Abstract
Salmonella species are Enterobacteriaceae associated with typhoid fever. In this study, the distribution of broad-spectrum β-lactamase regulatory genes and genetic relatedness of isolates was determined. Stool samples (400) were collected from patients with fever in Dalhatu Araf Specialist Hospital (DASH), Lafia, Nigeria, between March 2020 and April 2021. Salmonella species were isolated and extended-spectrum β-lactamase distribution was determined among resistant isolates using polymerase chain reaction (PCR). Genetic relatedness of Salmonella species resistant to the 10 first-line antibiotics administered was determined among S typhi isolated. Of the 60 isolates that were confirmed to belong to the genus Salmonella, 12 (20.0%) isolates with bla SHV genes were the most prevalent, blaOXA-1 and blaCTX-M-9 were present in 5 isolates each, while blaCTX-M-4 and blaTEM genes with a prevalence of 1.7% each were the least obtained in the isolates. Two isolates had a multidrug-resistant index (MDRI) of 1, and 2 others were positive with the S typhi staG gene. Sequencing to determine their diversity showed that isolates ST36 and ST138, respectively, had MDRI = 1 and are clustered in a group with a similarity coefficient of 0.00634. The 2 isolates had the highest genetic similarity, which indicates that the genetic diversity between the isolates is low, while Salmonella strain ST313L2 had a high level of genetic distance from the other isolates. The most resistant isolates are closely related which calls for concern.
Introduction
Gram-negative bacteria of the family Enterobacteriaceae are a concern with the ease at which they acquire resistance to known antibiotics. 1 Enterobacteriaceae are known extended-spectrum β-lactamases (ESBLs) producing bacteria accounting for huge morbidity and attendant cost burden on patients and health facilities.2,3 With challenges reported in the production and introduction of new antibiotics to reduce the demand placed on currently administered drugs, failing efficacies recorded to β-lactam antibiotics is due to the production of enzyme lactamases which bind and hydrolyze the beta-lactam ring to induce resistance. 4 The enzyme acts on most antibiotics except carbapenems, colistin, tigecycline, and clavulanic acid, which halts its activity.5,6
There are more than 150 types of ESBL enzymes produced by resistant members of Enterobacteriaceae. The ESBL enzymes are transferable among the members of the Enterobacteriaceae being plasmid-mediated. 7 The production of the enzymes is suspected to be triggered by the misuse or overuse of antibiotics. Infection with ESBL-producing bacteria is most common in health care settings where it prolongs the stay of hospitalized patients. The β-lactam hydrolyzing enzymes override the β-lactam antibiotics, which traditionally prevent the synthesis of the cell wall in bacteria. Bacterial species, geographical location, health institution, infection type, and prolonged abuse of antibiotics all affect the prevalence of β-lactamase producers.8,9
In Nigeria and other developing nations, the challenge posed by ESBLs producing bacteria is not well quantified due to poor health facilities availability, crowding in hospitals, poverty, poor hygiene practices, ignorance, inappropriate use of antibiotics, and consumption of animal products. These factors drive the transmission of multidrug-resistant bacteria and especially ESBL-producing bacteria. Nigeria has the highest incidence of intestinal perforation from typhoid fever caused by Salmonella. 10 Salmonella species are enteric pathogens associated with infectious diseases resulting from poor water supply and sanitation. Salmonella enterica serovar Typhi, Paratyphi A and C cause typhoid fever. This study was to ascertain the population of blaSHV, blaOXA-1, blaCTX-M-4, blaCTX-M-9, and blaTEM ESBL regulatory genes from Salmonella species in patients with fever in Dalhatu Araf Specialist Hospital (DASH), Lafia, Nigeria.
Materials and Methods
Sample collection
Four hundred samples (fecal) were obtained from patients with symptoms of fever in DASH, Nasarawa State for a whole year starting from March 2020. The samples were inoculated on selenite F broth for 24 hours at 37°C. The growing colonies were inoculated on Salmonella Shigella Agar (SSA), and the procedure was repeated. Observed colonies were purified on Nutrient agar after 24 hours incubation at 37°C. Isolates were identified using morphology, gram reaction, and biochemical tests. 11 Ethical clearance for the study was from DASH Ethics and Research Committee, Lafia.
