Abstract
Antibiotic-resistance genes carried by coliforms in drinking water is a concerning issue for public health in Bangladesh. This research was carried out to identify coliforms in drinking water and to understand the importance of the
Introduction
Availability of drinking water is one of the main prerequisites for healthy life (Fawell & Nieuwenhuijsen, 2003) and safe drinking water is also considered a universal human right by the United Nations Conventions (Gorchev & Ozolins, 2011; World Health Organization, 2011). However, the availability of safe drinking water is still considered a dream for many people living in Asia, South America and Africa (Rubino et al., 2019). Access to clean, contamination-free water is also necessary to maintain an improved public health (Mina et al., 2018).
Generally, the population of Chattogram City, Bangladesh consumes drinking water as safe water in homes, hospitals, restaurants, vendors, educational institutions, offices, etc. But the presence of microorganisms in drinking water has attracted great attention worldwide because of public health problems (Olaoye & Onilude, 2009). According to the World Health Organization (WHO), approximately 1.1 million people globally consume impure water and 88% of diarrheal diseases in the world are the result of the consumption of contaminated water (World Health Organization, 2003). Ten major waterborne diseases have also reported to be responsible for more than 28 billion disease episodes yearly in developing countries (Suthar et al., 2009). It is very concerning that Chattogram city, Bangladesh, has also been suffering from various water-related problems along with insufficient drinking water supply (Zuthi et al., 2009).
Coliform has generally been used as an indicator of water quality, and its presence in drinking water is also used as an index of the presence of entero-pathogens in water (Rompré et al., 2002). The total number of fecal coliforms should be zero in drinking water (World Health Organization, 2004). Among coliforms,
The antibiotic resistant gene- integron, an integrase gene (
In developing countries like Bangladesh, antibiotic resistance is a severe threat due to the widespread misuse of antibiotics, inappropriate regulations, poor effectiveness of drugs, etc. (Ayukekbong et al., 2017). This resistance emerges in both pathogenic microbes and coliform like
Materials and Methods
Selective plating
The MacConky agar medium (HI media) was prepared in 250 ml conical flask according to the manufactured protocol. After the sterilization process was completed, the medium was cooled to around 40°C. Meanwhile, the water samples were properly vortexed and 1 ml of sample was poured into the Petri dish. The prepared MacConky media was poured into the same Petri dish and then the plates were stirred gently clockwise and anticlockwise with a hand, so that the samples were thoroughly mixed with the medium. The Petri dishes were allowed to solidify. After solidification, the plates were kept in the incubator in an inverted position at 37°C overnight. After incubation, a characteristic colony of coliforms was observed and the selected colony was picked and stored for biochemical confirmation. Same process was done for two times to get repetitive results.
Biochemical identification
The prepared TSI medium (Himedia, M021-500G) in conical flask was placed in a sterile test tube and the opening of the tube was closed with a cotton gauge. The test tube was then autoclaved. After autoclaving, the test tube was kept in a tilted position for 2 to 3 hours for solidification. By microwire sterile loop, the bacterial single colony was inoculated on TSI slant and loop was dipped into butt carefully. In the case of every suspected sample, the same procedure was followed. The TSI test tubes were incubated into an incubator at 37°C for overnight. The TSI test tubes within an acidic (yellow) slant and an acidic butt with gas were suspected for coliforms. These suspected samples were selected for DNA isolation and molecular identification.
DNA extraction and PCR
The genomic DNA extraction of the isolated coliforms was performed according to the previously published protocol (Queipo-Ortuño et al., 2008). PCR amplification of the target genes (Table 1) was performed using a thermal cycler (NyxTechnik) (Mehdi et al., 2018; Mina et al., 2018)). After amplification, the PCR products were analyzed by gel electrophoresis at 1.5% agarose.
Target Genes, Primers, Cyclic Condition, Composition of the PCR Mixture, and Amplicon Size.

