Abstract
Background:
Dengue virus (DENV) causes an important disease and directly affects public health, being the arbovirus that presents the highest number of infections and deaths in the Western Brazilian Amazon. This virus is divided into 4 serotypes that have already circulated in the region.
Methodology:
Molecular characterization of a cohort containing 841 samples collected from febrile patients between 2021 and 2023 was analyzed using a commercial kit to detect the main arboviruses circulating in Brazil: Zika, DENV-1, DENV-2, DENV-3, DENV-4 and, Chikungunya. Subsequently, Sanger sequencing was performed for positive samples.
Results:
The cohort detected 162 positive samples, 12 for DENV-1 and 150 identified as DENV-2, indicating co-circulation of serotypes. The samples were subjected to sequencing and the analysis of the sequences that obtained good quality revealed that 5 samples belonged to the V genotype of DENV-1 and 46 were characterized as DENV-2 Cosmopolitan genotype—lineage 5.
Conclusion:
The results allowed us to identify for the first time the Cosmopolitan genotype in Rondônia, Brazilian Western Amazon, and its fast spread dispersion.
Background
Dengue virus (DENV) belongs to the Flaviviridae family and is characterized as the arbovirus most widespread worldwide. 1 Dengue virus is an enveloped virus with a genome composed by a positive-sense single-stranded RNA with approximately 11.000 nucleotides,2,3 transmitted mainly through of blood repast by infected species of Aedes aegypti and A albopictus. 4 The virus is further classified according to the distinction of their antigenic and genetic characteristics into 4 serotypes 5 DENV-1, DENV-2, DENV-3, and DENV-4, each one being subdivided into genotypes.6,7
Dengue virus serotypes with the highest rates of reported infections in Brazil are DENV-1 and DENV-2. 8 The first one is subdivided in genotypes I, II, III, IV, V, and Wild, while DENV-2, is divided in Asian I and II, Asian-American, American, Cosmopolitan, and Wild. 9 In Brazil, considering DENV-2, genomics data show a higher prevalence of the DENV-2 Asian-American genotype and the sporadic detection of other genotypes, including the Cosmopolitan genotypes, which represents the bigger number of cases worldwide.4,10,11
The constant change in the phylogenetic profile of DENV-1, DENV-2, DENV-3, and DENV-4 in the Brazilian Amazon,2,12 has been sustained mainly by migratory factors.9,13 This study aims to characterize the DENV genotypes circulating at the state of Rondônia and in the municipality of Humaitá, Amazonas State, both located at the West Amazon.
Materials and Methods
Ethical statement
The project was evaluated and approved by the research ethics committees of Centro de Pesquisa em Medicina Tropical—CEPEM—Rondônia under number 1.474.102.
Study site and biological samples
The study was carried out at the Laboratório de Virologia Molecular of the Fundação Oswaldo Cruz—Rondônia (FIOCRUZ/RO). The retrospective cohort was composed of 841 human biological samples (serum) collected from December 2021 to May 2023, 756 from 37 municipalities of Rondônia state, received by Laboratório Central de Saúde Pública de Rondônia (LACEN/RO), or collected at the malaria outpatient clinic of the Centro de Pesquisa em Medicina Tropical—CEPEM and at the 2 Emergency Room—both located in Porto Velho, and 85 samples in a border municipality of Humaitá belonging to the Amazonas state.
Molecular analysis
Viral RNA extraction
Viral RNA was extracted from 140 μL of serum samples by QIAamp Viral RNA Mini Kit (QIAGEN, Germany), according to the manufacturer’s instructions. RNA was eluted in 60 μL of AVE buffer.
RT-qPCR for Zika, dengue, and Chikungunya
RNA samples were analyzed for the presence of Zika virus; DENV serotypes 1, 2, 3, and 4; and Chikungunya virus using real-time reverse transcription polymerase chain reaction (RT-PCR) kit ZDC Biomol (Instituto de Biologia Molecular do Paraná, Brazil), following the manufacturer’s instructions.
Reverse transcription
Positive samples were subjected to reverse transcription to generate complementary DNA (cDNA) synthesis using SuperScript III Reverse Transcriptase enzyme (Thermo Fisher Scientific, Massachusetts, USA) combined with 0.5 μg of random primer, according to the manufacturer’s instructions.