Molecular identification and genetic diversity of resistant Salmonella isolates
Identification to confirm the isolates were Salmonella was by using genes ttrF: 5′-CTCACCAGGAGATTACAACATGG-3′ and ttrR: 5′-AGCTCAGACCAAAAGTGACCATC-3′, respectively, while genes staGF: 5′-CGCGAAGTCAGAGTCGACATAG-3′ and staGR: 5′-AAGACCTCAACGCCGATCAC-3′ specific to S typhi. The bacterial cells were inoculated in Luria Bertani (LB) broth overnight and centrifuged (at 14 000 r/min for 3 min). The cells were suspended in normal saline (500 µL), heated at 95°C for 20 minutes, and cooled on ice, and the procedure was repeated. The obtained DNA was amplified using primers for ttr (detection of all Salmonella) and staG (detection of S typhi) based on the methods of Nair et al. 12
Extended-spectrum β-lactamase gene amplification
DNA samples were subjected to multiplex polymerase chain reaction (PCR) using primers given in Table 1 to detect the ESBL genes (blaTEM, blaSHV, blaCTX-M, blaCTX-M-1, blaCTX-M-9, blaCTX-M-4, and blaOXA-1). For the amplification, a Taq DNA polymerase (Inqaba Biotec, South Africa), 200 M of each of the 4 deoxynucleoside triphosphates (Inqaba Biotec, South Africa), and 0.6 M of each primer in 1 × PCR buffer were used in a 50 μL mixture in an ABI 9700 Applied Biosystems GeneAmp 9700 PCR thermal cycler. In total, 48 µL of the master mix was added to the template DNA (2 μL), which was then coated with mineral oil. The amplification process consists of a preliminary denaturation at 94°C for 2 minutes, followed by 35 cycles of primer annealing at 62°C for 1 minute, and extension at 72°C for 1 minute. A 10-minute extension step was performed at 72°C, and the products were stored at 4°C until use.
ESBL target genes with amplicon sizes for ESBL used.

Abbreviation: ESBL, extended-spectrum β-lactamase.
Sequencing of the staG and ttr genes
Sequencing to determine the genetic relatedness of resistant isolates was carried out on isolates that were positive with staG gene specific to S typhi and isolates S11 and S39 with multidrug-resistant index (MDRI) of 1 (Supplementary file 1). The PCR was carried out by a method earlier described by Antonio. 15 The sequencing reaction was prepared by mixing 5 μL of ddH20, 5.5 μL of bacterial DNA template, 2 μL each of staG and ttr primers, and 8 μL of Dideoxy terminator cycle sequencing (DTCS) quick start master mix (Beckman Coulter) in a 2.0 mL tube and reaction run for 30 cycles. The sequence reaction was transferred into a 0.5 mL sterile tube, containing a stop solution/glycogen mixture (5 μL) and mixed thoroughly. The mixture was then combined with 60 mL (60 L) of cold 95% ethanol and centrifuged for 15 minutes at 14 000 r/min. With the aid of a micropipette, the pellets were washed in 70% ethanol before being centrifuged at 14 000 r/min for 2 minutes. A micropipette was used to extract the supernatant, which was then vacuum-dried. A drop of mineral oil was applied after the sample was once more suspended in 40 L of loading solution and placed in the appropriate well of the sample plate. The instrument was run once the sample plate had been loaded. The Basic Local Alignment Search Tool (BLAST) of the National Center for Biotechnology Information (NCBI) was used to examine the sequencing findings. The neighbor-joining approach was used to infer the evolutionary history. The p-distance approach described by Nei and Kumar 16 was used to calculate the evolutionary distances in the number of base changes per site. Molecular Evolutionary Genetics Analysis (MEGA X) was used to undertake evolutionary analyses. 17
Results
Of the 400 samples collected, only 60 with a prevalence of 15% were positive for the presence of Salmonella species. staG genes specific to S typhi were found in 2 isolates S9 and S25 (prevalence 2.33%), while 14 isolates (S32, S42, S43, S44, S45, S46, S47, S48, S51, S52, S54, S58, S59, and S60) do not have the ttr gene in their genetic makeup (Supplemental material 2).