Forward primer; bReverse primer.
Antibiotic susceptibility test
Kirby-Bauer disk diffusion method with McFarland standard was followed for the antibiotic susceptibility test (Andrews, 2001; Wiegand et al., 2008).In this study, 0.5 McFarland turbidity standard contained 0.5 ml of 1.17% barium chloride and 9.5 ml 1% sulfuric acid (Andrews, 2001). A loopful of bacterium colony was taken into a 5 ml sterile double distilled water containing test tube. The bacterium turbidity concentration was compared with the McFarland turbidity standard. The cotton swab was then dipped in distilled water containing the bacteria and spread on Mueller Hinton agar plate. Three antibiotic disks (ampicillin, Ciprofloxacin, and co-trimoxazole) were placed on the culture plate carefully. Plates were kept in refrigerator at 4°C for 20 minutes so that the antibiotic could diffuse. The plates were then incubated for overnight at 37°C in the incubator. Finally, after incubation, the inhibition zone against each antibiotic disc was measured.
Results
Screening of coliforms in drinking water
This research project has been designed to investigate the occurrence of coliform bacteria in the drinking water of different public places in Chattogram city, Bangladesh (Table 2). To identify coliform in drinking water, collected water samples were cultured in MacConkey agar by the pour plate technique. Coliforms are a group of Lactose-fermenting bacteria and usually grow as pink colonies on the MacConkey agar plate. The pour plating of the water sample on MacConkey agar revealed the occurrence of coliform bacteria in 10 of the 31 drinking water samples collected as clear pink colonies were observed among the plates prepared by those samples. The samples numbers were S3, S6, S11, S12, S13, S15, S17, S23, S30, and S31 (Table 2, Figure 1A). To identify coliforms from suspected samples, a single pink colony of each plate positive for coliforms has been collected and performed the Triple Sugar Iron test (TSI). TSI agar is a differential medium that can assess the ability of a microbe to ferment lactose, which is used for the identification of coliform bacteria. In TSI, slant coliform only generated gas and the entire media remained acidic (yellow slant and yellow butt) indicating lactose fermentation. In the TSI test, suspected isolates were confirmed as coliform as they showed a specific coliform result (Table 2, Figure 1B). After the confirmation of selected isolates by TSI, the findings have also been confirmed by molecular analysis. For molecular confirmation, the presence of the
Summarized Information About the Screening of Coliforms From Drinking Water.

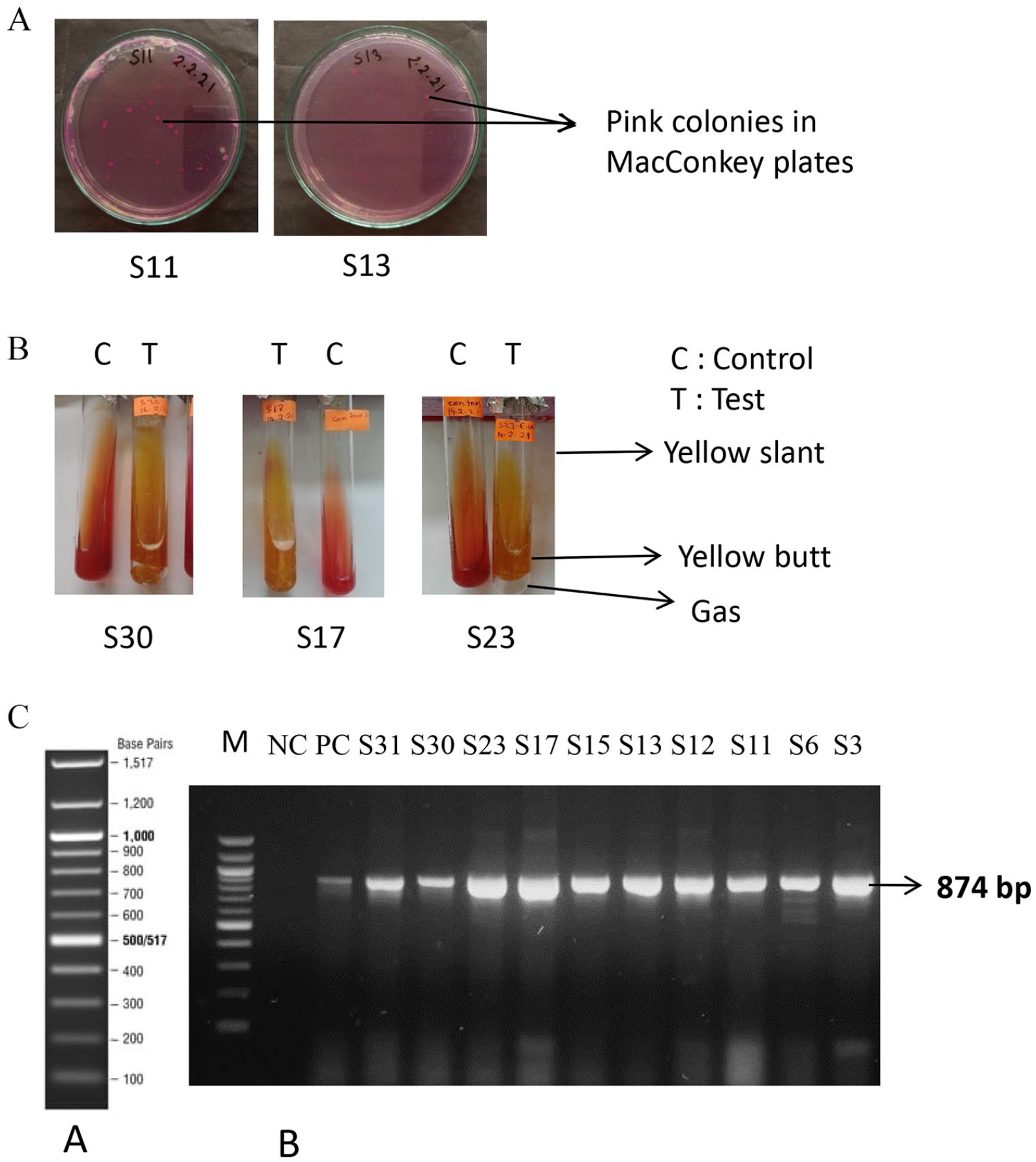
Screening of coliforms from drinking water: (A) selective plating of water sample in the MacConkey agar plate. Characteristic pink colonies of suspected coliforms indicated by arrows. Representative plates of some selective samples have been shown, (B) biochemical identification of suspected coliforms by TSI test. Pink colonies from MacConkey agar were inoculated on TSI slants. After incubation coliforms were identified through the observation of yellow (acidic) slant and yellow (acidic) butt with gas in the TSI slant. Representative TSI slants some selective coliforms have shown, (C) molecular confirmation of biochemically identified coliforms. A; Ladder, B: The target lacZ gene was amplified through PCR by using specific primers. After the completion of PCR, the PCR products were visualized through agarose gel electrophoresis. M = Marker; NC = Negative control; PC = positive control.
Investigation of fecal contamination in drinking water
Identifying coliform implies whether or not fecal contamination occurred in these water samples. Screening of fecal contamination was performed by amplification of the