Conventional polymerase chain reaction followed by sequencing
The reaction adapted using the protocol of Lanciotti et al 14 under the following conditions 10 µL of 2x Platinum SuperFi II PCR Master Mix supplemented with 1.5 mM MgCl2, 300 nM Primer D1 (5′-TCAATATGCTGAAACGCGCGAGAAACCG–3′), 300 nM Primer D2 (5′–TTGCACCAACAGTCAATGTCTTCAGGTTC–3′), and 2 μl of cDNA generated according to 2.3.3 item to amplify a 511 base pair (bp) PCR product. The cycling used for the reaction was as follows 98°C for 30 seconds for initial denaturation, followed by 40 cycles at 98°C for 15 seconds, 53°C for 30 seconds for annealing and 72°C for extension 40 seconds, ending with a final extension step at 72°C for 5 minute followed by PCR product purification with ExoSAP-IT PCR Product Cleanup (Applied Biosystems, California, USA). The BigDye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems, California, USA) was used according to the manufacturer’s instructions for sequencing. The reaction product was purified using BigDye XTerminator Purification Kit (Applied Biosystems, California, USA). Sequencing run was performed by the Rede de Plataformas Tecnológicas FIOCRUZ RPT09F—FIOCRUZ/RO using an automated Sanger sequencer Seqstudio (Applied Biosystems, Waltham, MA, USA). The electropherogram was edited and analyzed using MEGA11-Molecular Evolutionary Genetic Analysis software. 15
Dengue virus-2 complete genomes were obtained using Illumina Viral Surveillance Panel (VSP w ILMN RNA Prep w Enrich), which is a hybrid capture method using biotinylated probes. The VSP libraries were then submitted to nucleotide sequencing on an MiSeq instrument using V3 cartridges on 2 × 150 cycles paired-end run (Illumina). FastQ reads were assembled against the GenBank DENV-2 RefSeq NC_001474 using BBDuk (trimming) and BBMap v38.84 embedded in Geneious Prime 2023.0.4 software.
Phylogenetic analysis
The sequences generated in this study were aligned with representative sequences of the DENV-1 (Dataset 1) and DENV-2 (Dataset 2) genotypes collected through the GenBank public database available at the National Center for Biotechnology Information—NCBI, 16 all representatives with available collection date and locality information were collected. Multiple sequence alignment was performed using online software MAFFT v.7. 17 A maximum likelihood (ML) phylogenetic tree was generated in IQ-TREE v.2.2.2.6 18 using the best fit nucleotide substitution model by the ModelFinder tool and edited using FigTree v.1.4.4. software. 19 The branch support values obtained were estimated using the bootstrap in 1000 replicates.
Bayesian analysis and spatiotemporal reconstruction
For the inference of the Bayesian phylogenetic tree, only sequences from the study that obtained a complete genome (n = 20) were included. The existence of a temporal signal was checked using TempEst v.1.5.3. 20 To create the .xml file in BEAUti v.1.10.1 software, the dataset included the collection date information. The Bayesian tree was inferred using the nucleotide substitution GTR + F + I + G4 as the best fit model by the ModelFinder, lognormal relaxed molecular clock and a Bayesian Skyline coalescence model. The length of the Markov chain Monte Carlo (MCMC) was set at 100 million generations, and the convergence of the parameters was assessed by calculating the effective sample size (ESS) using Tracer v1.7.1. 21 The phylogenetic tree was summarized using TreeAnnotator v.1.10 excluding 10% of the samples as burn-in, creating a Maximum Clade Credibility (MCC) tree. The final tree was edited and visualized by the ggtree package 22 using R v4.3.2. 23
Results
The cohort of 841 samples was tested for Zika virus, DENV 1-4 serotypes and Chikungunya virus using the commercial kit that resulted in 81% negative (679/841) and 19% positive (162/841), being 93% DENV-2 (150/162) and 7% DENV-1 (12/162) in the period from December 2021 to May 2023 (Figure 1). All positive samples for DENV were subjected to sequencing, which 31% of the positives for DENV-2 (46/150) and 42% of positives for DENV-1 (05/12) obtained enough quality for subsequent analysis.

Temporal distribution of confirmed cases. Time series graph showing the number of cases of DENV-1, DENV-2, and total cases in each month.
The state of Rondônia showed 92% (149/162) of the cases, 56% (90/162) in the Porto Velho, followed by Presidente Médici with 10% (16/162), and the other cases were dispersed among the municipalities of this state, which contain between 1 and 6 cases each, being represented in Figure 2. The municipality of Humaitá, the only one belonging to the state of Amazonas identified in this study, housed 8% (13/162) of the cases.