The distribution of the ESBL regulatory genes among the 60 isolates showed that blaSHV had the highest prevalence of 12 (20.0%). Genes BlaCTX-M-9 and blaOXA-1 were positive in 5 isolates each with a prevalence of 8.30%, respectively, while blaCTX-M-4 and blaTEM isolated in 1 isolate each with a prevalence of 1.7%, respectively.
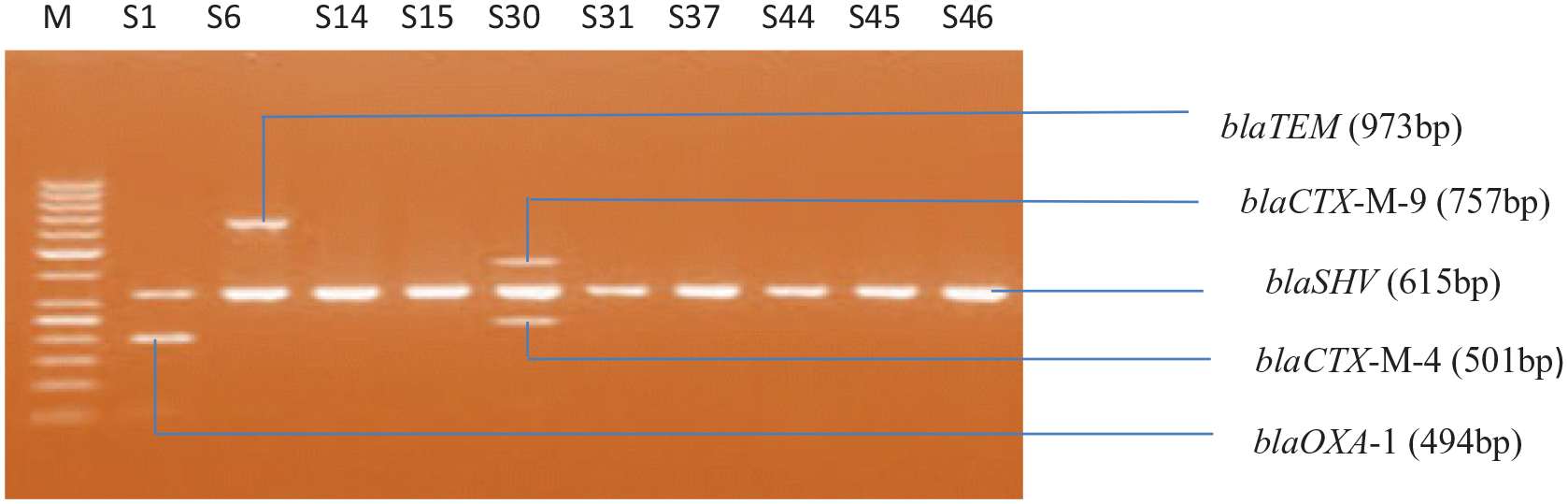

Salmonella species isolates’ amplified ESBL genes. Lanes S1, S6, S14, S15, S30, S31, S37, S44, S45, and S46 represent blaSHV (615 bp); Lane S1 represents blaOXA-1 (494 bp); Lane S30 represents blaCTX-M-4 (501 bp) and blaCTX-M-9 (757 bp); and Lane S6 represents blaTEM (973 bp). Lane M represents DNA ladder. Isolate S1 harbors 2 ESBL gene blaSHV and blaOXA-1, isolate S6 has both blaTEM and blaSHV genes while isolate S30 has 3 of the ESBL genes, blaCTX-M-9, blaSHV, and blaCTX-M-4.
