Abstract
The Microchimerism Literature Atlas (MCLA) is a comprehensive online dataset to facilitate the investigation of microchimerism (MC), condition where individuals harbor cells from another individual of the same species. The MCLA provides access to more than 15 000 references from MC research, covering peer-reviewed articles and reviews from 1970 to the present. Key features include a multidimensional search function and logical operators for assembling search queries. The MCLA dataset offers a clearly structured data table view, combined with dynamic graphical data representation and visual citation analysis, aiding in the investigation and identification of research trends and patterns. The MCLA supports data export in various formats and receives regular updates. The MCLA is being developed as an essential resource for the MC research community while its framework is easily adaptable for custom literature datasets, enabling its use in other research fields.
Keywords
Introduction
Microchimerism (MC) is the presence of non-self cells in an individual naturally obtained during pregnancy, e.g., maternal cells in the offspring or fetal cells in their mothers. 1 The definitions among the MC community vary and may include non-cellular MC such as fetal extracellular vesicles or cell-free fetal DNA, probably most elegantly summarized in the umbrella term “microchiome.”2,3 In humans, MC cells presently appear to be mainly transferred through pregnancy, nursing or intercourse, or artificially; i.e., via medical interventions, such as blood transfusions or organ transplants, though other as yet undiscovered routes of transmission may be possible. 4 Microchimerism has been associated with a diverse range of effects on health and disease and is often considered a biomarker in the context of other research topics including cancer, autoimmune disease, and tissue repair. 1 This diversity reflects the minor common basis for MC research which is a challenge MC research faces. To unite and support the MC research community, we have created the Microchimerism Literature Atlas (MCLA, https://literature-atlas.microchimerism.info), a comprehensive knowledge dataset currently comprising 15 000 references from MC research that covers peer-reviewed articles and reviews from 1970 to the present.
Papers from the late 1960s and early 1970s start to analyze microchimerism
The first report of fetal cells in maternal circulation dates back to 1893, when a German pathologist detected trophoblast cells in lung capillary tissue of women who had died from eclampsia.5,6 Later in 1945, Owen published the first report of chimerism in animals showing cattle harboring their twin’s blood and in humans, Booth and colleagues reported chimerism in a pair of twins.7,8 The first reports of low numbers of chimeric cells were those of Walknowska and colleagues, who identified male cells in lymphocyte cultures obtained from pregnant women, and later the actual detection of Y-chromosome-positive microchimeric cells using flow cytometry by Schröder’s group.9,10 Although these cells were not termed microchimeric at the time, we decided to start our literature examination as of 1970 because these early papers address pregnancy as the source for MC and reflect the rarity of these cells. In the following decades, technologies to analyze microchimeric cells improved considerably (e.g., the use of fluorescent in situ hybridization [FISH], immunohistochemistry [IHC], fluorescence-activated cell sorting [FACS]/cytometry,11-15 microscopy,16-28 and polymerase chain reaction [PCR]29-32 [digital PCR allows for the detection of as little as one microchimeric trait against a background of 1 000 000 host signatures]). 33 Microchimerism research has evolved in parallel with the technical development toward digital PCR. For example, after the first reports detecting male cells in the female blood of pregnant women,10,34 researchers tried to isolate circulating fetal cells for non-invasive prenatal diagnosis. 28 ,35-38
Methods
The following sections outline only the key components of the methods used. Given their technical nature, the full methodological details are presented in the Supplementary Materials.
Data preprocessing was done in R: in addition to downloading the metadata of the articles as a result of the previously defined search queries using the easyPubMed package (see Results—PubMed queries for MCLA and Supplemental Material Table 1), the metadata were encoded in xml format and distributed across several folders. These were then converted into a tabular form using the XML package. For all articles whose metadata contain the PubMed IDs (PMIDs) of the articles they cite, the citation network was computed: a table with nodes (title, authors, journal, and article type) and a table with assignments (PMID A cites PMID B). To minimize the server load, a pre-filtering step was performed for defined terms related to microchimerism. All tables were then updated and deployed to the Shiny App. The R packages used during preprocessing were: DOParallel, DOsnO, dplyr, easyPubMed, foreach, methods, parallel, pbapply, readxl, rcrossref, rvest, stringi, stringr, tictoc, utils, writexl, xlsx, and XML.
