Abstract
Breast cancer is the most commonly diagnosed cancer globally, with the highest incidence of breast cancer occurring in Asian countries including Indonesia. Among the types of breast cancer, the estrogen receptor (ER)-positive subtype which is prominent with estrogen receptor alpha (ERα) and heat shock protein 90 (HSP90) overexpression genes becomes the most prevalent than the others, approximately 75% of all breast cancer cases. ERα and HSP90 play a role in breast cancer activities including breast tumor growth, invasion, and metastasis mechanism. Propolis, a natural bee product, has been explored for its anticancer activity. However, there is lack of studies that evaluated the potential inhibitor from propolis compounds to the ERα and HSP90 proteins. Therefore, this article focuses on examining the correlation between ERα and HSP90’s role in breast cancer and investigating the potential of 93 unique propolis compositions in inhibiting these genes in breast cancer using in silico approaches. This study revealed the positive correlation between ERα and HSP90 genes in breast cancer disease development. Furthermore, we also found novel potential bioactive compounds of propolis against breast cancer through binding with ERα and HSP90; they were 3’,4’,7-trihydroxyisoflavone and baicalein-7-O-β-D glucopyranoside, respectively. Further research on these compounds is needed to elucidate deeper mechanisms and activity in the real biological system to develop new breast cancer drug treatments.
Introduction
Breast cancer is the most commonly diagnosed cancer globally with the highest incidence of breast cancer and mortality rates occurring in Asian countries including Indonesia. 1 According to the World Health Organization (WHO), there were 2.3 million women diagnosed with breast cancer and 685 000 deaths globally in 2020. In 2021, it is predicted that the rate of breast cancer will rise to 85 cases per 100 000 women. The incidence of breast cancer in the United States is 1 in 8 women, whereas in Asia, 1 woman has breast cancer out of 35 people. 2 The standard age of incidence of breast cancer in Indonesia is 44 years and 15.3% per 100 000 population, where most breast cancers diagnosed occur at an advanced stage with a low survival rate. 1
Breast cancer is caused by gene mutations of mammary gland epithelial cells by various factors such as the presence of radiation exposure, injury at the cellular level, and aging of DNA or RNA, as a result of which cell proliferation is not well controlled.3,4 The development of breast cancer is influenced by various elements, one of the important parameters is the immune cells from the tumor microenvironment. Less immune infiltration can imply a weaker immune response against tumor activity resulting in poor prognosis and low survival rate. Several studies reported that estrogen receptor alpha (ERα) and heat shock protein 90 (HSP90) genes decrease immune activity in the tumor environment.5-7 Furthermore, Rimsza et al 8 and Brown et al 9 showed that loss of major histocompability complex (MHC) II in the ER+ tumor environment promoted to lack of B-cell activation. B-cells are involved in tumor immunology through antibody production, antigen presentation, cytokine and chemokine production, and other immunoregulatory mechanisms. 10 Recently, combinatorial therapeutics that focused on immunotherapy, by targeting B-cells and related receptors, were suggested to be more developed by researchers. 11
In addition, ERα and HSP90 play a crucial role in breast cancer development. ERα induces tumor growth in breast cancer through its binding to estrogen, thereby increasing the proliferation of breast cancer. 12 Moreover, estrogen stimulation causes ERα to translocate within the nucleus and regulate a certain set of gene expressions through its interaction with estrogen response elements (EREs). 13 As for HSP90, it can promote invasion, and migration of cancer cells which further leads to tumor metastasis. 14 HSP90 also interacts with the c-Myc transcription factor to prevent proteolytic degradation and promotes tumor progression, interacts with HIF-1α (a transcription factor that works when oxygen is reduced), and plays a role in metabolic reprogram control. This control will help the cancer cells to maintain their growth and development through some crucial processes such as glycolytic metabolism. 14
The therapeutic approaches generally varied ranging from hormonal therapy, cytotoxic chemotherapy, radiotherapy, and targeted therapies such as the development of combination treatments. 15 Generally, hormonal therapy becomes the main therapy to treat ER-positive breast cancer complemented by surgery in most patients. 16 Hormonal therapy aims to lower hormone levels involved in the growth of breast cancer and inhibit the growth of such cancer. However, the therapy still causes side effects including hot flashes, venous thromboembolism, an increased risk of endometrial cancer, fatigue, poor sleep, and difficulty concentrating which significantly impacts the women’s quality of life. 17 Therefore, new treatments using safer medications that are derived from natural substances need to be explored for new breast cancer treatment.
Propolis also known as bee glue is a resinous substance collected by bees from various types of plants, which is used to seal holes (cracks) and reconstruct hives. 18 The content of propolis compounds depends on plant sources and bee species and will generally consist of polyphenol compounds (flavonoids, phenolic acids, and esters), aldehydes, and fenol ketones. 19 The active compound propolis is rich in benefits, such as having antioxidant, anti-inflammatory, and antimicrobial activity. 20 The antitumor activity of propolis in human breast cancer has also been studied. 21
Some previous studies have explored the anticancer activity of propolis extract. Caffeic acid phenyl ester (CAPE), artepillin C, and chrysin become the key compounds of propolis that are responsible for anticancer potential activity. 22 Caffeic acid phenyl ester significantly increases endothelial and inducible nitric oxide synthase in breast cancer cell lines experiment, thereby it exhibits the antiproliferative and cytotoxic effects of propolis against breast cancer. 23 Caffeic acid phenyl ester can promote apoptosis in breast cancer cell lines MCF-7 (human breast cancer ER[+]) and MDA-MB-231 (human breast cancer ER[−]). 21 In melanoma and carcinoma cells, artepillin C causes cytotoxicity by triggering apoptosis and inhibiting mitosis. Moreover, major compounds of propolis, flavonoids, and polyphenol were found to have chemopreventive and anticancer effects. Previous studies found that propolis compounds can induce apoptotic and inhibit cell proliferation through multiple signaling pathways, such as mitogen-activated protein kinase (MAPK), phosphoinositide 3-kinase (PI3K)/akt, tumor necrosis factor α (TNFα)/nuclear factor kappa B (NF-kB), signal transducer and activator of transcription 3 (STAT-3)/polo like kinase 1 (PLK-1), and extrinsic and intrinsic apoptotic pathways. 22 Besides, limited studies still evaluated the potential inhibitor from propolis compounds to the ERα and HSP90 proteins which are prominently involved in breast cancer. Kamiya et al have investigated ethanol extract of Brazilian red propolis and found that it can induce MCF-7 cell apoptotic through ER stress-related pathway. 24 Some previous studies, using in silico methods such as network pharmacology and molecular docking, have found that Nigerian, Egyptian, and Sulawesi propolis compounds can potentially target neither ERα, HSP90, or other receptors related to anticancer activity.25-27 As propolis composition varies greatly depending on its origin, plant source, bee type, and extraction process, this affects the diversity of propolis biological activity from one another. 28 Therefore, to gain more knowledge and fill the gap in current research, this article will investigate the potential of 93 unique propolis compositions for inhibiting protein receptors of ERα, and HSP90 in breast cancer using in silico approaches. This study aims to reveal the novel potential bioactive compounds of propolis against breast cancer through binding with ERα and HSP90.
Materials and Methods
Gene expression level of ESR1 and HSP90 analysis among cancer types
TIMER database (Tumor Immune Estimation Resource) (https://cistrome.shinyapps.io/timer/) was used to analyze the gene expression analysis among cancer types compared with normal tissue. In the TIMER database, the DiffExp module was selected for the option to conduct this analysis. Then, ESR1 and HSP90AA1 were input to assess the expression pattern in the cancer disease.
Gene expression and survival related to ESR1 and HSP90 genes in breast cancer patients compared with normal condition
UALCAN database (https://ualcan.path.uab.edu/) was used to analyze gene expression and survival data from breast cancer patients compared with normal people. TCGA (The Cancer Genome Atlas) gene and breast invasive carcinoma were selected to analyze the interest gene. Next, expression and survival options were shown to get the result.
Correlation analysis between ESR1 and HSP90 genes to immune cells infiltration
To further analyze ERα and HSP90 genes in breast cancer, the TIMER database was used to show the relationship between the interest genes and immune cells infiltration level. Gene, survival, and correlation modules were selected with breast invasive carcinoma as the type of cancer. The rest parameters were kept default.
Kyoto Encyclopedia of Genes and Genomes cancer pathway analysis of ESR1 and HSP90 genes
The Database for Annotation, Visualization, and Integrated Discovery (https://david.ncifcrf.gov/home.jsp) was used to investigate the role of ERα and HSP90 in the cancer pathway. ERα and HPS90 were input using the gene official symbol (ESR1 and HSP90AA1) and started analysis which Homo sapiens as the species target. Then, Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway was selected to show the cancer pathway regarding the interest genes.
Protein structure preparation
All protein structures used were obtained from the Research Collaboratory for Structural Bioinformatics Protein Data Bank (RCSB PDB) database (https://www.rcsb.org/). The PDB ID used is a human protein and fulfills the following criteria: (1) does not have mutations in the sequence of amino acid residues forming a protein; (2) the experimental method for obtaining the 3-dimensional structure is X-ray diffraction; (3) has a main ligand in the form of a small molecule (native ligand); and (4) has published papers. We selected 3UUC for ERα protein and 5J64 for HSP90 protein. Then, the downloaded protein structure was prepared using AutoDockTools version 4.2 program (Scripps Research Institute, La Jolla, CA, USA) including adding polar hydrogens, merging nonpolar hydrogens, and adding a charge. Parameter values were set to default.
Ligand structure preparation
We selected 93 propolis compounds from our previous study that was prepared to simulate their interactions with each target protein. 29 The compounds are downloaded from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) and created manually with ChemSketch if not available in PubChem. SMILES and additional information are also taken from PubChem. All ligand preparations used AutoDockTools version 4.2 program (Scripps Research Institute) with default parameters including adding polar hydrogens, merging nonpolar hydrogens, and adding a charge.
Ligand evaluation
ADMET evaluation was carried out to predict the pharmacokinetic properties and toxicity of each ligand, including evaluation based on Lipinski’s Rule of Five with SwissADME, prediction of the oral rat LD50 in Hodge & Sterner scale with T.E.S.T software, prediction of hepatotoxicity with the Protox-II web server, and Toxtree to predict the mutagenic and carcinogenic properties of ligand. The 4 software uses SMILES from the ligands as input. The mutagenic and carcinogenic properties of ligands in the Toxtree software were evaluated based on the Benigni-Bossa rule, which predicts the carcinogenic and mutagenic properties based on the presence of potentially mutagenic or carcinogenic fragments or functional groups of ligands. 30
Molecular docking
Molecular docking validation was carried out first to determine the grid box size, indicating the protein’s active site. Validation was performed by redocking each protein with its native ligand (redocking cocrystallized ligand method) using AutoDock4 in PyRx software. The validation parameter used was root mean square deviation (RMSD) < 2 Å which was analyzed using the LigRMSD web server (https://ligrmsd.appsbio.utalca.cl/). Then, molecular docking of all ligands with each target protein was performed using AutoDock4 in PyRx. The docking site used the size and location of the grid box as native ligand-binding site from the docking validation step with default values (Table 1). PyMol and BIOVIA Discovery Studio Visualizer were also used to visualize molecular docking results.
Grid selection of ERα and HSP90 proteins.

