Abstract
The rapid development of high-performance technologies has greatly promoted studies of molecular oncology producing large amounts of data. Even if these data are publicly available, they need to be processed and studied to extract information useful to better understand mechanisms of pathogenesis of complex diseases, such as tumors. In this article, we illustrated a procedure for mining biologically meaningful biomarkers from microarray datasets of different tumor histotypes. The proposed methodology allows to automatically identify a subset of potentially informative genes from microarray data matrices, which differs either in the number of rows (genes) and of columns (patients). The methodology integrates nonnegative matrix factorization method, a functional enrichment analysis web tool with a properly designed gene extraction procedure to allow the analysis of omics input data with different row size. The proposed methodology has been used to mine microarray of solid tumors of different embryonic origin to verify the presence of common genes characterizing the heterogeneity of cancer-associated fibroblasts. These automatically extracted biomarkers could be used to suggest appropriate therapies to inactivate the state of active fibroblasts, thus avoiding their action on tumor progression.
Introduction
Complex human diseases are caused by the interactions of many genetic, environmental, and behavioral factors. Development of high-performance technologies greatly promoted studies on molecular oncology producing large amounts of omic data. The availability of massive volume of experimental data based on cancer researches requires the development of mathematical, statistical, and computational techniques, which automatically extract valuable information useful for a better understanding of pathogenesis mechanisms of complex diseases, such as tumors. Identifying marker genes potentially involved into the development of tumors may facilitate the understanding of cause of disease, thus contributing to the advancement of diagnostic tools and/or to the evaluation of more efficient clinical strategies.
In most of the publicly available data sets, the number of genes is significantly larger than the number of samples. This high dimensionality represents a major problem for an automatic gene array–based cancer analysis. In this scenario, dimensionality reduction methods became indispensable as they can eliminate irrelevant and redundant information, thus reducing the dimensionality as well as the complexity of the original problem, with significant benefits in terms of computational efficiency, model interpretability, and data understanding.
Among linear dimensionality reduction methods, low rank matrix decomposition algorithms have been successfully exploited as alternative approaches for studying various types of high-dimensional biological data, including gene expression data. 1 The application of these algorithms relies on the assumption that large-scale biological data have an intrinsic low-dimensional representation, with the dimension often corresponding to the number of latent information embedded into the original data. These methods transform the space of original data into a lower dimensional more discriminating (informative) space that makes the subsequent analysis more efficient.
Among low-rank reduction mechanisms, nonnegative matrix factorizations (NMFs) emerge as useful approaches for the analysis of microarray data. The intrinsic non-negativity property of these techniques, in fact, produces more intuitive results as many biological measurements are represented by positive values. Nonnegative matrix factorizations demonstrated their ability in a number of tasks, including the identification of sets of genes co-operating in a relatively tightly regulated manner; 2 the discovery of potential relationships in large biological data samples and link genes to these patterns; 3 the detection of distinct genomic subtypes in cancer patients; 4 the inspection of expression data sets including time evolution of the gene expression profile in different samples; 5 and the extraction of gene expression profiles from fibroblasts of cancer blood diseases. 6
In this article, we present a gene extraction methodology, which integrates a NMF method7,8 with the functional enrichment analysis web tool WebGestalt 9 and a gene extraction procedure designed ad hoc to automatically mine different microarray cancer data sets to extract a reduced subset of genes to be further investigated from a biological point of view. An approach for mining multi-omics cell line data with the same row size by joint nonnegative matrix factorization (JNMF) and pathway signature analyses was recently proposed in Fujita et al. 10 The methodology presented in this article instead allows to analyze microarray data matrices, which differ either in the number of rows (genes) and in the number of columns (patients) to verify the presence of common genes characterizing the heterogeneity of different cancer datasets. This proposal enriches the panorama of large-scale data-driven computational methods based on matrix factorization algorithms, which are able to extract concise and useful pieces of information from existing disease-associated data sets.1,11,12 Particularly, through the NMF method, our methodology mines the metagenes, which are the most representative of the information embedded into different tumor datasets and then, by simple intersection set operations allow to extract genes in a natural way to obtain interpretable and useful knowledge readily usable from biologists and analyzed, thanks to functional and visualization approaches based on the WebGestalt tool. The proposed methodology has been used to mine microarray of solid tumors of different embryonic origin to verify the presence of biomarkers characterizing the heterogeneity of cancer-associated fibroblasts (CAF) thus being applicable in clinical practice.
