Abstract
RNF20 is a RING-domain E3 ubiquitin ligase that mediates monoubiquitination of histone H2B at lysine 120 and plays important roles in transcriptional elongation, repair of DNA double-strand breakages, chromatin homeostasis as well as tumor suppression. Even though RNF20 has been investigated in some cancer types, its global biological functions and other related mechanisms in diverse cancers are yet to be fully understood. To investigate the systematical evaluation of the pan-cancer expression pattern of RNF20 and its potential to serve as a diagnostic and prognostic biomarker, we first pulled together the information of several publicly available databases and bioinformatics tools. We compared association between RNF20 expression and important clinical and molecular variables—including clinical survival outcomes, immunomodulatory molecular, tumor stemness index, prognostic use, immunomodulatory molecular, genomic and immunological variables, immunotherapy response, response to chemotherapeutic agents, and functional enrichment profiles. The levels of RNF20 were also substantially greater in tumor tissues relative to identical non-tumor tissues; furthermore, the high levels of RNF20 in some cancer tissues are associated with the good prognostic outcome. The subsequent studies demonstrated that RNF20 is highly correlated with the immune checkpoint molecules, immunomodulatory genes, prognostic biomarkers, tumor stemness scores, as well as immune cell infiltration in various tumors. The SN-38 and talazoparib were identified as possible RNF20-targeting agents through drug susceptibility analysis. It was revealed that missense mutations are most frequent type of variations, which is revealed by functional enrichment analysis, mutation profiling, RNA modification mapping and analysis of genomic heterogeneity of the RNF20 related genes within the framework of glioblastoma, glioma, low-grade glioma in addition to thyroid cancer. Collectively, these results support the exculpatory role of RNF20 in the treatment of different types of cancer, which could help to certify the usage of this biological molecule as a predictive biomarker of treatment response.
Keywords
Introduction
According to the International Agency for Research on Cancer (IARC) estimates, the world might have recorded 19.96 million new cancer cases and 9.73 million cancer-related deaths in 2022 contributing to the global cancer burden. Approximately 14.5% of deaths in the globe are due to cancer. 1 Regardless of the current improvements in the diagnostic and treatment plans of cancer the survival rate in general with respect to cancers is low, 2 and the prognostic response does not meet the expectations in the clinical context. 3 Based on that, it is crucial that new methods of cancer treatment and diagnosis are designed as soon as possible. Currently, the main cancer treatment methods include the surgical intervention, chemotherapeutic agents, radiotherapy, molecular targeted therapy as well as immunotherapeutic treatment. Cancer biomarkers have received immense interest, and an increasing number is being discovered. 4
RNF20, which is a sub-unit of the hBRE1/RNF20-RNF40 E3 ubiquitin ligase complex, is the methionine of histone H2B that is specifically monoubiquitinated at lysine 120 (H2Bub1).5,6 RNF20 through this histone modification regulates DNA damage response, chromatin remodeling and transcriptional regulation- which is crucial in the establishment of the genomic stability.7,8 RNF20 silence has also been demonstrated to disable the DNA damage repair ability of lung adenocarcinoma cell. 9 RNF20 and RNF40 high expression are associated with an increased invasiveness of cancer cells and unfavorable patient outcome in cervical cancer, which is mediated by ferroptosis resistance. 10 Even though literature on RNF20 has concentrated on single types of cancer, there are no studies that are comparing different types of cancer. Pan-cancer analysis Pan-cancer analysis provides a global framework of cancer-related data analysis through the analysis of patient genomes in many different cancers, to recognize common and unique features of molecular profiles. In the last 10 years, immune cell infiltration has become an important predictor when it comes to determining both prognostic factors and effectiveness of treatment of cancer patients. 11 Some forms of cancer are defined by the expression of immune checkpoints- the major molecular processes, which regulate immune processes. 12
To thoroughly evaluate the level of this role of RNF20 in a wide range of types of cancer, we used a pan-cancer study through repositories like the Cancer Genome Atlas (TCGA) and Genotype-Tissue Expression (GTEx) project. The extent of RNF20 expression, its impact on the prognosis, and its possible relationship with its genetic variation and tumorigenesis were evaluated by us. To further study the biological role of RNF20 to tumor response to immunotherapy, we conducted a group of specific analysis experiments. Namely, we paid attention to the relationships between RNF20 and various prominent biological biomarkers such as immune cell infiltration, immune checkpoint-related genes, immune scoring measures, and drug sensitivity. To identify the regulatory mechanisms behind enhancement of tumor development by the RNF20 family, we used the STRING database to conduct gene functional enrichment analysis and protein-protein interaction (PPI) network analysis. The abbreviated forms of tumor names are provided in Table 1.
Abbreviation of tumor name.

