Abstract
Contamination of maize adversely affects maize quality, yield, and export creating a gap in the attainment of food security, which is a millenium development goal in Nigeria. The study determined fungal abundance, genetic variability, and prevalence of toxigenic fungi in maize grains consumed in North Central, Nigeria. Sixty composite stored maize samples were collected and fungi were isolated and identified after which a multiplex polymerase chain reaction was used to confirm the presence of mycotoxin regulatory genes in suspected toxigenic fungi. The genetic relationship among the toxigenic fungi was determined and the genetic correlation between Aspergillus isolates was established through Restriction fragment length polymorphism (RFLP) analysis. About 389 (64.83%) of the total maize samples collected had fungal species belonging to the genera Aspergillus, Mucor, Penicillium, Fusarium, Trichophyton, and Talaromyces associated with them. Among the regions surveyed, Kogi State exhibited the highest maize contamination rate at 89 samples, accounting for 22.9% of the total samples collected. In Benue and Kogi, the genus Aspergillus exhibited the highest relative abundance, with percentages of 76.6% and 76.3%, respectively. Among its species, Aspergillus flavus and A. niger were the most predominant. Kwara State had the highest fungal diversity with a value of 1.711 (P < .05). Benue State had 11 isolates (4.6%) with genes encoding for mycotoxin production, the highest recorded. Conversely, Nasarawa and Niger States each had the lowest count, with 4 isolates possessing such genes. Out of the 238 fungi suspected to be mycotoxigenic that were isolated, 39 have genes that encode for mycotoxin synthesis. Low divergence existed between toxigenic fungal species using the alpha diversity index. This study confirmed that the grains were contaminated with closely related fungal strains, and concluded that maize grains consumed in North Central Nigeria showed high association with fungal microbiota, including species capable of contaminating the grains with mycotoxins
Introduction
Maize grains contain nutrients that encourage fungal growth and colonization. These fungi damaging crops are categorized into pre-harvest fungi that colonize crops before harvesting, and the post-harvest fungal species which invade crops after harvesting through to the storage and processing stages. Contamination of crops with mycotoxins along the food chain threatens food security and it is a problem in sub-Saharan Africa where food demand is high and supply is grossly inadequate. Maize is the staple food for the vast majority of Africans and provides an excellent source of carbohydrates, proteins, vitamins, and minerals to the population. It is the mainstay in animal feed and is needed for various industrial applications. Maize has a genetic plasticity that has made it the most widely grown crop in Nigeria. Maize grows under a wide range of climatic conditions (temperature 21°C to 27°C and rainfall 480 to 880 mm). The problem of fungal and mycotoxin contaminations reduces income, yield, and quality of agricultural production, and introduces animal and human diseases when such food is consumed. 1 Nigeria with over 200 million people relies heavily on cereals and grains as food sources and feed for animals. The importation of grains has continued to make up for the deficit supply as a result of the dependence on essential grains (wheat, corn, and rice).
The International Grains Council (IGC) projected the total grain production of Nigeria for the 2021 to 2022 season to be around 21.5 million tons, with maize accounting for 12.5 million tons. However, maize production in 2022 decreased to 12.1 million tons, representing a 5% decline from the previous year. 2 The adverse effects of climate change and its consequential impact resulted in a significant decline in agricultural production, yield, and distribution. Human activities, such as overgrazing, desertification, soil erosion, and deforestation, also contribute to the unsustainable agricultural goals and targets set by the government and the United Nations. The prevalence of mycotoxins continues to be a challenge and recent findings suggest a change in fungal diversity and mycotoxin profile in some regions of Africa as new mycotoxins are emerging. 3 Variations in the climatic conditions, planting period, and agricultural practices engaged can impact the distribution of fungal species and infection, and the mycotoxin contamination that is associated with grains.4,5 Despite an increase in the amount of land allocated to grain cultivation, this expansion has not translated into higher yields due to the detrimental impact of environmental factors and inadequate crop management practices, and lack of will from the government.
