Abstract
Non-typhoidal Salmonella (NTS) infections occur globally with high morbidity and mortality. The public health challenge caused is exacerbated by increasing rate of antibiotic resistance and absence of NTS vaccine. In this study, we characterized the outer membrane protein C (OmpC) serovars isolated from different food animals and predicted antigenicity. ompC of 27 NTS serovars were amplified by polymerase chain reaction (PCR) and sequenced. Sequence data were analysed and B-cell epitope prediction was done by BepiPred tool. T-cell epitope prediction was done by determining peptide-binding affinities of major histocompatibility complex (MHC) classes I and II using NetMHC pan 2.8 and NetMHC-II pan 3.2, respectively. ompC sequence analysis revealed conserved region among ompCs of Salmonella Serovars. A total of 66.7% of ompCs were stable with instability index value < 40 and molecular weight that ranged from 27 745.47 to 32 714.32 kDa. All ompCs were thermostable and hydrophilic with the exception of S. Pomona (14p) isolate that had ompC with GRAVY value of 0.028 making it hydrophobic. Linear B-cell epitope prediction revealed ability of ompC to elicit humoral immunity. Multiple B-cell epitopes that were exposed and buried were observed on several positions on the ompC sequences. T-cell epitope prediction revealed epitopes with strong binding affinity to MHC–I and -II. Strong binding to human leukocyte antigen (HLA-A) ligands, including HLA-A03:1, HLA-A24:02 and HLA-A26:01 in the case of MHC-I were observed. While binding affinity to H-2 IAs, H-2 IAq and H-2 IAu (H-2 mouse molecules) were strongest in the case of MHC-II. ompCs of NTS serovars isolated from different food animal sources indicated ability to elicit humoral and cell-mediated immunity. Hence, ompCs of NTS serovars are potential candidate for production of NTS vaccines.
Introduction
Salmonella enterica remains a pathogen of concern globally, since it continually causes infections that results in high morbidity and mortality. 1 Depending on clinical outcomes, serotypes of Salmonella are broadly grouped into typhoidal Salmonella (TS) and non-typhoidal Salmonella (NTS). 2 While TS serovars are human redistricted, NTS serovars cause infection in birds, mammals and humans. 3 Although NTS serovars cause self-limiting diarrhoea in individuals, infection in infants and immunocompromised persons could be severe or fatal. 4 It is estimated that NTS serovars cause 93 million infections with 155 000 associated deaths per year globally. 5 In 2022 alone, several outbreaks of NTS infections have been reported in Europe (S. Enteritidis and S. Typhimurium) and the United States (S. Enteritidis, S. Typhimurium, S. Hadar, S. Infantis, S. Indiana, 1 4,[5], 12:1 and S. Mbandaka) resulting in hospitalization and deaths.6-9 Treatment of bacterial infections, including NTS infections are becoming difficult as a result of antimicrobial resistance (AMR). The World Health Organization (WHO) has categorized fluoroquinolone (FQ)-resistant Salmonella spp. as priority pathogen urgently in need of new antibiotics. 10 This underscores the need for NTS vaccines. There is currently no NTS vaccine in use, however, several vaccine candidates, including live-attenuated, glycoconjugate, outer membrane vesicle vaccines and many more are in various stages of development and clinical trials. 11 Salmonella live-attenuated vaccines (SLAVs) have the potential to elicit both humoral and cell-mediated responses, however, their safety in immunocompromised individuals at high risk of coming down with invasive non-typhoidal Salmonella (iNTS) has been brought to question. 5 Hence, the need to explore other cell components of NTS serovars for vaccine production is imperative. Returning to normal life during this COVID-19 pandemic could in part be attributed to the prompt production of vaccines especially the mRNA vaccine that initiates the production of viral spike proteins that in turn elicit immune response to severe acute respiratory syndrome coronavirus 2 (SARS-Cov-2) virus. 