Abstract
The global rise of antimicrobial resistance in Escherichia coli poses critical public health challenges, especially in resource-limited settings. This study aimed to investigate the role of untreated hospital sewage and industrially polluted river water as reservoirs for antimicrobial-resistant E. coli. A total of 40 environmental samples were collected, yielding 75 E. coli isolates identified using phenotypic methods. Due to time and resource limitations, 50 isolates were subjected to antimicrobial susceptibility testing by the Kirby-Bauer disk diffusion method against 11 antimicrobial agents. These antimicrobials were selected based on their clinical significance, availability, dominance in resistance mechanism, and utilization trends in Ethiopia. Notably, E. coli isolates from the Yerer River downstream of industrial waste discharge exhibited the highest multi-drug resistance rate (90%), while upstream isolates were fully susceptible to all tested antimicrobials. The Xadacha River isolates showed moderate multi-drug resistance (66.67%). Hospital sewage isolates displayed escalating multi-drug resistance rates across the 3 departments, including the intensive care unit (62.5%), open patient department (91.67%), and inpatient department (100%). Overall, 86.67% of hospital sewage-derived isolates and 65% of river isolates demonstrated multi-drug resistance. Polymerase chain reaction confirmed antimicrobial resistance-associated genes (tetA: 83.33%; blaTEM: 57.14% of the resistant isolates), underscoring hospital sewage and polluted water as critical reservoirs for antimicrobial resistance gene dissemination. The 2 genes were selected based on their availability, phenotypic resistance profile, and nature as sentinel markers for high-use antimicrobial classes in Ethiopia. These findings highlight the urgent need for comprehensive wastewater treatment systems, stricter antimicrobial stewardship, and integrated One Health surveillance to mitigate AMR risks to human, animal, and environmental health.
Introduction
Antimicrobial resistance (AMR) occurs when pathogens such as bacteria, viruses, or fungi evolve to resist antimicrobial treatments, rendering infections increasingly difficult to manage. Recognized as a critical global public health threat, AMR disproportionately impacts low- and middle-income countries (LMICs), including Ethiopia, due to systemic vulnerabilities in healthcare infrastructure and resource limitations. 1 Without urgent intervention, AMR could lead to 10 million annual deaths by 2050, underscoring the need for coordinated interdisciplinary action. 1 Escherichia coli is commensal gram-negative bacteria which causes different infections both in human and animal. Several studies confirmed the potential of E. coli to develop resistance to common antibiotics through natural process as well as anthropogenic factors. 2 In 2023, high AMR profile of E. coli was confirmed in human, animal, and the environment they shared. Environmental samples of Hospital are the key reservoirs for drug resistant bacteria as stated in the comprehensive study conducted in Zambia. 3
The emergence and spread of AMR is already severe according to a study by Murray et al 4 . It attributed 4.95 million global deaths to AMR in 2019, with 1.27 million directly caused by resistant infections. The key drivers of AMR include misuse of antimicrobials (eg, over prescription, under dosing), lack of rapid diagnostic tools, lack of awareness, inadequate sanitation, and poor infection control. Environmental pathways, such as untreated hospital waste and industrial runoff, further increase resistance by dispersing antimicrobial residues and resistant genes into ecosystems. Likewise, livestock production, particularly the use of antimicrobials in farming, boost this risk, enabling b zoonotic pathogens like multidrug-resistant E. coli to transfer resistance genes across species. 4
Environmental waste, particularly from pharmaceuticals, agriculture, and healthcare, plays a critical role in the global emergence and spread of antimicrobial resistance (AMR) microbials. In Ethiopia, poor waste management and untreated effluents from hospitals and farms introduce antimicrobial residues and resistant microbes into water and soil ecosystems. This accelerates the selection and spread of resistant pathogens, posing serious threats to both human and animal health. AMR pathogens lead to increased morbidity, mortality, and treatment costs. Animals exposed to contaminated environments may act as reservoirs, facilitating zoonotic transmission. Addressing environmental AMR drivers is crucial for a One Health response. Antimicrobials can exert selective pressure on bacteria, leading to the development of antimicrobial resistance. Tiwari et al 5 reported that MDR bacteria have been found in untreated hospital effluents and wastewater treatment plants (WWTPs). The continuous exposure of bacteria to antimicrobials contributes to the emergence of resistance. Wastewater systems harbor a reservoir of antimicrobial resistance genes (ARGs). These genes can be inherent, exchanged among bacteria, or acquired from the reservoir.5 -7
Diagnostic stewardship ensures appropriate antimicrobial use by integrating accurate testing into clinical decision-making. Coupled with vaccination, which reduces infection incidence and antimicrobial demand, these strategies offer a proactive defense. Vaccines prevent the transmission of resistant strains and decrease secondary infections, providing a cost-effective solution for marginalized populations.8,9 Addressing AMR demands a holistic strategy that bridges human, animal, and environmental health. The One Health framework highlights how resistance genes circulate among pathogens in diverse ecosystems, necessitating policies to regulate antimicrobial use in agriculture, improve waste management, and foster cross-sector collaboration. Combating AMR requires innovation in diagnostics, vaccine development, and environmental governance. Prioritizing the One Health approach, alongside global partnerships and equitable resource allocation, will safeguard antimicrobial efficacy and protect vulnerable populations. Without such efforts, the world risks a post-antibiotic era where common infections become lethal threats10,11 (Figure 1). While AMR in E. coli is widely studied globally, this research addresses critical gaps in low resource, high-pollution setting country, Ethiopia, where several rivers are used for drinking, irrigation, and recreation. There is dramatic increase in industrialization and concomitant expansion in public infrastructures like hospitals in Ethiopia. The hospital and industrial runoff liquids carry several chemicals, heavy metals, and antimicrobial residues. Particularly, hospital sewage and rivers polluted with pharmaceutical industry waste water are the major contributors as they are contaminated with diverse sub-inhibitory concentration of antibiotics. This leads antimicrobial deposition in environment including rivers which in turn enhance the emergence and spread of resistant genes. Once polluted liquid discharged to environment, they create a selective pressure to microbial where mutation and horizontal gene transfer (HGT) increase. Moreover, no prior data exists on AMR linkages between hospital sewage and industrial pollution in this study area. 12 Therefore, The specific aims of this study were to address the following 3 key points: (1) to assess antimicrobial resistance (AMR) patterns in E. coli from untreated hospital sewage and industrially polluted river water, (2) to identify genetic determinants (tetA and blaTEM), and (3) to evaluate public health implications. Data were critically analyzed using phenotypic and molecular methods, with statistical validation (Using Excel and SPSS) to ensure robustness. Results were compared with global trends, confirming high multi-drug resistance rates and the role of environmental reservoirs in AMR dissemination. The study provides a rigorous, evidence-based analysis, addressing the need for improved wastewater treatment and antimicrobial stewardship in resource-limited settings.