Abbreviations: ERα, estrogen receptor alpha; HSP90, heat shock protein 90.
Conserved amino acid and biological activity analysis
Conserved amino acid analysis was investigated using the ConSurf web server (https://consurf.tau.ac.il/). 3UUC and 5J64 were input to the web server and selected chain A as the structure analysis. The rest parameters were kept default. The result of the conserved amino acid was shown by the gradation scaling color in each amino acid sequence.
Meanwhile, the Prediction of Activity Spectra for Substances (PASS) was conducted using the Way2Drug web server (http://www.way2drug.com/). To assess the PASS value, the top 3 ligands based on their binding affinity were selected and input to the web server using their SMILES which were obtained from the PubChem database. The result was selected that has a Pa value of more than 0.3.
Molecular dynamics
Molecular dynamics simulations were carried out to determine the stability of protein-ligand interactions in a dynamic environment. GROMACS software is used to form a simulation system, equilibration, and MD production, until the analysis step. The simulation was carried out for 25 ns with a simulated environment similar to the cytoplasmic conditions in human body cells. PyMol was also used to visualize protein-ligand interactions during the simulation. 31
Results
Gene expression and patient survival related to ESR1 and HSP90 genes among cancer types
ERα and HSP90 show the different abundance in the different types of cancer (Figure 1). From Figure 1, HSP90 has a higher expression level than ERα among cancer and normal condition. It indicates that HSP90 is more abundant in the various tissue of the human body than ERα, as ERα expresses a specific hormone. Furthermore, Figure 1 also reveals that ERα and HSP90 have the highest expression level in breast cancer among other cancer types with a significant P value indicated by an asterisk sign compared with normal conditions. Hence, targeting these genes is the crucial step to treating cancer further.

