Abstract
Tumor microenvironment is characterized by the occurrence of significant changes due to disrupted signaling pathways that affect a broad spectrum of cellular activities such as proliferation, differentiation, signaling, invasiveness, migration, and apoptosis. Similarly, a downregulated suppressor of cytokine signaling 3 (SOCS3) promotes increased JAK/STAT function due to aberrant cytokine signaling, which results in increased cell proliferation, differentiation, and migration. Multiple carcinomas including breast cancer, prostate cancer, hepatocellular carcinoma, pancreatic cancer, and colorectal cancer involve the disruption of SOCS3 expression due to microRNA overexpression. MicroRNAs are small, conserved, and non-coding RNA molecules that regulate gene expression through post-transcriptional inhibition and mRNA destabilization. The aim of this study was to identify putative microRNAs that interact with SOCS3 and downregulate its expression. In this study, miRWalk, TargetScan, and miRDB were used to identify microRNAs that interact with SOCS3, whereas RNA22 was utilized to identify the binding sites of 238 significant microRNAs. The tertiary structures of shortlisted microRNAs and SOCS3 regions were predicted through MC Sym and RNAComposer, respectively. For molecular docking, HDOCK was used, which predicted 80 microRNA-messengerRNA complexes and the interactions of the top 5 shortlisted complexes were assessed. The complexes were shortlisted on the basis of least binding affinity score and maximum confidence score. This study identifies the interactions of known (miR-203a-5p) and novel (miR-6756-5p, miR-6732-5p, miR-1203, miR-6887-5p) microRNAs with SOCS3 regions due to their maximum interactions. Identifying the interactions of these microRNAs with SOCS3 will significantly advance the understanding of oncomiRs (miRNAs that are associated with cancer development) in tumor development due to their influence on SOCS3 expression. These insights will assist in future studies to understand the significance of miRNA-SOCS3-associated tumor development and progression.
Introduction
The uncontrolled growth and division of abnormal cells promotes cancer development as a consequence of cellular changes. 1 Cancerous cells form heterogeneous tumors that cause dysfunction of the immune system and prevention of apoptosis. 2 Breast cancer (BC) is characterized by the unusual growth and division of cells in the breast, becoming an abnormal mass of tissue called tumor. 3 It remains a global health challenge as it is the most commonly diagnosed cancer with an estimated annual incidence rate of 2.26 million cases worldwide according to GLOBOCAN 2020 global cancer statistics. 4 It is a heterogeneous disease due to presence or absence of receptors that can attach to the hormones, estrogen or progesterone and causes the cells to grow abnormally. 5 On the contrary, colorectal cancer (CC), colon cancer or rectal cancer, is a type of cancer that affects the large intestine or rectum. It develops in the polyps, termed as the clumps of cells present on the inner lining of the bowel. 6 According to GLOBOCAN 2020, the estimated annual occurrence of CC globally was 19.3 million with the highest incidence rate of 52.3% in Asia. 4
Suppressor of cytokine signaling 3 (SOCS3) is an inhibitor of JAK/STAT signaling pathway. The pathway is involved in the regulation of multiple key functions such as homeostasis, immune cell development, hematopoiesis, and stem cell maintenance. 7 SOCS3 regulates STAT3 function, which is induced by cytokines, by inhibiting the kinase activity of JAKs. 8 (p3) Downregulation of SOCS3 stimulates unregulated JAK/STAT signaling, which leads to increased cell proliferation, differentiation, and migration, a hallmark of cancer. 9 Decreased SOCS3 expression elicits aberrant inflammatory signaling pathways including JAK2/STAT3, which leads to BC development and progression. 10 The overexpression of STAT3 due to interleukin 6 (IL-6), an inflammatory cytokine, promotes hypermethylation of SOCS3, which results in growth and metastasis of CC. 11