Nigeria needs an estimated 50% increase in maize production to meet growing demand over the next few years. The highest production of maize is from the Northern region of Nigeria, where the savannas have the favorable conditions required for maize growth.6,7 Efforts made to ensure food security in developing countries need to be assessed in terms of their impact on food security and the reduction of poverty. The climatic conditions in Nigeria, characterized by high temperatures and humidity, provide an environment conducive to the growth of toxigenic fungi in staple foods, particularly grains. 8 It is estimated that approximately 25% of cereals consumed globally are infected by these fungi, and recent research suggests that this percentage may be higher. 9
To maintain the quality of grain and grain products for consumption, it is crucial to have surveillance and monitoring systems in place for mycotoxigenic fungal proliferation and mycotoxin contamination. 10 With annual losses due to mycotoxins estimated at 25% of total production, reducing these losses would have a significant impact on the economy. While existing literature has extensively explored the mechanisms of action of mycotoxins within cells, the pathogenesis of the toxins in susceptible hosts, and the cumulative effects in animals, 11 limited information is available regarding the prevalence of toxigenic fungi in the Central-Northern States of Nigeria, which are responsible for the highest production volume of cereal crops in Nigeria. The research identified the predominant fungi found in maize from the maize growing belt of Nigeria, their relative abundance, and the community structure of the fungal population; and determined the genetic relatedness among the toxigenic isolates. The study of the diversity and relationship of fungi is important to gain a deeper understanding of fungal species destroying crops in Nigeria.
Materials and Methods
Collection of maize samples and isolation of fungi from grains
Maize grains under storage (4-8 weeks) in bags totaling 600 (306 white maize and 294 yellow maize) samples were collected from 10 local government council areas in Nasarawa, Kogi, Benue, Plateau, Niger, and Kwara States respectively. Sixty composite samples of 10 sub-samples of 2 to 3 kg grains were randomly collected. Four grains were inoculated at equal distances on Potato Dextrose Agar plate at 25°C between 3 and 5 days. Phenotypic characteristics of observed colonies were recorded, after which cultures were purified by subculturing, and fungal mycelia were viewed under the microscope to check for conidia formation and septation of the hyphae.
Extraction and identification of fungi from maize grains
A sterile toothpick was used to add a small piece of mycelia from a young culture to a 1.5-mL Eppendorf tube containing 500 µL of lysis buffer (composed of 400 mM Tris-HCl [pH 8.0], 60 mM ethylenediaminetetraacetic acid [EDTA] [pH 8.0], 150 mM NaCl, and 1% sodium dodecyl sulfate). The mycelia lump was disrupted and the tube was left at room temperature for 10 minutes. About 150 µL of potassium acetate (pH 4.8) was added to the tube, which was prepared by combining 60 µL of 5 M potassium acetate, 11.5 µL of glacial acetic acid, and 28.5 µL of distilled water. The tube was vortexed and then subjected to centrifugation at 10 000g for 1 minute. The resulting supernatant was carefully transferred to another 1.5-mL Eppendorf tube and subjected to another round of centrifugation using the same parameters as before. Once the supernatant had been transferred to a fresh 1.5-mL Eppendorf tube, an equivalent amount of isopropyl alcohol was added. The tube was then mixed by inverting it and spun at 10 000g for 2 minutes after which the liquid portion was removed and discarded. After centrifuging the resultant DNA pellet at 10 000 rpm for 1 minute, it was washed in 300 mL of 70% ethanol and the supernatant was discarded. The DNA pellets were air-dried and reconstituted with 50 mL of deionized H2O. The purified DNA (1 mL) was used in 25 to 50 mL of PCR mixture.
