Abstract
In this article we try to discuss nonparametric linkage (NPL) score functions within a broad and quite general framework. The main focus of the paper is the structure, derivation principles and interpretations of the score function entity itself. We define and discuss several families of one-locus score function definitions, i.e. the implicit, explicit and optimal ones. Some generalizations and comments to the two-locus, unconditional and conditional, cases are included as well. Although this article mainly aims at serving as an overview, where the concept of score functions are put into a covering context, we generalize the noncentrality parameter (NCP) optimal score functions in Ängquist et al. (2007) to facilitate—through weighting—for incorporation of several plausible distinct genetic models. Since the genetic model itself most oftenly is to some extent unknown this facilitates weaker prior assumptions with respect to plausible true disease models without loosing the property of NCP-optimality.
Moreover, we discuss general assumptions and properties of score functions in the above sense. For instance, the concept of identical by descent (IBD) sharing structures and score function equivalence are discussed in some detail.
Keywords
Introduction
In

A pedigree set example consisting of 5 distinct pedigrees of different structures and phenotype settings.
In practise the genotypes are observed as well-defined
As a restriction or application, in
In this context the prime quantity of central importance to the actual statistical analysis-procedure is the process of
For a single pedigree, at locus

where
A
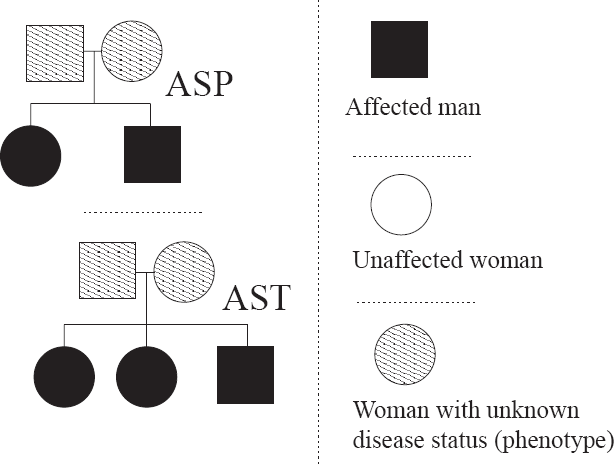
The pedigree structures corresponding to affected sib-pair (ASP) and affected sib-trio (AST) pedigrees.
In the same manner as (1), but somewhat more obscure, one may summarize inheritance through
In other words, they are both inherited instances of
To numerically facilitate analysis of inheritance and phenotype-genotype dependence one may introduce a
One may note that this notion of a score function may be seen as adopting a data-mining perspective where such functions are used for scoring patterns (Hand et al. 2001). In this case one observes and scores inheritance patterns.
Or more generally within phenotype-groups.

Hence considering a pedigree with

assuming some order of inheritance vectors. 5 In this setting some scores will according to symmetry, and in some cases by—explicitly or implicitly—definition, be numerically equal. Using the context of IBD-sharing structures one may reformulate (2) as
For instance using the standard decimal interpretation (conversion) of the binary zero-one inheritance vector with length

where the index corresponds to the by-score-ordered set of IBD-sharing structures, i.e. a natural restriction (order) is given by assuming
According to the fact that the pedigree construction excluded farther (earlier) generations.
For further information on equivalent IBD-sharing structures consider Appendix A.
Our primary goal with this paper is to, in such a generally accessible way as possible, formalize and discuss the structure of nonparametric linkage score functions. Oftenly, in published works, these functions are either directly applied using some of the standard instances or derived in an ad hoc or highly theoretical, or non-intuitive, fashion.
Having this in mind, the text to follow is not a complete summary of suggested and published score function variants, or the most theoretical exposition out there. Rather, it aims at being a review-like overview discussing the underlying structure, contexts of derivations and interpretations (and to some extent performance) of certain families of NPL score functions.
In
Approaches to Score Function Definitions
For an underlying disease to be genetically inheritable, i.e. to include a

where
Now, to define a score function one basically has to instantiate the numerical scores corresponding to (2) or (3). This may be done in several distinct ways, which is furtherly discussed below. What truly is the core question with respect to such definitions is the evidential performance of the corresponding score function. (Most likely in the form of
A score function performing well under a wide range of different λ ∈ Λ, where Λ is the set of all possible disease models, is termed a
Vaguely speaking, as noted above, at a true disease locus, the IBD-sharing within phenotypes should be expected to increase. This makes it possible to define functions, depending on pedigree IBD-sharing only, meeting this requirement (property). Since such functions implicitly instantiate (2) and (3) through the higher-level sharing-based function definition we call them
Traditional score functions
Firstly,

where
Including the ordered affected individuals
Secondly,

where
Each specific selection

Both functions (5) and (6) are defined, given the inheritance vector
In Ängquist (2006) several extensions to traditional score functions are given. Now, assume a traditional instance