MicroRNAs (miRNAs) are small, approximately 22 nucleotides (nt) in length, non-coding RNAs that regulate the gene expression post-transcriptionally by binding to a specific site in the target gene. 12 The miRNAs may play as tumor suppressor or oncomiRs (miRNAs that are associated with cancer development) in different conditions and their dysregulation leads to the suppression of certain genes that may lead to cancer development. 13 Cellular differentiation, proliferation, migration, and apoptosis are among the key functions that are regulated by miRNAs, ultimately leading toward disease. 14 Recent research has reported the downregulation of an miRNA, miR-502-5p, which in turn upregulates its target gene, DNMT3a. As a DNA methyltransferase, DNMT3a transfers methyl group to SOCS3 and causes its downregulation, which accelerates JAK2/STAT3 signaling pathway in hepatocellular carcinoma (HCC). 15 Several studies have reported the suppression of SOCS3 due to miRNA binding in various cancers such as BC, CC, ovarian cancer, prostate cancer, HCC, and lung cancer. The oncomiRs that may cause SOCS3 inhibition in these cancers include miR-196b-5p, miR-222-3p, miR-4449, miR-221, miR-155, and miR-1203. 16 - 18 In addition, miR-203 is highly associated with BC promotion due to increased expression of pStat3 and c-Myc. 19 Moreover, the stimulation of metastasis as a result of differentiation of monocytes to M2-TAMs due to miR-203 overexpression is observed in CC. 20 MYC, a proto-oncogene activates miR-9 that suppresses a metastasis suppressing protein, E-cadherin, which in turn activates β-catenin and increases expression of vascular endothelial growth factor that leads to tumor angiogenesis. 21 Suppression of glycogen synthase kinase-3 beta (GSK3β) and secreted frizzled-related protein 2 (SFRP2) by miR-224 in CC leads to increased cell proliferation, differentiation and cancer-related inflammation due to activation of Wnt/β-catenin signaling. 22
The study of interaction of miRNAs with their target genes provide insights into binding sites (3-UTR, 5-UTR, and CDS), free energy and conformation with which these miRNAs bind to a certain region of the gene and cause its regulation and silencing. 23 This research focuses on the interaction of putative miRNAs with SOCS3, a tumor suppressor gene, in order to identify their preferred binding orientation, interaction region, affinity and strength of these oncomiRs and their role in BC and CC development, progression, and metastasis. The identification of miRNAs that show strong binding affinity with SOCS3, can help in targeted therapies due to their eminent role in tumor suppressor gene silencing in cancer.
Methodology
Overview
The interactions of miRNAs with SOCS3 were analyzed to identify the potential binding regions and the affinity of miRNA-mRNA complexes. The overall strategy followed in this research study is demonstrated in Figure 1.

Workflow representing binding site prediction analysis of miRNA-SOCS3.
Prediction of miRNA binding sites on SOCS3 gene
To identify the binding sites of miRNAs on SOCS3, miRWalk (http://mirwalk.umm.uni-heidelberg.de/), TargetScan (v8.0) (https://www.targetscan.org/vert_80/) and miRDB (v6.0) (https://mirdb.org/) were employed to predict the miRNA-mRNA interactions. MiRWalk predicts interactions using machine learning techniques and includes miRNA-target interactions that are experimentally verified. 24 TargetScan predicts miRNA targets by searching the conserved sites of miRNAs, while miRDB elucidates targets by analyzing miRNA-target interactions that were obtained from high-throughput sequencing experiments.25,26 The predicted interactions were filtered using the filter function of dplyr (v1.0.10) package in R (v4.2.2) to separate the miRNAs with more stable and strong interaction scores with SOCS3. The filtration criteria of binding probability 1, energy ⩽ –30 (indicates high binding strength) and AU content < 0.6 (AU-rich elements of the transcript that are present 30 nucleotides upstream and downstream of predicted site) was applied. 27 The common miRNA-SOCS3 interactions, produced by all databases, were considered for the miRNA-mRNA alignment.
Alignment between shortlisted miRNAs and SOCS3
To align the shortlisted miRNAs against SOCS3 mRNA, RNA22 (v2.0) (https://cm.jefferson.edu/rna22/Interactive/) was employed. RNA22 implements a pattern-based approach combined with estimation of the folding energy to predict the putative miRNA target sites. It first analyzes the sequences of known mature miRNAs and then based on pattern information from the miRNAs, predicts the putative target sites, and then identifies the miRNAs that are likely to bind to the predicted target sites. 28 Sensitivity and specificity values were kept at 63% and 61%, respectively and seed size of 7 was selected with one unpaired base allowed in the seed region without limiting the maximum number of G:U wobbles in the seed region. Minimum number of paired-up bases in heteroduplex was kept at 10 while the maximum folding energy was kept at −12 kcal/mol. 29 The nucleotide sequence of SOCS3 was extracted from NCBI (accession: NG_016851). The sequences of shortlisted miRNAs were retrieved from miRBase database (v22; https://www.mirbase.org/). 30 The sequences of shortlisted miRNAs along with SOCS3 gene sequence, were given as input to the server. In order to retrieve the significant alignments from RNA22 results, threshold of P-value < .05 was applied using the filter function of dplyr as described earlier.