The 18SrRNA/ITS sequence was used for the identification of fungal isolates, adopting the modified methods of Orole et al. 12 For Aspergillus, Penicillium, and Talaromyces, the multiplex polymerase chain reaction (mPCR) was carried out in a reaction mix of about 45 µL, while identification of Fusarium was in a reaction mix of 35 µL. The ABI 9700 Applied Biosystems thermal cycler was used for the mPCR. The reaction mixture contained MgCl2 (50 mM) (2.5 µL), 10× buffer (2.5 µL), primer (2.7 µM)—0.9 µL, dNTPs (8 mM), 0.5 µL Taq DNA polymerase, and 8, 5, and 3 µL of DNA template for Aspergillus, Fusarium, and Penicillium, respectively (Table S1: Supplemental File). The amplification procedure took 5 minutes at 95°C (initial denaturation) for step 1, 1 minute at 94°C (35 denaturation cycles) for step 2, annealing for 30 seconds at 57°C, 65°C, and 64°C for Aspergillus, Fusarium, and Penicillium, respectively, and extension at 69°C for 30 seconds. The amplification ended with a final extension step at 69°C for 7 minutes for Aspergillus and Fusarium, and 10 minutes for Penicillium. For PCR amplification, 1 µL of purified DNA was added to a PCR mixture containing 25 to 50 µL (Table S2: Supplemental File). Amplification was carried out with primers ITS1 (5′-CTTGGTCATTTAGAGGAAGTAA-3′) and ITS2 (5′-GCATCGATGAAGAACGCAGC-3′). In addition, the Euk1a (5′-CTGGTTGATCCTGCCAG-3′) and Euk516r (5′-ACCAGACTTGCCCTCC-3′) primers were employed to PCR amplify the DNA. The bioinformatics software Geneious, version 8.1, was then used to edit, align, and sequence the DNA.
Determination of toxigenic regulatory genes in fungal isolates
The multiplex polymerase chain reaction (mPCR) was carried out in a 50 µL reaction mix using the ABI 9700 Applied Biosystems thermal cycler. The reaction mix contains 0.9 µL of each primer (2.7 µM) (Table 1), 2.5 µL MgCl2 (50 mM), Taq DNA polymerase (0.5 µL), 10× buffer (2.5 µL), 8 mM dNTPs, and 3 µL DNA template. Amplification was at an initial denaturation of 95°C for 5 minutes, followed by 35 denaturation cycles at 94°C for 1 minute, annealing at 63°C for 30 seconds and extension at 68°C for 30 seconds, and a final extension step at 68°C for 10 minutes (Tables S1 and S2: Supplemental File).
Primer and oligonucleotide sequences for the identification of fungal species isolated from maize grains and toxigenic isolates.

Phylogenetic relationship among toxigenic fungal species
The evolutionary distances between the toxigenic fungi were computed using the Maximum Composite Likelihood method and described in the units of the number of base substitutions per site. All ambiguous positions were removed for each sequence pair (pairwise deletion option). Evolutionary analyses were conducted in MEGA X. 13
Restriction fragment length polymorphism
Restriction endonucleases target different nucleotide sequences in a DNA strand and therefore cut at different sites into short pieces with fingerprints specific to a particular isolate or strain. Restriction Fragment Length Polymorphism (RFLP) treatment was carried out on 63 Aspergillus and one Fusarium strains using restriction enzymes (PstI, XhoI, XbaI, EcoRI, SmaI, and NedI) of each PCR amplicon (ITS, 18SrRNA - Euk1a). The enzyme cuts were performed for 16 hours at 37°C. Following the cutting process, the products were electrophoresed in 2% agarose gel and visualized in the UV system. One percent (1%) agarose gel was used to resolve DNA fragments produced by digesting the DNA with a restriction enzyme. This was prepared by combining 2 g agarose in 10 times concentration of tris-borate ethylene diamine tetraacetate (10 mL 10XTB-EDTA) buffer and 90 mL sterile distilled water in 250 mL beaker flask and heating in a microwave for 2 minutes until the agarose was dissolved. Exactly 0.7 µL of ethidium bromide was added to the dissolved agarose solution with swirling to mix. The DNA was loaded onto the wells of the gel at a concentration of 10 µL, the mini horizontal electrophoresis gel setup was covered, and electrodes were connected. Electrophoresis was carried out at 100 to 200 mA for 1 hour. After electrophoresis, the gel was removed from the buffer visualized under UV light, and documented. 14
Results
Fungal population in maize samples collected from the North Central States of Nigeria
A total of 389 maize samples (64.83%) harbored fungal species (Table 2). Kogi State had the highest number of fungal contamination (89%) of the maize grains. White maize samples had 59.5% fungal infection, while yellow maize seeds of 294 samples had 207 maize seeds (70.41%) contaminated with fungi. Yellow maize from Kogi State had all samples collected infected (100.0%), while the least infected were samples collected in Benue State (50.0%). White maize seeds were most infected in Plateau State (84.2%) and the least infection was from Niger State with 35.3% seed samples. Samples collected from Benue and Nasarawa States respectively were not statistically significant at P ⩽ .680 and P ⩽ .112 respectively.