Geographical location of each city with identification of DENV cases. The cities were marked using a variety of colors. In orange is characterized the municipality belonging to the state of Amazonas. The map was assembled using QGIS 3.18.1-Zurich software.
The study population (n = 162) had a mean age of 33.8 years with a higher proportion in the age group of 21 to 40 years. The Table 1 presents epidemiologic data referring to the individuals included in the study. Regarding DENV positive samples, 48.77% (79/162) were male and 51.23% (83/162) were female. Around 56.17% (91/162) reported symptomatology with fever and headache both at 45.05% (41/91) being the most common reported symptom, followed by back pain at 29.67% (27/91) and myalgia at 25.27% (23/91).
Epidemiological data and symptoms obtained from the application of the survey.

fisher’s exact test.
A total of 46 sequenced DENV-2 samples from the study were classified into the Lineage 5 cluster of the cosmopolitan genotype, as shown in Figure 3.
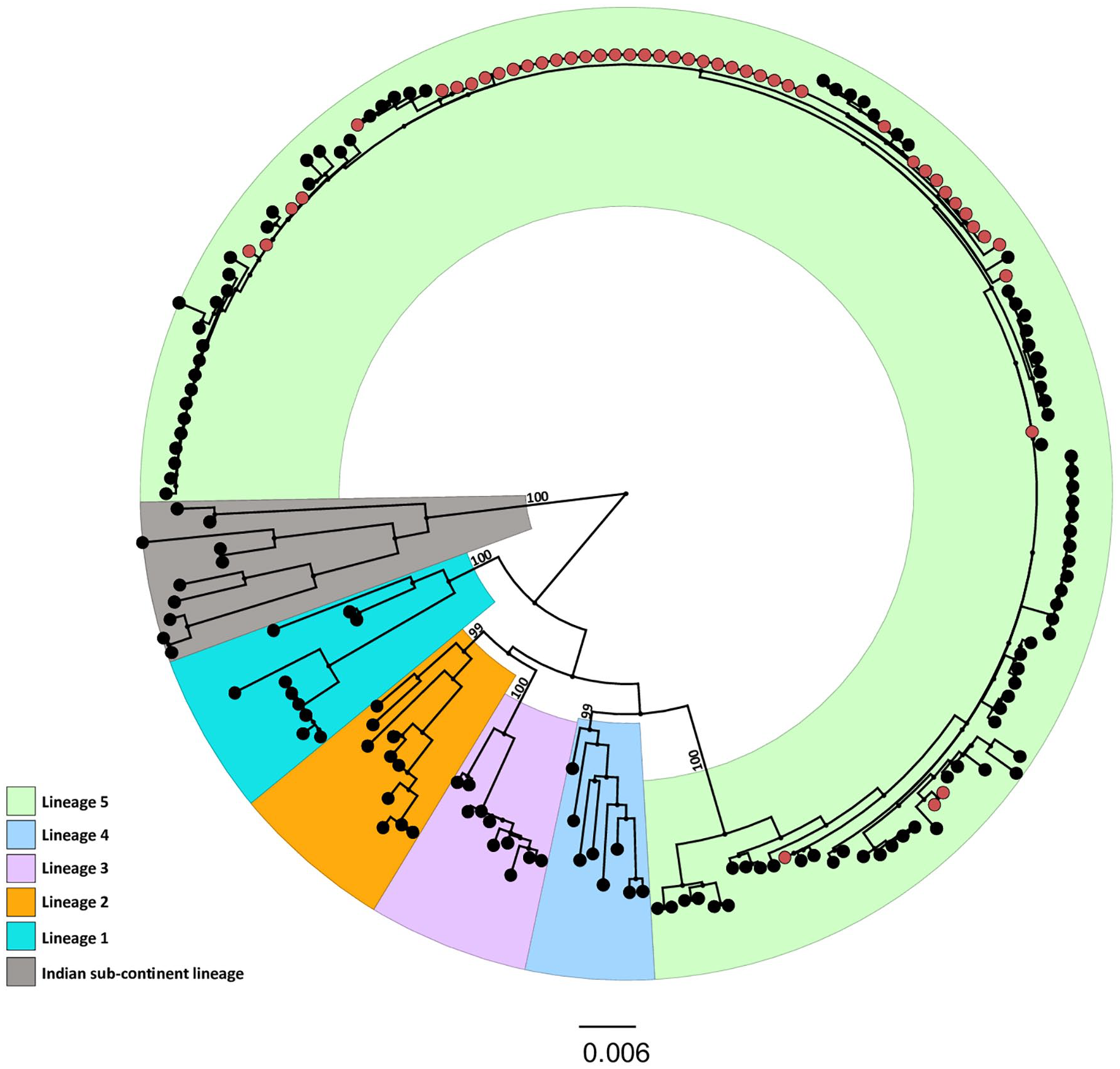
Maximum likelihood phylogenetic tree composed of 46 sequences from the study and 141 sequences retrieved from GenBank representing the lineages (Lineage 1, Lineage 2, Lineage 3, Lineage 4, Lineage 5, and Indian Subcontinent Lineage) of the cosmopolitan genotype of DENV-2. The study samples are highlighted as red circles, and support values (bootstrap) are indicated on the main branches. TIM2 + F + I + G4 substitution model chosen according to Bayesian Information Criterion (BIC).
