Abstract
The kinetochore is a multiprotein structure that attaches at one end to DNA in the centromere and at the other end to microtubules in the mitotic spindle. By connecting centromere and spindle, the kinetochore controls the migration of chromosomes during cell division. The exact position where the kinetochore assembles on each centromere was uncertain because large sections of centromeric DNA had not been sequenced due to highly repetitive alpha-satellite arrays. Embedded in the arrays is a 17 bp consensus sequence, the so-called CENP-B box, which binds the CENP-B protein, the only protein that binds directly to centromeric DNA. Recently, the Telomere-to-Telomere Consortium published the complete centromeric DNA sequences of all chromosomes including their epigenetic modifications in the T2T-CHM13 map. I used data from the T2T-CHM13 map to locate the CENP-B boxes in the centromeres as anchor of kinetochores. Most of the CENP-B boxes in centromeric DNA are methylated with the exception of the so-called centromere dip region (CDR), where CENP-B protein dimers bind to adjacent unmethylated CENP-B boxes and interact with CENP-A and CENP-C proteins to assemble the kinetochore. The centromeres of all chromosomes combined have a size of 407 Mb of which the kinetochores account for 5.0 Mb or 1.2%. There is no correlation between centromere and kinetochore size (P = .77). While the number of CENP-B boxes varies 4-fold between chromosomes, their density (number/Kb) varies less than 2-fold with a mean of 2.61 ± 0.33. The narrow range ensures a uniform pull of the spindle on the centromeres. I illustrate the findings in a model of the human kinetochore anchored at unmethylated CENP-B boxes in the CDR and present circos plots of chromosomes to show the location of kinetochores in their respective centromeres.
Introduction
When the Human Genome Project was officially declared complete in 2003, large sections of the genome had not been sequenced because of highly repetitive segments of DNA that were difficult to align with the rest. There were more than 300 gaps, mainly involving the centromeres and their alpha satellite DNA containing repeated 171 bp sequences. Successive versions of the human genome reference have been published since the original draft, reducing the number of gaps. In 2022, the Telomere-to-Telomere Consortium finally filled in that remaining DNA, providing the first complete, gapless genome sequence, named the T2T-CHM13 map. 1 The availability of the T2T-CHM13 map offers the opportunity for computational biologists to analyze the centromere in greater detail.
Centromeres are built on DNA repeats of 171 base pairs, named alpha-satellites, which span several Mb. 2 The centromeric chromatin, which serves as the foundation of the kinetochore, contains a core of 3 proteins, namely Centromere Protein A, B, and C (CENP-A, CENP-B, CENP-C). Centromere Protein A exists in a complex with CENP-B and CENP-C, which aggregates with 4 additional modules of centromere proteins (CENP-HIKM, CENP-LN, CENP-OPQUR, CENP-TWSX) to form the constitutive centromere associated network (CCAN) of the inner kinetochore.3-5 The inner kinetochore, in turn, recruits the outer kinetochore KMN network, which comprises the KNL1C, MIS12C, and NDC80C complexes. 6 Finally, the KMN network attaches to the microtubules in the mitotic spindle. By connecting centromere and spindle, the kinetochore controls the migration of chromosomes during cell division.
