthcare system. However, the AMR problem is severe in low-income countries due to weak healthcare systems, poor sanitation and hygiene standards, and inadequate waste management systems. Antimicrobial resistance arising from commonly used drugs at the human–animal–water interface is a major driver of AMR spread and poses a significant public health threat. 22 This study provides baseline data on Enterobacteriaceae antimicrobial resistance in surface waters.
The higher proportion (63.6%) of Enterobacteriaceae isolates from waste-receiving streams compared to Lake Tana (36.4%) may be due to the greater disposal of solid and liquid waste into the streams rather than the lake. The Enterobacteriaceae most frequently isolated from all water samples were P. alcalifaciens, followed by Citrobacter spp., E. aerogenes, E. coli, E. cloacae, and K. pneumoniae. This composition of Enterobacteriaceae aligns with findings from other studies conducted on surface water in the Akaki River 23 and urban rivers in Addis Ababa, Ethiopia. 24
Overall, most of the AMR Enterobacteriaceae were isolated from waste-receiving stream water rather than lake water. Here, we emphasized discussing the cephalosporin resistance levels of environmental Enterobacteriaceae isolates rather than their resistance to other antimicrobials. 5 Overall, in the present study, 44% to 68% of Enterobacteriaceae exhibited resistance to first-generation cephalosporins, which aligns with the average resistance of 23% to 70% in Enterobacteriaceae found in urban rivers in Addis Ababa, 24 Ethiopia, and 50% to 100% resistance in the Akaki River. 23 The overall 10% third-generation cephalosporin resistance (ceftriaxone) in this study aligns with the 10.8% resistance to ceftriaxone observed in waterborne Enterobacteriaceae in South Africa. 24 Specifically, the resistance levels of E. coli (30%) to third-generation cephalosporin were lower compared to the 70% E. coli resistance in urban rivers in Addis Ababa, 24 and 52.4% E. coli resistance in hospital wastewater in Ethiopia. 24 Similarly, Citrobacter spp. and Klebsiella spp. showed significant levels of resistance against third-generation cephalosporins.23 -26 These findings indicate that surface waters act as reservoirs for antimicrobial-resistant Enterobacteriaceae, presenting significant public health risks.
In the current study, the overall 50% to 68% resistance of Enterobacteriaceae to ampicillin is consistent with study findings reporting 69% resistance in the Addis Ababa River, 23 and 63.1% in Romania. 27 Furthermore, a recent meta-analysis from Africa among surface water isolates reported a high pooled prevalence of resistance to penicillin and ampicillin resistance pooled estimate was 69.4% that aligns well with the present ampicillin resistance. 28 However, it was lower than the 84% resistance along the Akaki River in Addis Ababa 23 and 92.5% resistance in the aquatic environment in Turkey. 29
Enterobacteriaceae isolated from surface water exhibited MDR phenotype to widely used antibiotics. Water-associated E. coli isolates from low- and middle-income countries (LMICs), including several African studies, reported a pooled MDR prevalence of 43.7% among E. coli. 30 The MDR rate of 44.8% observed in the present study among surface-water Enterobacteriaceae is therefore consistent with these pooled estimates for LMIC aquatic environments and highlights the potential risk for human exposure. In comparison to other findings from Ethiopian studies, the 44.8% MDR level observed in this study is lower than the 64% MDR found in polluted urban rivers in Addis Ababa, 24 but higher than the 28% MDR in the Little Akaki River 22 and 23.6% in hospital wastewater. 28 The most common Enterobacteriaceae that showed MDR from Ethiopian water sources were E. coli, which revealed 33.3%, 36.4%, and 78% MDR23 -25, Citrobacter spp., which showed 13% to 39%, and 60% MDR, and K. pneumoniae, which showed 16.8% to 39%.23,26 Further research is needed to determine the corresponding resistance genes in Enterobacteriaceae from surface water.
Conclusion
This study demonstrated that antimicrobial-resistant Enterobacteriaceae were more frequently isolated from waste-receiving stream water than from lake water. Notably, significant resistance to third-generation cephalosporins was detected in opportunistic pathogens such as Escherichia coli, Citrobacter spp., and Klebsiella spp. In addition, surface water Enterobacteriaceae exhibited high levels of MDR phenotype. These findings underscore the critical role of surface waters in the dissemination of AMR Enterobacteriaceae and highlight the importance of implementing systematic monitoring and surveillance programs. Strengthening public awareness on proper waste management, ensuring adequate treatment of waste before environmental discharge, and continuous water quality evaluation should be prioritized to mitigate the spread of AMR bacteria in the environment.
Supplemental Material
sj-doc-1-ehi-10.1177_11786302261419983 – Supplemental material for Antimicrobial Resistance Profiles of Enterobacteriaceae in Surface Water From Streams and the Southern Gulf of Lake Tana, Ethiopia
Supplemental material, sj-doc-1-ehi-10.1177_11786302261419983 for Antimicrobial Resistance Profiles of Enterobacteriaceae in Surface Water From Streams and the Southern Gulf of Lake Tana, Ethiopia by Mole Yibeltal, Alemale Admass, Michael Getie, Abebech Beyene, Belay Bezabih and Bayeh Abera in Environmental Health Insights
Footnotes
Acknowledgements
We thank the Amhara Public Health Institute (APHI), especially the Microbiology staff, for providing laboratory space and equipment, as well as antibiotic disks and control strains for conducting the laboratory work.
Author Contributions
B.A. and M.Y. conceived the research idea and scientific write-up; A.A., M.G., B.B., M.Y. material acquisition, methodology design, supervision and review of the manuscript, laboratory work, data analysis and interpretation, and writing and review of the manuscript. A.B. drew the study map and edited and wrote the manuscript and B.A. designed the methodology, wrote the manuscript, and interpreted data.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request. Corresponding author (Dr. Bayeh Abera), email:
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