For spatiotemporal reconstruction of the study samples, a total of 20 DENV-2 complete genome samples were submitted to a Bayesian phylogeographic analysis with a time-scale approach. It can be observed that the lineage 5 cosmopolitan possibly originated from sequences in Bangladesh and that there was a later introduction in Peru and in different Brazilian states, respectively (Figure 4). In addition, 2 clusters of the study samples (Rondônia and Amazonas state) were found, phylogenetically related to samples from other Brazilian states. The first (smaller cluster), with the common ancestor dated August 14, 2021 (HPD 95% December 31st 2020 to December 28, 2021), was closely related to a sample from the state of Santa Catarina (SC). The second (larger cluster), with the common ancestor dated February 10, 2021 (HPD 95% August 12, 2020 to July 02, 2021), was similar to samples from the states of Parana (PR), Rio Grande do Sul (RS), São Paulo (SP), and SC.

Bayesian phylogenetic tree containing samples from DENV-2 cosmopolitan lineage 5 clade.
To compose the data set, all the available complete genomes of lineage 5 of the cosmopolitan genotype were downloaded from GenBank. Tips are colored according to location, and the study samples are indicated as triangles. The support values are indicated on branches as posterior probability > 0.9.
Regarding the positive cases for DENV-1 serotype, 41.66% (5/12) were characterized as genotype V, as represented in Figure 5. Among these, 4 of the identified isolates are from the municipality of Porto Velho, capital of the state, and 1 from the municipality of Buritis, located 319 km from the capital. Considering the current circulation profile, DENV-1 was the nondominant serotype within the state.

Maximum likelihood phylogenetic tree for DENV-1 genotypes. The phylogeny was composed of 63 sequences retrieved from GenBank representing DENV-1 genotypes (GI, GII, GIII, GIV, and GV) and 5 strains generated in this study (highlighted in red). To improve visualization of the results, clades containing GI, GII, GIII, and GIV were collapsed and highlighted (Blue—GI; Orange—GII; Green—GIII; Yellow—GIV). Support values (bootstrap) are indicated on the branches. Best-fit model: TIM2 + F + I + G4 chosen according to BIC.
Discussion
The retrospective cohort describes the introduction of the cosmopolitan DENV-2 genotype in Rondônia and in a border municipality located in the state of Amazonas, far western Amazon, and the current replacement of the dominant serotype in the region, DENV-1. In the state of Rondônia, the 4 serotypes of DENV have already circulated, however, without detection of the cosmopolitan genotype. 24
Dengue virus-1 isolates belong to genotype V that circulated widely in previous outbreaks in the state of Rondônia.12,25,26 Suggested that DENV-2 strains were maintained at baseline levels in recent DENV outbreaks in some regions of the country 26 which may justify the high number of DENV-2 cases found in this study and the current substitution of genotypes in the state.