12 The outer membrane proteins (omps) of Salmonella, such as ompC, ompD and ompF, are known antigens that are highly immunogenic and elicit cellular immunity and efficient protective antibody. 13 OmpCs of Salmonella have been indicated as a potential broad spectrum Salmonella vaccine candidate as several highly conserved amino acid sequences located in the transmembrane β-sheet have been identified in both typhoidal and non-typhoidal serovars. 13 Carreño et al 14 in a study demonstrated that OmpC of S. Typhi elicited specific B-cell responses and polyfunctional CD4+ T-cell responses that evolved over a period of 2 months post-administration of live-attenuated vaccine. Similarly, OmpC of S. Typhimurium has been shown to elicit cell-mediated and humoral immune responses that conferred absolute protection against S. Typhimurium in rats. 15 Although omps of Salmonella are promising vaccine candidates, available data tend to centre majorly on S. Typhimurium that is just one among other numerous NTS serovars that have been implicated in outbreaks and invasive infections. Questions on spectrum of activity (broad spectrum) of NTS omps base vaccines have been raised. For instance, ompD has been shown to mediate immune response that limits clinical outcome of invasive S. Typhimurium infection. 11 However, Ashton et al 16 reported the absence of ompD locus in S. Typhimurium (ST313) strain U60 isolated in the United Kingdom. The need to study the omps of other NTS, including rear serovars for vaccine prospect is therefore imperative. This study aimed to molecularly characterize OmpC of NTS serovars isolated from different sources and predict their antigenicity.
Materials and Methods
Salmonella isolates
Thirty previously isolated Salmonella Serovars from animal sources (chicken, sheep and cattle) stored in the stock culture of Molecular Epidemiology Unit of the Molecular Biology and Biotechnology Department of Nigerian Institute of Medical Research (NIMR) Yaba Lagos were retrieved. To resuscitate the isolates, a loop full of the isolates in glycerol stock were inoculated into 1 mL of brain heart infusion broth (Oxoid, Basingstoke, UK) and incubated for 18 hour at 37°C after which they were streaked on nutrient agar plate and incubated for 24 hours at 37°C. Isolates were then stored in agar slants for further use.
DNA Extraction
Isolates were inoculated into 1 mL of brain heart infusion broth (Oxoid, Basingstoke, UK) and incubated for 18 hours at 37°C. Cells were harvested by centrifuging at 14 rpm for 3 minutes and supernatant was discarded retaining the cell pellet. Genomic DNA was extracted with bacterial genomic DNA isolation Kit (NorgenBiotek Corp., Canada) according to the manufacturer’s instructions. Two hundred and fifty microliter of re-suspension solution A was added to cell pellet and vortexed to make a cell suspension to which 12 µL of lysozyme (400 mg/mL) was added and mixed. Then, 250 µL of lysis buffer P and 12 µL of proteinase K were added to the cell suspension vortexed and incubated at 37°C for 2 hours. Five hundred microliter was added to the lysate and mixed well with gentle vortexing after which 500 µL of 100% ethanol was added and mixed again by gentle vortexing. Spin columns were assembled into which lysate mixtures were dispensed and centrifuged at 14 000 rpm for 1 minute. After centrifugation, flow-through was discarded and spin column was reassembled. Five hundred microliter of wash solution A was added to spin column and centrifuged for 1 minute at 14 000 rpm. Flow-through was discarded and the washing process was repeated once more. After the second wash and discarding of the flow-through, the column was centrifuged for 2 minutes to ensure the column was thoroughly dried. The spin column was reassembled with a new 1.7 mL collection tube and 200 µL of elution buffer B was added to the centre of the column which was then centrifuged for 2 minute at 14 000 rpm and pure genomic DNA obtained was stored at −20°C for further use.