Gene expression of ERα (A) and HSP90 (B) in various cancer diseases.
Next, we use the UALCAN database to investigate the gene expression for only breast cancer and its survival data. From Figure 2A and B, it was confirmed that ERα and HSP90 were upregulated compared with the normal condition in which HSP90 has more gene expression level than ERα. Then, from Figure 2C and D, it was clear that cancer patient who has higher ERα and HSP90 expression intended to have a higher risk of relapse than those with lower expression genes.

Clinical analysis results of the relationship between the interest genes and breast cancer from the database. Expression level of ERα (A) and HSP90 (B). Survival analysis of ERα (C) and HSP90 (D) related genes.
Gene expression related to immune cells infiltration level
TIMER database is a web server that contains several modules to conduct analysis of the correlation of immune cells infiltration with various cancer types. From the result of this web server, ERα and HSP90 gene expression showed almost similar patterns relating to immune cell infiltration. From Figure 3, these genes have a negative correlation with immune cell activity as more genes are expressed toward less immune infiltrated. Furthermore, in Figure 4, it was shown that the expression of ERα and HSP90 genes is positively correlated.

Interest genes related to the immune system in the breast cancer model.
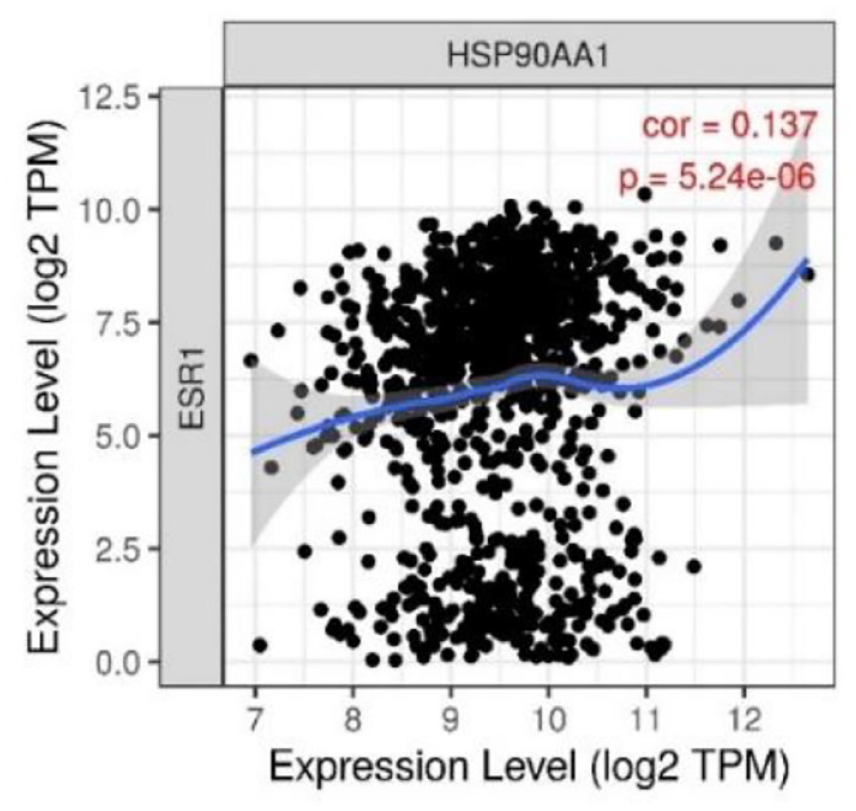
Expression correlation between ERα and HSP90 in the breast cancer model.
Propolis ligands evaluation
Propolis screening from our previous study using liquid chromatography-mass spectrometry (LC-MS/MS) revealed 93 compounds. Major of these compounds include to the flavonoid family which is divided into the most prominent classes such are flavone, flavonol, isoflavone, and flavanone (Figure 5 and Supplemental Table S1).