The cosmopolitan genotype of DENV-2 is widespread in several countries worldwide, with 6 identified lineages. Lineage 5 of the cosmopolitan genotype was previously described in several countries in Asia,4,27,28 and identified in the Americas during the described dengue infections in Madre de Dios, Peru, in 2019. 28
In Brazil, its introduction was first described from a case at the state of Goiás, in November 2021. 29 Then, the analysis of a retrospective cohort, collected from February to March 2021, showed 5 cases infected with this genotype in the state of Acre. 30 The wide dispersion of the cosmopolitan genotype has been demonstrated in other Brazilian states, including the Amazonas State, in a recent study showing multiple introductions of the cosmopolitan DENV-2 in Brazil. 11
Starting in December 2021, we observed the raise of dengue cases in Rondônia. Our results showed that the DENV-2 sequences grouped with sequences from the states of the south region (SC, PR, and RS) and southeast Brazil (SP). However, more detailed analyses of transmission routes are still underestimated due to the distance of these regions from the study site and the reduced sampling from other Brazilian states. To support the data analyzed, an increase in DENV-2 cases compared to DENV-1 was already reported in several states since 2022, according to the National System of Diseases of Compulsory Notification (SINAN), which allowed for the consolidation of this serotype in different Brazilian regions. 11
The emergence of the Cosmopolitan genotype in the state of Rondônia and the consequent increase in the number of severe cases in the region has significant implications for the susceptibility of the population. This is not only limited to viral serotypes as demonstrated in recent studies 31 but also to the Cosmopolitan genotype itself, which has shown genetic variations over the years that impact on pathogenicity, transmissibility, and competition between DENV2 genotypes.4,32,33
Conclusions
This study identified the simultaneous circulation of DENV-1 genotype 5 and DENV-2 Cosmopolitan genotype in different municipalities of the Brazilian Western Amazon, highlighting the emergence and dispersion of the Cosmopolitan genotype of DENV-2 in the state of Rondônia and in the municipality of Humaitá, Amazonas. The results obtained by our research provide pertinent information about the multiple introductions of the Cosmopolitan genotype that have occurred in the state of Rondônia and contribute to understanding the dynamics of Dengue dispersal in the Brazilian Amazon. Considering the wide dissemination and evolution of the Cosmopolitan genotype around the world, and now in the Americas, we emphasize the need for further studies into the dynamics of DENV transmission.
Footnotes
Acknowledgements
The study has been developed by a group of researchers from the Laboratório de Virologia Molecular of the Fundação Oswaldo Cruz Rondônia. The Fundação Rondônia de Amparo ao Desenvolvimento das Ações Científicas e Tecnológicas e à Pesquisa do Estado de Rondônia—FAPERO, as well as the Instituto Nacional de Ciência e Tecnologia de Epidemiologia da Amazônia Ocidental—INCT—EpiAmo they have been important contributors to scientific development in the Amazon region. Collaboration from the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), from whom some authors received financial support (scholarships) during the production of this study, the Instituto de Biologia Molecular do Paraná (IBMP), the Laboratório Central de Saúde Pública de (LACEN/RO), Coordenação de Vigilância em Saúde e Laboratórios de Referência, the Rede de Plataformas Tecnológicas FIOCRUZ RPT09 F-FIOCRUZ/RO, Rede Genômica da Fundação Oswaldo; Agência Estadual de Vigilância Sanitária de Rondônia (AGEVISA-RO), and the Secretaria Municipal de Saúde (SEMUSA). They were essential for the development of the study.
Author Contributions
Conceptualization: JRR, TPR, GSC, AMPS, and RCPR; Data curation: JRR, TPR, GSC, and AMPS Formal analysis: JRR, TPR, and AMPS; Funding Acquisition: DVS and FGN; Investigation: JRR, TPR, GSC, and AMPS; Methodology: GSC, JRR, TPR, AMPS, ALFS, HMM, and KST; Project Administration: DV and RAR; Supervision: RCPR and DVS; Writing—original draft: GSC, JRR, TPR, AMPS, KST, HMM, CABL, AOS, JMVS, VAN, and VCS; Proofreading and editing of the text: RCPR, DVS, FGN, and MAK. All authors have read and agreed to the published version of the manuscript.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was financed by the Foundation Oswaldo Cruz of Rondônia—FIOCRUZ/RO, Fundação Rondônia de Amparo ao Desenvolvimento das Ações Científicas e Tecnológicas e à Pesquisa do Estado de Rondônia—FAPERO (Process: VPPIS-003-FIO-20-2-67—INOVA AMAZÔNIA; Process: 35562.558.20485.10082022—INICIATIVA AMAZONIA + 10) and by the Instituto Nacional de Ciência e Tecnologia de Epidemiologia da Amazônia Ocidental—INCT EpiAmO.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