Three-dimensional structure prediction of SOCS3 mRNA
The binding sites and folding energy of the predicted miRNA-SOCS3 were plotted on a scatter plot using the scatter method of seaborn (v0.11.2) library in Python (v3.9.12). The nucleotide sequence regions showing maximum interaction with miRNAs were extracted from mRNA sequence and provided to RNAComposer (v1.0) for model prediction (https://rnacomposer.cs.put.poznan.pl/). 31 It is a manually curated database of RNA three-dimensional (3D) structures and prediction of tertiary structure. It takes a maximum of 500 nucleotides as input. Hence, the regions greater than 500 nucleotides were split into smaller regions. Moreover, CONTRAfold was selected as the secondary structure prediction method.
3D structure prediction of shortlisted miRNAs
The miRNAs showing maximum interaction against the shortlisted regions were separated and their sequences were given as input to MC Fold (https://www.major.iric.ca/MC-Fold/) for secondary structure prediction. MC Fold predicts multiple secondary structures, which is based on the decomposition of RNA structure into loops. 32 The MC Fold output (secondary structures) having minimum energy was submitted to MC Sym (https://www.major.iric.ca/MC-Sym/) for the prediction of stable tertiary structures of the miRNAs. MC Sym examines the conformational search space of RNA and generates structures that satisfy the input. 33
Molecular docking between shortlisted miRNAs and SOCS3
HDOCK (v2021; http://hdock.phys.hust.edu.cn/), a template-based modeling and ab initio docking web server was used to perform docking between shortlisted miRNAs and SOCS3 regions to identify the complexes with optimized conformation and lowest binding affinity. 34 The tertiary structures of shortlisted miRNAs and SOCS3 regions with default parameters were provided as input. For each docked miRNA-mRNA model, 10 complexes were generated and only 1 complex with least binding affinity score and high confidence score of > 0.7 was selected. Out of all the predicted complexes, five were shortlisted based on lowest binding affinity and high confidence, which indicates strong interactions. PyMol (v2.5.4; https://pymol.org/2/), was used to identify the interactions between the miRNA and mRNA nucleotides of the shortlisted complexes. PyMol is a molecular graphics tool, used for 3D visualization of proteins, nucleic acids, trajectories, electron densities, surfaces, and small molecules. 35
Results
Prediction of interaction sites between miRNAs and SOCS3 gene
Through miRWalk, a total of 2101 miRNAs were observed to be interacting with SOCS3. Furthermore, filtration of miRNAs that have strong and stable interaction with SOCS3 resulted in 47 interactions. Moreover, 21 interactions were found to be common after validation of filtered interactions from TargetScan and miRDB. Out of 21 interactions, 3 were found for miR-6732-5p and 2 for miR-6805-5p because of different binding positions. Hence, a total of 18 unique miRNAs were shortlisted for alignment against SOCS3 (see Table 1). Despite its unavailability in other databases, miR-203 was retained because it was experimentally determined to downregulate SOCS3 in various studies (see Table 2).19,36- 38
Prediction of miRNAs that interact with SOCS3 through miRWalk, TargetScan, and miRDB.