The web app is implemented using the R-based Shiny framework: the dashboard interface was created using the shinydashboard and shinydashboardPlus packages. After restarting the Shiny app, the updated tables are loaded and rendered in tabular form using the DT package. For graphical representation of information as a word cloud, the wordcloud2 package was used. For the citation network visualization, the visNetwork package was employed, with the number of nodes being limited for clarity. Features not available in the packages, such as the search function across all columns, were implemented as JavaScript functions and added to the Shiny app using shinyjqui. Additional widgets and effects were incorporated using the shinyWidgets and shinyEffects packages.
The R packages used in the web app include DT, dplyr, igraph, network, readxl, shiny, shinyEffects, shinyWidgets, shinydashboard, shinydashboardPlus, shinycssloaders, shinyjqui, stringr, tidyr, visNetwork, wordcloud2, and xlsx.
Results
The Microchimerism Literature Atlas
The MCLA dataset was built by downloading and combining the results of multiple PubMed archives (https://pubmed.ncbi.nlm.nih.gov) queries and contains the essential metadata (i.e, title, abstract, authors, publication year, journal, article type, and DOI) extracted from it. Quality measures, including the comparison of references from 3 highly cited review papers from various decades,39-41 showed that 94% of the published MC literature within this period is covered by the dataset (see Results—Quality measures). In addition to an intuitive graphical user interface (GUI), the MCLA features a modular structure that allows users to navigate through different sections such as the “Data Table,” “Filter Settings,” “Chart View,” “Network View,” and “Info.” In the “Data Table” section, the literature data is displayed in a tabular format. The “Filter Settings” section includes predefined search terms commonly used in the MC field. The “Chart View” provides graphical representations of the dataset content, and the “Network View” section shows the embedding of references within a citation network. Compared with literature archives such as PubMed, the MCLA has the advantage of displaying the relevant literature information in a clearly structured data tabular format in the “Data Table” section. Citations can be expanded to display their abstracts (Figure 1) or directly accessed via the digital object identifier (DOI).

The graphical user interface (GUI) of the MCLA is structured modularly, with individual sections selectable through buttons. The main section “Data Table” displays essential information from the literature dataset in a clearly structured table that can expand to show abstracts, i.e., Shree et al. 31 An advanced multidimensional search function allows for precise searches, i.e., the whole dataset for microchimerism NOT “chimerism” and the author’s column for “Nelson.” The obtained results can then be exported in various formats.
PubMed queries for MCLA
The dataset can be filtered with an advanced search function that supports word part searches, phrase matching, and logical operators (AND, OR, NOT) to assemble precise queries (see Supplementary Material Table 1).
The literature metadata for building the dataset (papers, reviews, editorials, comments, etc) was retrieved from PubMed (https://pubmed.ncbi.nlm.nih.gov) using the MeSH (Medical Subject Headings) terms and/or keywords listed in Supplemental Table 1. In addition, the publication period of the papers is limited to the period from 1970 to the present, adding the string “AND (1970 [PDAT]:2024 [PDAT])” to each of the PubMed search queries.
Publications deemed relevant to MC were selected from the PubMed dataset using MeSH terms and tokens/keywords, which identify literature dealing mainly with pregnancy-associated MC but also other forms (e.g., transplantation-associated MC). MeSH was used as it represents a National Library of Medicine controlled vocabulary thesaurus that is used for indexing PubMed citations and helbgped to identify relevant literature. In addition, we used specific text (i.e., keywords or a sequence of text, also known as a token) known to be present in MC-related literature.
The search function also provides a multidimensional search option, allowing for individual column searches as well as an all-encompassing search function, enhancing the precision and efficiency of finding relevant literature. As shown in Figure 1, the search term microchimerism NOT “chimerism” has been used to search the entire dataset, with further restriction by using the term “Nelson” in the author column to refine the findings. However, the search results from the MCLA and PubMed cannot be directly compared for performance evaluation purposes, as they are implemented differently (Figure 2).

Comparison of references from search queries in the MCLA (green) and on the PubMed archive (orange). Numbers in the overlapping areas indicate references found in both datasets, while the numbers in the non-overlapping areas indicate references exclusively found in the MCLA (green) and PubMed (orange) archive, respectively. (A) MCLA query microchimerism NOT “chimerism” yielded a higher number of references compared with the PubMed query “microchimerism” NOT “chimerism.” (B) Restricting the query in the PubMed archive to title and abstracts (TIAB) improves the hits in PubMed.