2D structure of flavonoid backbone compound: (A) flavone, (B) flavonol, (C) isoflavone, and (D) flavanone.
Molecular docking and ADMET evaluation
Molecular docking is important to know the capability of our interest compounds to bind to our protein’s target. The docking protocol was validated using the redocking method which is docking the cocrystalized ligand to its receptor. We use 4,4’-(2,2-dichloroethene-1,1-diyl) diphenol (0D1) as the native ligand for ERα and 5-(2,4-dihydroxy phenyl)-4-(2-fluorophenyl)-2,4-dihydro-3H-1,2,4-triazol-3-one (6G7) as the native ligand for HSP90 protein. From Table 2, the RMSD value was 0.54 Å for 0D1 to ESR1 and 0.5 Å for 6G7 to HSP90. To more give detail, the visualization of the ligand’s pose was shown in which the yellow structure represents the native ligand before the validation process. Hence, the docking validation was successful as the RMSD value was below 2 Å.
Redocking native ligand result.

Abbreviations: HSP90, heat shock protein 90; RMSD, root mean square deviation.
According to the docking result (Tables 3 and 4), the top 5 highest binding affinities ranged from −9.2 until −8.39 kcal/mol from ERα and −9.4 until −9.21 kcal/mol for HSP90 protein. Meanwhile, the control and native ligand for ESR1 protein obtained −6.03 and −8.37 kcal/mol, whereas the control and native ligand for HSP90 protein obtained −9.18 and −8.24 kcal/mol. Interestingly, fawcettiine showed the highest binding affinity for ERα and HSP90 protein which gained −9.2 and −9.4 kcal/mol, respectively. All top 5 ligands for ERα protein revealed no violation of Lipinski’s rule; however, only fawcettiine and wogonin 7-O glucoside in the HSP90 protein which has no Lipinski’s rule violation whereas the remained ligands have only 1 violation.
Top 5 highest binding affinity ligands from ERα protein.

Abbreviation: ERα, estrogen receptor alpha.
Drug standard used as control.
Native ligand of protein.
Top 5 highest binding affinity ligands from HSP90 protein.

Abbreviation: HSP90, heat shock protein 90.
Drug standard used as control.
Native ligand of protein.
Furthermore, we also assess the toxicity characteristic of the top 5 ligands from each protein (Tables 5 and 6). The LD50 value for ERα best ligands ranged from 534.63 to 2097.67 kg/mol whereas for HSP90 ranged from 1052.77 to 1544.83 kg/mol. Thus, some particular toxicity parameters are also important regarding drug design such as hepatotoxicity, mutagenicity, and carcinogenicity. All ligands from both proteins showed negative activity from these parameters except for adenosine in ERα protein which has predicted active activity in mutagenicity and carcinogenicity.
Toxicity evaluation of top 5 highest binding affinity ligands from ERα protein.

Abbreviation: ERα, estrogen receptor alpha.
Toxicity evaluation of top 5 highest binding affinity ligands from HSP90 protein.

Abbreviation: HSP90, heat shock protein 90.
Conserved amino acids analysis
Tables 7 and 8 showed the detailed interaction from the top 5 ligands of each protein whereas the 2-dimensional (2D) interaction was shown in Supplemental Figures S1 and S2. Bold residues shown in the table are the catalytic residue based on the original literature that published the crystal ligand on the RCSB website. According to Delfosse et al, 32 Glu353, Arg394, and His524 act as important residues on the end of the active pocket in the ERα protein. Based on Table 6, Naringenin and Adenosine became the ligand that fully interacted with the 3 important residues whereas the other ligands built interaction with 2 important residues. Furthermore, all top ligands in the ESR1 protein also built various interactions with the ERα protein such as hydrogen, hydrophobic, anion, and van der Waals bonds.
Docking interaction of top 5 highest binding affinity ligands from ERα protein.

Abbreviation: ERα, estrogen receptor alpha.
Bold letter means key active residues of the protein.
Docking interaction of top 5 highest binding affinity ligands from HSP90 protein.