Materials and Methods
Gene expression analysis
We obtained normalized pan-cancer RNA-seq and clinical data in the University of California Santa Cruz (UCSC) Pan-Cancer Atlas. TIMER2.0 (http://timer.cistrome.org/)13,14 provided us with the level of expression of RNF20 in different malignant tumors. In addition, to examine variations in its expression between normal and tumor tissues, we performed further analyses on data provided by The Cancer Genome Atlas (TCGA; http://cancergenome.nih.gov) 15 and Genotype-Tissue Expression (GTEx) Program (http://commonfund.nih.gov/GTEx/). 16
Pathological staging and prognostic study
We explored the association between RNF20 and pathological staging. In such a way, we used the Gene Expression Profiling Interactive Analysis 2 (GEPIA2) platform (http://gepia2.cancer-pku.cn / index). 17 The UCSC Xena Functional Genomics Explorer (https://xenabrowser.net/), provided us with the normalized, pan-cancer, gene expression data (PANCAN), which contained 10 535 samples and 60 499 genes. We filtered out the expression values of RNF20 values per sample, and only those were eligible that were defined as primary tumors, or peripheral blood-derived primary hematologic malignancies. To transform these expression data to normalize, we transformed the data using log2 (x + 0.001) formula. To retain an analytical strength, we narrowed down to those types of cancer that had over 3 samples, and expression data in 14 different types of cancer was retained. Pan-cancer overall survival (OS), disease-specific survival (DSS), progression-free interval (PFI), and disease-free interval (DFI) analyses of RNF20 by SangerBox 3.0 were done (http://sangerbox.com/home.html).18,19
Correlation analysis of tumor immune microenvironment
We have examined the expression data of RNF20 and 150 immune-associated signature genes in UCSC, SangerBox 3.0, and R software. The genes were organized into 5 categories of the immune pathways which are; chemokines (41), immune receptors (18), major histocompatibility complex (MHC, 21), immunoinhibitors (24) and immunostimulators (46). We made normalization of the expression values after eliminating normal tissue samples as earlier mentioned. Subsequently we determined the pair wise Pearson correlation coefficients of RNF20 and the marker genes of the 5 immune pathways. Moreover, the RNF20 measurements, as well as DNA methylation data, were merged together to calculate the levels of stemness of each sample in tumors. 20
Immune infiltration analysis
Each individual tumor profile of RNF20 expression was obtained out of the set of TCGA cross-cancers accessed when using the UCSC data repository. The profile of gene expression was matched against Gene Symbols and read using the R based tool ESTIMATE to produce matrix, immune, and composite ESTIMATE scores on each patient.20,21 These scores were used to provide inferences of the extent of immune infiltration on 10179 tumor samples on 44 tumor types. Therefore, to compare the gene expression and immune infiltration, the corr.test R package of the psych (2.1.6) package was used to compute Pearson correlation coefficients between the expression of RNF20 and the immune scores in individual tumor types. TIMER2 platform was used to test the correlation between the RNF20 expression and the infiltration rates of the different cell types of immune cells in all TCGA cancers. Our paper selected a set of algorithms (xCell, EPIC, MCPCOUNTER, TIDE, CIBERSort and CIBERSort-ABS) to ensure the study results.
Immunotherapy analysis
The association between the expression of RNF20 and immune checkpoint blockade (ICB) therapy response were to be analyzed using the KMplot (https://kmplot.com/analysis/). 22 In the meantime, we used TISIDB (http://cis.hku.hk/TISIDB/index.php) 23 to search correlation between RNF20 expression and ICB and cytokine-based therapies.
Drug susceptibility association and molecular docking studies
The Genomics of Drug Susceptibility in Cancer (GDSC) data repository, which is a repository created by the Sanger Institute, was used to obtain drug susceptibility data. This platform provides tumor cellular reactivity profiles of several drugs 24 and supports the identification of therapeutic targets of genomic changes. The degree of susceptibility was determined by calculating half-maximal inhibitory concentration (IC50) values by using the Prophet R package. Gene-drug interactions correlated with susceptibility to the top 30 drugs in the GDSC pan-cancer dataset were summarized and further analyzed those that passed the false discovery rate cutoff (<0.05). The positive associations denote that high levels of expression of gene can foster drug resistance whereas negative relationships imply that genes can be vulnerable to drugs. 25 It was retrieved from the Protein Data Bank (PDB; www.rcsb.org/pdb/home.do) 26 to obtain the 3D structure of the RNF20 protein and received PubChem (https://www.ncbi.nlm.nih.gov/pccompound) 27 to retrieve the 3-dimensional structure of the drugs. The main steps of molecular docking included the pre-processing and pre-refinement of receptor proteins and small-molecule ligands, the generation of grids, and the docking of target compounds. In this study, molecular docking simulations were conducted through the AutoDockTools-1.5.7 program, in accordance with the Schrodinger Glide standard protocol of docking. The most desirable docking conformation put forward had then been selected and ranked in terms of hydrogen bonding, electrostatic forces, and hydrophobic interactions in the formation of ligand-receptor complexes. The docking results were evaluated using the core evaluation index based on the calculated values of binding energy (kcal/mol). Finally, the hydrogen bonding modes, ligand-receptor binding affinities, the amino acid residues that play a relevant role in the interaction and the 3-dimensional structure of the ligand-receptor complex were given using PyMOL software visualization. 28
Pathway and functional enrichment analysis of RNF20-related genes
The STRING online tool (https://string-db.org/)29,30 was used to identify RNF20-binding proteins and conduct PPI network analysis on the related genes. Cytoscape software (v3.9.1) was used to visualize the PPI network and conduct follow-up analyses. Functional enrichment analysis based on Gene Ontology (GO) and pathway analysis using the Kyoto Encyclopedia of Genes and Genomes (KEGG) 31 were performed for RNF20-related genes. Leveraging the “SimilarGeneDetection” bioinformatics tool integrated within the GEPIA2 database, we initially identified and prioritized the top 100 putative target genes of RNF20 exhibiting the most robust correlations. Subsequently, expression profiling data of the aforementioned top 10 key core target genes across various tumor subtypes were retrieved via the TIMER2.0 online bioinformatics platform, with their expression distribution patterns visually delineated through heatmap visualization.