Fibroblasts constitute the most heterogeneous and abundant population of mesenchymal cells in tumor microenvironment (TME). Their presence goes from tumor formation up to the final stage of metastatic diffusion, but their precise functional role in tumor is not fully understood yet. 13 Also, it is not clear how different subtypes of CAF could exert distinct paracrine actions affecting specific tumor oncogenesis.14,15 Therefore, we adopt the proposed procedure to analyze gene expression profiles of CAFs belonging to primary cultures of 3 distinct tumor histotypes (ie, colon of endodermal origin, breast carcinoma of ectodermal origin and ovary of mesodermal origin) to select common biomarkers and characterize activated fibroblast phenotype. Moreover, as it is known that bone marrow (BM) is a CAF recruitment source, 16 we aim at investigating also the existence of common genes among CAF gene expression profiles of selected solid tumors and those of BM of patients with multiple myeloma (MM) and monoclonal gammopathy of undetermined significance (MGUS; MGUS is a benign pathological condition characterized by the proliferation of a plasma cell clone that rarely evolves into a malignant neoplasm. 17 The latter, being a benign pathological condition, simulates a well-differentiated CAF phenotype compared to those of patients with MM.). Using the proposed methodology, we highlight the existence of biomarkers among CAFs of different embryonic origin that uniquely identify them and could be useful for developing or evaluating more efficient clinical strategies.
This article is organized as follows. First, the pipeline of the whole gene extraction methodology is presented, illustrating a general way to pre-process this kind of data, its analysis core (based on the NMF method), and the ad hoc gene landscape extraction procedure. The subsequent section describes the specific biological problem we investigated, as well as the databases of CAF populations used in the experimental session. Then, the functional and pathway analysis performed during the experiments are deeply discussed and the obtained results are highlighted from a biological point of view. Some conclusive remarks conclude the article.
Methods
This section discusses in some detail the main architecture, data pre-processing, and basic NMF approach used in this study.
The ensemble gene extraction methodology
The proposed methodology is based on different operations that integrate the acquisition of data, their mathematical analysis, and the biological exploration of the obtained results.
Figure 1 illustrates the work-flow and the main operations performed when selected tumor data sets are provided.

Work flow of the NMF-based methodology for extraction of common genes between different tumor gene expression datasets.
The methodology is mainly based on the integration of 4 parts: the data preparation module, the core module computing NMF of given data matrices, the Gene Landscape extractor devoted to the extraction of common genes between processed results, and the Functional and Visual Analysis module. The first 3 modules combine R/BioConductor platform and the NMF library, which are well established in bioinformatics research,18-20 whereas the biological analysis of the latter module is performed using the well-known web tool WebGestalt.
Data preparation and pre-processing
Working on microarray data requires data sets are imported in the virtual environment as a single different expression set, in which probeset represents gene interrogating a

Possible cases supported by the pre-processing phase.
The pre-processing operations implemented in this module are based on the adjusted median absolute deviation (MAD) of the gene expression values. The use of the MAD (a scale factor is used to consider the MAD as a consistent estimator for the estimation of data standard deviation), which computes the variability of the data from the median, makes the process more robust to outliers. In particular, a tuple probeset id, gene symbol, and the associated expression value are uniquely identified assuming that greater is the MAD higher is the goodness of the hybridization in the experiment. The code and dataset related to data preparation and pre-processing module have been made available at GitHub at https://github.com/flaespo/NMF-for-GSE.
Nonnegative matrix factorization for genetic data sets
After data pre-processing is performed, most omic data sets can be represented as a matrix in which each element contains the measurement of a single molecule in a single experimental condition. In the case of expression data, the resulting high-dimensional data set is re-formulated as a numerical nonnegative matrix with rows being genes and columns representing samples, for example, tissues of various patients (as in this study), development stages, or treatments. This matrix is called “gene expression matrix” and its elements
Nonnegative matrix factorization can be applied to reduce data dimensionality as it decomposes a gene expression matrix

for
From an algebraic point of view, NMF finds 2 nonnegative matrices, the
When NMFs are applied to produce clusters of genes, the rank value
Computationally, metagenes and expression profile values can be obtained solving a non-linear constrained optimization problem over the cone of nonnegative matrices

where

is used, which corresponds to the maximum likelihood estimation under an independent Poisson assumption.
