Abstract
Empirical Bayes is a choice framework for differential expression (DE) analysis for multi-group RNA-seq count data. Its characteristic ability to compute posterior probabilities for predefined expression patterns allows users to assign the pattern with the highest value to the gene under consideration. However, current Bayesian methods such as baySeq and EBSeq can be improved, especially with respect to normalization. Two
Introduction
RNA-seq is a common tool to obtain expression data.1,2 It is popularly applied to identify differentially expressed genes (DEGs) or transcripts under different groups or conditions.3,4 Accurate identification of these DEGs is crucial for multiple purposes; for example, they may serve as potential biomarkers for clinical diagnosis.5,6 So far, many methods have been developed for analysis of RNA-seq data7–20 and several evaluation studies have also been performed.21–25 Of these, two
The two packages commonly employ a generalized linear model (GLM) framework. When comparing a multi-group data (eg G1 vs G2 vs G3), the main output is an analysis of variance (ANOVA)-like
However, constructing a complete expression pattern based on the results of three two-group comparisons can be difficult. For example, two possible patterns (G1 ≠ G2 = G3 or G1 = G2 = G3) can be constructed if the results of the three two-group comparisons were G1 ≠ G2, G1 = G3, and G2 = G3. Furthermore, results of the three two-group comparison themselves (ie G1 ≠ G2, G1 = G3, and G2 = G3) can vary depending on both the multiple comparison procedure and the significance level. As the number of groups to be compared increases, construction of the complete expression pattern across all groups based on the GLM framework becomes more difficult.
Different from the GLM framework, an empirical Bayesian framework implemented in baySeq
10
and EBSeq
12
returns one posterior probability (PP) for each of the predefined expression pattern for each gene; thus, when considering a number of patterns for a particular gene, it can assign the pattern with the highest PP to the gene under consideration. In other words, the Bayesian framework does not require subsequent analysis such as the
We here focus on the improvement of the Bayesian framework. We demonstrate that a robust normalization strategy provided in TCC 13 can be adapted in the analysis pipeline. Although the Bayesian framework is generally inferior to the GLM framework when the overall degree of DE is evaluated,21,22 accurate assignment of expression patterns to individual genes obtained from the Bayesian framework with TCC can be independently valuable.
Materials and Methods
All analyses were performed using
DE analysis pipelines
In general, DE analysis consists of two steps (data normalization
According to the previous notation,
22
we refer to this pipeline as
The DE analysis typically starts with a so-called “count matrix,” where each row indicates the gene, each column indicates the sample, and each cell indicates the number of reads mapped to the gene in the sample. The
For the default normalization methods
Simulation data
In this study, to perform the multi-group comparison as simply as possible, we focused on the three-group data (G1 vs G2 vs G3) with equal numbers of biological replicates (ie 3 or 9 replicates per group;
The simulation framework and evaluation metric are the same as our previous study,
22
enabling the comparison of the current results with the previous ones. The simulation conditions were as follows: the total number of genes was 10 000 (
Real data
A total of three count data sets were analyzed. The first data set was originally sequenced from the three species (ie the three-group data): humans (G1), chimpanzees (G2), and rhesus macaques (G3). 31 Briefly, Blekhman et al studied expression levels of liver samples from three males and three females from each species, giving a total of six different individuals (ie six biological replicates) for each species. Since they performed duplicate experiments for each individual (ie two technical replicates), the publicly available raw count matrix consists of 20 689 genes × 36 samples (=3 species × 6 biological replicates × 2 technical replicates). To correctly estimate the biological variation and make the assumed structure of input data, we summed and collapsed the count data of technical replicates, giving a reduced number of columns in the count matrix (ie 18 samples).
The second and third data sets were derived from a study of human brain (SRP056477). 32 The count data consisting of 58 037 genes × 52 samples were obtained from the recount2 database. 33 The samples were divided into two source types: 25 cerebellum (CER) and 27 frontal cortex (FCX) samples, and further subdivided into three case types: healthy, sporadic amyotrophic lateral sclerosis (sALS), and ALS caused by a repeat expansion in C9orf72 (c9ALS). Accordingly, we performed two three-group comparisons (G1 vs G2 vs G3 as healthy vs sALS vs c9ALS) for these data. For simplicity, we excluded one sample (SRR1927053) from the CER data set and three samples (SRR1927071, SRR1927052, and SRR1927054) from the FCX data set such that each group had eight replicates.