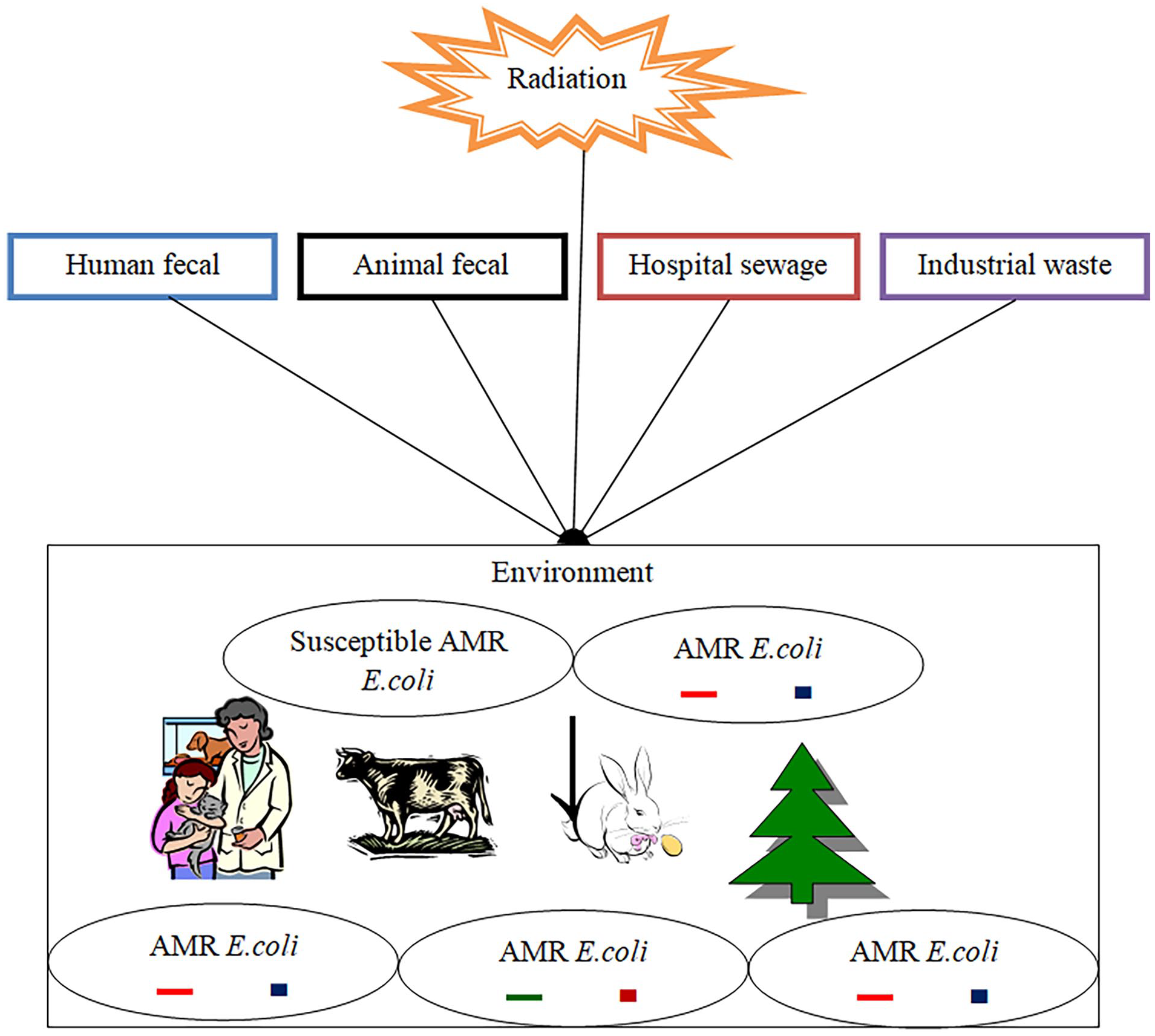
Reservoirs of AMR genes and their possible spreading and emerging mechanism. Human and animal feces, untreated hospital sewage, and industrial waste all contribute to the spread and emergence of AMR. Discharging these wastes into the environment can cause mutation and horizontal gene transfer, resulting in AMR E. coli.
Materials and Methods
Study Area
This study was conducted in the eastern Shawa vicinity of Adama and Dukem in the Oromia region of Ethiopia (Figure 2).
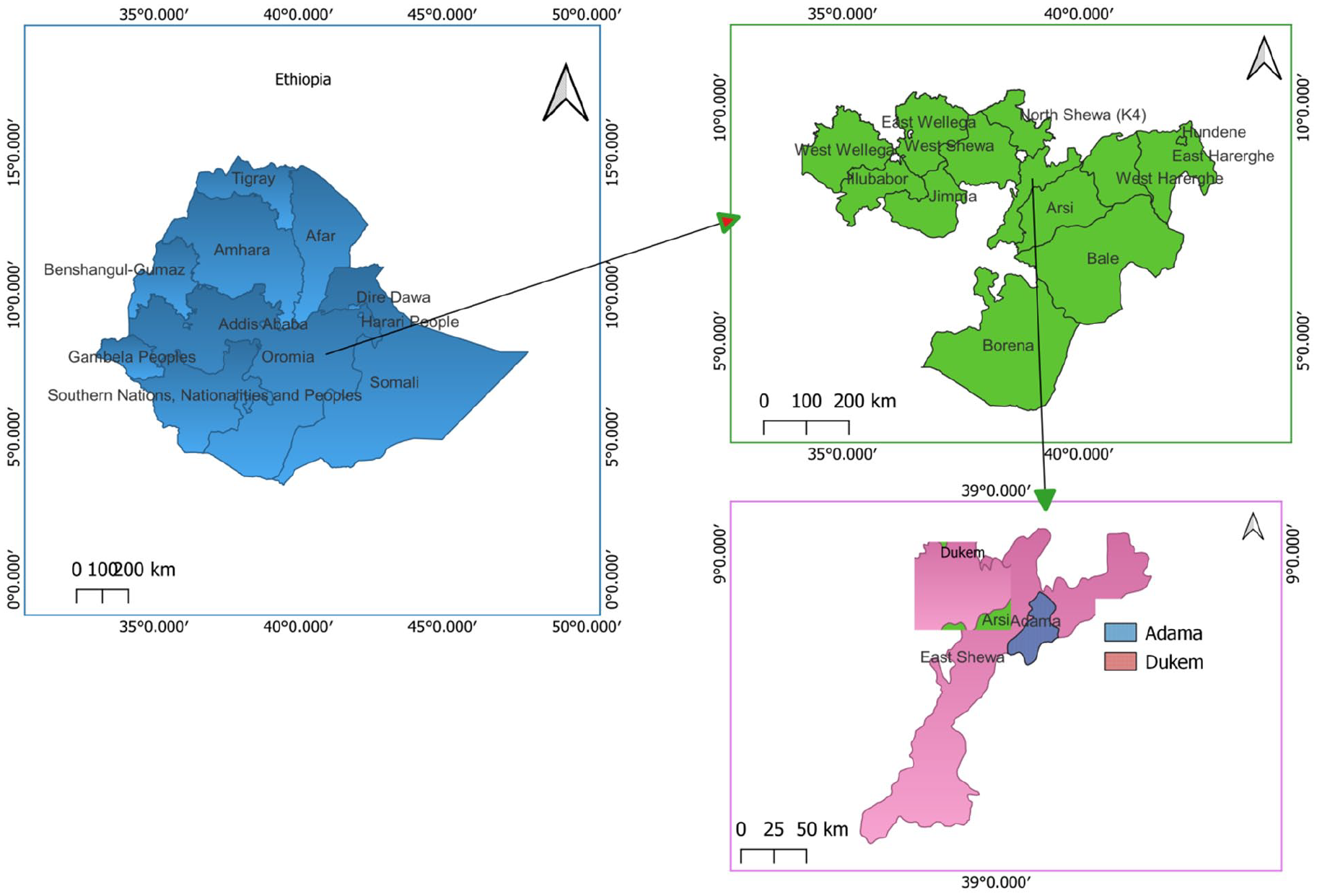
A map of sample sites (Adama Hospital Medical College, YRBI, YRAI, and XR) generated using QGIS version 3.24 software.