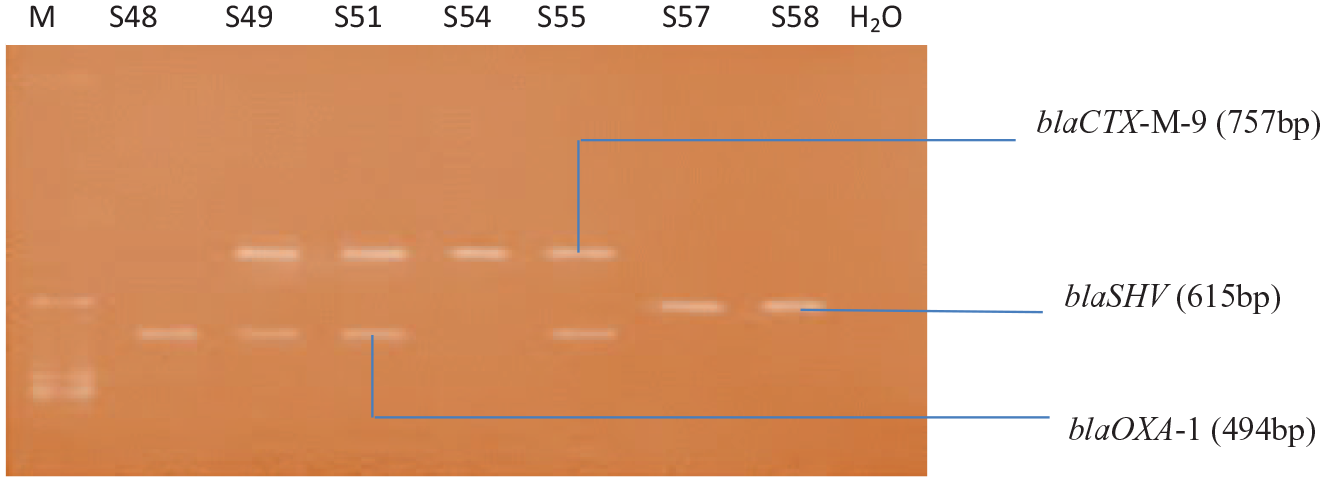
Agarose gel electrophoresis of the amplified ESBL genes from Salmonella species isolates. Lanes S48, S49, S51, and S55 represent blaOXA-1 (494 bp), Lanes S49, S51, S54, and S55 represent blaCTX-M-9 (757 bp) and Lanes S57 and S58 represent blaSHV (615 bp) while H2O (DNA-free water) represents a negative control and Lane M represents DNA ladder. Isolates S49, S51, and S55 harbor 2 ESBL genes blaOXA-1 and blaCTX-M-9, respectively.
Inferred ancestral sequences
Based on the predicted likelihood at site 1, the tree displays a set of potential nucleotides (states) at each ancestral node (Figure 1). The MEGA X software’s Maximum Composite Likelihood model was used for evolutionary analysis that resulted in 2 major clusters with isolates S25 (ST313L2) and S9 (ST313L1) as outliers (Figure 1). Isolates S11 (ST36) and S39 (ST138) with MDRI = 1 were clustered in a group with a similarity coefficient of 0.00634 (Table 2). The sum of 1915 positions was obtained in the final data set (Table 1).

Dendrogram of the relationship between Salmonella with MDRI = 1 (S11 and S39) and those with staG genes specific to S typhi (S9 and S25).
Quantification of evolutionary divergence.

The maximum composite likelihood model was used for the analyses.
Discussion
blaCTX-M, blaTEM, blaSHV, and blaOXA genes harbored by self-transmissible conjugative plasmids, and horizontally shared by the same and sometimes different species of bacteria, mediate ESBL enzyme production. Similar patterns of antibiotic susceptibility are observed when a particular type of resistant gene is responsible for the resistance experienced. The incidence of ESBL producers in Salmonella isolates seen in this research was 28.3% which is close to the 26.0%, 22.2%, and 26.3% reported by Mohamed et al, 18 Ogbolu et al, 19 and Fody et al 20 , respectively, and higher than 16.5% reported by Ahmed et al. 21 The population of blaSHV gene was the highest as opposed to the report that CTX-M-15 is the most frequently encountered ESBL regulatory gene responsible for human infections.22,23 The prevalence of the 5 genes examined in this study was higher than the report presented by Hijazi et al. 24 This research demonstrates that ESBL-containing isolates are more resistant to drugs.
Extended-spectrum β-lactamase producers in Salmonella are reported to be resistant to ceftazidime and cefotaxime. The study confirmed that not all the Salmonella isolates resistant to cefotaxime and ceftazidime were ESBL producers, which agrees with the report by Tsaku et al 25 in Keffi, Nigeria, and Nkene et al. 26 These isolates would have deployed other mechanisms of achieving resistance to the antibiotics tested. CTX-M, (plasmid-mediated cefotaximases), SHV (sulf-hydryl variable), and TEM (Temoniera) are class A ESBLs horizontally transferable enzymes. They are inactivated by clavulanic acid and can conveniently spread across epidemiological niches. They are spread also when consumed in contaminated water, meat, pork, and fish.27,28 blaTEM is a plasmid-mediated β-lactamase and has been described as the most prevalent β-lactamase in gram-negative bacteria which disagreed with findings in this study. The lactamase enzyme can evolve to another analog with a single amino acid substitution of Gln39Lys and conferring resistance to bacteria against penicillin and cephalosporins antibiotics.