Normalization
A total of five normalization methods (
Expression patterns
In this study, a total of five possible expression patterns were considered when performing the Bayesian methods: non-DEG pattern (say “
DE analysis with baySeq
The baySeq was performed using the
DE analysis with EBSeq
The EBSeq was performed using the
Evaluation metrics
The evaluation was performed using the rank information of PPs assigned for
In case of the binary (0 for non-DEG and 1 for DEG) classification problem, genes predicted as DEG by the pipeline were labeled as either true positive (TP) or false positive (FP). Genes labeled as TP correspond to those correctly predicted as DEG (ie the truth is DEG) and genes labeled as FP correspond to those falsely predicted as DEG (ie the truth is non-DEG). Similarly, genes predicted as non-DEG by the pipeline can be labeled as either true negative (TN) or false negative (FN). Genes labeled as TN correspond to those correctly predicted as non-DEG (ie the truth is non-DEG) and genes labeled as FN correspond to those falsely predicted as non-DEG (ie the truth is DEG). The accuracy was calculated as (TP + TN)/(TP + TN + FP + FN), where the denominator corresponds to the total number of genes (=10 000) and the numerator corresponds to the total number of correctly predicted genes. The actual FDR, sensitivity, and specificity were calculated as FP/(FP + TP), TP/(TP + FN), and TN/(TN + FP), respectively.
In this study, the Bayesian methods (baySeq and EBSeq) were performed by considering four or five possible expression patterns. When considering four expression patterns (ie
Results and Discussion
Simulation results when considering four expression patterns (Nrep = 3)
We assessed the performance of a total of eight DE pipelines when four possible expression patterns (
Average AUC values for simulation data (
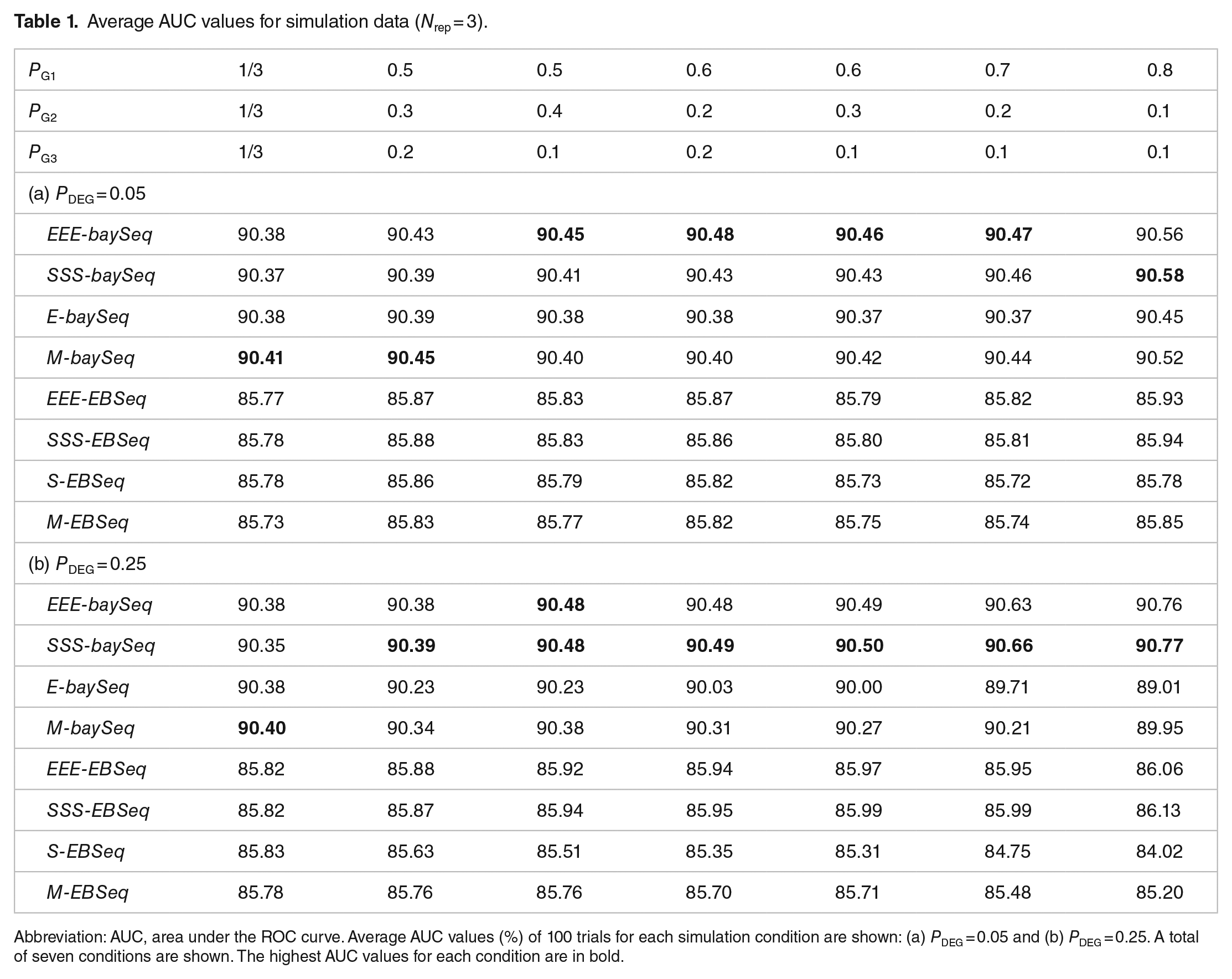
Abbreviation: AUC, area under the ROC curve. Average AUC values (%) of 100 trials for each simulation condition are shown: (a)
When we compared four baySeq-related pipelines, the AUC values for the baySeq with TCC (ie

Boxplots of AUC values for simulation data (
Surprisingly,
While the AUC values shown in Table 1 were comparable to those in our previous study (ie Table 1 by Tang et al
22
), the values for
Despite the different numbers of possible patterns to be considered, we observed similar AUC values between the current and previous studies. This is probably because these values are calculated based on the PPs assigned for the
In practice, it is important to control FDR when identifying DEGs. Remember we regarded genes satisfying
It should be noted that the AUC values (<91%) of the best performing pipeline (ie
Simulation results when considering four expression patterns (Nrep = 9)
We previously reported that the relative performances for the original EBSeq pipeline (ie
Simulation results when considering five expression patterns
Remember that the simulation data have four expression patterns (
Overall, the number of genes assigned to the
Results for real data
We analyzed a real data set consisting of 20 689 genes × 18 liver samples for three-group comparison: six humans (G1), six chimpanzees (G2), and six rhesus macaques (G3).