The Research Design
This study employed a cross-sectional design to examine AMR patterns of E. coli in hospital sewage and polluted river samples. A cross-sectional study design assesses a specific issue at a single time point across a period of weeks or months of sampling. The study approach was chosen to evaluate the AMR profile of E. coli isolates from 2 environmental sample sources since it is both quick and inexpensive. Furthermore, it uses a culture-based AST method rather than the more expensive metagenomic or genomic approaches. These factors made it critical to evaluate AMR patterns in countries with limited resources.
Sample Collection and Transportation
Liquid samples were collected from the Adama Hospital Medical College, Yerer River (polluted river), and Xadacha River (control sample) using the purposive sampling method. Following wastewater sampling guidelines, 13 samples were taken from the OPD, IPD, and ICU rooms of the hospital and the YRAI, YRBI, and XR river sites over 5 days. The YRAI was the targeted river site that was polluted with different industrial wastes whereas YRBI and XR samples were used as controls since there is no known industrial pollution discharged to these sites. The collected samples were labeled and transported in an icebox to Addis Ababa University, Institute of Biotechnology, Molecular Biotechnology Lab room 413. The samples were stored at 4°C and antimicrobial resistance was assessed from February 2023 to June 2024.
Isolation, Identification and Characterization of the E. coli Isolates From the Collected Samples
The study followed standardized microbiological methods for the isolation and identification of E. coli strains, aligned with guidelines from the Clinical and Laboratory Standards Institute (CLSI) for water quality. Phenotypic characterization was used to isolate and identify E. coli in hospital sewage and river water samples. Each sample was enriched with tryptone soy broth (TSB) and cultured at 37°C for 24 hours. Following enrichment, overnight cultures (0.5 mL) were inoculated on eosin methylene blue (EMB) agar and incubated at 37°C for 24 hours. Un-inoculated sterilized EMB agar media was used to confirm quality assurance. The streak plating technique was used to isolate E. coli, utilizing EMB and MacConkey agar. After the initial incubation, the streak plate method was used to further purify the isolates by transferring each colony from the overnight culture to a fresh medium. E. coli isolates were identified by their distinctive dark red to purple coloration, with a metallic green sheen on EMB agar. Isolates were further purified on MacConkey agar to differentiate between lactose-fermenting (pink colonies) and non-lactose-fermenting (pale colonies) E. coli strains. Well-separated colonies were cultured on nutrient agar to maintain a pure culture for further experiments. Standard biochemical characterizations, including indole, methyl red, Voges–Proskauer, citrate, catalase, and triple sugar iron (TSI) tests, were used to identify the isolates as E.coli. 14
Antimicrobial Susceptibility Testing (AST)
AST is an experimental protocol that determines the efficiency of a particular antimicrobial against microbial. The AMR of the E. coli profile from hospital sewage and river water samples was determined using the Kirby–Bauer disc diffusion method, which is the most widely used and economical method for determining antimicrobial susceptibility on Mueller–Hinton agar. The tests were conducted using the guidelines of the Clinical and Laboratory Standards Institute’s (CLSI) guidelines. E. coli isolates were tested for resistance to the most widely used antimicrobial agents, including ampicillin (10 μg), ciprofloxacin (5 μg), chloramphenicol (30 μg), sulphonamides (300 μg), erythromycin (10 μg), gentamycin (10 μg), norfloxacin (10 μg), tetracycline (30 μg), vancomycin (10 μg), streptomycin (10 μg), and Nalidixic acid (5 μg; Thermo Scientific™ Oxoid™, UK). 15 The Kirby–Bauer method involves inoculating E. coli on Mueller–Hinton agar with antimicrobial-impregnated disks, incubating for 24 hours and measuring inhibition zones to determine resistance. These results will guide physicians in selecting effective treatments and assessing the emergence and spread of AMR genes. The antimicrobial disks were carefully placed on the surface of the inoculation plates at a distance to avoid overlapping the inhibition zones and then incubated at 37°C for 24 hours according to the CLSI guidelines. 16 For erythromycin (macrolide) and vancomycin (glycopeptide), the break points were determined based on the previous works done by17,18 (Table 1). The E. coli ATCC 25 922 was used as a positive control in all AMR tests conducted in this study.
Inhibition zone diameter interpretive standards chart for AMR tests as determined by CLSI and other guidelines. 18 .

E. coli isolates resistant to 3 or more selected antimicrobials were considered as Multiple Drug Resistance (MDR). MDR is when a bacterium, E. coli, in our case, becomes resistant to 3 and above antimicrobials. The MAR index, a tool used to determine the risk level of AMR E. coli sources, was calculated as follows:

MAR index above 0.2 indicates the source is at high risk, suggesting environments with antimicrobial abuse.19,20
Molecular Confirmation of the Isolates for Resistance Gene
The study involved inoculating E. coli isolates with phenotypically confirmed AMR in TSB and incubating at 37°C for 24 hours. The cells were extracted and pelleted by centrifugation at 14 000 rpm for 5 minutes, then boiled and chilled. The quality and concentration of the DNA extracts were examined by gel electrophoresis using a nanodrop spectrophotometer. The gel was prepared by dissolving 0.45% agarose in 1 × TAE buffer, ethidium bromide, and loading dye. Gel electrophoresis was performed for 60 minutes and visualized using UV light. 21
In this study, PCR amplification was used to identify and characterize AMR E. coli at the molecular level as previously described. 15 PCR is an important molecular biology technique used to amplify small DNA sequences to generate millions of DNA fragments to determine the presence or absence of specific genes. The DNA extracts of phenotypic AMR-confirmed E. coli isolates were subjected to a polymerase chain reaction (PCR) to assess the presence of the selected target AMR genes, namely tetA and blaTEM genes.22 -24 Specific primer pairs were used for amplification.