CTX-M enzymes are most frequently linked to resistance among ESBLs. Genetic relatedness between CTX-M-4 and CTX-M-9 sublineages is >94% for amino acid similarity and 90% for the similarity between groups, and biological differences (diversification—a difference in amino acid positions). 29 This explains the differences recorded in the prevalence of the 2 enzymes in the study. Rahman et al 29 posited that the increase in the number of SHV and CTX-M-9 sublineages is a result of certain clones and epidemic plasmids spreading throughout the population and medical facilities. The CTX-M-type β-lactamase enzymes are known to be both plasmid-mediated and epidemic, and they also have a zoonotic nature. The research conducted by Falgenhauer et al 30 indicates that the geographical characteristics of a certain area facilitate the transmission of these enzymes from humans to animals.
The dendrogram depicted the genetic relationships between the 4 Salmonella species (Isolates S9, S11, S25, and S39). The Salmonella isolates were in 2 main cluster groups. Analysis of genetic distances among the isolates on the basis of source (patients) shows that isolates S11 (ST36), S39 (ST138), and S9 (ST313L1) are closely related genetically, while the strain S25 (ST313L2) from patients at the same time have a high level of genetic distance among samples isolated from different sources. The highest genetic similarity was observed between the isolates S25 (ST313L2) and S9 (ST313L1). The observed genotypic diversity is due to random mutations and environmental variation and stress. Hedrick 31 explained that genetic diversity ensures the survival and continuity of species.
The gravity of the challenge posed by resistant bacteria to health care is substantial due to the presence of both chromosomal and plasmid-mediated beta-lactamases within the bacteria. These beta-lactamases, which are transmissible in the community, contribute to the narrowing range of available treatment options for ESBL-producing bacteria. To prevent the transmission and dissemination of resistant isolates in health care settings, routine screening for ESBL is crucial. The increasing prevalence of TEM and CTX-M types of ESBLs can be attributed to the proliferation of clones or clonal groups and epidemic plasmids in both public and hospital environments. In addition, co-selection with other resistant factors, particularly fluoroquinolones, aminoglycosides, and sulfonamides, has further exacerbated the situation. 29
Conclusions
The study concluded that isolates show more similarity and less distance. The bacterial species are resident in the area, which calls for concern. The study recommends further research involving a cross-section of the adjoining communities to determine the carriage level of the resistant genes.
Supplemental Material
sj-docx-1-bbi-10.1177_11779322231220194 – Supplemental material for Phylogenetic Characterization of Resistant Salmonella Strains in Typhoid Fever Patients in Nigeria
Supplemental material, sj-docx-1-bbi-10.1177_11779322231220194 for Phylogenetic Characterization of Resistant Salmonella Strains in Typhoid Fever Patients in Nigeria by Olukayode Olugbenga Orole, Jebes Ngolo Lamini and Aleruchi Chuku in Bioinformatics and Biology Insights
Supplemental Material
sj-docx-2-bbi-10.1177_11779322231220194 – Supplemental material for Phylogenetic Characterization of Resistant Salmonella Strains in Typhoid Fever Patients in Nigeria
Supplemental material, sj-docx-2-bbi-10.1177_11779322231220194 for Phylogenetic Characterization of Resistant Salmonella Strains in Typhoid Fever Patients in Nigeria by Olukayode Olugbenga Orole, Jebes Ngolo Lamini and Aleruchi Chuku in Bioinformatics and Biology Insights
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
LJN, who composed the original draft and performed the statistical analysis, designed the study. OOO managed the literature searches and wrote the protocol for the study. AC and OOO managed analysis in the study, while the final article was approved for submission by all authors.
Ethical Approval
DASH Lafia, Nasarawa State’s Ethics, and Research Committee gave ethical approval for the study
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