RNF20 mutation profile
Using cBioPortal (http//www.cBioPortal.org/) which features multidimensional genome data on all tumors covered in TCGA32,33 we analyzed the mutation characteristics of RNF20 across different cancers. The categories and frequencies of RNF20 mutations across all tumor types were compiled using bioinformatics analysis tools. RNF20 and 44 marker genes’ expression signatures, among which are genes associated with 3 distinct RNA modification types [N1-methyladenosine(m1A; 10), 5-methylcytosine (m5C; 13), and N6-methyladenosine (m6A; 21)], were retrieved from the UCSC Xena platform. Expression values were normalized via the log2(x + 0.001) transformation, and Pearson correlation coefficients were computed to assess the association between RNF20 and immune pathway-related marker genes. Single-nucleotide variant (SNV) data from TCGA were analyzed using MuTect2 software, and tumor mutational burden (TMB) was computed with the TMB function integrated in the R package maftools. TMB values were then integrated with gene expression data. Figure 1 provides our detailed workflow. The report of this study was written strictly in accordance with the STROBE statement. The details of the relevant compliance verification and the item comparison table are detailed in Attachment.

RNF20 workflow diagram. (A) The left half of the figure labels the site of occurrence for each cancer. The right half delves into 5 aspects of each cancer type: genomic data analysis of the RNF20 gene, clinical analysis, immune characteristics analysis, drug and molecular mechanism research, and gene function and mutation characteristics analysis. The emphasis is on the analysis of differential gene expression, prognostic relationships, tumor immunity, molecular docking simulations, and genomic factors.
Results
RNF20 is overexpressed in various human malignancies
Initially, we evaluated RNF20 expression in multiple cancer types utilizing the TCGA’s comprehensive database (Figure 2A). For identifying differential expression in tumor tissues, we matched normal tissue specimens obtained from GTEx and TCGA’s comprehensive database. RNF20 mRNA exhibited significantly elevated expression in 21 tumor types, including GBM, GBMLGG, LGG, BRCA, LUAD, ESCA, STES, COAD, COADREAD, PRAD, STAD, HNSC, LUSC, LIHC, WT, SKCM, THCA, PAAD, ALL, LAML, and CHOL (Figure 2B). Significantly decreased expression of RNF20 was detected in 8 tumor types: UCEC, KIRP, KICH, BLCA, OV, TGCT, UCS, and KIPAN (Figure 2C).

RNF20 expression in tumor tissues. (A–C) Analysis of RNF20 expression across tumor types in TCGA (*P < .05; **P < .01; ***P < .001) (A), showing significant upregulation in the TCGA + GTEx database (B) and significant downregulation in select tumor types (*P < .05) (C).
RNF20 high levels might be prognostic in isolated types of cancers
The level of RNF20 expression was strongly associated with clinical stage in LIHC, THCA and KIRC (Figure 3A to C). We used UCSC database to provide RNF20 expression in various stages of pathology of various cancers. R software (version 3.6.4) was applied to measure variations of RNF20 expression between subsets by clinical stage in each of the individual tumor types and significant positive correlations found in KIPAN, HNSC and KIRC tumors (Figure 3D). In Table 2, the detailed presentation of the results and supporting data is provided. Also, we performed OS-centered survival analysis and revealed that high expression of RNF20 was also associated with poor prognosis in LAML, LIHC, BLCA, and ACC, and low expression in GBMLGG, KIPAN and KIRC (Figure 3E). DSS analysis also indicated that high levels of RNF20 expression unit were associated with low prognosis in PRAD, BLCA, ACC and KICH, and low levels were predictive of poor prognosis in GBMLGG, KIPAN and KIRC (Figure 3F). DFI revealed that high RNF20 expression was associated with bad prognosis in LUSC, LIHC and ACC, but low RNF20 expression meant poor clinical prognoses in STAD (Figure 3G). PFI analysis PFI indicated that high levels of RNF20 expression were associated with poor clinical outcome in LIHC, BLCA and ACC, whereas low levels of RNF20 expression indicated poor clinical outcome in GBMLGG, UCEC, KIPAN and KIRC (Figure 3H).

Effect of RNF20 expression on pathological stage and prognostic analysis. (A–C) Correlation analysis of RNF20 expression and pathological stage in LIHC, THCA, and KIRC tumors in TCGA. (D) Correlation between RNF20 expression and pathological stage across all TCGA cancers (*P < .05; **P < .01; ***P < .001; ****P < .0001). (E–H) Pan-cancer correlation analysis of RNF20 expression and OS (E), DSS (F), DFI (G), and PFI (H).
Expression data of 14 types of cancer.