Fungal population of stored maize in the collection state, North Central Nigeria.
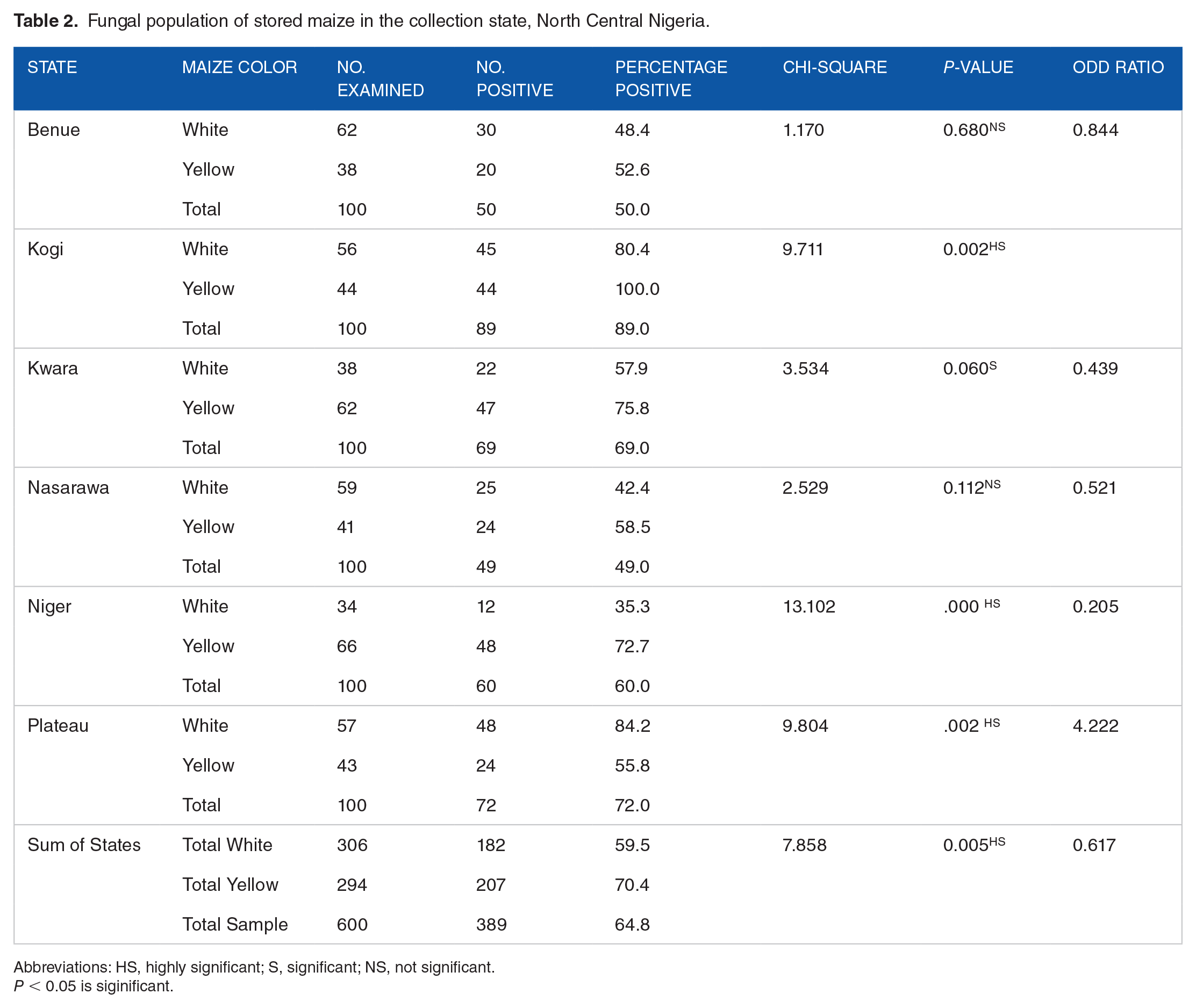
Abbreviations: HS, highly significant; S, significant; NS, not significant.