Centromere Protein B is unique among the centromeric proteins because it is the only one that binds directly to DNA in a sequence-dependent manner. The protein contains a DNA-binding domain at the N-terminus and a dimerization domain at the C-terminus. 7 The DNA-binding domain recognizes and binds a 17 bp sequence, called CENP-B box, in the centromeric alpha-satellite DNA (YTTCGTTGGAARCGGGA; Y = Cytosine/Thymine; R = Guanine/Adenine) (Figure 1a). 8 The dimerization domain of CENP-B (Figure 1b) juxtaposes 2 distant CENP-B boxes through its dimer formation and DNA binding ability. 9 In addition, CENP-B binds directly to both CENP-A and CENP-C. 10 Thus, the CENP-B box serves as an anchor where binding of CENP-B triggers the attachment of CENP-A, which, in turn, leads to the stable assembly of CENP-C and additional inner and outer kinetochore components. 11 The pivotal role of the CENP-B box is influenced by 2 additional factors, namely epigenetic modification and density, that is, number of CENP-B boxes/Kb. The CENP-B box contains 2 CpG dinucleotides, which can become methylated (Figure 1a). However, CENP-B preferentially binds to the unmethylated CENP-B box DNA. 12 Thus, epigenetic modification of the CENB-P box reduces CENB-B binding and thereby recruitment of CENP-A and CENP-C. Importantly, the density of CENP-B boxes in the centromeric alpha-satellite DNA has also been correlated with stronger CENP-A enrichment, which, in turn, is a marker of kinetochore positioning.13,14 Both the epigenetic map of CENP-B boxes as well as the precise number and position of CENP-B boxes, that is, the density, in the centromeres were uncertain prior to the T2T-CHM13 map. Because of this double uncertainty, the exact place within each centromere where the kinetochore attaches to the chromosome has not been identified. Here, I analyzed CENP-B box data from the recently published T2T-CHM13 map to precisely localize the kinetochore attachment site within each centromere. I also present a model centered on CENP-B boxes as anchor of kinetochores in centromeres of human chromosomes.
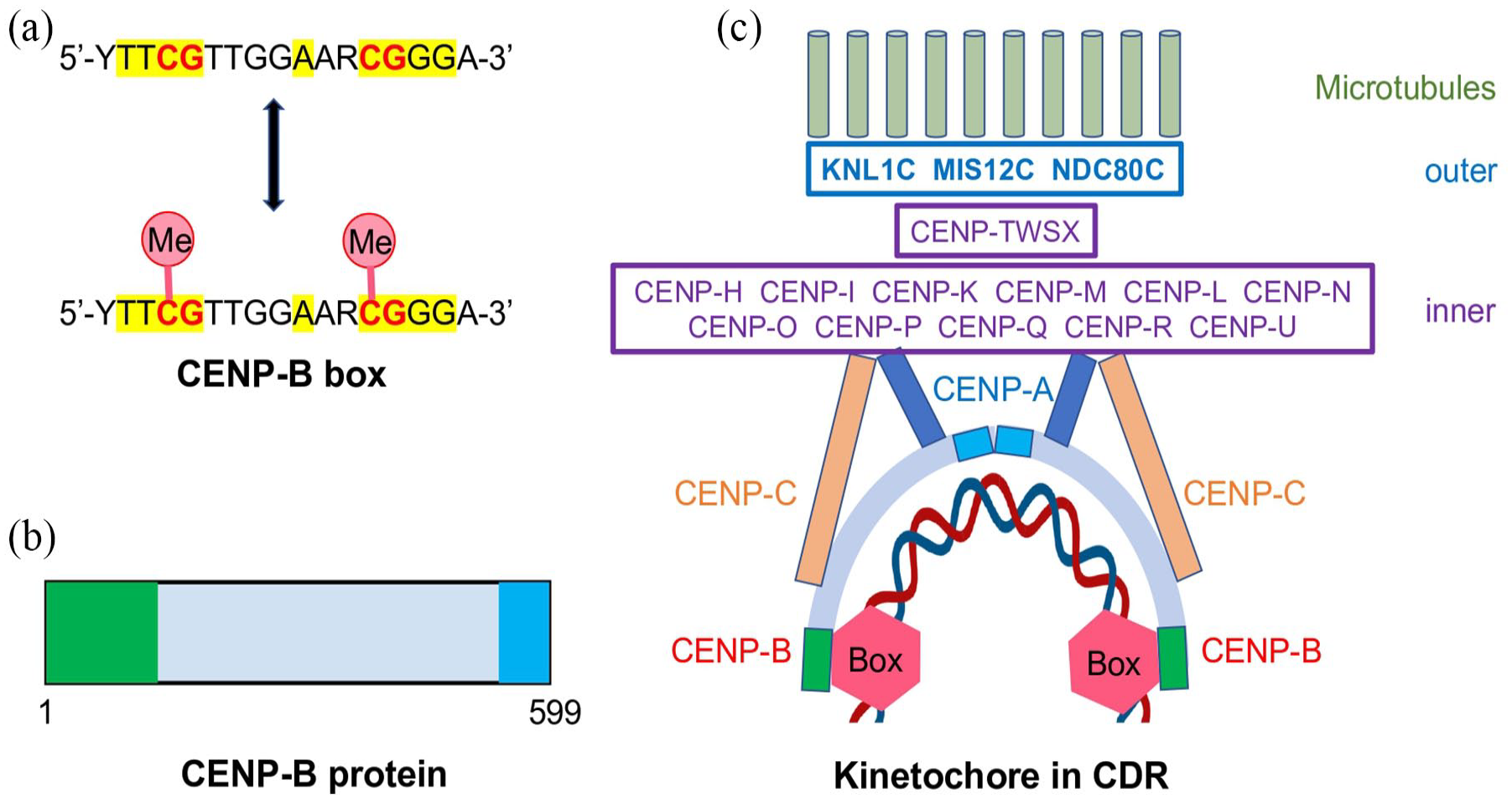
Diagram of CENP-B box, CENP-B protein, and kinetochore anchored on CENP-B boxes. (A) The CENP-B box consists of a 17 bp motif in which Y = Cytosine/Thymine and R = Guanine/Adenine. The 9 nucleotides highlighted in yellow form the core recognition sequence. The CpG dinucleotides indicated in red can undergo epigenetic modification in form of cytosine methylation (Me). (B) The human CENP-B protein contains a DNA-binding domain (amino acids 1-129) in green and a dimerization domain (amino acids 540-599) in blue. (C) Model of human kinetochore with anchor of unmethylated CENP-B boxes in CDR. A CENP-B dimer binds to 2 adjacent CENP-B boxes (red hexagon) and interacts with CENP-A and CENP-C to assemble the multicomponent inner kinetochore (purple boxes), which recruits the multicomponent outer kinetochore (blue boxes) to support the microtubules of the mitotic spindle.