24
The minimization problem (1) for the KL divergence function is solved applying the following multiplicative update rules for

Because of the non-negativity constraint, solutions to NMF are only unique up to scaling and rotation, but appropriately scaling and rotating the columns of
To do this, the
Metagenes in
Gene Landscape Extraction procedure
Referring to Figure 1, let
To compare gene expression matrices derived from different tumor histotypes (in this work from their associated CAF cultures), the most representative metagene
To identify common genes among those extracted by different tumor histotypes, an intersection set operation was performed on the identification labels of genes
Figure 3 provides a logical view of the Gene Landscape Extraction procedure, which represents the novel proposal to extract common genes from microarray matrices which differ in their sizes. Particularly, the microarray matrices
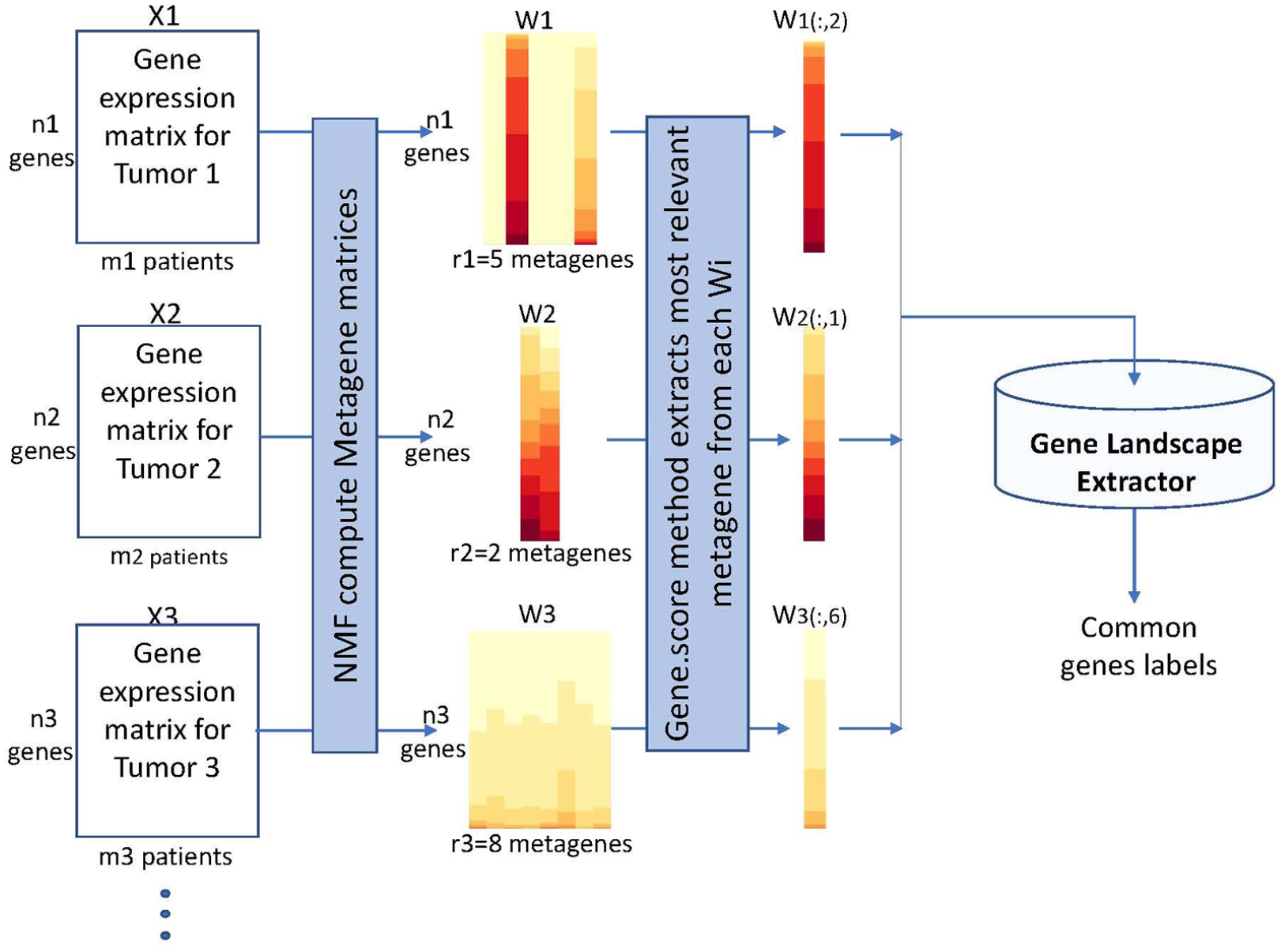
Logical view of the gene landscape extraction. Preprocessed expression data matrices related to 3 different tumors (
Functional and visual analysis
The obtained subset of genes is then analyzed by integrative bioinformatic tools, that is, the WebGestalt tool, 26 to discover the role they cover into the biological process under investigation.