P < 0.05 is siginificant.
Relative abundance and diversity of the fungal isolates
Comparing the fungal community structure, fungal genera relative abundance (%) was higher in Benue and Kogi States respectively. The fungal genera relative abundance (Figure 1) revealed that Aspergillus was the commonest genus followed by Mucor in the 6 States where samples were collected. Aspergillus was 76.6% in Benue State, 76.3% in Kogi State, and 53.1% in Niger State, while Plateau State had the least relative abundance of 16.3%. In Nasarawa State, 37% of the species isolates belong to the genus Trichophyton. In Figure 2 which showed the relative abundance of the fungal species, Aspergillus flavus and A. niger were the most predominant isolates obtained from the maize grains across all the States.

Genus bar chart of the relative abundance (%) of the predominant fungi in maize grains.

Relative abundance (%) at the species level of the predominant fungi in maize grains.
The alpha diversity analysis (Shannon diversity index) revealed that Kwara State (1.711) had the highest fungal diversity (Figure 3), while the lowest was at Kogi State with 0.693 (P < .05, student t-test).

Boxplot of the Shannon Diversity Index across the sample collection States.
Toxigenic fungal species isolated from maize seeds in north central Nigerian States
In Benue State, 11 of the isolates (4.6%) had genes that encode mycotoxin production (Table 3), while Nasarawa and Niger States respectively had the least number of toxigenic fungi (4 each). Of the 238 suspected mycotoxigenic fungi isolated, 39 of them were confirmed to have genes that encode mycotoxin synthesis.
Distribution of toxigenic fungi isolated from maize grains from the collection states.

Abbreviations: n, number of suspected toxigenic fungi from each state; N, total number of suspected toxigenic fungi.
Genetic relationship between the toxigenic fungi
The phylogenetic tree (Figure 4) showed 9 clades of fungal species and 4 outgroups. Clades 8 and 9 showed minimum sequence divergence, while clades 6 had no divergence with a bootstrap value of 100%. This analysis involved 28 nucleotide sequences. There were a total of 1008 positions in the final dataset.

Molecular phylogenetic analysis of the 18SrRNA gene by the Maximum Composite Likelihood method.
The individual genetic diversity of the different Aspergillus species was studied using the RFLP pattern. The software allowed for cluster analysis and to create a similarity matrix which was presented as a dendrogram. The tree showed similarities of the RFLP patterns in the analysis by grouping the fingerprints of the Aspergillus species sharing common bands into a subset. Patterns (dendrogram derived by MegaX software from RFLP) analysis of 62 Aspergillus species PCR amplicons following agarose gel electrophoresis (Figure 5 and Supplemental File 1—Figure S1). The isolates were numbered from 1 to 62 of specific species as identified by the 18SrRNA ITS region. Six major clusters were identified by the MegaX software according to different patterns of enzyme digestion.

Dendrogram of the individual genetic diversity of the different Aspergillus species.
Discussion
Maize is an important crop worldwide; and contains a large number of substrates that are necessary for fungal metabolism and development. This study, conducted in the North Central States of Nigeria identified 7 major fungi genera isolated from stored maize seeds (Figure 1). The findings are consistent with previous studies conducted by Giorni et al, 15 Harish et al, 16 Katati et al, 17 and Price et al 3 which reported the presence of Fusarium, Aspergillus, and Penicillium as the primary genera isolated from maize grain. The growth of these isolated fungi is influenced by such factors as the temperature of storage and storage duration, and the degree of fungal contamination before storage, as well as insects and mites’ activities all have an impact on the development of these fungi. 18 Poor and unhygienic storage conditions encourage the colonization of maize grains by fungi which leads to seed deterioration and loss of viability. The ability to colonize and infect different parts of the plant and host susceptibility is influenced by water activity (aw), 19 co-inoculum, and inoculum temperature. All these contribute to determining the frequency and type of fungi present in maize seeds. These fungal genera are capable of producing mycotoxins and are active both at the pre-harvest and post-harvest stages along the food chain.