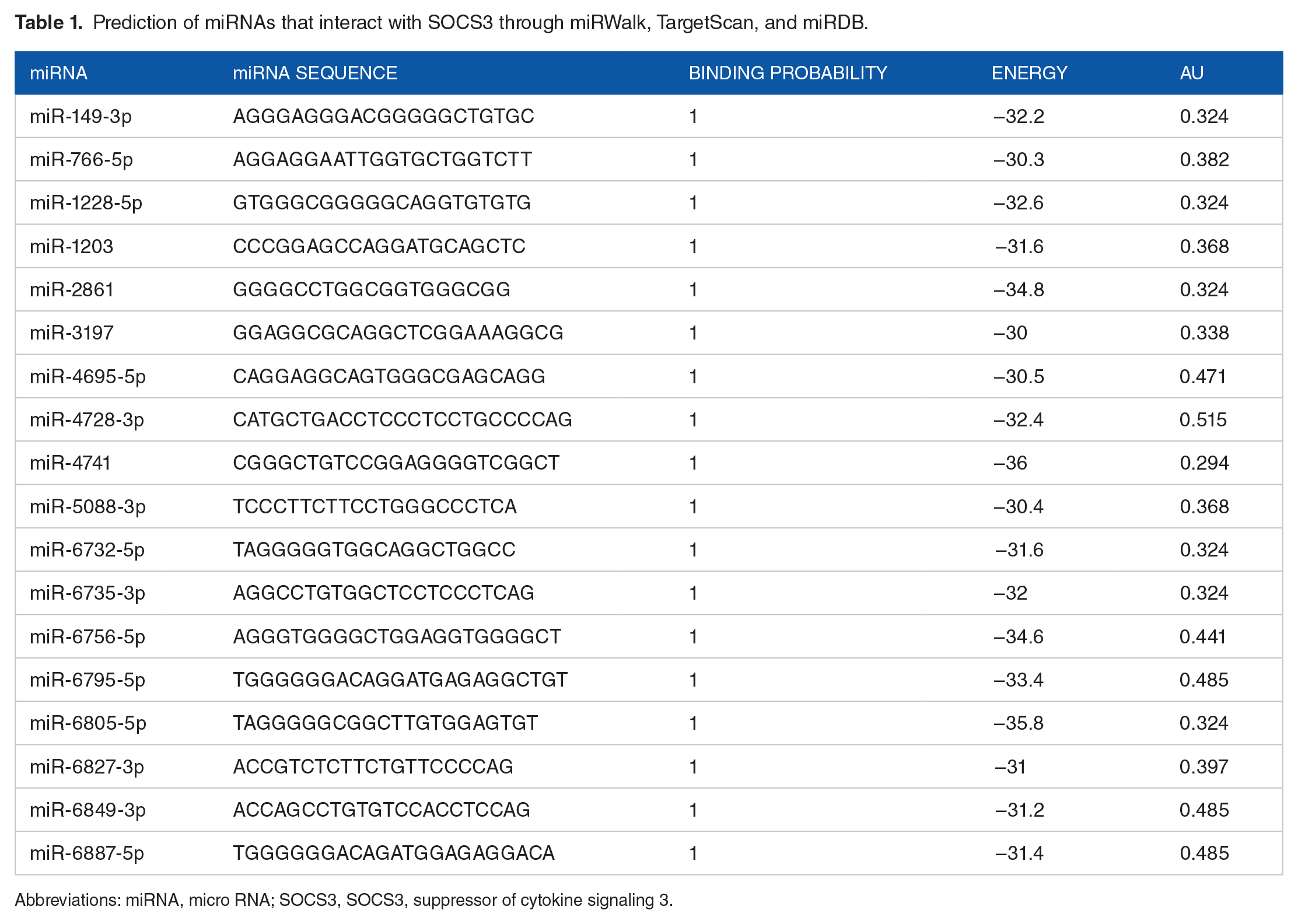
Abbreviations: miRNA, micro RNA; SOCS3, SOCS3, suppressor of cytokine signaling 3.
Prediction of miR-203a interaction with SOCS3 through TargetScan.

Abbreviations: miRNA, micro RNA; SOCS3, SOCS3, suppressor of cytokine signaling 3.
Significant alignments between miRNAs and SOCS3
RNA22 predicted 503 alignments between shortlisted miRNAs and SOCS3. In order to extract the alignments that were significant, a threshold of P-value < .05 was applied, which resulted in 238 significant alignments. The alignment output denoted the miRNA binding sites on the target gene and the binding energy represented the strength of the resulting miRNA-mRNA complex.
Shortlisting of SOCS3 regions
A scatterplot was generated in order to identify the regions of SOCS3 at which maximum miRNAs showed interactions with minimum energy (see Figure 2). Two regions, from 676nt to 959nt and 7217nt to 9072nt, that represented maximum and stable miRNA interactions were shortlisted. Due to the large size of region 2, it was split into four parts (2:1, 2:2, 2:3, 2:4) as shown in Table 3. A total of 29 shortlisted miRNAs showed maximum interactions with selected SOCS3 regions (see Table 4). Out of 29 miRNAs, 12 showed interactions with region 1, while 7 showed interactions with region 2:1, 9 with region 2:2 and 1 with region 2:3.