Detection and Sequencing of ompC
Salmonella ompC gene was amplified by PCR using primer set F-ATGAAAGTTAAAGTACTGTCCCTC and R-TTAGAACTGGTAAACCAGACCC. 17 A 30 µL PCR mix was used. The reaction mix contained 16.2 µL nuclease-free water, 0.9 µL forward primer, 0.9 µL reverse primer, 6 µL DNA template and 6 µL of 5X PCR Master Mix (Solis BioDyne, Estonia). PCR programming parameter consisted of initial DNA denaturation at 95°C for 5 minutes, followed by 30 cycles of denaturation at 95°C for 30 seconds, annealing at 58°C for 40 seconds and extension at 72°C for 2 minutes, followed by a final extension at 72°C for 10 minutes. Electrophoresis was done to visualize amplicons. Gel electrophoresis was ran at 100 V for 60 minutes and visualized under a trans-illuminator (Cleaver Scientific Ltd). A 100 bp DNA ladder (Solis BioDyne, Estonia) was used as a molecular weight marker. Fifteen microliter of PCR products, and 10 µL of forward and reverse primers were aliquoted into a micro-centrifuge tube and sent to Inqaba Biotec, South Africa, for Sanger sequencing. Sequences obtained in this study have been deposited in the National Centre for Biotechnology Information (NCBI) database with accession numbers ON190078–ON190104.
Comparative Analysis of ompC Gene Sequences and Prediction of Antigenicity
Protein sequence analysis and phylogenetic tree construction
Nucleotide sequences of ompC were base called on Bioedit software 18 and exported as FASTA to note pad. The Basic Local Alignment Search Tool (BLAST) analysis was done on NCBI database (https://blast.ncbi.nlm.nih.gov/Blast.cgi) to determine sequence homology with reference sequences. Nucleotide sequences were translated to amino acid sequences using the sorted six-frame translation tool of Bioedit software and BLAST analysis was done on UniProt database (https://www.uniprot.org/blast) to confirm protein identity. Multiple sequence alignment was done using CLUSTALW and phylogenetic tree was constructed using the maximum likelihood joining tree in MEGA 11.0 software 19 and variation were analysed. Protein 3D structure of ompC was determined from the PDBsum pictorial database in the protein data bank (http://www.ebi.ac.uk/thornton-srv/databases/cgi-bin/pdbsum).
Prediction of antigenicity
Properties of proteins were determined using ProtParam tool on the ExPASy platform and properties, such as number of amino acids, molecular weight, instability index, aliphatic index and theoretical isoelectric point (pI) was recorded. B-cell epitope prediction to determine B-cell epitope regions of ompC protein was done with BepiPred prediction tool on the cbs.dtu.dk server (http://www.cbs.dtu.dk/services/BepiPred/). 20 Scores of epitopes were recorded and peptides with 10 or more amino acids in length were considered as good B-cell epitopes. T-cell epitope prediction was done by determining peptide-binding affinities of MHC classes I and II by calculating epitope database analysis and ranking them according to their percentile using NetMHCpan 2.8 (https://services.healthtech.dtu.dk/service.php?NetMHCpan-2.8) and NetMHCIIpan 3.2 (https://services.healthtech.dtu.dk/service.php?NetMHCIIpan-3.2) servers for MHC-I and -II, respectively. 21 For MHC-I, affinity threshold for strong binding peptides was set at 5000 and affinity threshold for weak binding peptides was set at 500 000. While rank threshold for strong binding peptides was set at 0.500 and for weak binding peptides 2.00. On the other hand, rank threshold for strong and weak binding peptides for MHC-II was set at 2 and 10, respectively.
Results
Protein sequence analysis and properties of protein
ompC was detected in all (100%) the Salmonella isolates as the ompC gene of 1103 bp gene was amplified as shown in Figure 1. However, ompC of three of the Salmonella isolates failed sequencing quality control (QC) and could not be sequenced. ompC sequence analysis from multiple sequence alignment showed conserved regions among ompCs of Salmonella Serovars as shown in Figure 2. However, phylogenetic analysis of protein sequence revealed that three major clades (

Gel image of amplified ompC gene in Salmonella isolates.

Multiple sequence alignment of ompC protein sequence showing conserved glycine (G) and proline (P) among Salmonella Serovars.