The tetA gene encodes a membrane protein that facilitates efflux pumps and supports microbial resistance to tetracycline families by expelling antimicrobials from the cell, allowing E. coli to survive in its presence. 25 The blaTEM gene encodes the TEM β-lactamase enzyme, which breaks the β-lactam ring of antimicrobials such as ampicillin and penicillin and is crucial for their antibacterial activity.25,26
The PCR was performed in a total reaction volume of 50 µL containing 0.5 µL each of the forward and reverse primers of each blaTEM and tetA genes, 2.5 µL standard buffer B and MgCl2 each, 0.5 µL dNTPs, 0.25 µL Taq DNA polymerase (Delta Biotechnology PLC, Ethiopia), and 1 µL of DNA template. Nuclease-free water was used as a negative control while a sample previously confirmed to have resistant genes was used as a positive control. Amplification was performed under the following modified conditions: initial denaturation at 94°C for 5 minutes, 30 cycles of denaturation at 94°C for 30 seconds, annealing at 62°C for 30 seconds, and extension at 72°C for 30 seconds, followed by a final extension at 72°C for 10 minutes. 27 The PCR products were then run on a 1.3% agarose gel stained with ethidium bromide (10 mg/mL) in the presence of a negative control (no DNA) to check for possible contamination in the master mix. The gel was subjected to 100 V for 50 minutes in 1 × TAE buffer at pH 8. The buffer was prepared using 3 chemical compositions: tris hydrochloride (40 mM), EDTA (1 mM), and glacial acetic acid (20 mM). PCR products were visualized under UV light using a gel documentation machine (Bio-Rad, USA). The band sizes were estimated and recorded in the presence of a 1000 bp DNA ladder (Soli BioDyne).
Data Analysis
The combination of qualitative and quantitative data analyses provided a full view of the AMR E. coli profile, allowing for the extensive phenotypic and genetic characterization of resistant E. coli isolates and their prevalence. Among the many statistical software systems available today, IBM Corporation’s Statistical Package for the Social Sciences (SPSS) version 26, was used. SPSS software is commonly used and generates a full report on a computer screen that can be copied and pasted into another document format.
Results
In this study, 75 isolates were identified as presumptive E. coli. Among the 75 isolates, 50 (66.67%) E. coli isolates were randomly selected and subjected to AST against 11 antimicrobials based on the CLSI guidelines. The results showed widespread resistance among the isolates, with the highest resistance rates observed against erythromycin (76%), vancomycin (72%), tetracycline (72%), streptomycin (60%), ampicillin (56%), Nalidixic acid (54%), and sulphonamide (52%). Chloramphenicol (30%), ciprofloxacin (28%), gentamycin (28%), and norfloxacin (8%) were used (Figure 3). Notably, E. coli isolates exhibited particularly strong resistance to erythromycin, tetracycline, and vancomycin, compared to the other antimicrobials tested. The AMR patterns of E. coli isolates among the 6 sample sites were also evaluated, with OPD taking the first place for high AMR E. coli followed by YRAI and IPD (Figure 4). The MDR index was calculated based on the 3 most resistant antimicrobials listed above, to determine the proportion of isolates resistant to these 3 antimicrobials. This index underscores significant multidrug resistance among the studied E. coli strains, highlighting the urgent need for effective antimicrobial stewardship and surveillance measures. Of the total 50 isolates, 44 (88%) have more than 0.2 while only 6 (12%) have less than 0.2 MDR index (Figure 5). This study provides informative insights into the AMR profiles of E. coli strains in environmental settings, emphasizing the critical importance of monitoring and managing antimicrobial resistance in public health strategies. 16 There was no statistically significant difference among the AMR patterns of E. coli from the hospital sample groups (P > .05), whereas there was a difference between those from the river sample groups (P < .05; Table 2).

AMR profile of E. coli Isolates against the selected antimicrobials. The E. coli isolates from hospital sewage and river waters in Adama and Dukem, respectively, were subjected to 11 antimicrobials. Namely: Ampicillin, Chloramphenicol, Ciprofloxacin, Compound Sulphonamides, Erythromycin, Gentamycin, NaldicAcid, Norfloxacin, Streptomycin, Tetracycline, and Vancomycin.

The AMR patterns of E. coli isolates among the 6 sample groups. This figure represented the order of sample groups from the largest AMR E. coli profile source to the smallest: OPD → YRAI →IPD→ICU→XR→YRBI.
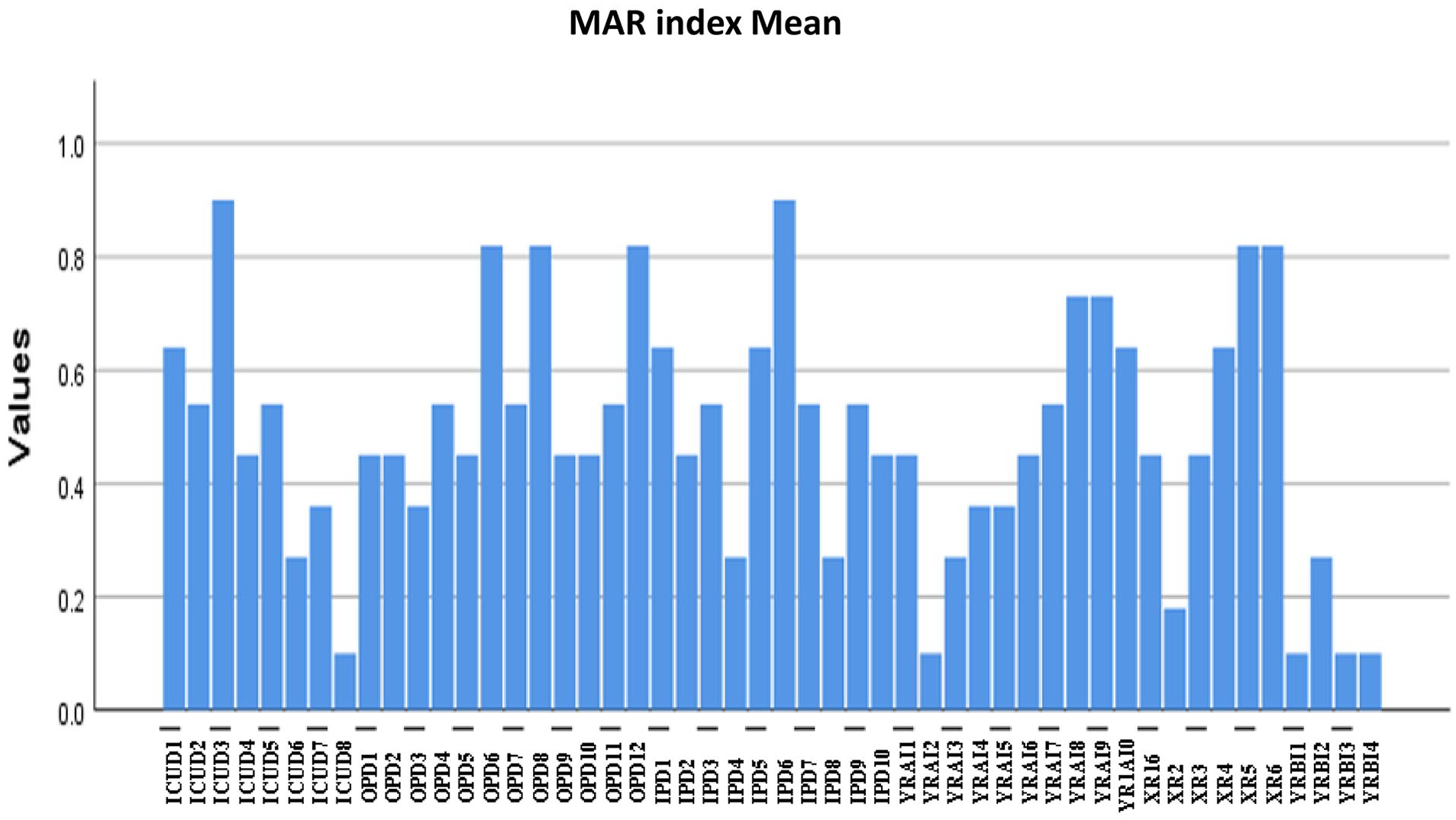
Multiple antimicrobial resistance (MAR) index of each identified AMR E. coli isolates. From the total 50 isolates, 44(88%) have more than 0.2 while only 6(12%) have less than 0.2 MAR index.