Correlational relationship of RNF20 actions with immune-regulatory venerables
We analyzed the relationships between RNF20 and immune checkpoints, tumor stemness indices, and immunomodulatory pathways as a way to investigate immune regulatory capacity of RNF20 in tumors. We tested the level of RNF20 and 150 marker genes that represent 5 categories of immune pathways (chemokine (41), receptor (18), MHC (21), Immunoinhibitor (24), Immunostimulator (46)) and calculated Pearson correlation coefficients in all samples using the UCSC pan-cancer data set. RNF20 had a positive association with a significant percentage of immunomodulatory genes (Figure 4A). We also compared it to 60 genes of inhibitory (24) and stimulatory (36) immune checkpoint pathways and identified significant correlations with most of the checkpoint markers (Figure 4B). After conducting Pearson correlation analysis on each type of tumor, we have found that the RNF20 exhibited a significant positive correlation with tumor stemness scores via THYM but a very strong negative correlation via GBMLGG, BRCA, ESCA, KIRP, TGCT, and BLCA (Figure 4C).
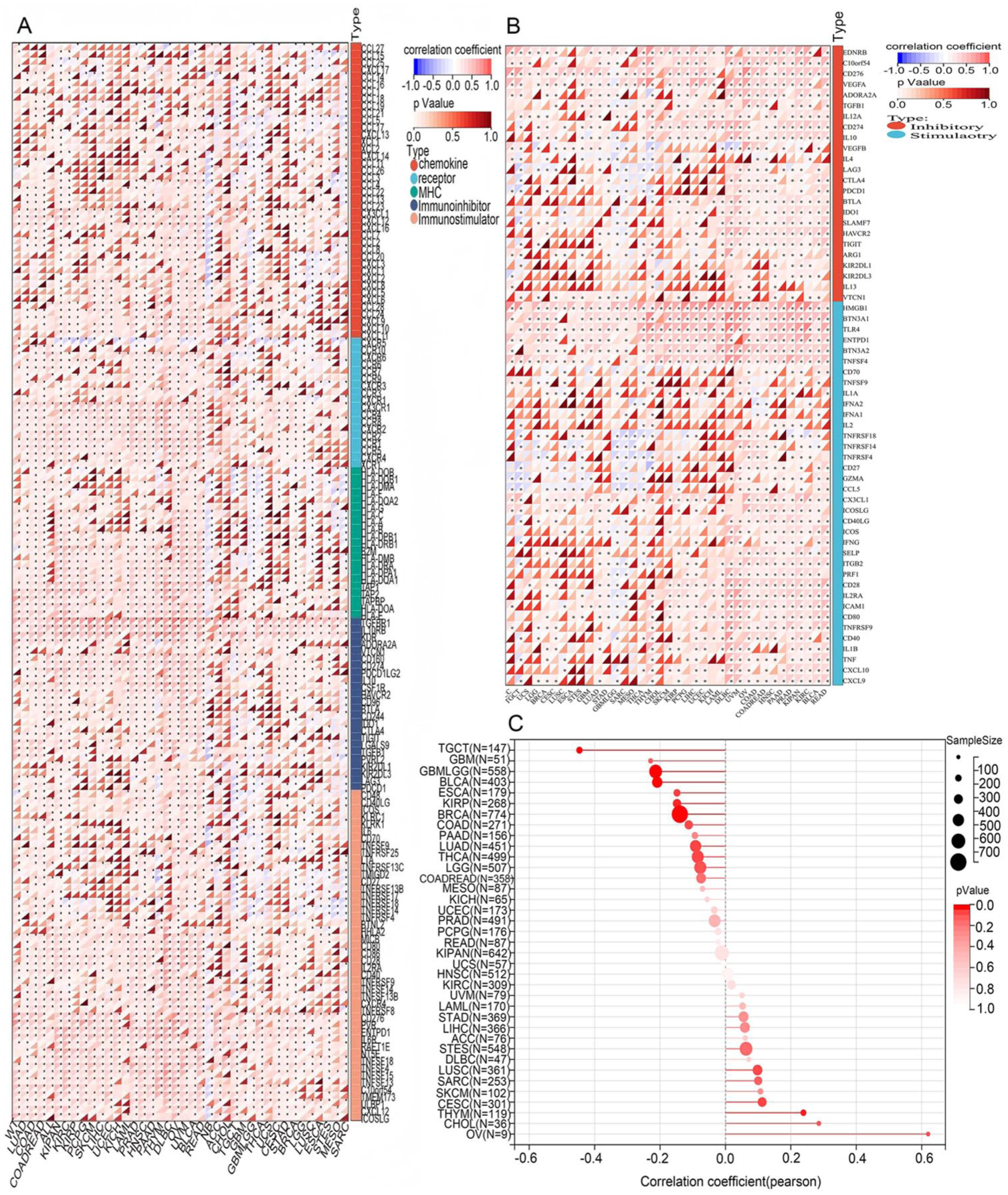
Correlation analysis of RNF20 expression with immunomodulatory genes, immune checkpoints, and tumor stemness scores. (A–C) Correlation analysis of RNF20 expression with immunomodulatory genes (A), known immune checkpoints across all TCGA cancers (B), and tumor stemness scores (C).