Visualization of predicted binding regions of miRNAs on SOCS3.
SOCS3 shortlisted regions.

Abbreviation: SOCS3, SOCS3, suppressor of cytokine signaling 3.
Interactions of shortlisted miRNAs and SOCS3 regions predicted through RNA22.

Abbreviations: miRNA, micro RNA; SOCS3, SOCS3, suppressor of cytokine signaling 3.
Molecular docking analysis between miRNAs and SOCS3
To study the interaction of shortlisted 20 miRNAs with shortlisted SOCS3 regions, HDOCK was employed for molecular docking. A total of 10 complexes for each miRNA against each shortlisted region were generated. Based on the binding affinity score, the top complex was selected for each miRNA-mRNA interaction that resulted in a total of 80 docked complexes. Out of 80 complexes, the top 5 were shortlisted with maximum confidence score and minimum binding affinity (see Table 5). The strength of association between miRNA and mRNA depends on binding affinity and confidence score, which represents that a complex is stable, in a preferred orientation and is more likely to bind. The miR-6756-5p-mRNA (Region 2:3) complex was shown to have least binding affinity (–701.78) and highest confidence score (1) while the complex miR-6887-5p-mRNA (Region 2:2) have maximum binding affinity and least confidence score (0.9999).
Shortlisted docked complexes through HDOCK.

Abbreviation: mRNA, messenger RNA.
Visualization of shortlisted complexes
The shortlisted complexes were visualized through PyMol to identify the nucleotides involved in interaction between miRNA-mRNA complexes. As shown in Figure 3, miR-203a-5p interacts within the region 2:2. The miRNA interacts with mRNA at 21 different sites having binding