Abbreviation: HSP90, heat shock protein 90.
Bold letter means key active residues of the protein.
On the other side, according to Abbasi et al, 33 Leu48, Ser52, Asp93, and Thr184 are important residues in the catalytic pocket for the HSP90 protein especially involved in ATPase activity. Based on Table 8, the top 3 ligands in the HSP90 protein fully interacted with all these important residues whereas wogonin 7-O-glucoside interacted with 2 important residues and apigenin-7-ogalactopyranoside only interacted with 1 important residue. All ligands also built various chemical interactions with HSP90 protein from hydrogen, hydrophobic, anion, and van der Waals bonds except Fawcettiine which did not build anion interaction.
Conserved amino acid analysis was conducted to examine the structure sequence of ERα and HSP90 regarding important residues and validated these residues stated from the literature. From the result of the ConSurf web server, scaling conserved amino acid sequences from both proteins were displayed in Figures 6 and 7. From Figure 6, it was shown that Glu353, Arg394, and His524 are highly conserved regions and act as functional residues. Similar to Figure 6, highly conserved amino acids also were possessed by Leu48, Ser52, Asp93, and Thr184 as shown in Figure 7.

Conserved amino acids analysis from ERα protein. Black round for Glu353, red round for Arg394, and green round for His524. ERα indicates estrogen receptor alpha.

Conserved amino acids analysis from HSP90 protein. Black round for Leu48, red round for Ser52, green round for Asp93, and purple round for Thr184.
Molecular dynamics results
After conducting a 25 ns molecular dynamics simulation, RMSD, root mean square fluctuation (RMSF), solvent-accessible surface area (SASA), and gyration values were obtained from each ligand to each protein. From Figure 8, all forms showed almost similar patterns that have low fluctuation, except for Naringenin in the RMSF value which had more fluctuating residue. Moreover, Figure 9, also showed a similar pattern for all molecular dynamics results. However, RMSD results in HSP90 protein showed slightly different from each other. Baicalein-7-O-β-D glucopyranoside and onalespib showed the lowest and most stable RMSD results than others that interacted with the HSP90 protein.

Molecular dynamics results from ERα protein. RMSD (A), RMSF (B), radius of gyration (C) and SASA result (D).

Molecular dynamics results from HSP90 protein. RMSD (A), RMSF (B), radius of gyration (C) and SASA result (D).
Furthermore, Table 9 shows that 3’,4’,7-trihydroxyisoflavone became the ligand that had successfully kept all important residues from 0 ns until 25 ns and 1 of them built the hydrogen bond constantly to Glu353 at a close distance. Then, Table 10 shows that only 7-O-α-L rhamnopyranosyl kaempferol could not maintain full interaction with important residues during 25 ns. Besides, baicalein-7-O-β-D glucopyranoside successfully built the hydrogen bonds to Asp93 and Thr184 with a distance below 2.5 Å.
Important amino acid interaction during molecular dynamic (MD) simulation between ligands and ERα protein.

Abbreviation: ERα, estrogen receptor alpha.
Important amino acid interaction during molecular dynamic (MD) simulation between ligands and HSP90 protein.

Abbreviation: HSP90, heat shock protein 90.
Biological activity prediction
To validate the potential active ability of anticancer and other potential activities from the best-selected ligands, we used the PASS examination based on the similarity from the active structure contained in the data set. Therefore, it is designed to evaluate the general biological function of an organic drug-like molecule based on its structure. From Supplemental Tables S2 until S6, the most prominent biological activity among selected ligands was membrane permeability inhibitor, whose Pa value ranged from 0.7 to 0.9. Furthermore, from Table 11, all selected ligands showed anticancer potential activity.
Anticancer activity predictions of best ligands from both proteins.