Immune infiltration
RNF20 showed a strong correlation with immune infiltration in 22 cancer types (positively correlated in 9 (KIPAN, COAD, COADREAD, KIRC, BLCA, READ, PAAD, TGCT, LAML) and negatively correlated in 13 (GBM, GBMLGG, LGG, UCEC, STES, SARC, LUSC, LIHC, WT, SKCM, THCA, NB, ACC) (Figure 5A to V). The level of immune cell infiltration was evaluated with the help of several computational algorithms, which allowed noting that the infiltration levels of various tumor types depends positively on RNF20 expression but negatively on the infiltration levels of natural killer (NK) cells, cancer-associated fibroblasts, endothelial cells (xCELL), and T cell γdelta and T follicular helper cells (CIBERSORT) (Figure 5W). These results improve the comprehension of tumor-associated immunomodulatory pathways, which is a major factor to consider as a means of improving the efficacy of immunomodulatory events.

Immune infiltration analysis. (A–V) Correlation analysis of RNF20 expression with immune infiltration scores in 22 cancer types. TIMER2 algorithmic analyses of immune infiltration and RNF20 expression in all TCGA cancers (W).
RNF20 expression associated with outcomes of immunotherapy
To determine the therapeutic potential of RNF20 as an immunotherapeutic candidate target used in treating cancer, we compared the OS outcomes between patients with a high or a low expression of RNF20 receiving anti-programmed cell death protein 1 (PD-1), anti-programmed death-ligand 1(PD-L1), or anti-cytotoxic T-lymphocyte-associated protein 4 (CTLA-4) inhibitors. Those who possess upregulated RNF20 expression levels showed remarkably better OS than their counterparts with downregulated RNF20 expression levels (Figure 6A). RNF20 enhanced OS in patients treated with immune checkpoint inhibitors, among them anti-PD-1 monotherapy (nivolumab) and anti-PD-L1 (atezolizumab) (Figure 6B-F). Conversely, patients with elevated RNF20 expression do not respond to anti-CTAL4 therapy (ie, ipilimumab) when compared with low-expressing patients (Figure 6G and H). These results show that the high levels of RNF20 can serve as a predictor of the high efficiency of anti-PD-1 and anti-PD-L1 immunotherapies.

Association of RNF20 expression in cancer with ICB therapy or cytokine therapy.
Drug susceptibility and molecular docking studies
Drug susceptibility remains a long-standing central concern in the context of personalized cancer chemotherapy. Specifically, determining the response of tumors to specific pharmacological agents is fundamental to personalizing therapeutic plans for individual patients and propelling innovations in precision medicine. Nevertheless, considerable variability in drug responsiveness among individuals means relying exclusively on restricted research resources frequently proves insufficient—thus, identifying molecular determinants governing drug sensitivity is essential to improving therapeutic strategies. Currently the largest public repository of drug Susceptibility data and molecular tumor markers, the GDSC integrates datasets from multiple studies to conduct systematic reanalysis of drug Susceptibility and cellular response at the cellular, pharmacologic, and molecular levels. Through our analysis, we found that RNF20 expression was inversely correlated with tumor cell sensitivity to bleomycin (50 µM), BMS-754807, camptothecin, docetaxel, FK866, MLN4924, olaparib, SN-38, talazoparib, and YK 4-279, but positively associated with Susceptibility to A-770041, AS605240, BHG712, bryostatin 1, dasatinib, LY317615, NG-25, nilotinib, sunitinib, THZ-2-49, TL-1-85, and TL-2-105 (Figure 7A). Combined bioinformatics analysis and molecular docking simulations predicted possible interactions between RNF20 and the selected drugs (Figure 7B). Among these, RNF20 exhibited the weakest binding affinity for SN–38, with a binding energy of −7.2 kcal/mol. By comparison, the RNF20–talazoparib complex underwent stabilization through 2 hydrogen bonds mediated by residues threonine 921, lysine 917, and glycine 975, with a computed binding energy of –6.7 kcal/mol. Our findings suggest that SN-38 and talazoparib may have promising potential as therapeutic inhibitors of RNF20. However, we failed to obtain reliable molecular docking results for bleomycin (50 µM), camptothecin, or docetaxel. Complete molecular docking data for the remaining drugs are provided in Table 3.

Drug sensitivity. Based on GDSC data, the correlation between RNF20 expression and drug sensitivity (top 30 drugs) across various cancers is shown (A). A positive correlation suggests that higher gene expression may confer drug resistance, while a negative correlation suggests that higher gene expression may enhance drug sensitivity. Molecular docking of RNF20 with drug molecules is visualized in (B).
The molecular docking outcomes for the different tablets with RNF20.

Functional enrichment analyses and RNF20-PPI network of genes
To further comprehend the molecular mechanisms underlying tumorigenesis of RNF20, we sought to determine the interactions between RNF20 and proteins and RNF20 expression-linked genes these proteins were then enriched by pathway analysis. The RNF20-binding proteins with the help of the STRING tool were identified (n = 55), and the experimental results and co-expression profiles confirmed them. The outcome was a PPI network (Figure 8A; Table 4) with 55 nodes and 714 edges and the average node degree was 26. RNF20-binding proteins were over-represented in signaling pathways that relate to systemic lupus erythematosus, alcoholism, neutrophil extracellular traps and viral carcinogenesis (KEGG analysis: Figure 8B). GO term enrichment analysis revealed that the binding proteins of RNF20 were enriched with the following categories: biological processes (BP) involving chromatin assembly, proteinDNA complex assembly, and chromatin assembly or disassembly; cellular components (CC) such as nucleosomes, DNA packaging complexes and proteinDNA complexes; and molecular functions (MF) such as protein heterodimerization activity (Figure 8C). There were 100 candidate target genes that were ranked as the top 100 associated with RNF20 that were retrieved using the GEPIA2 platform (Table 5). Interestingly, the top 10 of these candidate target genes show a high-positive association with the expression pattern of various types of cancers (Figure 8D).