Methodology
The scientists engaged in the T2T consortium published their results in a series of articles. I extracted data from the following articles.1,15,16 In particular, the centromere coordinates were obtained from STable 5, 16 the epigenetic map of CENP-B boxes from STable 11, 1 and the position and number of CENP-B boxes from database S15. 1
Circos plots of chromosomes, centromeres, and kinetochores were created with Circa (http://omgenomics.com/circa).
Results
In Table 1, I combined data extracted from 3 different sources.1,15,16 Both chromosomes and centromeres vary in size, ranging from 47 to 249 Mb and 12 to 42 Mb, respectively (Table 1; Figure 2). There is no correlation between chromosome and centromere size (P = .22). The T2T consortium generated epigenetic maps of all centromeres and found a region of reduced CpG methylation called centromere dip region (CDR) in each centromere.1,15 CUT&RUN experiments of CENP-A and CENP-B data showed that the hypomethylation colocalized with CENP-A enrichment and CENP-B binding at the CDR. Since CENP-A enrichment is a marker of kinetochore positioning, 13 I used the CDR coordinates to localize the kinetochore sites (Table 1). Finally, I combined STable 11 and database S15 to determine the number and density of CENP-B boxes in each CDR (Table 1). 1 The centromeres of all chromosomes combined have a size of 407 Mb of which the kinetochores account for 5.0 Mb or 1.2% (Table 1; Figure 2). There is no correlation between centromere and kinetochore (P = .77) or chromosome and kinetochore (P = .88) size. While the number of CENP-B boxes in the CDR varies 4-fold from 288 (chromo X) to 1158 (chromo 19), their density varies less than 2-fold, ranging from 1.82 in chromo X to 3.10 in chromo 15 (Table 1). The mean density of CENB-P boxes was 2.61 ± 0.33 number/Kb.
Summary of chromosomes, centromeres, centromere dip regions (CDRs), and CENP-B boxes.


Circos plots showing chromosomes (outer track), centromeres (middle track), and kinetochores (inner track).
Circos plots of all chromosomes, centromeres, and kinetochores are presented in Figure 2. There is no consistent pattern in the position of centromeres in their respective chromosomes nor in the position of kinetochores in their respective centromeres. However, examination of acrocentric chromosomes (13-15, 21, 22) reveals that the kinetochores are positioned off-center in their respective centromeres and shifted away from the telomeres.