Phylogenetic analysis of ompC protein sequences of Salmonella Serovars. Three main clades
Table 1 shows the physicochemical properties of ompCs of various Salmonella Serovars. Total number of negatively charged residues (Asp + Glu) in some of the ompCs of some of the serovars were low with concomitant high instability index resulting in unstable proteins. However, 66.7% of the proteins were stable with instability index values < 40. Molecular weight of ompCs ranged from 27 745.47 to 32 714.32 kDa. While number of amino acids ranged from 259 to 298. pI of ompCs of Salmonella Serovars ranged from 7.12 to 11.94. Hydrophobicity or hydrophilicity measured as grand average of hydropathicity (GRAVY) values of ompCs indicated they were all hydrophilic with the exception of the ompC of isolate 14p (S. Pomona) that had a GRAVY value of 0.028 making it hydrophobic. Aliphatic index (thermostability of protein) indicated that ompCs of the Salmonella isolates were thermostable.
Various properties of ompCs of Salmonella Serovars, including number of amino acids, molecular weight, theoretical pI, instability index, aliphatic index and hydropathicity.

Abbreviations: AA, amino acid; MW, molecular weight; pI, isoelectric point.
Prediction of B-cell epitopes
Linear B-cell epitope prediction was high in several positions of ompC sequence of Salmonella Serovars as depicted in Figure 4. Some of the predicted epitopes were buried. However, majority of the others were exposed indicating relative surface accessibility as predicted by NetSurfP’s default threshold. 8P_S. Anatum had the highest number of predicted B-cell epitopes (218) of which 151 (69.27%) were exposed while 67 (30.73%) were buried; 18P gera had 134 predicted B-cell epitopes making it the least among epitopes possessed by other serovars as shown in Table 2. The number of predicted B-cell epitopes and their orientation (exposed or buried) among the same serovars varied. 12P_S. Enteritidis possessed 212 epitopes of which 142 (66.98%) were exposed while 21P_S. Enteritidis had 188 epitopes with 141 (75.00%) exposed. Although, 11P_S. Ealing possessed 204 predicted B-cell epitopes, only 81 (39.71) were exposed making the serovar with the least exposed epitopes.

Result output of MHC 2.0 prediction tool showing positions of predicted epitopes that are buried or exposed form NetsurP’s default threshold, structure of proteins and protein sequences illustrating BepiPred-2.0 prediction.
Number of predicted B-cell epitopes and surface orientation in Salmonella Serovars.

Prediction of T-Cell Epitopes
MHC-I
Affinity and ranks of representative epitopes with the strongest binding affinity is shown in Table 3. Mapped set of 9mer peptides sequences of each isolate ranged from 246 to 290. Isolate 23P (S. Gera) had 19 the highest number of peptides that were strong binders followed by isolate 6P (S. Muenster) and 20P (S. Gera) that had 17 strong binding peptides each. Weak binding peptides were highest in isolate 3P with 51 weak binders followed by isolate 16P (S. Blockley). Not all peptides 9mers had binding affinity. More affinity to HLA-B ligands were recorded with rank ranging from 0.001 to 0.5 (Table 3). Although, strong binding affinity to HLA-A ligands, HLA-A03:1, HLA-A24:02 and HLA-A26:01 had more binders recorded compared to HLA-A01:01 and HLA-A02:01. No binding to any of the HLA alleles was recorded for isolate 21P (S. Enteritidis), however, 12P also an S. Enteritidis isolate had strong binders to only HLA-A03:1 and HLA-A24:02 of the HLA ligands.
Affinity and ranks of strongest binding representatives of peptides to various alleles of HLA-A and HLA-B ligands, including total numbers of 9mers peptides of various Salmonella isolates.