affinity score of –656.07. In addition, Figure 3 represents interaction of miR-6887-5p with mRNA in the region 2:2. The complex has the least number of interactions, that is, 13 which indicates less binding affinity of –625.77. Similarly, miR-6756-5p interaction within the region 2:3 is as indicated in Figure 4. The miR-6756-5p has 15 different interactions with the mRNA. The miR-1203-mRNA complex has 15 different interactions within the region 2:3 (see Figure 4). Furthermore, the miR-6732-5p-mRNA has the second highest number of interactions that is, 18, within the region 1, which represents a stable complex with high bonding probability (see Figure 5).

Interaction of miR-203a-5p and miR-6887-5p with mRNA (Region 2:2): (A) Surface representation of the interaction of miR-6887-5p-mRNA complex. (C) Surface representation of the interaction of miR-203a-5p-mRNA complex. (B) and (D) represent the interactions of miRNA-mRNA where

Interaction of miR-6756-5p with mRNA (Region 2:3). (A) Surface representation of the interaction of miR-6756-5p-mRNA complex. (C) Surface representation of the interaction of miR-1203-mRNA complex. (B) and (D) represent the interactions of miRNA-mRNA where
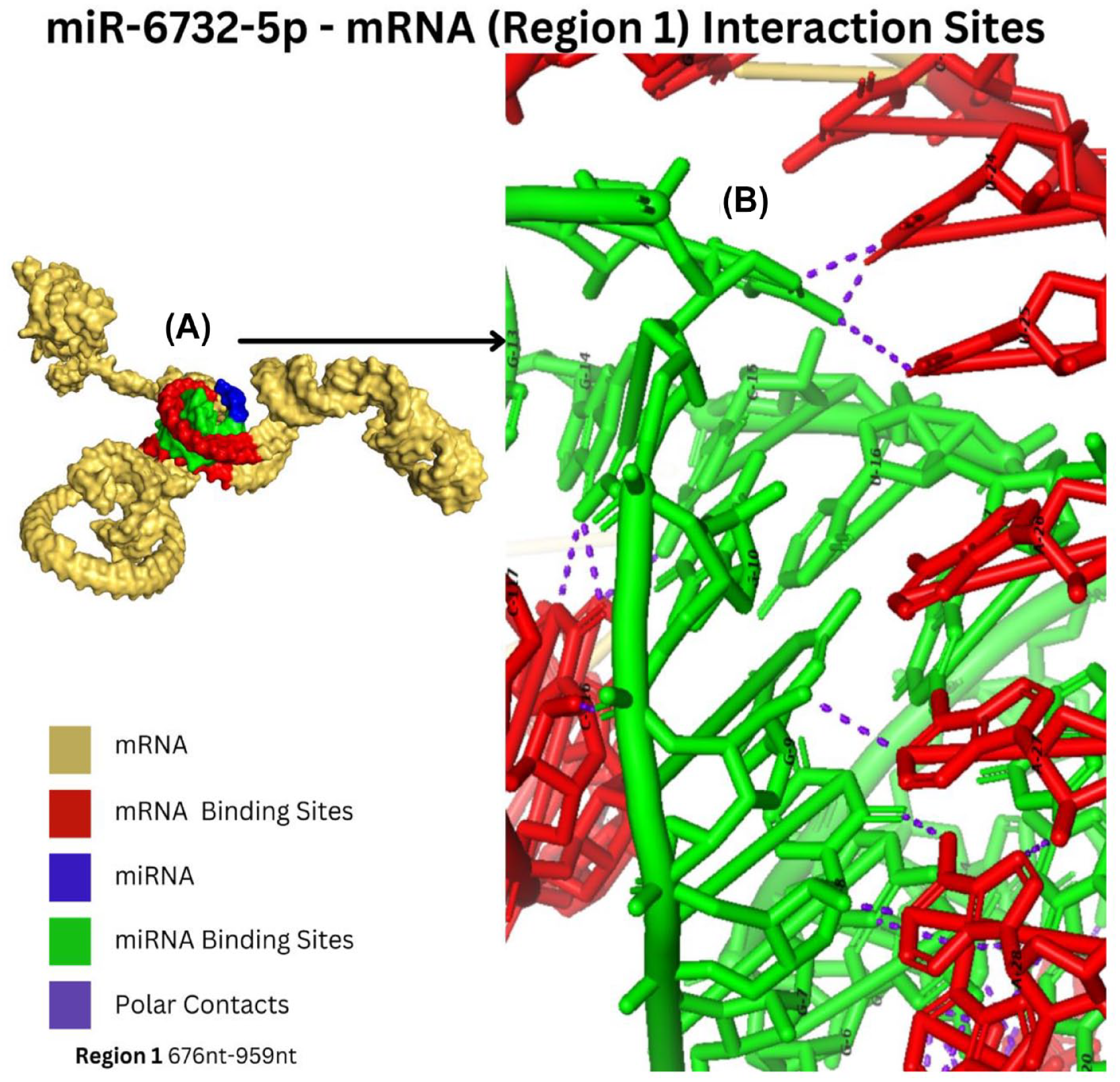
Interaction of miR-6732-5p with mRNA (Region 1). (A) Surface representation of the interaction of miR-6732-5p-mRNA complex. (B) The yellow-orange color represents the region (receptor), while the red color indicates the binding site of the receptor. Similarly, the miRNA (ligand) is denoted by blue color and green indicates its binding site. The dotted lines (violet color) represent the polar interactions between the receptor and the ligand.