The Cross-tabulation of the Sample groups with AMR patterns. It reveals distinctive AMR patterns across the sample sites and sample groups. The highest resistance rates were found in OPD (73 cases) and YRAI (66 cases), while the ICU had the highest susceptibility rate (29 instances). Intermediate patterns were more common in OPD (36 cases) and YRBI (28 cases). (AMR = Antimicrobial resistance, ICU = Intensive care unit, IPD = Inpatient department, OPD = outpatient department, XR = Xadacha river, YRAI = Yerer river after industrial waste discharge site, YRBI = Yerer river before industrial waste discharge site).

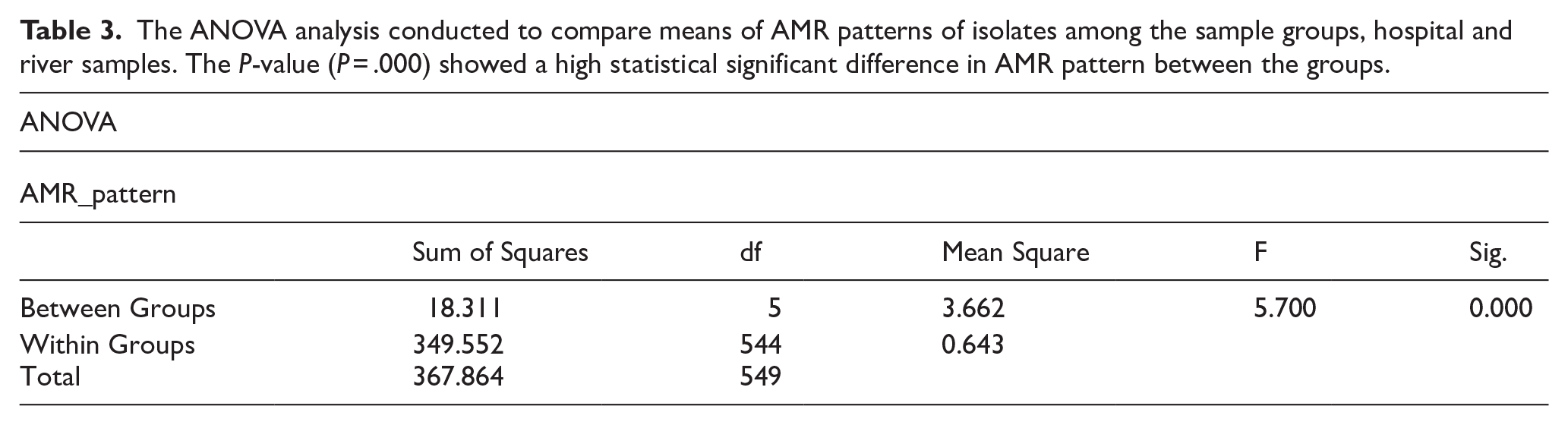
Analysis of AMR pattern variations between AMR E. coli isolated from hospital sewage and river water samples. The analysis confirmed the presence of a statistically significant difference between sample sites (P < .05; Table 3).
The ANOVA analysis conducted to compare means of AMR patterns of isolates among the sample groups, hospital and river samples. The P-value (P = .000) showed a high statistical significant difference in AMR pattern between the groups.
The association among the variables was evaluated using the chi-squared test, and the result depicted a statistically significant association between the variables with a 57.967 Pearson chi-squared value and P < .05 at 10 degrees of freedom. This indicated that the observed variations were not due to chance. Both the likelihood ratio test and linear-by-linear association test results supported the Pearson chi-squared results (Table 4).
Summarized Chi-Square Tests for AMR pattern among the isolates from hospital samples and river samples.
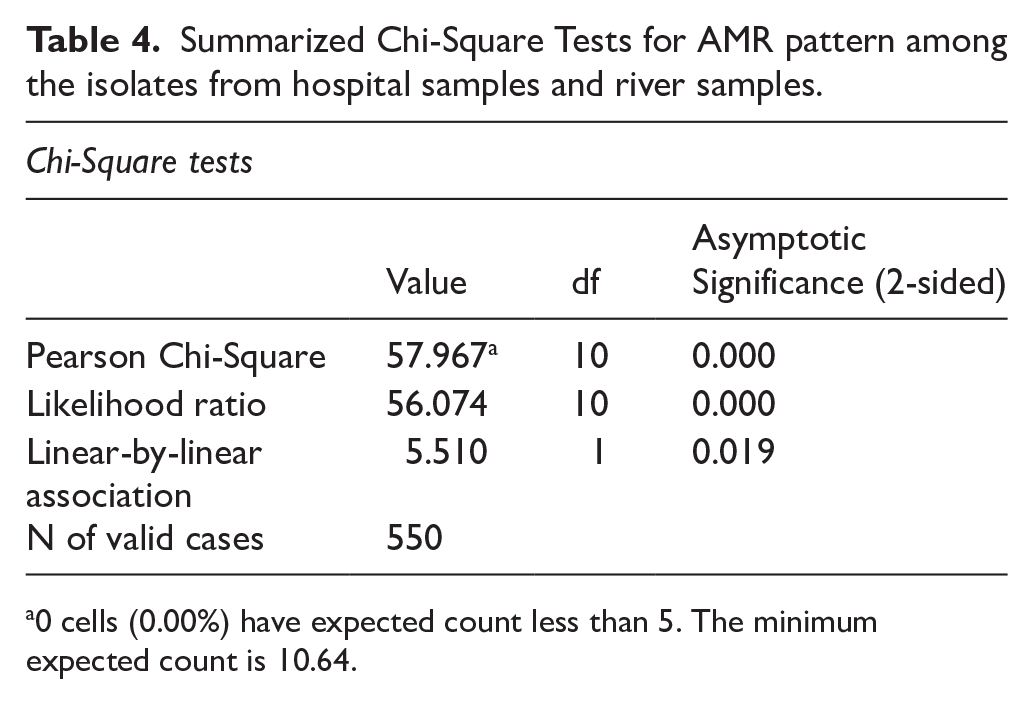
0 cells (0.00%) have expected count less than 5. The minimum expected count is 10.64.