Discussion
The team of scientists engaged in the T2T genome project accomplished 2 goals. They closed the remaining gaps in the DNA sequence and thereby provide a truly complete DNA sequence of the human genome. On top, they elucidated the epigenetic modifications of these newly completed sequences. Both accomplishments were essential for this study. While the sequence of the CENP-B box (Figure 1a) has been known for several years, 8 the precise number and exact location of CENP-B boxes that function as high-affinity sites for the CENP-B protein could only be determined now. Most of the CENP-B boxes in centromeric DNA are methylated with the exception of the CDR, where the CENP-B box is unmethylated.1,15 The identification of the CDR is functionally relevant because the CENP-B protein preferentially binds to the unmethylated CENP-B box DNA. Competition analyses revealed that the affinity of CENP-B for the CENP-B box DNA is reduced nearly to the level of nonspecific DNA binding by CpG methylation. 12 Thus, epigenetic methylation of the CENB-P box reduces CENB-B binding and thereby impairs kinetochore assembly. In addition to the DNA-binding domain, the CENP-B protein contains a dimerization domain (Figure 1b), which bundles 2 distant CENP-B boxes through its dimer formation and DNA-binding ability, thereby creating a higher-order structure in the centromere by juxtaposing 2 CENP-B boxes. 9 The DNA-protein structure becomes enlarged because CENP-B binds directly to both CENP-A and CENP-C. 10 Thus, the unmethylated CENP-B boxes serve as broad-based anchor in the centromeric DNA where binding of dimeric CENP-B triggers the attachment of CENP-A, which, in turn, leads to the stable assembly of CENP-C and additional inner and outer kinetochore components. 11 In Figure 1c, I present a model centered on unmethylated CENP-B boxes in the CDR as anchor of kinetochores in human chromosomes.
The pivotal role of the CENP-B box is further illustrated by its density, that is, the number of CENP-B boxes/Kb. The density of CENP-B boxes in the centromeric alpha-satellite DNA has been correlated with stronger CENP-A enrichment, which, in turn, is a marker of kinetochore positioning.13,14 While the number of CENP-B boxes in the CDR varies 4-fold from 288 (chromo X) to 1158 (chromo 19), their density varies less than 2-fold, from 1.82 in chromo X to 3.10 in chromo 15 (Table 1). Since the CENP-B boxes serve as anchor of kinetochore assembly, their narrow density range (mean 2.61 ± 0.33 number/Kb) signifies an even support of microtubules. The narrow range is important to ensure a uniform pull of the spindle on the centromeres.6,17
The size (surface area) variability of human kinetochores was first shown by morphological studies. In agreement with this study, the immunofluorescence intensity of anti-kinetochore antibodies varied 4-fold between chromosomes. 18 An analysis of serial-section electron micrographs determined the average surface area of kinetochores and found that kinetochore microtubules are packed more densely on smaller kinetochores. 19 While these earlier morphological studies provided detailed measurements, they could not identify individual chromosomes but only divide them into small and large ones. In contrast, the molecular data derived in this study from the T2T genome project allowed the actual determination of kinetochore numbers in the centromere of each individual chromosome. Thus, I could correlate the sizes of chromosomes, centromeres, and kinetochores (Table 1).
Interestingly, functional centromere studies have shown that chromosome segregation fidelity depends mainly on CENP-B bound to centromeric DNA as the sole source of centromere/kinetochore interaction. 10 A recent study reported that alpha-satellite arrays with more CENP-B boxes and thus greater CENP-B binding were less likely to mis-segregate. 20 Specifically, in CENP-A depleted RPE-1 cells there was a statistically significant negative correlation between the rate of chromosome mis-segregation and the abundance of CENP-B boxes.
In conclusion, unmethylated CENP-B boxes located in the CDRs of centromeres serve as anchors for kinetochore assembly. A CENP-B dimer binds to 2 adjacent CENP-B boxes and interacts with CENP-A and CENP-C to assemble the multicomponent kinetochores, which support the microtubules of the mitotic spindle. The narrow density range of CENP-B boxes ensures a uniform pull of the spindle on the centromeres.
Footnotes
Funding:
The author received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Not applicable.