Discussion
Breast cancer is the most prevalent cancer among other cancer types in Indonesia. 34 Furthermore, GLOBOCAN revealed 2.3 million new cases (11.7%) of breast cancer and a mortality rate of 6.9% which showed a high rate. 35 According to the data, developing breast cancer therapy is urgently needed especially in Indonesia. Some receptors have been linked to breast cancer control including ERα and HSP90 which are prevalent among breast cancer patients.
In Figures 1 and 2, there was a clear pattern among selected genes, expression level, and survival rate in breast cancer patients. ERα and HSP90 play a vital role in breast cancer disease as the high expression of both genes decreases the survival rate of the patient. These findings were supported by other research which stated both genes are upregulated in breast cancer.36-39 Pick et al 40 stated that high expression of HSP90 in breast cancer is related to high-risk cancer patients and its inhibition will decrease the tumor growth and promote the apoptosis mechanism. Wang et al 41 also reported that an inhibitor drug for HSP90 successfully decreased MCF-7 cell growth and induced the apoptosis mechanism. It implied that HSP90 plays a vital role in the evading apoptosis mechanism in the cancer cells as supported by our data in Supplemental Figure S3. Therefore, the high expression of the gene can maintain the malignant phenotype and metastasis and promote treatment resistance under cancer therapy. 42 On the contrary, ERα also has essential biological activity in breast cancer progression. High expression of this gene will promote the proliferation of cancer cells driving them to a more dangerous cancer state because activating the ER pathway forces the cell from the G1 phase to the S phase. 39 Supplemental Figure S4 from KEGG data showed that ERα plays a direct role in the cancer proliferation cells through the estrogen signaling pathway.
The development of breast cancer is influenced by various elements besides the gene expression level. One of the important parameters is the immune cells from the tumor microenvironment. An understanding of immune cell infiltration in breast cancer disease has essential implications for targeting and developing cancer therapy. From Figure 3, both ERα and HSP90 showed immunosuppressive activity because these genes have a negative correlation with immune cell infiltration levels. These data may reveal a correlation with Figure 2 which low survival patients in with overexpressed interest genes. Less immune infiltration can imply a weaker immune response against tumor activity resulting in poor prognosis and making the patient harder to be treated. Several studies also reported that ERα and HSP90 genes involve in decreasing immune activity in the tumor environment,5,6 specifically, Mostafa et al 5 revealed the decreased MHC-II in the ER+ tumor environment. Rimsza et al 8 and Brown et al 9 showed that loss of MHC-II promoted to lack of B-cell activation and it was clear regarding Figure 3 that was a lack of B-cell infiltration.
According to Figure 3, B-cells and dendritic cells were the most negative correlation to ERα whereas B-cells and CD4+ T-cells were the most negative to HSP90. From these findings, B-cells showed the poorest infiltration level regarding the high expression of ERα and HSP90. B-cells are involved in tumor immunology through antibody production, antigen presentation, cytokine and chemokine production, and other immunoregulatory mechanisms. 10 Furthermore, a recent study revealed that better immunotherapy outcomes from cancer patients were associated with B-cells.43-45 Therefore, the lack of B-cell infiltration in breast cancer regarding overexpression from ERα and HSP90 promoted to poor-survival rates in the patients.
As ERα and HSP90 are important for targeting cancer therapy, we continue to assess the relationship between these genes. From Figure 4, our analysis showed a positive correlation between the expression of ERα and HSP90 in breast cancer as supported by the result from Figures 1 and 2. In another study, Zhang et al reported the increased expression of HSP90 promoted poor survival in ER+ breast cancer patients due to the development of metastasis mechanism. Furthermore, the study also stated that the higher risk of recurrence in ER+ breast cancer also was influenced by the high expression of the HSP90 gene. This may be a result because HSP90 plays a vital role in cancer growth and survival, such as ER, progesterone receptor (PR), and essential components of human epidermal growth factor receptor 2 (HER2) signaling.46,47 Among the types of breast cancer, the ER-positive subtype which is prominent with ERα and HSP90 overexpression genes becomes the most prevalent than the others, approximately 75% of all breast cancer cases. 7 Therefore, discovering an alternative drug to treat ERα and HSP90 protein related is important research to cope the breast cancer disease, especially in Indonesia.
Propolis becomes one of the potential natural sources to treat many types of cancer. We have investigated the potential biological activity of propolis in our previous studies. Our previous study assessed our propolis content using LC-MS/MS method. 29 We found that neobavaisoflavone, methylophiopogonone A, 3’-methoxydaidzin, and genistin as novel candidates for anti-SARS CoV-2 infection through inhibition of S2 and Mpro, 48 whereas (RR)-(+)-30-senecioylkhellactone and pinobanksin-3-O-butyrate as novel aldose reductase inhibitors in the treatment of type-2 diabetes mellitus. 31 Moreover, diosmetin, cosmosiin, genistin, and 3’-methoxy-5’-hydroxyisoflavone-7-O-β-D revealed as novel candidates for inhibition of Streptococcus mutans activity in caries. 29 We also found that isoetin poses strong binding interaction to AChE, Caspase3, MAPK14, NF-kB, and TNF-α that related to neuroinflammation. 49 Besides, numerous studies reported the potential antibreast cancer activity of propolis that depend on their active compounds, such as CAPE, artepillin C, and chrysin. These compounds showed apoptotic, cytotoxic, and antiproliferative effects on breast cancer. 22 However, in this study, there were 93 unique flavonoid and polyphenol compounds of propolis that rarely investigated their antibreast cancer activity, especially in targeting ERα and HSP90.