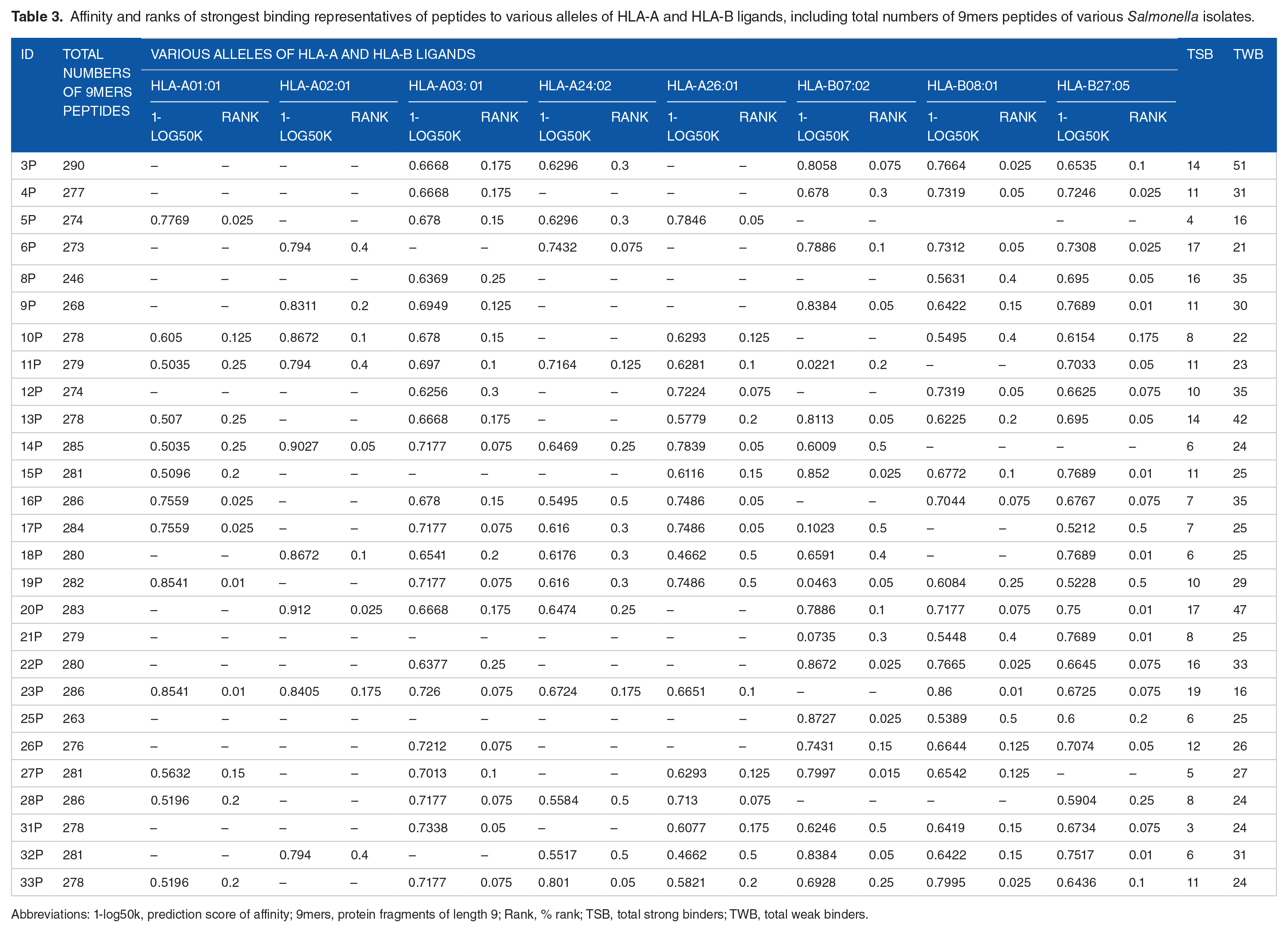
Abbreviations: 1-log50k, prediction score of affinity; 9mers, protein fragments of length 9; Rank, % rank; TSB, total strong binders; TWB, total weak binders.
MHC-II
Predicted MHC–peptide-binding affinity of the various H-2 mouse molecules revealed varied numbers of strong and weak binders. Molecules H-2 IAs had a total of 371 strong binders among Salmonella servers followed by molecule H-2 IAq with 348 strong binders and H-2 IAu with a total of 284 strong binders among the serovars (Figure 5). A total of 246 and 283 strong binders were detected in molecule H-2 IEd and H-2 IEk, respectively, among the serovars (Figure 6). The least number of binders among the serovars was found in the H-2 IAk molecule with 166 strong binders as shown in Figure 7.