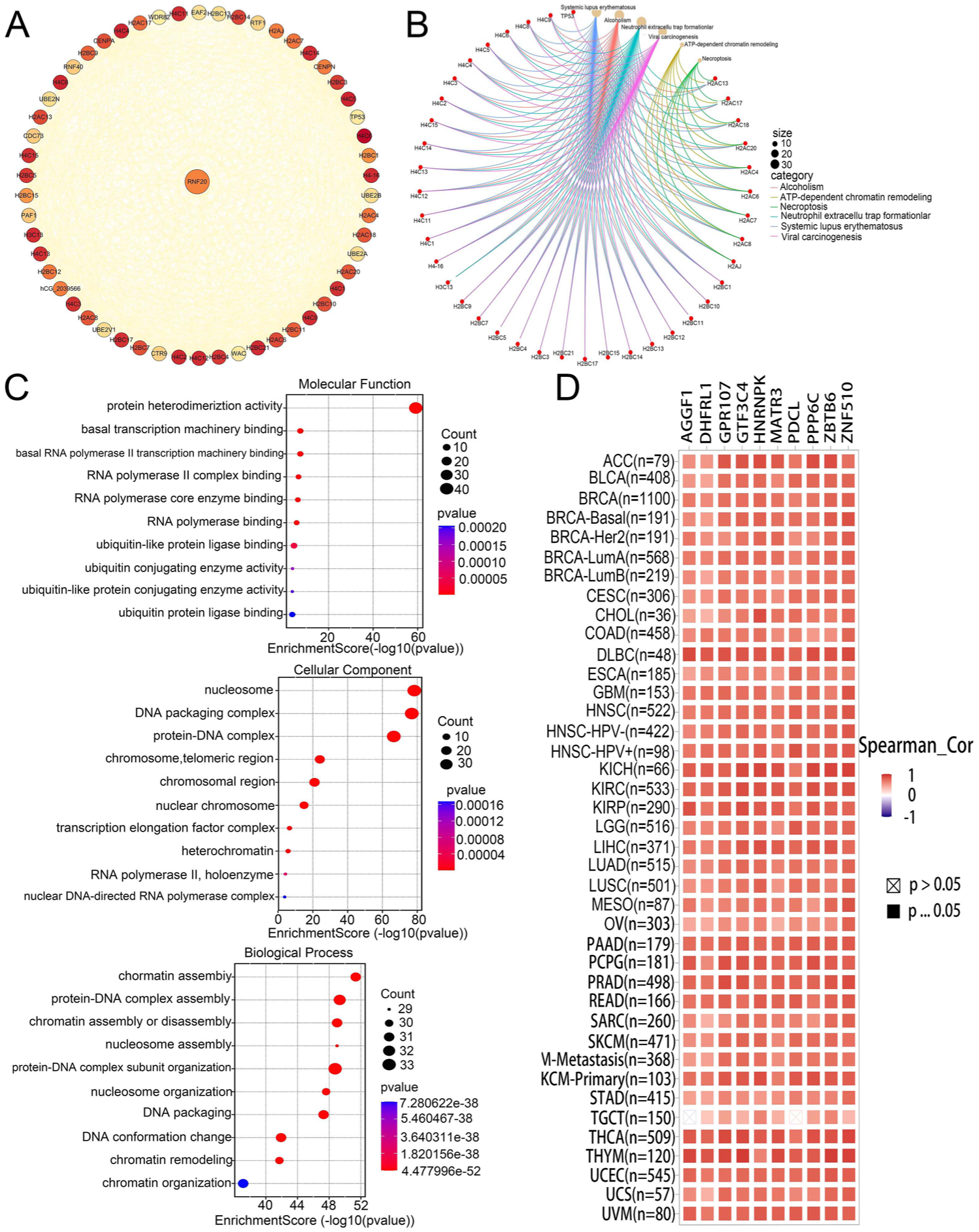
RNF20-related gene enrichment analysis. (A) PPI network of RNF20-binding proteins. (B) KEGG pathway analysis of RNF20-related genes. (C) GO analysis of RNF20-related genes. (D) Expression levels of the top 10 RNF20-related target genes in cancer.
The PPI network data of proteins bound to RNF20.