This module performs either functional and pathway enrichment analysis of the genes in
Application to the Analysis of Gene Expression Profiles of CAFs
In this section, we sequentially applied the main modules of the proposed methodology for automatically extract genes from gene expression profiles of CAFs.
Fibroblasts are the most heterogeneous and abundant population of mesenchymal cells in the TME, 27 but their precise functional role in tumor is not fully understood yet. 13 During initial phases of oncogenesis, fibroblasts are activated giving rise to fibroblasts associated with the tumor (CAF), which play a key role in generating a specific extracellular matrix (ECM) in TME. 28 The kinetics of changes in CAF actions might be different in the various types of tumor, partly because of organ-specific transcriptomic profiles of resident fibroblasts 29 and partly as different subtypes of CAF could exert distinct paracrine actions affecting specific tumor oncogenesis.14,15 Currently, either the number of CAF subpopulations present in the tumor stroma and the role assumed by the presence of an individual population or different cell types into tumor initial development stages are unknown.6,16 Some characteristics distinguish CAFs from quiescent fibroblasts as, for instance, metabolic adaptations supporting their need for advanced proliferation and biosynthesis activities. 13 Furthermore, a potentially controversial area of research on CAFs is focused on their origin. In fact, to define and identify origin of fibroblasts, it is fundamental to consider that CAFs are “activated fibroblasts” and unlike the non-activated (quiescent) fibroblasts residing in the tissue, they are an expansion of the cell population proliferating “in situ” or are recruited in the tumor. 30 Recruitment of BM mesenchymal stem cells, differentiation from adipose stem cells, or conversion from endothelial cells through an epithelial-endothelium-mesenchymal transition process are potential origins of CAFs. 13 However, the best documented source of CAFs is the activation of “normal” resident fibroblasts which, with their heterogeneity, imply the existence of different subsets. This heterogeneity may reflect the variability with respect to phenotypic state of both cell and tissue of origin and therefore also the reporting mediators and the mechanisms to be activated.14,31 In fact, CAFs can be distinguished from other types of cells within the tumor by means of exclusion criteria defined by their morphological characteristics and by a lack of expression of non-mesenchymal markers, such as those expressed by endothelial, epithelial, immune, and neuronal cells even if none of these has an absolute specificity. 32 A challenging aspect in CAFs studies is the precise definition of heterogeneous CAF populations in distinct phases of tumor progression through markers informing on their functions.6,33
In this study, we used the proposed NMF-based gene extractor methodology to investigate CAFs heterogeneity to identify the possible presences of genes, which can be biomarkers and characterize activated fibroblast phenotype. Gene expression profiles of CAFs belonging to primary cultures of 3 distinct solid tumor histotypes from patients (ie, colon of endodermal origin, breast carcinoma of ectodermal origin and ovary of mesodermal origin) were considered. Moreover, we also verified the existence of common genes among CAF gene expression profiles of selected solid tumors and those of BM of patients with multiple myeloma (MM) 6 and MGUS. 17
Identification of CAF populations and download of transcriptomic profiles
International Agency for Research on Cancer (IARC) estimates 18.1 million (17.0 million excluding non-melanoma skin cancer) new cancer cases in 2018, causing about 9.6 million of deaths (9.5 million excluding non-melanoma skin cancer). 34
For studying the heterogeneous CAF population, which represents the most abundant cellular component of the TME, representative tumor histotypes with high rate of mortality have been selected: colon carcinoma, breast cancer, ovarian cancer, and MM. The gene expression profiles of CAFs were downloaded from the Gene Expression Omnibus database (GEO) NCBI (a functional public genomic database that supports the submission of MIAME-compliant data) directly from the original publications of each series (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GEO series). 35 Data sets were selected according to the following standards: (a) the GEO platform (GPL) and (b) the number of samples with labels to identify the fibroblasts associated with the tumor (being ⩾7 the total amount of samples for each representative cancer). Table 1 reports GSE series, bibliographical references, GPL, GSM labels, and the numbers of samples and expressed genes for each data set and finally, the rank values adopted to obtain NMF decomposition.