The average means of the MAR index for the hospital, Yerer River (polluted), Xadacha River (control), and Yerer River samples before the industrial waste entry site (control) were 0.53, 0.6, 0.33, and 0.14, respectively. Therefore, both hospital and YRAI samples had a high MAR index in our study.
Molecular analysis confirmed the presence of the AMR gene in E. coli isolates previously identified through morphological and biochemical tests (Figure 6). Of the 36 isolates phenotypically confirmed to have AMR to tetracycline, 30 (83.33%) harbored TetrA (Figure 7). Similarly, 16 (57.14%) of the 28 ampicillin-resistant isolates tested positive for blaTEM. This study underscores the utility of PCR for molecular characterization of AMR profiles in bacterial isolates, providing insights into the genetic mechanisms underlying antimicrobial resistance.
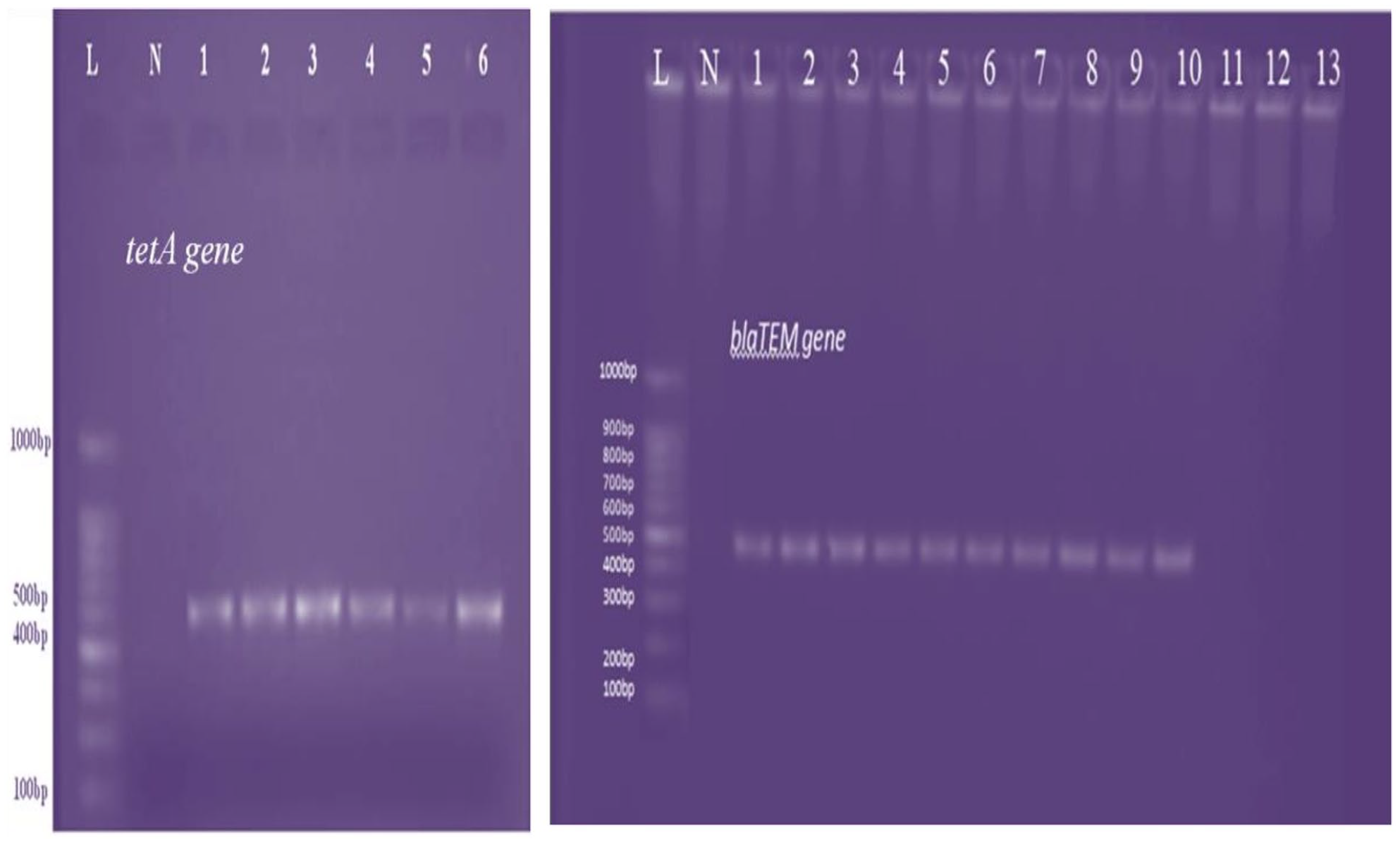
Gel electrophoresis results of PCR products: This picture shows the confirmed molecular gel image of the amplified product of the TetrA gene and blaTEM gene of some of the identified positive E. coli isolates. Where: L = 1 kb ladder, N = Negative control (Master mix solution without DNA template), Lane3-Lane8 represents positive isolates with an estimated band size of 550 bp. In the case of the blaTEM gene, Lane3 up to Lane10 represents positive isolates with an estimated band size of 534 bp while Lane 11, 12, and 13 were blaTEM gene negative.