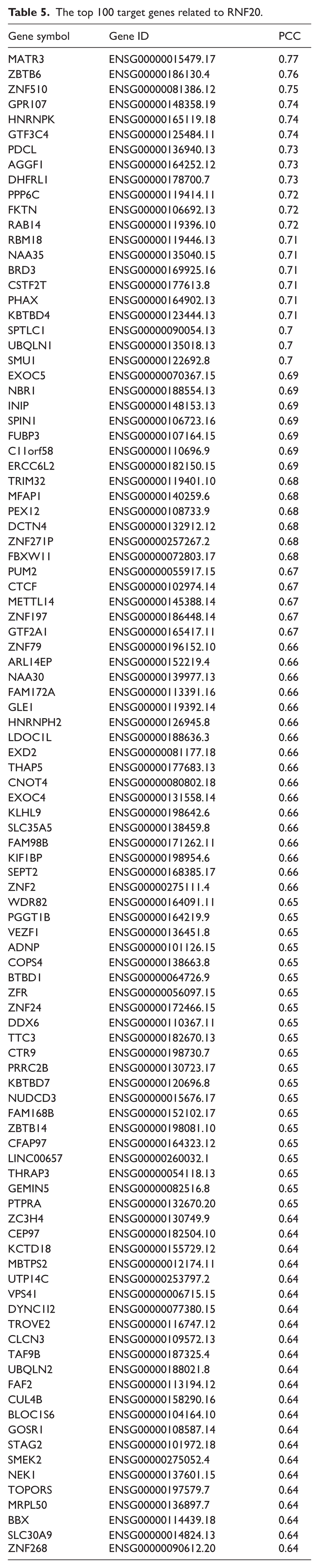
The top 100 target genes related to RNF20.
RNF20 mutational characteristics of the TCGA pan-cancer cohort
We conducted an analysis of the genetic changes in RNF20 to determine whether genetic changes in this protein might cause tumorigenesis. location of mutations, types of RNA modifications and genomic heterogeneity among different types of cancer. RNF20 changes were observed in 32 types of cancer with the highest rate (6.81) being identified. in UCEC (Figure 9A). The results described RNF20 mutations by their types, loci, and the number of cases, and the most popular form of RNF20 gene variation was a missense mutation (Figure 9B; Table 6). In the effort to develop better insights into the role of RNF20 in tumors, genomic heterogeneity and RNF20 expression were studied. The correlation of TMB with RNF20 in 37 tumors revealed significant relationships in 3 tumors, which include GBM, GBMLGG, and THCA (Figure 9 C). To ascertain additional insights into the molecular pathways through which RNF20 interact with cancer we compared its expression with 44 marker gene expression patterns related to class III RNA modification, ie, m1A (10 genes), m5C (13 genes), and m6A (21 genes) (Figure 9D).

Distinct genomic profiles associated with RNF20 expression and integrative analysis of cancer genomic and clinical profiles. (A) Genetic alteration types of RNF20 (mutation, structural variant, amplification, and deep deletion) across 32 different tumors, analyzed using TCGA data via the cBioPortal tool. (B) RNF20 mutation sites and corresponding case counts. (C) Analysis of genomic heterogeneity in relation to RNF20 expression. (D) Co-expression patterns of RNF20 and 44 marker genes associated with 3 class III RNA modification types (m1A, 10 genes; m5C, 13 genes; m6A, 21 genes).
The genetic mutation form in RNF20.

Discussion
As a ubiquitously expressed E3 ubiquitin ligase, RNF20 interacts with RNF40 to form homodimeric or heterodimeric complexes, and this RNF20–RNF40 complex modulates H2Bub1 and gene expression. 34 Numerous investigations have indicated that RNF20 and RNF40 govern β-cell gene expression and insulin secretion, forging an association between the Isl1 complex and pan-cellular epigenetic mechanisms. 35 RNF20 maintains systemic insulin homeostasis by regulating the expression of Slc2a4. 36 Abrogation of RNF20 and RNF40 causes downregulation of the expression of peroxisome-related genes, leading to ferroptosis. 10 RNF20-mediated H2Bub regulates chromatin dynamics to protect replicating genomes. 37 Both the target genes of the p53 tumor suppressor and many histone H2A and H2B genes are positively regulated by RNF20, while the protein can also suppress gene expression by blocking the binding of transcription factor IIS (TFIIS) to RNA polymerase II. 38 Interestingly, the RNF20–RNF40 E3 ligase complex mediates H2B monoubiquitination at DNA damage loci, 6 which is indispensable for the recruitment of double-strand break repair proteins and the promotion of timely DNA repair.39,40 In addition, H2Bub1 mediated by RNF20 enhances transcriptional elongation cooperatively with the FACT complex (facilitates chromatin transcription complex), which serves to impair nucleosome stability. 41 Alternatively, the labile promoters of oncogenic transcription RNF20 can inhibit the TFIIS recruitment, which is necessary to allow RNA polymerase II to become active in elongation of oncogenic genes, silencing oncogenic transcription programs. 38 Deficiency of RNF20 and a reduction of H2Bub1 have been reported in several cancers and was allied with harsh phenotypic characteristics. 42 RNF20 has been shown to be associated with impaired tumor suppressor function and enhancement of malignancy -which could be the heightened ability of cancer cells to foster epigenetic alterations and cancer signaling in lung adenocarcinoma. 43 Conversely, in pancreatic cancer, RNF20 is an oncogenic protein that can inhibit which may have anti-cancer properties. These findings indicate that RNF20 can either favor tumor proliferation or metabolic mal adjustment under different circumstances and inhibiting RNF20 might have therapeutic potential. 44 Changes in RNF20 expression may also reflect physiological conditions, e.g. the estrus cycle, in which its activity is more likely to be associated with developmental and differentiation control than with cancer pathways. 45 This feature that is context-dependent outlines how the targeting of RNF20 to reduce possible side effects should consider cell type and disease context. Nonetheless, the low-level correlation between RNF20 and tumor formation/progression has not been well defined in the research.