Discussion
Cancer is developed as a consequence of growth and division of abnormal cells that have the ability to infiltrate surrounding normal tissues. 39 The uncontrolled growth and division of abnormal cells in the breast tissues is termed as BC. 40 It is classified into three major subtypes, that is, triple negative BC, hormone receptor positive BC, and human epidermal growth factor receptor 2 (HER2) positive BC, based on the presence of estrogen or progesterone receptors and HER2. 41 CC is the conversion of non-cancerous clumps of cells called polyps into cancerous cells that are present on the inner lining of the colon or rectum. 42 Based on the site where the tumor grows, CC is majorly classified into colorectal adenocarcinoma and gastrointestinal carcinoid tumors. 43 It has been reported by multiple studies that individuals with primary BC are more likely to develop CC.44,45 Several studies suggest that mutations in genes involved in BC (BRCA1 and BRCA2) are also susceptible to cause CC.46,47 Moreover, ROBO1 and CHEK2 mutations are also observed in both BC and CC. 48 SOCS3, a tumor suppressor gene, is associated with BC and CC development and progression due to its downregulation that results in dysregulation of JAK/STAT pathway. 49 The SOCS family consists of eight proteins, which are responsible for regulating cytokine signaling. 50 Cytokines are small proteins that regulate innate and adaptive immunity by controlling the activity of immune cells and enable them to signal over long distances. 51 SOCS3 regulates cytokine signaling by directly binding to the cytokine receptor and JAK domain, which promotes STAT inhibition. Furthermore, SOCS3 might also negatively regulate JAK/STAT signaling by proteasome degradation or ubiquitination of cytokine receptors, thus controlling the immune homeostasis in pathological and physiological conditions. 52 MicroRNAs (miR-19, miR-221, miR-155) inhibit SOCS3 function by binding to its 3′-untranslated region (3′-UTR), thus promoting tumor development by increased cell proliferation, differentiation, invasion, and metastasis. 16
In order to identify the miRNAs that interact with SOCS3 to inhibit its function, miRWalk, TargetScan, and miRDB were used. These interactions may help in identifying miRNAs that bind to SOCS3 regions and repress the mechanism of protein synthesis. 53 Out of multiple interactions, only 20 unique miRNA-mRNA interactions having binding probability of 1, energy ⩽ –30 and AU content < 0.6 were shortlisted. RNA22 was employed for the identification of binding sites of the shortlisted miRNAs. A similar research employed the standard RNA22 input parameters, which predicted 242 miRNAs suggesting possible binding sites against the influenza C virus genome. 29 In this study, RNA22 predicted the binding sites of 503 miRNAs against SOCS3. The results were filtered based on the P-value criteria of < .05 which yielded 238 significant alignments. The significantly aligned miRNAs were identified to be associated with cancerous characteristics including miR-2861, miR-149-3p, miR-1228-5p, and miR-4741. It has been reported that upregulated miR-2861 is responsible for lymph node metastasis in papillary thyroid carcinoma. 54 Upregulated miR-149-3p causes disabled-2 interacting protein (DAB2IP) downregulation, which promotes cell aggressiveness and viability in various cancers such as prostate cancer, lung cancer, bladder cancer, BC, and endometrial cancer. 55 A study reported the upregulation of miR-1228-5p in non-small cell lung cancer (NSCLC) individuals. It downregulates a tumor suppressor gene, LRP1, which leads to tumor progression and invasion through negative regulation of transcription. 56 Moreover, miR-4741 upregulation was observed in gastric cancer individuals, which promotes increased cell proliferation, invasion, and metastasis through suppression of e-cadherin, a glycoprotein involved in cell-cell adhesion functions.57,58
For the prediction of tertiary structures, mRNA regions and miRNAs were shortlisted on the basis of miRNA-mRNA maximum interaction. This binding depends on multiple factors, that is, abundance of target mRNAs and miRNAs, subcellular locations of miRNAs, and the binding affinity of miRNA-mRNA interactions. 23 Two mRNA regions (676nt-959nt and 7217nt-9072nt) showed maximum interactions with 29 unique miRNAs. HDOCK was used to perform the molecular docking to identify the preferred orientation, affinity and interaction of the miRNA on the mRNA binding site. Molecular docking of all the predicted structures resulted in 80 complexes and only 5 were shortlisted based on least binding affinity score and high confidence score that represents stable and strong interaction. Out of five shortlisted miRNAs, four (miR-6756-5p, miR-203a-5p, miR-1203, and miR-6887-5p) showed interaction at 3′-UTR region of SOCS3, while only miR-6732-5p showed interaction with 5′-UTR region. The miRNA typically binds to the 3′-UTR of mRNAs and causes its destabilization through translational inhibition and degradation. 59 The 3′-UTR region of mRNA plays a crucial role in gene expression regulation via orchestrating interaction between mRNA cis and trans regulatory elements. Alterations in any of these elements can lead to several pathologies including cancer development and progression. 