Dataset information: GSE series used, bibliographical references, GEO platforms (GPL570, GPL6244, and GPL2136 indicate Affymetrix Human Genome U133 Plus_2.0 Array, Affymetrix Human Gene 1.0 ST Array, and Micro-CRIBI Human Oligo Array [Operon V2.0], respectively), fibroblast sample labels, representative cancer, number of genes, number of samples, and NMF rank value

Results
Functional analysis
Nonnegative matrix factorization–based methodology identified a subset of 108 genes common to colon, breast, and ovarian tumors. The biological analysis these genes underwent during the functional and visual analysis phase is detailed in the following. Table 2 summarizes the Gene Ontology functional analysis, Table 3 the Pathway enrichment analysis, whereas in Table 4 are detailed all the 108 genes that underwent the biological analysis. Furthermore, considering that the population of CAF in TME is also recruited by the BM, contributing with important percentages on the total CAF population, the gene expression profiles of primary CAFs of MGUS and MM were used to select the presence of genes, which are common among the CAF “in situ” of the analyzed tumors and CAF recruited by the BM.
Gene ontology functional analysis: based on the parameters used, 10 categories are identified as enriched categories and all are shown in this table. 9
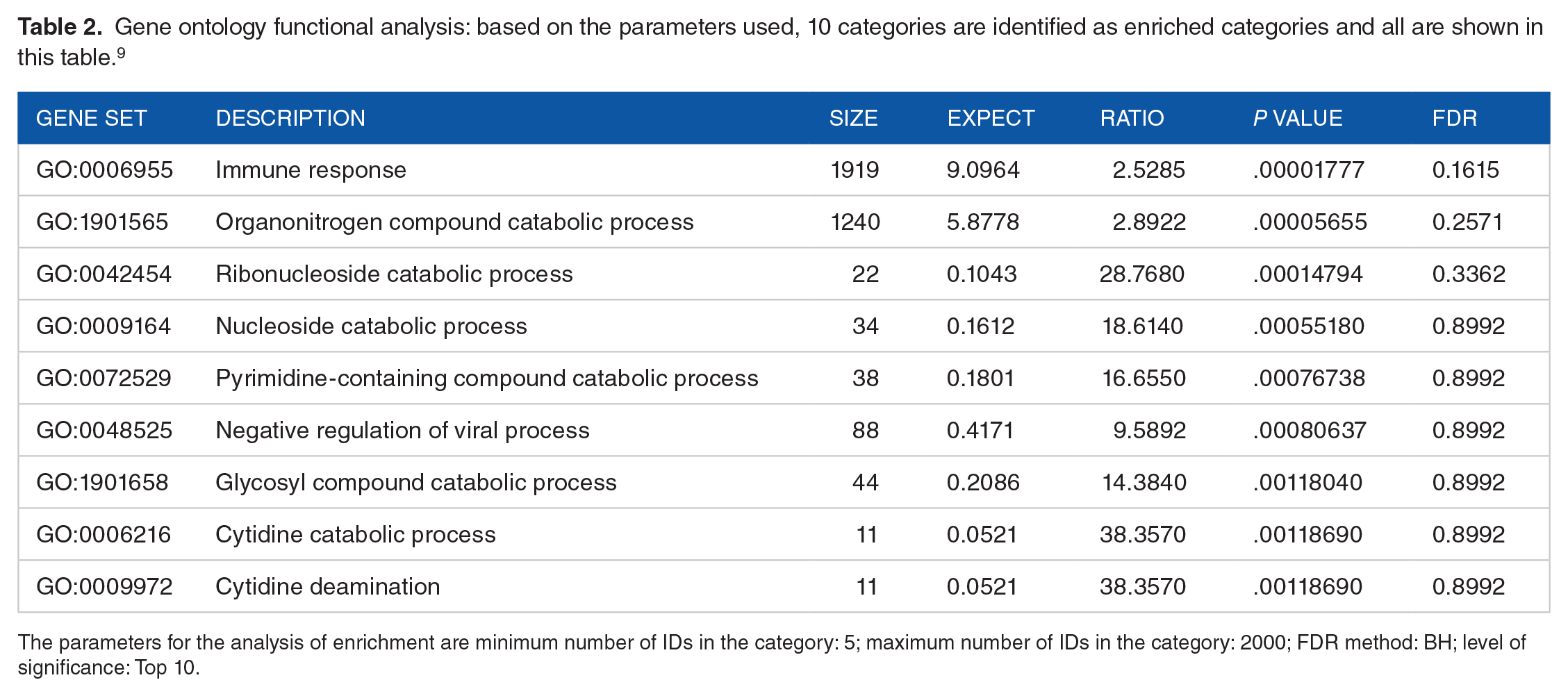
The parameters for the analysis of enrichment are minimum number of IDs in the category: 5; maximum number of IDs in the category: 2000; FDR method: BH; level of significance: Top 10.
Pathway enrichment analysis: based on the parameters used, 10 categories are identified as enriched categories and all are shown in this table. 9
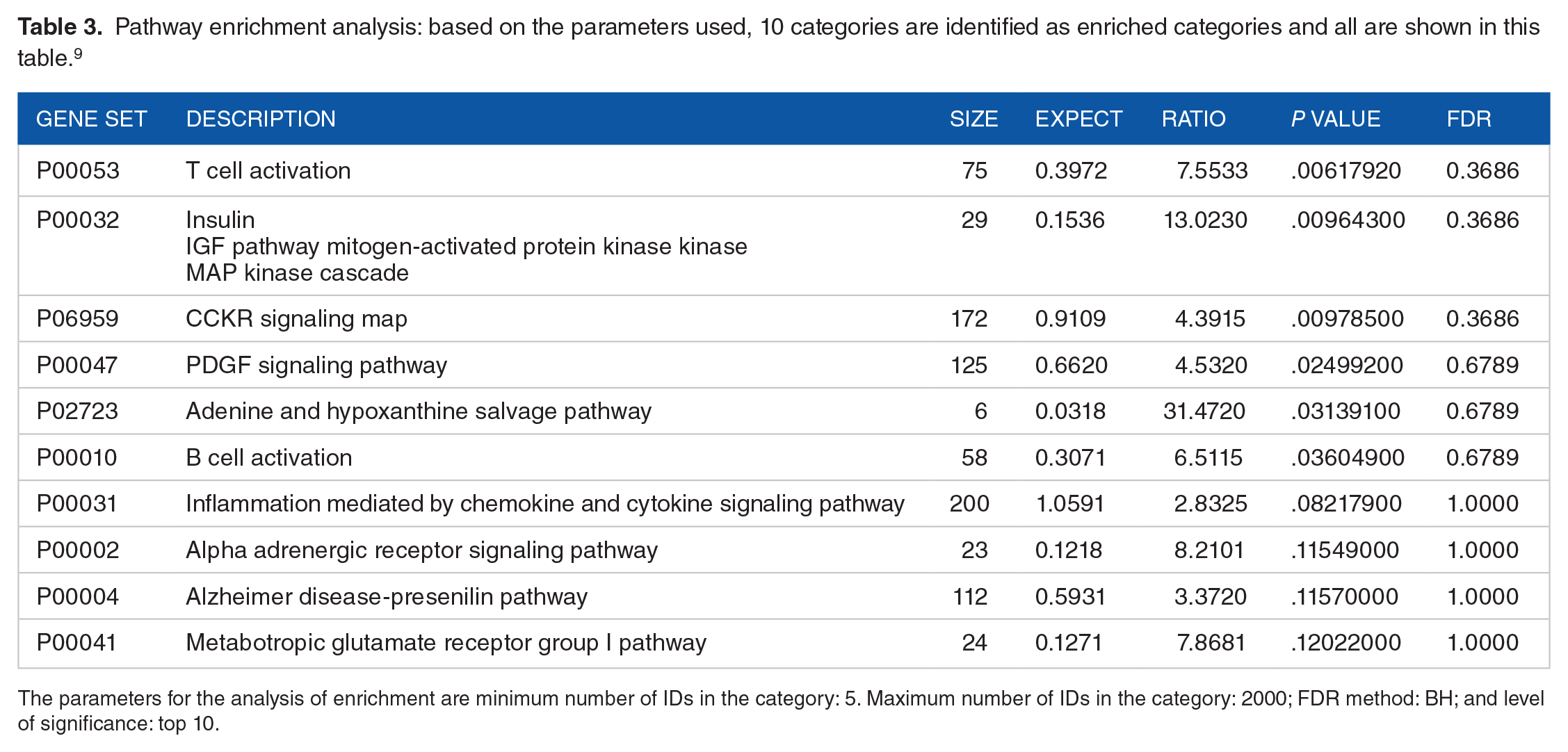
The parameters for the analysis of enrichment are minimum number of IDs in the category: 5. Maximum number of IDs in the category: 2000; FDR method: BH; and level of significance: top 10.