Under this study, we have compared the expression of RNF20 in 34 tumor types and their normal counterparts. RNF20 differed between different types of cancer: it was overexpressed in 21 types and underexpressed in 8, namely UCEC, KIRP, KIPAN, BLCA, OV, TGCT, UCS and KICH (Figure 2). Additional examination showed that there were considerable correlations between the expression of RNF20 and the clinical stage in LIHC, THCA and KIRC. RNF20 expression at a high level in tumor was highly correlated with low survival in all of the survival outcomes of the analysis of LIHC, BLCA, and ACC, while low RNF20 expression was strictly correlated with negative results in GBMLGG, KIPAN, and KIRC in the analysis of OS, DSS and PFI (Figure 3). All the results mentioned above indicate that RNF20 is both a prognostic factor as well as a critical mediator of the tumor progression. It is interesting to note that RNF20 expression was closely linked with immune checkpoint molecules, immunomodulatory genes, tumor stemness characteristics, and tumor immune microenvironment infiltration, which underscores its significance in controlling tumor immunity. We identified strong possible interactions among RNF20 and RNF40, H2BC21, H4 C6 and H2BC11 via protein-protein interaction (PPI) network analysis (Figure 8). The analysis of pathway enrichment revealed that the genes most related to RNF20 belong to unique signaling cascades and biological functions, whereas RNF20 exhibits a typical mutation pattern in various types of cancer. High levels of the tumor RNF20 response positively favored the survival of patients; therefore, they gained more survival benefits with immunotherapy response. Conversely, the association between low RNF20 expression and improved clinical outcomes was especially observed in patients undergoing anti-CTLA-4 therapy based on ipilimumab. This finding attests to the idea that the expression levels of RNF20 and their interaction with certain immunotherapeutic interventions deserve to be considered when developing treatment plans tailored to each particular case.
Conclusions
Overall, it can be stated that RNF20 is often overexpressed in many types of tumors, and its level is tightly correlated with clinical staging and prognostic values (e.g. OS, DSS) in other cancers. Further investigations have also found that RNF20 demonstrates high positive correlations with immune checkpoint proteins, immunomodulatory factors, prognostic biomarkers, immunoregulatory genes, and tumor stemness scores and tumor infiltration of the immune microenvironment. The most frequent type of variation of RNF20 is the missense mutations which have a vital functional impact in certain cancers like GBMLGG and THCA. Also, the RNF20 targeted inhibitors SN-38 and talazoparib have a possibility of being used. This research provides a theoretical basis through the systematic discovery of the possible RNF20 functions in tumor initiation and immunotherapy response, to formulate new RNF20-based therapeutic interventions.
Limitations
In our article, we propose that RNF20 has a potential in terms of pan-cancer prognostic predictor and therapeutic decision-making; however, a number of limitations of the research must be noted. First, much of our information was obtained in open databases–most of which rely on the retrospective cohorts of patients and are therefore susceptible to intrinsic selection biases. This may invalidate the generalizability and reproducibility of our results due to these biases.
Second, the systematic analysis of issues such as tumor microenvironment, immune regulatory systems or drug responses, is largely an analysis of predictive inference through bioinformatics algorithms with no systematic functional validation being performed through in vitro cellular analysis or in vivo animal model studies. Moreover, all results of the molecular docking are simply the products of the computer-based simulation, and the binding affinity between the drug and RNF20 and its inhibitory effect should be additionally confirmed in experimental studies. In addition, the innate heterogeneity of many types of cancer, patient groups, and tissues can potentially trigger changes in the expression pattern of RNF20 that can then result in the lack of sample consistency and reliability of its prognostic value.
Footnotes
Acknowledgements
We acknowledge the contributions from UCSC, HPA, TIMER2.0, TCGA, GTEx, GEPIA2, SangerBox 3.0, KMplot, TISMO, GDSC, PDB, PubChem, STRING and cBioPortal databases. In addition, we acknowledge Michelle Kahmeyer-Gabbe, PhD, from Liwen Bianji (Edanz) (![]() ) for editing the English text of a draft of this manuscript.
) for editing the English text of a draft of this manuscript.
Ethical Considerations
The current study was conducted using de-identified public data and was categorized as minimal risk research. Given that the original data collection protocols had already received prior approval from the relevant institutional review boards (IRBs), separate ethical approval from an institutional review board was not required for this study.
Consent to Participate
This retrospective pan-cancer analysis used anonymized aggregate data from the open-access TCGA and GTEx databases for secondary evaluation, with no participant identification or intervention. Informed consent was obtained in the original studies; thus, additional consent was not required.
Author Contributions
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Research Project of the Shanxi Provincial Science and Technology Department (202203021221279) and the Science and Education Cultivation Fund Project (SD2023032) for Doctoral and Master Supervisors of the National Cancer Regional Medical Center, Shanxi Province Cancer Hospital.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The datasets analyzed during the current study are available in public repositories, as detailed in the Materials and methods section. All data pertaining to this study can be obtained from the corresponding author upon reasonable request.