Flavonoids are secondary metabolites in plants that have phenolic structures in many forms. The most prominent class in flavonoid groups are flavone, flavonol, isoflavone, and flavanone which are also contained in our propolis. 29 The diverse structures of flavonoids make them have the board pharmacologic activities including antioxidant, antimutagenic, antibacterial, antiangiogenic, anti-inflammatory, antiallergic, enzyme modulation, and anticancer.50,51 The hydroxyl groups in the structure of the flavonoid may increase the free radical scavenging ability, and it will interfere with redox activity in the cells. Regarding that capability, flavonoids can inhibit the production of reactive oxygen species (ROS) in macrophages resulting in an immune-modulating process.52-56 Furthermore, Xu et al 57 and Ishak et al 58 reported that the flavonoids compound capable to induce CD8, CD4, and CD19 lymphocytes demonstrated immune-stimulating effects. Thus, the effect of propolis containing flavonoids on immune mechanisms is important and related to breast cancer patients, especially in treating the overexpressed ERα and HSP90 genes which results in immunosuppressed effects.
The common active substance of natural products found in our propolis (Supplemental Table S1), namely, ferulic acid, genistin, naringenin, and pinostrobin. Interestingly, these compounds have potential antibreast cancer activity in previous reports. Ferulic acid downregulated ER stress pathway that implicates its anti-inflammatory activity in breast cancer. 59 Genistin has been reported its promising antibreast cancer activity through cytotoxic and antiproliferative activity against MCF7 and MDA-MB-231 breast cancer cell lines. 60 Naringenin, from Citrus L extract, poses antibreast cancer activity by its mechanism in estrogen signaling, apoptotic, and inhibition of metastasis. 61 Pinostrobin, from Finger Root (Boesenbergia rotunda) extract, shows inhibition activity to breast cancer cell growth by targeting Matrix metalloproteinase (MMP) genes, vascular endothelial growth factor (VEGFR) genes, JAK3, CDK4, and KCNA3. 62 Therefore, using a computational approach, this study will provide an in-depth prediction of the antibreast cancer mechanism by the unique composition of propolis.
To assess the capability of propolis contents regarding for inhibiting ERα and HSP90 protein, we conduct molecular docking and molecular dynamics analysis. Before conducting docking to 93 propolis ligands, we took a docking validation process. Docking validation is used to ensure the docking procedure can be done. 63 Table 2 showed the molecular validation result in which the RMSD value was below 2 Å and the ligand pose after docking was very close to the original pose indicating the reliability of the docking protocols so the docking process can be processed further. 64 From Tables 3 and 4, we chose the top 5 highest binding affinity ligands which are also higher than native and standard drugs from each protein. Furthermore, we also examine the ligand characteristics and toxicities (Tables 3 to 6). All ligands showed favorable drug-like properties, except for adenosine which may be harmful in mutagenicity and carcinogenicity. Interestingly, from both proteins, fawcettiine showed the highest binding affinity. Fawcettiine has the most lipophilic characteristic than the other ligands (Tables 4 and 5). Lipophilicity means by the octanol-water partition coefficient. 65 This parameter is important for improving biological activity in drug design, especially for oral drugs. 66 The higher lipophilicity promoted easier passive diffusion in membranes, such as cell membranes that have a hydrophobic environment. According to Stocks, 67 lipophilicity between 1 and 3 has moderate solubility, permeability moderate, balanced volume of distribution, and potential for good absorption and bioavailability. Not only contribute to its biological characteristic but lipophilicity also influence the binding receptor-ligand ability. Previous studies68-70 revealed that large lipophilic groups with hydrophobic interactions can increase the binding affinity of the ligand. Furthermore, Wang et al have shown that there is a positive correlation between the lipophilicity of the iodinated compounds and the binding affinity. However, they also found that higher lipophilicity resulted in low drug clearance in the brain.
We continue to analyze the interaction of each selected ligand with its protein. From Tables 7 and 8, all selected ligands built hydrogen, hydrophobic, and van der Waals interactions except for electrostatic interaction as pi-sulfur which was only built by 7-O-α-L rhamnopyranosyl kaempferol and Baicalein-7-O-β-D glucopyranoside to HSP90 protein. According to Delfosse et al, 33 Glu353, Arg394, and His524 are the important residues in ERα protein because these residues are located at the end of the binding pocket from the protein. Meanwhile, Abbasi et al 34 reported that Leu48, Ser52, Asp93, and Thr184 are the important residues in the HSP90 protein. They are involved in HSP90 activity to form direct interaction with a ligand in the bottom of the binding pocket. According to Amitai et al, 71 Ma et al, 72 and Giaccia et al, 73 the important residues have a crucial role in drug development, targeting, or drug interaction studies and are generally well conserved in the structure. These important roles are gained because the conserved region generally has crucial structural and functional protein activity. 74 Figures 6 and 7 showed that all important residues from both proteins were predicted to have a high conservation value.
After conducting the molecular docking process, we were looking further for molecular dynamics simulation for 25 ns. In contrast to molecular docking, molecular dynamics simulates the protein-ligand interaction within certain solvents and time to show its conformational change. 75 From the molecular dynamics results, we collected RMSD, RMSF, SASA, and radius gyration (RG).
Root mean square deviation means the changing of atomic sequences in the entire protein at a certain time which is usually used to measure the difference confirmation to its reference structure.76,77 The average RMSD value from all forms in the ERα protein was around 0.05 Å. There were slight fluctuations during MD simulation in all forms; however, they kept fluctuating below 0.2 Å suggesting that the complex was stable (Figure 8A). On the other side, RMSD from HSP90 protein showed more fluctuation than ERα protein. The average RMSD was similar at 0.2 Å except for baicalein-7-O-β-D glucopyranoside and onalespib as a standard drug with almost all forms gained equilibrated after approximately 10 ns. In the HSP90 protein, baicalein-7-O-β-D glucopyranoside showed the lowest RMSD value than the other ligands (Figure 9A).
Root mean square fluctuation measures each residue of amino acid in the entire protein in a certain period. 78 Each residue in the protein will give influence its entire structure regarding the RMSF value. Figure 8B showed almost all similar RMSF patterns in all forms except for the fawcettiine ligand that had the highest fluctuation around residues 310 until 350, indicating a more change in conformation with the binding of ERα protein. All complexes were kept stable below 0.5 Å. Meanwhile, RMSF from HSP90 showed more fluctuations than in the ERα complexes; however, the average RMSF value was below 0.4 Å (Figure 9B). According to the figure, residue numbers between 97 and 140 have more fluctuation, suggesting the fluctuating sequence in this region in HSP90.