List of the 108 genes common to colon, breast, and ovarian tumors identified by NMF-based methodology described in this article.

Gene Ontology functional analysis
Gene Ontology (GO) was used to classify common genes; classification is obtained according to biological processes, molecular functions, or cellular components. Some genes were identified as related to processes significantly representative of the immune response and associated with nucleic acids metabolism. In particular,
23 (21.3%) and 13 (12%) genes are involved in the “immune response” and “innate immune response” biological processes with false discovery rate (FDR) values 0.1615 and 0.3362, respectively.
17 genes (15.7%) are involved in the catabolic “organonitrogen compound catabolic process” with an FDR value of 0.2571; they are
The 3 major biological processes “immune response,” “innate immune response,” and “organonitrogen compound catabolic process” characterizing the GO share 3 genes:
The
The genes that differentiate the “immune response” from the “innate immune response” are
The
Pathway enrichment analysis
PANTHER reference database (Protein Analysis Through Evolutionary Relationships) was used to evaluate with the WebGestalt tool the selected 108 genes during the pathway enrichment analysis. The selected 10 pathways were reported in Table 3. The first 6 pathways are characterized by an FDR below 1 and a
Figure 4 illustrates the most frequent genes and the pathway they belong to using a variation of the UpSet plot. Height and width of bars are proportional to the relative frequencies and to the number of genes selected in each set of pathway.

Variation of the UpSet plot
59
of the most frequent genes:
The
Identification of genes common among CAFs in colon, breast, and ovarian tumors and MGUS
The subgroup of the 108 genes obtained from the intersection of genes belonging to the representative metagenes of the CAF gene profiles related to colon, ovary, and breast tumors has been intersected with the metagene representative of the MGUS CAF gene profiles. The data sets containing gene subgroups are made available as supplement materials.
The profile intersection produced a subgroup of 9 genes:
Identification of common genes among CAFs in colon, breast, ovary, and MM tumors
Similarly, the subgroup of the 108 genes obtained from the intersection of genes belonging to the metagenes representative of the gene profiles of CAFs related to colon, ovary, and breast tumors was intersected with the representative metagene of the gene profiles of MM CAF. The profile intersection produced only one gene: ENPP2 (ectonucleotide pyrophosphatase/phosphodiesterase or autotaxin-ATX; illustration is provided in Figure 5). Autotaxin is an exo-enzyme originally identified as an autocrine motility factor of the cancer cell. ATX is unique among nucleotide pyrophosphatase/phosphodiesterase (NPPs) as it functions primarily as lysophospholipase D, converting lysophosphatidylcholine into the lipid mediator of LPA. Lysophosphatidic acid acts on specific G protein–coupled receptors to elicit a wide range of cellular responses, ranging from cell proliferation and migration to the production of cytokines.75,76

ECM (extracellular matrix). The ENPP2 protein, common to the CAFs of the 4 tumors analyzed, catalyzes the LPC hydrolysis in LPA, activating its local LPA receptors and the corresponding G proteins. LPA signals through its receptors to induce proliferation, survival, and invasion in tumor cells and cancer stem cells. LPA signaling also induces the recruitment of CAFS cells and a wide range of cellular responses and also reduces the cytotoxic immune response.