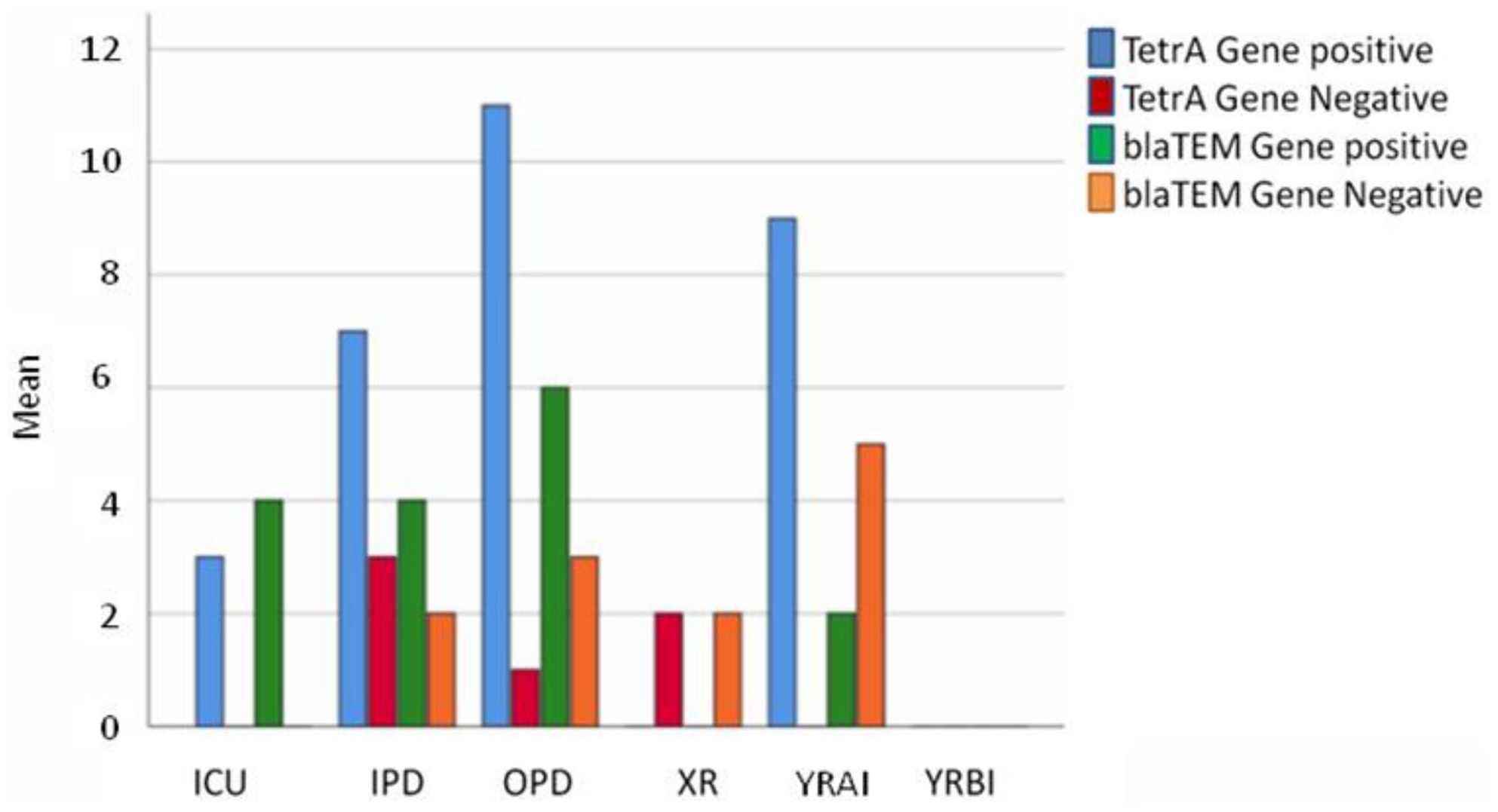
The different sample sources and the antimicrobial resistant genes, tetA and blaTEM genes, distribution patterns among the sample sites. The results depicted significant difference of the selected AMR gene distribution among the sample groups.
Molecular confirmation of the targeted AMR genes revealed the status and spread of these 2 AMR genes. This was conducted after careful isolation, identification, and determination of the E. coli load and phenotypic AMR profile of E. coli. The abundance of E. coli in the liquid samples was determined to evaluate the most likely routes for AMR gene spread.
This study revealed a significant difference in the antimicrobial resistance (AMR) profile and abundance of E. coli among the sampling sites. The number of AMR E. coli at the hospital was higher than that at the river. Likewise, the AMR E. coli profile was higher at YRAI compared to samples taken from YRBI and XR (P = .004).
Discussion
This study focused on assessing the AMR profile of E. coli isolated from hospital sewage and polluted river water. Many rivers in Ethiopia, including those flowing through Adama and Dukem, are heavily polluted by various pollutants, including industrial waste, clinical sewage, household, and agricultural waste. Almost all the sites assessed along the different rivers had a water quality of class I, which no longer met the primary water quality standard. 28 Liquid waste is a highly suspected vehicle for the emergence and spread of AMR pathogens in humans, animals, and the environment. 3 Therefore, effective regulatory policies and sustainable interdisciplinary monitoring are needed for waste management and environmental protection to curb the emergence and spread of AMR genes. However, to the best of our knowledge, there has been no previous research on the AMR profile of E. coli at selected study sites. Susceptibility tests were used to determine the effectiveness of the antimicrobials against bacterial isolates from different sources. 29 Our findings revealed a higher distribution of E. coli isolates identified in hospital sewage samples than in river water samples. This disparity suggests that hospital sewage may serve as a more prominent reservoir for E. coli compared to river water in the studied context. This could be attributed to several factors including the presence of AMR strains originating from clinical settings, the concentration of human waste, environmental factors for the bacteria to survive, and/or the potential for the hospital effluents to act as a hotspot for bacterial dissemination.
The MAR index of most isolates was greater than 0.2, which is similar to the results of a study conducted in Nigeria. 30 The MAR index of all samples indicated that the targeted sample sources were in the high-risk category, except for YRBI, which showed a MAR index of 0.14. MDR isolates showed resistance to multiple antimicrobials, 3 or more in this case, during AST. 31 This study revealed a dramatic increase in resistance to erythromycin, the most resistant, indicating frequent use of antimicrobials around the sampling sites. E. coli and other Enterobacteriaceae are intrinsically resistant to erythromycin. However, the primary aim of this study was on the resistance mechanism of the isolates. The diameter of the inhibition zone was measured based on previous works.17,18 Generally, the diameter of the zone of inhibition was measurement with caliper and reported in millimeters. The results were compared to CLSI guidelines to determine if the bacteria are susceptible, resistant, or intermediate to the antibiotic. The study contextualized the inherent resistance of Enterobacteriaceae to macrolides including this antimicrobial. This result was lower than that of a study conducted by Yitayew and his colleagues. 32 The tetracycline resistance rate was the second most abundant resistance which was greater than 23 (42.56%) the report in Bahir Dar. 33 Tetracycline is one of the most frequently used antimicrobials in African countries, where agriculture is the primary source of income. 34 The predominance of tetracycline resistance (72%) and tetA gene detection (83.33%) mirrors trends in agricultural regions, where tetracycline is extensively used in livestock. However, this finding exceeds resistance rates reported in Tanzania (42.56%) and the Philippines (39.6%), 20 suggesting region-specific factors such as unregulated antimicrobial sales and limited wastewater infrastructures in Ethiopia.
The high MDR rates observed, particularly in hospital inpatient department (IPD) sewage (100%) and downstream Yerer River sites (YRAI) sample (90%) reflect the compounding effect of clinical antimicrobial misuse and industrial pollution on resistance selection. This percentage is greater than the results reported in Tanzania, 86.76% (177/204). 35 The observed difference between this finding and those of previous studies could indeed be attributed to several factors, including differences in sampling techniques and sample sizes, evolution of pathogens, culturing techniques, and environmental conditions.