The obtained search results can be exported as CSV, TXT, or XLSX files for further analysis. In addition to user-defined search queries, the “Filter Settings” section allows users to quickly assemble complex search queries by combining predefined terms commonly used in MC references, such as organ, tissue, and MC types, detection techniques, and specific diseases. In addition, users can select the publication period and type, i.e., article or review. This search criteria refinement allows users to comprehensively narrow their searches, making it easier to pinpoint specific information within the MCLA. The “Chart View” section graphically summarizes important information about the literature in the dataset. Bar plots and word clouds highlight the number of publications per year, the most frequently used title words, the most active authors, the most published-in journals, and the most common MeSH (“Medical Subject Headings”) terms and keywords by their respective frequencies, aiding users in identifying patterns. Furthermore, temporal heatmaps, which include detection techniques and highlight the most frequently used journals for publication, may assist in recognizing emerging trends in MC research. Moreover, the information displayed in the charts dynamically updates according to the results of applied search queries. The results for the search term microchimerism NOT “chimerism” are displayed as heatmaps, indicating citation trends across all techniques (Figure 3A) or normalized to every individual technique (Figure 3B) with the latter showing more details on the individual use of the respective technique. In addition, the most common authors can be shown in a bar graph (Figure 3C) or word cloud (Figure 3D).

The charts in the “Chart View” section dynamically update based on the used query, i.e., microchimerism NOT “chimerism.” (A) Citation trends of various detection techniques are indicated by colors highlighting the temporal changes across all techniques showing the in situ techniques to be used most often. (B) The trends can be normalized to each technique now indicating the citation peak of every technique itself showing that in microchimerism research there has been a shift from using the in situ techniques in the 2010s to rather PCR-based techniques thereafter. The authors of the respective references are listed according to their frequency either as bar chart (C) or word cloud (D). Clicking the data area allows switching between the views of A and B and between C and D, respectively.
This dynamic allows users to tailor their analysis to their specific needs (see Supplementary Material—MCLA search function). The “Network View” section allows users to visually explore the embedding of a specific paper in the citation network with up to 500 references (nodes). The network has been derived from the references included in the literature metadata, showing relations to papers within and beyond this dataset. An example is shown for Shree et al 31 (Figure 4A), where each node represents an article or review, distinguished by different colors and shapes. Hovering the cursor over a node provides information such as the title, authors, journal name, and publication year of the paper while clicking on a node highlights its connections to other papers (Figure 4B).
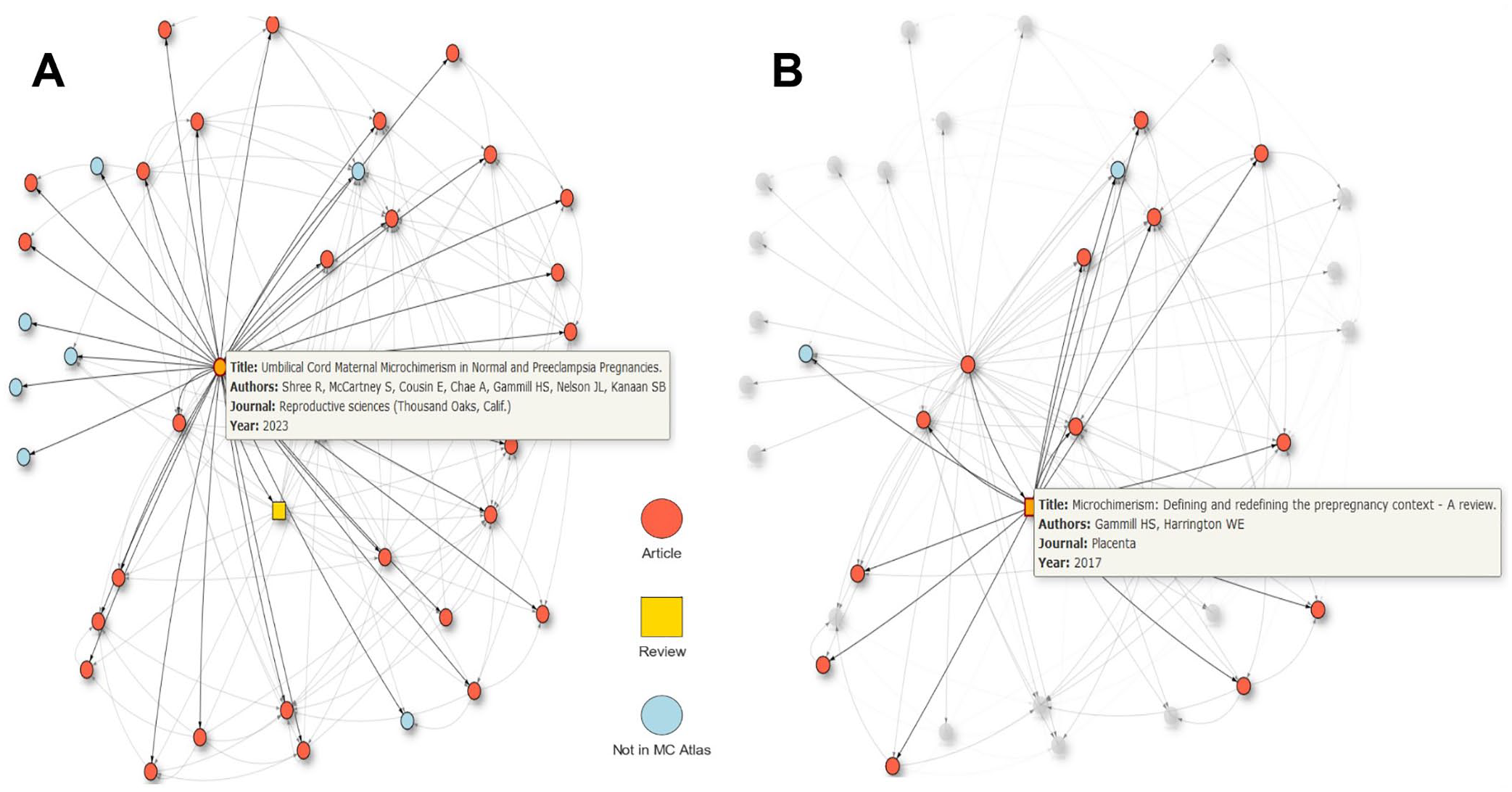