60 It has been reported that overexpression of miR-6756-5p in prostate cancer suppresses IL-24, which leads to eukaryotic initiation factor 2 alpha (eIF2α) phosphorylation through endoplasmic reticulum (ER) stress pathway, which promotes ternary complex (the nascent protein, the messenger RNA, and the ribosome) depletion and inhibition of translational initiation. 61 Overexpression of miR-6732-5p leads to FOXA1 downregulation in BC and CC, which induces cancer stem cell properties through increased expression of IL-6.62,63 Increased IL-6 expression promotes differentiation and proliferation of multiple non-immune cells. 64 Several researchers reported the overexpression of miR-203a-5p in multiple cancers. Upregulation of miR-203 family suppresses SOCS3, which in turn promotes cell proliferation, differentiation, and transformation in BC MCF-7 cells. 65 Moreover, a metallo-hydrolase, PDE4D, which degrades cAMP, is suppressed by miR-203a in CC. This downregulation disrupts normal cellular functions and leads to increased cell growth, differentiation, gene transcription, and protein expression. 66 In a similar study, miR-1203 suppresses SOCS3 in HCC by binding to its 3′-UTR region, which promotes over-phosphorylation of STAT3, increased HCC cell proliferation and invasion, hepatic fibrosis, and tumor aggressiveness. 67 Furthermore, miR-6887-5p overexpression causes suppression of signal transduction in transforming growth factor beta (TGF-β), mediated by Smad3. This leads to dysregulation of diverse cellular processes such as cell proliferation, differentiation, migration and apoptosis which promotes the development of multiple cancers such as CC, BC, ovarian cancer, and pancreatic cancer. 68
In this study, it was identified that miR-203a-5p shows binding affinity of -656.07 and confidence of 1 with SOCS3 and interacts at 21 different sites. Although miR-6756-5p had the least binding affinity score (–701.78) and highest confidence score (1), it showed less number of interactions (15). Large number of strong interactions of the miRNAs on the 3′-UTR region of SOCS3 indicates a hotspot region on the gene structure where miRNAs may bind to downregulate the gene expression.69,70 These strong binding complexes between miR-203a-5p, miR-1203, and miR-6756-5p suggest the 3′-UTR region to be highly targeted by the different miRNAs.
This study shows highly significant regions of the SOCS3 mRNA that could be targeted by oncomiRs (miR-203a-5p, miR-6756-5p, miR-203a-5p, miR-1203, miR-6732-5p, and miR-6887-5p). These miRNAs-mRNA binding may lead to the dysregulation of the gene through increased expression of these miRNAs and ultimately promoting increased JAK/STAT signaling, cancer development and progression through dysregulated cell proliferation, differentiation, and migration due to increased cytokine signaling at the tumor microenvironment. To the best of our knowledge, this is the first study to identify significant novel miRNAs (miR-6732-5p, miR-6756-5p, miR-1203, miR-6887-5p) that may have the potential to cause BC and CC through downregulation of SOCS3 gene expression. These miRNAs have been reported previously to cause various different cancers such as, HCC, prostate cancer, pancreatic cancer, and ovarian cancer.61,67,68,71 Hence, it is suggested that anti-miR drugs should be developed that can target these oncomiRs and downregulate their expression, which may lead to normal expression of the SOCS3. 72
Conclusions
The uncontrolled growth and division of abnormal cells in the breast and colon or rectum leads to BC and CC development and progression, respectively. It has been observed that mutations in the same genes (BRCA1, BRCA2, ROBO1, and CHEK2) could lead to incidence of multiple primary neoplasms. Dysregulation of SOCS3 in BC and CC promotes disruption in JAK/STAT pathway due to downregulation of cytokine signaling. In this study, 238 significant miRNAs were found to be interacting with SOCS3 at the 3′-UTR region. Out of the significantly aligned 238 miRNAs, based on molecular docking, top 5 miRNAs were shortlisted due to least binding affinity at different SOCS3 regions. It was found that miR-6756-5p, miR-203a-5p, miR-1203, and miR-6887-5p interacted with SOCS3 at 3′-UTR region and showed 15, 21, 15, and 13 interactions, respectively, whereas only miR-6732-5p interacted at 5′-UTR with 18 interactions. This study identified oncomiRs that could target hotspot SOCS3 mRNA regions. The binding of miRNA-mRNA may lead to increased JAK/STAT function through abnormal cytokine signaling due to dysregulation of the SOCS3 gene. In this research, significant binding regions and interactions of 4 novel miRNAs (miR-6732-5p, miR-6756-5p, miR-1203, miR-6887-5p) and one known (miR-203a-5p) have been identified that can suppress SOCS3 expression to promote BC and CC. These miRNAs have been reported previously to cause multiple carcinomas including HCC, prostate cancer, pancreatic cancer, and ovarian cancer. Thus, targeting miRNAs that sustain cancer can present a new paradigm for miRNA-based cancer therapy in several carcinomas, particularly BC and CC.
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