The growth and dissemination of fungi during the pre- and post-harvest periods are conditioned by the environment. The species of fungi that dominate crops are influenced by several critical factors including water activity and temperature. 20 These factors have a direct consequence on the growth of fungi, as well as their production of mycotoxins. 21 Fungi tend to multiply rapidly in the presence of high temperature and humidity. 22 Unhygienic methods of drying seeds on floors also provide a gateway for fungi to infect the seeds. Furthermore, late harvesting can aid the spread of fungi, as the conditions on the farm such as diurnal changes in temperature and humidity are conducive to their rapid growth.23,24 The co-existence of fungal species on maize grains was confirmed in this study. The production of mycotoxins is an associated response to the shared relationship between co-existing fungal species. Furthermore, it was found that the growth and production of toxins by these fungi have a direct impact on microbial competition and fungal pathogenicity toward the host.25 -27 The study noted that yellow maize samples were more prone to fungal contamination compared to white maize (Table 2). This could be attributed to either their higher sugar content and/or greater susceptibility to infection or both. The predominant fungal genera found in stored maize were Aspergillus, Fusarium, Penicillium, and Mucor. These fungi are known to contaminate cereals, fruits, and vegetable crops under unfavorable storage conditions. 28 The storage methods used by farmers might have led to an increase in the number of fungal contaminants identified as producers of mycotoxins. 29
The mycotoxin regulatory genes were located in the DNA of the fungi. The regulation of these genes could be triggered when the fungi are under stress leading to the production of mycotoxins. The presence of the regulatory gene is interpreted to mean that the fungal species are implicated in the production of mycotoxins (though in varying amounts) in the fungi-infected seeds. This is a concern because mycotoxins such as aflatoxins, ochratoxin A, and fumonisins are known carcinogenic substances, zearalenone and its metabolites have estrogenic effects, and hematogenic metabolites include trichothecenes such as deoxynivalenol (DON), T-2 toxin, and HT-2 toxin as classified by the International Agency for Research on Cancer. 30 Prevention of fungal proliferation and toxin production depends on the application of good post-harvest practices that can significantly reduce fungal colonization and maintain crop quality, nutritional status, and integrity.31,32 When it comes to identifying Aspergillus and other medically significant fungi, the PCR-RFLP (Polymerase Chain Reaction-Restriction Fragment Length Polymorphism) technique is the preferred option, though the study recognizes that morphological identification is not only complementary but an essential step preceding molecular studies of microorganisms. This is due to several disadvantages associated with morphological identification. These include the delayed production or absence of unique morphological traits on culture media, the absence of sexual reproductive cycles in certain Aspergillus species, which can make them difficult to identify, and the presence of similar microscopic or macroscopic features at the species level. The study revealed the distinctiveness of mycotoxins type produced in the North Central States of Nigeria (Benue, Niger, Nasarawa, Kogi, Kwara, and Plateau States respectively) and their associated pathogenic fungi. Though the species richness was high across the States together (Aspergillus—210, Mucor—114, Fusarium—45, Trichophyton—37, Penicillium—18, and Talaromyces—2), the diversity was generally low (Shannon Diversity Index 0.519-1.711) of the fungal community from the States (Figure 3). Aspergillus niger and A. flavus were the 2 most common fungal species reported in this study which disagreed with the report by Joshi et al 33 who had Fusarium species as the most prevalent in their study. Globally in 2019, data from January to December showed the most prevalent mycotoxins were Fumonisins (FUM) (70%) and Deoxynivalenol (DON) (68%) 34 produced by Fusarium species
This study used restriction fragment length polymorphism (RFLP) analysis of the ITS, actin, and β-tubulin regions of PCR-amplified ribosomal DNA to determine the relationship among the toxigenic fungi. Restriction enzymes recognize a set of nucleotides at a restriction site and cleave DNA at that site, exhibiting an RFLP pattern during electrophoretic separation of digested DNA and producing cut fragments of varying lengths characteristic of the DNA sequence. The findings of this research were congruent with those of previous studies that were tailored to the PCR-RFLP approach by employing the restriction enzyme MwoI to differentiate medically significant Aspergillus species among 153 (75%) environmental strains and 52 (25%) clinical strains. The data we have gathered corroborates the notion that Aspergillus species are prevalent and widespread in various environments, including soil, water, air, seeds, and even food. 35 The presence of Aspergillus species as the most prevalent fungi from the states calls for concern. The fungi remain the primary cause of invasive aspergillosis, and species like Aspergillus flavus, Aspergillus terreus, Aspergillus niger, Aspergillus niger, A. ustus, and A. versicolor have been linked to human diseases.36,37 As such, it is crucial to accurately and promptly identify the species present for effective impact assessment management, as well as for surveillance and epidemiological purposes. Aspergillus species can be found across the globe as they can grow in any climate.