The “Network View” section allows for a graphical investigation of a selected paper’s embedding within the citation network. (A) When selecting a paper from the “Data Table,” its network of references is shown in the “Network View” with the selected reference being in the center (e.g., Shree et al.). 31 Articles are shown as red-filled, reviews as yellow-filled rectangles. If a reference is not part of the MCLA (i.e., not related to microchimerism), it is symbolized with a blue-filled circle. Hovering the cursor over this node displays information from the data table. (B) Selecting another node (Gammill & Harrington, 2017) 42 in the network changes the view accordingly now showing the network of the newly selected reference (within the initially selected references).
This feature provides users with a broad view of the MC research landscape, facilitating the identification of especially influential works. Finally, it should be mentioned that the dataset content of the MCLA is updated on a daily basis, to include newly published literature and ensure that users have access to the most current research findings in the field of MC. The MCLA can be accessed via the web application covering the literature on microchimerism (https://literature-atlas.microchimerism.info). In addition, the research content can be adapted to whatever field of research. Therefore, the MCLA source files can be downloaded from GitHub. For details, see the Supplementary Materials—Software setup and customization of the MCLA.
Quality measures
The quality of the MCLA was evaluated by the coverage of the MC literature and the accuracy of the papers dealing with the topic MC. To determine coverage, 3 highly cited review papers (Sahota et al. 39 ; Gammill and Nelson 40 ; Johnson et al. 41 i.e.) were chosen from various decades, and the papers cited therein were used as references to calculate the degree of coverage. The coverage of the cited papers that occur also in the MCLA are 100%, 89%, and 97%, respectively, resulting in a total weighted coverage of 94%. Although this coverage is quite high, there is still room for improvement. It should be noted that some papers were removed from the dataset during the cleaning process because of incomplete information of at least one of the essential metadata (DOI, publication year, journal). For the accuracy calculation, 100 papers were randomly selected from the MCLA dataset and presented to 3 MC researchers, who individually assessed their thematic relevance by answering the simple question “is the selected paper about MC?”—revealed an accuracy of 12%. The low value can be explained by the overlapping topics (Microchimeric cells are found in maternal and fetal tissues after pregnancy1,43 and seem to be associated with both beneficial and harmful effects in the host including cancer,44-49 autoimmune diseases,2,50 and tissue repair 51 and immune educative effects. 52 Thus, there are multiple questions the understanding of MC research can be applied to, covering a wide range of associations with sometimes contradictory results.) of multiple research fields with MC, such as cancer, chimerism, immunology, or pregnancy, so that papers dealing with those fields were often given the same tags as MC papers. On the one hand, achieving a high coverage while maintaining high accuracy is challenging due to the inverse relationship. For the content of the MCLA dataset, emphasis has been placed on achieving high coverage, which inherently resulted in lower accuracy.