Predicted MHC-II–peptide-binding affinity of H2-IAs, H2-IAq and H2-IAu showing numbers of strong and weak binders.

Predicted MHC-II–peptide-binding affinity of H2-IEd and H2-IEk showing numbers of strong and weak binders.

Predicted MHC-II–peptide-binding affinity of H2-IAb, H2-IAd and H2-IAk showing numbers of strong and weak binders.
Discussion
The genus Salmonella consist of six subspecies with over 2600 phenotypically, ecologically and genetically diverse serovars. 22 All NTS serovars in this study isolated from different food animal sources were positive for OmpC. This implies that OmpC is largely distributed among Salmonella Serovars. Jawad and Al-Charrakh 23 similarly reported the PCR amplification of OmpC in all Salmonella strains isolated from human and animal samples in Iraq. Serovars in this study showed genetic interrelatedness with similar serovars clustering with unrelated serovars inferred from phylogenetic analysis of OmpC sequences. Although variation existed in protein sequences of serovars there was a high level of sequence homology with identified conserved regions. Valero-Pacheco et al 13 in a previous study have reported highly conserved amino acid sequences of OmpC among both TS and NTS serovars. The need for prospecting for NTS vaccine is important considering the increasing number of iNTS infections coupled with the high burden of AMR. 24 Vaccine potential of the OmpC of NTS as earlier highlighted tends to be a resource that needs to be explored and exploited to solve the problem. Beyond being conserved among Salmonella Serovars, 25 OmpC of NTS possess physicochemical parameters that make them ideal antigens. Functional and physicochemical properties of vaccine antigens are core determinants of the level of antigen-adjuvant interaction and vaccine response. 26 In this study, a large percentage of the OmpCs of serovars, including S. Enteritidis were stable with instability index of < 40. They had aliphatic index that indicated they were thermostable and GRAVY values that showed they were hydrophilic. Rabbani et al 27 in a previous study have demonstrated the conformation and thermostability of omp of S. Typhi to different pH conditions. Similar physicochemical properties have been reported for ompC and ompF of S. Typhimurium by Jha et al 28 and Soman et al, 29 respectively.
Functional properties of OmpC as an ideal vaccine antigen have been demonstrated in previous studies. However, most studies have only evaluated and reported on the OmpCs of S. Typhimurium and S. Typhi.13-15,28 Concerns have been raised on the limitation of the use of certain omps, such as ompD, as broad spectrum vaccine against NTS infections. 11 ompCs of Salmonella Serovars investigated in this study all possessed linear B-cell epitopes on several positions with majority exposed capable of evoking immune response. This is in line with the findings of Jha et al 28 who reported B-cell epitopes with antigenic index of 1.7 from the mapping of the OmpC of S. Typhimurium. Studies have highlighted the importance of the accessibility of epitopes to MHC-I and MHC-II to elicit optimal immune response. 30 Immunogenic potential of OmpC for MHC-I predicted in this study showed strong binders with more affinity to HLA-B ligand. On the other hand immunogenic potential of OmpC for MHC-II showed strong binding with high preference to H-2 IAs a variant of H-2 mouse molecule. Valero-Pacheco et al 13 in their study reported conserved OmpC amino acid sequences with immunogenic potential for MHC-II binding.
Conclusion
In conclusion, findings from our study revealed that OmpC of various Salmonella Serovars possessed B-cell and T-cell epitopes that have the potential of evoking immune response. Hence, OmpC of these serovars can be considered for design of broad spectrum vaccine against NTS.
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
MY, AA and EUU contributed to data collection; MY, AA and EUU contributed to data analysis and preparation of draft; AA, AIA and SIS contributed to conceptualization of study; and OO, AIA and SIS supervision and correction of draft. All authors read and approved final draft.