The dendrogram pattern showed that different Aspergillus species had identical RFLP patterns (Figure 5). Despite this diversity in the Aspergillus species, they tended to cluster not only based on the same species or state of origin. This study showed the presence of remarkable similarity among the different Aspergillus species. Six of these fragments, namely PstI, XhoI, XbaI, EcoRI, SmaI, and NedI, were found to possess a significant degree of nucleotide-level similarity to A. fumigatus. The reason for the heightened precision of these 6 fragments in contrast to the 1,500 bp DNA ladder EcoRI can be attributed to their distinct positioning within the retrotransposon of the enzyme. This is due to the presence of some stop codons within the sequence. The RFLP restriction sites observed in this study were separate and dissimilar.
A. welwitschiae BW1 (NR111348.1) from Benue State and A. niger KN1 (OQ476657.1) from Kogi State had the least sequence divergence among the species. The most evolutionary distant strains were clade one sister fungi made of A. niger NN1 from Niger State and A. flavus BF2 from Benue State (Figure 4). These organisms are more likely not to share epidemiological associations with the other Aspergillus strains because they are phylogenetically distant from the others. Evolutionary distance allows for modifications and changes responsible for diversity. The results in the study suggest that most of the recurrent clusters are due to competition for substrate and pathogenicity, and are likely due to endogenous colonizing strains. Little is known about the establishment patterns of Aspergillus species. This study indicates that the grains were contaminated with genetically unrelated strains. Analysis of genetic distances among the isolates based on location shows that isolates are closely related genetically. The observed genotypic diversity is due to random mutations and environmental variation and stress. Genetic diversity ensures the survival and continuity of species.
Conclusion
The isolated fungal species have the potential to introduce mycotoxins into the maize grains in North Central Nigeria. As one of the world’s most crucial food crops, maize is highly susceptible to contamination by mycotoxin-producing fungi during storage and in the field. This presents a grave threat to food safety. The study’s discoveries indicate that the recorded fungal contamination might be a result of poor agricultural practices by middlemen, retailers, and farmers, who engage in mostly unsanitary methods. It is speculated that these contaminations could have occurred during the harvesting, processing, storage, or transportation of the maize. The study showed a phylogenic relationship existing between the fungal species isolated from maize across the states which is likely responsible for the multiple mycotoxins contaminations observed in the study. The study confirmed the presence of mycotoxin regulatory genes co-existing in some of the grains, which point to the need for improved post-harvest management practices by farmers, and government intervention through continued advocacy and enlightenment programs for the farmers. The study will serve as an early warning mechanism for maize producers and respective consumers in the sampling areas, which are the highly prone areas, to avert acute poisoning that, could lead to fatalities. The research showed there is a phylogenic relationship between fungi isolated from maize samples from the North Central state of Kogi, Kwara, Niger, Nasarawa, Plateau, and Benue Nigeria between 2021 and 2022. The study recommends that hygienic practices and HACCP be employed by farmers and marketers dealing with maize grains and their products.
Supplemental Material
sj-docx-1-ehi-10.1177_11786302241249858 – Supplemental material for Epidemiology and Genetic Relationship of Toxigenic Fungi in Maize Grains From North Central Nigerian States
Supplemental material, sj-docx-1-ehi-10.1177_11786302241249858 for Epidemiology and Genetic Relationship of Toxigenic Fungi in Maize Grains From North Central Nigerian States by Orole Olukayode Olugbenga and Mantu Eno Chongs in Environmental Health Insights
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Author OOO - Conceptualization, Project administration, Software, Supervision, Writing – original draft, Writing – review & editing; Author MEC - Funding acquisition, Formal analysis, Investigation, Resources, Validation, Writing – review.
Ethical Approval
Not applicable.
Availability of Data and Materials
Not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