Identification of fecal coliforms. (A) Ladder, (B) The target
Screening of antibiotic-resistant gene integron (int1 ) in coliforms
Antibiotic resistance is one of the main health concerns in Bangladesh. Therefore, the main objective of this study was the screening of antibiotic resistant gene

Detection of
Antibiotic resistance profiling of identified coliforms
Antibiotic resistance in different environments is quite common in Bangladesh. Furthermore, isolated coliforms carrying the

Antibiotic resistant profiling of isolated coliforms: (A) Antibiotic sensitivity test by Kirby-Bauer Disk diffusion method. Picture of culture plates showed zone of inhibition by
Discussion
Bangladesh is one of the most densely populated countries in the world and has a severe scarcity of safe drinking water. Most of the total population of Bangladesh has access to water, but the quality of water is always questionable. Coliform bacteria are considered the best indicator of bacteriological quality of drinking water. In order to understand bacterial contamination in drinking water in Chattogram City, Bangladesh, during this study the prevalence of coliform bacteria was determined by collecting 31 drinking water samples from hotels, restaurants and food stores (
Antibiotic resistance and its widespread occurence in and around Bangladesh is a major public health concern. Recently, a study has reported the occurrence of multi drug resistance coliforms in human sewage collected from different places of Chattogram city, Bangladesh (Akter et al., 2021). As a consequence, in this study we also investigate the contribution of coliform bacteria from drinking water to the occurrence of antibiotic resistance in Bangladesh. Surprisingly, in this study we found the presence of a highly important antibiotic resistance gene
Footnotes
Acknowledgements
The authors are thankful to the teachers and staffs of the Department of Genetic Engineering and Biotechnology, University of Chittagong, Bangladesh.
Author Contributions
A. M. M. A. C designed this study. SAM, AKMZH, P.D., M.ZH performed the experiments; A. M. M. A. C analyzed the data; A. M. M. A. C., S. A.M. and A.K.M. Z.H wrote the original draft. A. M. M. C and S. A. M reviewed the several versions of the manuscript. All authors read and approved the final manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Data Availability
All data analyzed during this study are included in this manuscript.