The significantly higher AMR burden in the YRAI (P = .004) aligns with studies linking heavy metals and pharmaceutical residues in industrial effluents to the co-selection of resistance genes. Such pollutants create selective pressures that favor resistant bacterium, E. coli in this case, even in the absence of direct antimicrobial exposure. Similarly, the 86.67% MDR rate in hospital sewage E. coli isolates correlates with the widespread use of broad-spectrum antimicrobials in healthcare settings, where inadequate wastewater treatment allows AMR strains to enter the ecosystem.
Furthermore, this study showed that E. coli isolates from the YRAI were more resistant to the antimicrobials than those from the YRBI and XR (P = .004). This indicates that industrial pollution significantly affects the emergence and spread of antimicrobial resistance. Industrial sewage often contains antimicrobials, heavy metals, and other pollutants that create a selective environment that favors the growth of AMR E. coli. The higher AMR profile at the polluted river site suggests that industrial pollution was a key driver of resistance in the river environment.
The PCR test confirmed that most of the isolates harbored tetA in 83.3% of the isolates tested, which was greater than (65.1%) the result reported by other group. 36 The high prevalence of the tetA gene highlights that most of the AMR E.coli isolates follow an efflux pump resistance mechanism. This actively transports the drugs out of the E.coli’s cell decreasing the impact of the antimicrobials enabling them to survive exposure to tetracycline, which is commonly used in both human medicine and agriculture.
The high prevalence of the blaTEM gene (57.1%), a gene associated with extended-spectrum beta-lactamase (ESBL) production, signals emerging resistance to ampicillin, penicillin, and cephalosporins, cornerstone drugs for treating Gram-negative infections. The prevalence of the blaTEM gene revealed the ability of the tested AMR isolates for beta-lactamase production, which is another important resistance mechanism. These resistance mechanisms underscore the urgency of curbing AMR reservoirs in vulnerable sites like hospitals and rivers contaminated with industrial discharges.
Populations relying on the Yerer River for drinking water, agriculture, and recreation face a heightened risk of exposure to antimicrobial-resistant bacteria. The spread of AMR genes from environmental sources to animals and humans can lead to infections that are difficult to treat and manage. 37
The disparities in MDR rates among the 3 hospital departments, IPD (100%) E. coli isolates compared with the ICU (62.5%) isolates likely depict differences in antimicrobial prescribing practice and patient exposure. IPD is the setting where prolonged horizontal and empiric broad-spectrum antimicrobial use is common. These cases may accelerate the selection of resistant strains. Conversely, the lower MDR rate in the ICU E. coli isolates could be attributed to stricter infection control protocols or targeted antimicrobial regimens for critically ill patients.
The isolates that showed phenotypic resistance were subjected to molecular testing to confirm the distribution of 2 selected genes, tetA and blaTEM. Genes were selected based on the phenotypic resistance patterns of the isolates to tetracycline and beta-lactam antimicrobial families.
This finding indicates that the AMR profile of E. coli isolates was high in hospital sewage samples and Yerer River water after the discharge of industrial sewage waste. Analysis of the results showed that there were almost no statistically significant differences between the 2 (P > .05). The outcomes of this study align with the findings in Nigeria and Tanzania, where clinical and industrial waste were linked to elevated MDR rates in aquatic ecosystems.
Despite the significance of the findings of this study, some limitations have to considered in the future. These limitations include a relatively small sample size of 50 isolates subjected to AST and the specific geographical region. However, this number is still sufficient for preliminary analysis. Further studies with larger sample sizes and diverse sampling sites would enhance the robustness of these findings. Additionally, the molecular characterization focused on 2 AMR genes, tetA and blaTEM only. While these genes were prevalent among resistant isolates, other suspected genes would be added together with whole-genome sequencing or metagenomic analysis could provide a more comprehensive understanding of resistance mechanisms in future studies. Furthermore, the selective pressure exerted by antibiotic use differs across healthcare settings, which may explain variations in MDR rates among different hospital departments. For instance, the inpatient department (IPD) exhibited the highest MDR rate (100%), likely due to prolonged hospitalization and broad-spectrum antibiotic use, whereas the intensive care unit (ICU) showed a comparatively lower MDR rate (62.5%), potentially due to stricter infection control practices.
Those limitations include the sample size of 50 isolates subjected to AST due to time and resource constraints while sufficient for preliminary analysis. Furthermore, some tested antimicrobials, such as ciprofloxacin (28%), gentamycin (28%) norfloxacin (8%), displayed much lower resistance rates, which may warrant further investigation to understand underlying factors. Moreover, the statistical analysis revealed no significant differences among hospital sample groups, unlike river water sample groups which revealed distinct patterns emphasizing the complexity of AMR dynamics.
Conclusion and Recommendation
This study evaluated antimicrobial resistance (AMR) patterns of E. coli strains isolated from hospital sewage in Adama and polluted river water in Dukem. The results revealed high resistance profiles, particularly to tetracycline, ampicillin, and streptomycin. The targeted representative resistance genes, tetA and blaTEM, persist even in diluted environments. These findings confirm that untreated hospital and industrial wastes significantly contribute to the emergence and spread of AMR E. coli among humans, animals, and the ecosystem they share. The high MAR index values of the resistant isolates indicate serious public health risks to nearby communities. This calls for an urgent need for improved wastewater treatment, stricter antimicrobial regulations, and regular monitoring of antimicrobial susceptibility. To address this public health challenge, a coordinated One Health approach is highly required. Furthermore, future researchers should explore more resistance mechanisms and assess the broader ecological and health impacts of AMR E. coli to inform effective prevention and control strategies.
Ethical Considerations
This study aimed to assess the AMR profiles of E. coli isolates from hospital sewage and polluted river water in Adama and Dukem. A series of bacteriological methods have been used to isolate, identify, and evaluate AMR patterns of E. coli. The results of this study were compared with those of previous studies conducted in different parts of Ethiopia and followed the ethical principles and guidelines of the American Society for Microbiology. 38
This study posed a minimal risk to the participants, researchers, animals, and the environment, and included human and animal participants indirectly because they were the source of E. coli in hospital sewage samples and river water samples. The researchers did not collect any personal or identifiable information from the patients nor were they contacted for any reason. Informed consent was obtained from the researchers via hospital laboratory management before the sample collection. The confidentiality and privacy of the participants and managers were secured. This study was approved by the Institutional Research Board of the College of Natural and Computational Sciences of Addis Ababa University. Permission for hospital sewage water and river water sample collection was provided by the College of Natural and Computational Science Institutional Review Board (CMS-IRB).
Footnotes
Acknowledgements
The authors would like to acknowledge the support from Prof. Tesfaye Sisay Tessema and the thematic research grant to AGB funded by the Office of Vice President for Research and Technology Transfer (VPRTT) of Addis Ababa University.
Author Contributions
AGB and KFT: Conceptualization, methodology and analyzed the data, writing-review & editing, and drafted and revised the manuscript. AGB: Supervision. KFT: Data collection and fieldwork. All authors have read, reviewed, and approved the final manuscript.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