31
Table 2 shows the number of genes assigned to individual patterns when considering (1) five patterns, (2) four patterns, and (3) the common ones. Similar to the simulation results, the numbers of genes assigned to
Numbers of genes assigned to individual patterns.

Abbreviations: DE, differential expression; FDR, false discovery rate. Genes satisfying 5% nominal FDR (
Surprisingly, we observed considerably different results among the EBSeq-related pipelines with four patterns (Table 2). While three pipelines (
Conclusions
We evaluated a total of eight Bayesian-based DE pipelines using three-group RNA-seq count data. The Bayesian methods coupled with TCC normalization performed comparably or better than the Bayesian methods with the default normalization settings. In particular, the AUC values for
It is important to note that we do not recommend the use of
Supplemental Material
Additional1_xyz19222377f73da – Supplemental material for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data
Supplemental material, Additional1_xyz19222377f73da for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data by Takayuki Osabe, Kentaro Shimizu and Koji Kadota in Bioinformatics and Biology Insights
Supplemental Material
Additional2_xyz1922267354467 – Supplemental material for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data
Supplemental material, Additional2_xyz1922267354467 for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data by Takayuki Osabe, Kentaro Shimizu and Koji Kadota in Bioinformatics and Biology Insights
Supplemental Material
Additional3_xyz19222d459e214 – Supplemental material for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data
Supplemental material, Additional3_xyz19222d459e214 for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data by Takayuki Osabe, Kentaro Shimizu and Koji Kadota in Bioinformatics and Biology Insights
Supplemental Material
Additional4_xyz1922271719133 – Supplemental material for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data
Supplemental material, Additional4_xyz1922271719133 for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data by Takayuki Osabe, Kentaro Shimizu and Koji Kadota in Bioinformatics and Biology Insights
Supplemental Material
Additional5_xyz19222be3d8dc9 – Supplemental material for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data
Supplemental material, Additional5_xyz19222be3d8dc9 for Accurate Classification of Differential Expression Patterns in a Bayesian Framework With Robust Normalization for Multi-Group RNA-Seq Count Data by Takayuki Osabe, Kentaro Shimizu and Koji Kadota in Bioinformatics and Biology Insights
Footnotes
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by JSPS KAKENHI Grant Number JP18K11521.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
TO made the R codes, performed the analysis and drafted the manuscript. KS designed the study, supervised the critical discussion and revised the paper. KK confirmed the analysis, revised the paper and led this project to completion.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