Discussion
The transcriptome analysis of colon, ovary, and breast tumors selected a subgroup of 108 genes by NMF-based methodology identifying a CAF phenotype with morphological and functional characteristics regardless of embryonic origin. In fact, the
Conclusions
We proposed a mathematical methodology based on NMF, which has been originally assembled for automatically extracting common genes from different gene expression data. This computational mechanism has 2 main features: (1) through NMF method, metagenes which are the most representative of the information embedded into different tumor gene expression data are extracted; (2) through basic mathematical set operations (implemented into the gene landscape extraction module), genes common to different tumor histotypes are selected. These genes can then be easily analyzed from a biological point of view through to the functional and the visualization approaches based on the WebGestalt tool, which integrates the computational mechanisms.
The proposed mechanism demonstrated its usefulness in identifying genes which could be helpful to better understand the molecular mechanisms behind the activation of fibroblasts and their role in tumor progression. Different CAF subpopulations in the TME can be distinguished from one another by the expression of one or more specific markers. In general, CAFs have the common function of altering the TME; in fact, once activated, fibroblasts synthesize and deposit ECM components, release chemokines and cytokines in the stroma, and generate tension forces at the tissue level through their cytoskeletons, all are requirements key for tissue remodeling. The multifaceted nature of CAF is therefore linked to the TME in which the various pathways are activated as the mechanism based on the TGF-β signal and/or the bidirectional communication between the fibroblasts and the tumor cells.
Our study has shown that, even if there is phenotypic heterogeneity of CAF (APOBEC3B, APOBEC3G), in tumors of different embryonic origin, there are common biomarkers, characterizing the phenotype of activated fibroblasts. Furthermore, we were able to identify a phenotype of CAF as an anti-inflammatory (ANXA1, UBA6) likely associated with the first stages of tumor transformation and subsequently pro-inflammatory with the
Supplemental Material
Supplementary_File_1_xyz32442b9e8854b – Supplemental material for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations
Supplemental material, Supplementary_File_1_xyz32442b9e8854b for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations by Flavia Esposito, Angelina Boccarelli and Nicoletta Del Buono in Bioinformatics and Biology Insights
Supplemental Material
Supplementary_File_2_xyz32442e5519468 – Supplemental material for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations
Supplemental material, Supplementary_File_2_xyz32442e5519468 for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations by Flavia Esposito, Angelina Boccarelli and Nicoletta Del Buono in Bioinformatics and Biology Insights
Supplemental Material
Supplementary_File_3_xyz324423708d119 – Supplemental material for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations
Supplemental material, Supplementary_File_3_xyz324423708d119 for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations by Flavia Esposito, Angelina Boccarelli and Nicoletta Del Buono in Bioinformatics and Biology Insights
Supplemental Material
Supplementary_File_4_xyz324420b6b3b2c – Supplemental material for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations
Supplemental material, Supplementary_File_4_xyz324420b6b3b2c for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations by Flavia Esposito, Angelina Boccarelli and Nicoletta Del Buono in Bioinformatics and Biology Insights
Supplemental Material
Supplementary_File_5_xyz32442f879d720 – Supplemental material for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations
Supplemental material, Supplementary_File_5_xyz32442f879d720 for An NMF-Based Methodology for Selecting Biomarkers in the Landscape of Genes of Heterogeneous Cancer-Associated Fibroblast Populations by Flavia Esposito, Angelina Boccarelli and Nicoletta Del Buono in Bioinformatics and Biology Insights
Footnotes
Author Contributions
AB designed the research and analysed the numerical results. NDB and FE performed the mathematical analysis and the data preprocessing. FE designed and implemented the framework modules. AB, NDB and FE drafted and rieviewed the manuscript and did the critical revision of the final version of the paper. All the authors read and approved the manuscript in its final form.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by the GNCS-INDAM (Gruppo Nazionale per il Calcolo Scientifico of Istituto Nazionale di Alta Matematica) Francesco Severi, P.le Aldo Moro, Roma, Italy.
Supplemental material
Supplemental material for this article is available online.
*
INDAM Research Group GNCS.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