The radius of gyration was examined to evaluate the compactness of the protein structure. High compactness of the protein structure means that there is no significant change in the structure and stable. A weak intermolecular interaction introduced to the structure can promote the increase of RG value as the structure is less compact. 79 Figure 8C showed that ERα complexes were almost similar to RG value below 1.85 Å. Furthermore, the ERα protein only showed the lowest RG suggesting the introduction of a ligand makes some folding protein structure. A similar pattern was also shown in Figure 9C from the HSP90 protein. Although the average RG was below 1.75 Å lower than ERα complexes, the apo-protein form of HSP90 showed the lowest RG value which may show the role of the introduction from a ligand.
The solvent-accessible surface area (SASA) defines as the area of molecules that can be accessed by the environmental solvent. 79 Solvent-accessible surface area generally has a positive correlation with RG. The less compactness of a biomolecule promotes to increase in the molecular size, therefore the more surface from its molecule exposed to the environmental solvent. Both from Figures 8D and 9D, the original protein form without any ligand has the lowest SASA value, and it was supported by Figures 8C and 9D which from these figures, the RG value from the original form was also the lowest score than the others. The introduction of a ligand to a protein is essential for many biological functions. The introduction will change the protein shape, stabilizing some conformations and destabilizing others. Wankowicz et al 80 stated that the ligand introduction to certain proteins will affect its conformation with ligand-properties dependent. In some cases, the binding pocket of the protein will be more rigid than its side chain after ligand binding, and the other residues far from the pocket will tend to be more flexible, especially in nonsolvent-exposed areas.
To gain more detailed information from molecular dynamics simulation, we were looking forward to the important residue interactions during the simulation. Table 9 shows that 3’,4’,7-trihydroxyisoflavone was the only ligand that could maintain the full interaction with the ERα protein with all important residues during 25 ns with hydrogen bond, carbon-hydrogen bond, and van der Waals interactions. Furthermore, Glu353 was constantly bonded to the 3’,4’,7-trihydroxyisoflavone ligand with the hydrogen bond. These interactions were supported by the molecular dynamics result in Figure 8 which 3’,4,’7-trihydroxyisoflavone revealed a more stable simulation than the other complexes with ligands. On the contrary, Table 10 shows the interaction during molecular dynamics simulation in the HSP90 protein. P51 and p21 could maintain all interactions during simulation; however, baicalein-7-O-β-D glucopyranoside showed constant hydrogen bond interactions with Asp93 and Thr184 residues. These interactions suggested a more stable complex between baicalein-7-O-β-D glucopyranoside and HSP90 (Figure 9). Recent previous studies81-83 also reported that hydrogen bond plays a vital role in enhancing protein stability and affinity with the introduction of ligand binding. Furthermore, Ouassaf et al 81 and Imberty et al 84 reported that hydrogen interaction below 3 Å considered a strong interaction, and according to Tables 9 and 10, the hydrogen interaction was around below 3 Å.
Conclusions
To summarize our study, we have reported the crucial role of ERα and HSP90 in breast cancer disease. ERα and HSP90 are commonly involved in cell proliferation and avoiding immune response, respectively. Furthermore, these overexpressed genes in breast cancer were negatively correlated with immune activity. Then, to target ERα and HSP90, we examined the propolis content ability based on bioinformatics studies. We found 3,’4’,7-trihydroxyisoflavone from ERα protein and baicalein-7-O-β-D glucopyranoside from HSP90 protein were the best ligands to form the stable complex with the protein, based on molecular docking and dynamics studies also supported by the biological activity prediction value using PASS. Further research on these compounds is needed to elucidate deeper mechanisms and activity in the real biological system to develop breast cancer drug treatment.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322231224187 – Supplemental material for Revealing Propolis Potential Activity on Inhibiting Estrogen Receptor and Heat Shock Protein 90 Overexpressed in Breast Cancer by Bioinformatics Approaches
Supplemental material, sj-docx-1-bbi-10.1177_11779322231224187 for Revealing Propolis Potential Activity on Inhibiting Estrogen Receptor and Heat Shock Protein 90 Overexpressed in Breast Cancer by Bioinformatics Approaches by Masriana Vivi Simanjuntak, Muhammad Miftah Jauhar, Putri Hawa Syaifie, Adzani Gaisani Arda, Etik Mardliyati, Wervyan Shalannanda, Beni Rio Hermanto and Isa Anshori in Bioinformatics and Biology Insights
Footnotes
Acknowledgements
The authors thank School of Electrical Engineering and Informatics, Bandung Institute of Technology and Nano Center Indonesia for high performance computing and research facilities support. The authors express their gratitude toward PT Nano Herbaltama Internasional for providing propolis sample.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors thank a research grant from Bandung Institute of Technology under Riset Kolaborasi Indonesia 2023 program [project number LPPM.PN-6-36-2023 and contract number 1994/IT1.B07.1/TA.00/2023].
Competing interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
Masriana Vivi Simanjuntak: Performed bioinformatics analysis and writing original draft.
Muhammad Miftah Jauhar: Data interpretation, formal analysis, data curation, investigation, validation and writing original draft.
Putri Hawa Syaifie: Designed a conceptual framework, data curation, investigation, validation, review, editing, and project administration.
Adzani Gaisani Arda: Formal analysis and validation.
Etik Mardliyati: Acquisition of resources, data curation, investigation, validation, software, review, and editing.
Wervyan Shalannanda and Beni Rio Hermanto: Software.
Isa Anshori: Acquisition of resource, software, and investigation.
Ethics approval and consent to participate.
Not applicable.
Consent for publication
None.
Availability of data and materials
None.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