Discussion
The MCLA can be used to get an overview of the literature that is currently available on MC. It covers citations of MC before the phenomenon was named “microchimerism” and it specifically tries to avoid citations from transplantation medicine that describe chimerism generated due to tissue and organ transplantation—different in cause (artificially due to surgery) and frequency (higher) with potentially different biological effects (although similar). There is a need for every research field to screen what is known to draw conclusions and to falsify hypotheses in order to move on in science. This need becomes harder due to the growing body and quality differences of results published, potentially drawing wrong conclusions from their experiments. Thus, the MCLA will be the first step toward organizing the known literature on MC by providing a one-stop for researchers. The first approaches we will take to weigh citations based on the technology the authors used for detecting MC. For example, it is known that certain techniques yield false-positive results20,24; thus, citations using these techniques need to be critically checked to assess their scientific soundness. In addition to its primary purpose, the MCLA will be used with recent/advanced automation techniques such as natural language processing, text mining, and certain machine learning applications all of which offer significant advantages for extracting insights and identifying trends within large volumes of text in such a specific field. As such, the MCLA aims to facilitate cooperation and collaboration between researchers in the same field and represent a crystallization point for establishing a standard in microchimerism research. With the establishment of these common good practices, it has the potential to act as a central repository of information to contribute to the elimination of inconsistent findings resulting from different methodologies.
We foresee that other fields of research might also profit from an atlas like the MCLA, be it for finetuning their literature research, obtaining additional information from the metadata or displaying the most used techniques or main active groups in their fields, or be it for a start toward organizing their scientific work or research topic. Despite its current limitations (e.g., no contextual analysis), the MCLA will allow a first approach toward structured literature analysis. The MCLA software setup and customization can be found in the Supplementary Material.
Conclusions
We present the MCLA as a comprehensive and customizable online literature dataset designed to facilitate the investigation of MC by providing access to over 15 000 references. The MCLA offers multiple features, compared with traditional archives like PubMed, making it a complementary resource for the MC research community. The source code of the MCLA was designed for easy customization to any literature dataset and is hosted on GitHub where it can be accessed and adapted to serve other scientific communities to mine and analyze published data in their field of research.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322251324104 – Supplemental material for The Microchimerism Literature Atlas
Supplemental material, sj-docx-1-bbi-10.1177_11779322251324104 for The Microchimerism Literature Atlas by Michael Christian Gruber, Daniel Kummer, Katja Sallinger, Henderson James Cleaves, Arsev Umur Aydinoğlu and Thomas Kroneis in Bioinformatics and Biology Insights
Footnotes
Acknowledgements
All individuals listed as authors contributed substantially to this manuscript. M.C.G. set up the MCLA supported by D.K. T.K. and K.S. tested the MCLA. M.C.G., D.K., and A.U.A. contributed to the design of the MCLA, and all of the authors revised the initial and the submitted manuscript.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The MCLA was developed as part of the project “We All Are Multitudes: The Microchimerism, Human Health and Evolution Project” that is funded by the John Templeton foundation (Grant-ID 62214).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Availability of Data and Materials
This manuscript includes original computer code, which is essential for the work presented. The code is available for download and review on GitHub (![]() ">
https://github.com/microchimerism/MCLA
). Access to the repository is unrestricted and does not require any form of personal identification, ensuring anonymity for the reviewers. The repository and the program code include an overview and a detailed description of the program. In addition, the supplementary materials contain a chapter on how to set up the environment to run the code.
">
https://github.com/microchimerism/MCLA
). Access to the repository is unrestricted and does not require any form of personal identification, ensuring anonymity for the reviewers. The repository and the program code include an overview and a detailed description of the program. In addition, the supplementary materials contain a chapter on how to set up the environment to run the code.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