This aims at additionally searching for unusual IBD-sharing within the set of unaffecteds
Generally

A pedigree consisting of 4 siblings (two affecteds, two unaffecteds). The two distinct cases (left to right) display the corresponding phenotype-switching process involved in the definition of extended score functions.
A second-order extension may be formulated as

where UP denotes the set of individuals with unknown phenotype. Here one additionally corrects for the
This might be the case for, in some relative (to the disease) sense,
It is perfectly possible not to use a closed definition or high-level algorithm when calculating the vector of scores constituting the corresponding score function. We refer to such cases as
The construction of an explicit score function reduces to (explicitly) distributing scores to all present IBD-sharing structures, thus reflecting numerically the assumed connection between these sharing structures and evidence for a present disease locus. For instance, such an approach might be interesting if one can show by some real examples, or a priori assume, that certain combination of inheritance vector states are impossible or unlikely.
Explicit definitions, so to speak, implicitly make some (though quite vague) assumptions on the type of underlying disease structure. In this sense they are more strongly directed towards certain disease models than implicit definitions, but much less so than the family of definitions described below in Section 2.3. There explicit assumptions on true (plausible) genetic disease models λ under corresponding alternative hypotheses
Optimality defined versions
If having an explicit algorithm (as for implicitly defined versions) but where this algorithm is formulated with respect to, in some sense, an optimality criterion
Given a disease model λ, define the expected score at the disease locus under this model as

where
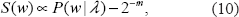
with
Note that
Hence one may note that the optimal score function (10) depends on the true genetic model and should be interpreted as, in this sense, the best possible result that the investigator might expect when the genetic model is correctly specified. In practice though, the genetic model is often unknown. Then in a natural way, for each choice of score function and for a range of different genetic models, (10) facilitates comparisons with optimality, leading to a quantification of the apparent loss of information. The optimal score function might also serve as a form of explicit score function with respect to certain assumptions or prior information.
Further, in Hössjer (2003)
As a way of enhancing interpretation one usually uses

where, for a pedigree with

are the mean and variance of
Note that we end up with the standardized properties
Equipped with the concept of standardization one may define
If two unstandardized score functions through standardization are transformed to equal 13 standardized score functions they are referred to as being equivalent. For more detailed information and corresponding equivalence-criterions, see Appendix B.
Two score functions
One may also note that actual numerical standardized scores corresponding to a specific score function (or several equivalent ones) are dependent on the score distribution
This follows since these settings uniquely define the standardization parameters μ and σ in (11).
Note that throughout this article we try to discuss score functions without explicitly mentioning the actual test statistics they are used in connection with when facing real and imperfect marker data MD. 15
In other words, when the complete inheritance process over corresponding loci is not known with probability one.
An exception is the use of standardization through (11) which implicitly refer to the practise of the ‘NPL score’ test statistic (Kruglyak et al. 1996; Ängquist, 2007).
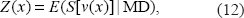
where the expected value, at locus
Note that (12) refer to a single pedigree (
Given imperfect data the variance of the NPL score
A textbook on HMMs is Cappé et al. (2005).
Replacing σ
2
in (11) with
On
However, note that although the choice of test statistic and possible standardization procedure are important from a testing and statistical significance perspective it is not particularily essential for the present discussion. Moreover, generally the interpretations and relative performances of the different score function variants will not change when dealing with imperfect data, hence this matter is only noted on in this specific subsection.
One may generalize the one-locus procedure above in order to simultaneously, or sequentially, search for two distinct disease loci on the genome. The former case is referred to as an
Implicitly defined score functions may in some cases be relatively easily generalized to the two-locus case, but in some cases the corresponding score-algorithm will be refrainingly more complex. As a positive example, one may generalize (5) into a two-locus score function. In Ängquist et al. (2007) the following, quite general, formulation is given

where IBD
For

which these authors also implemented into the analysis program GENEHUNTER-TWOLOCUS. In the applications of Ängquist et al. (2007) the case
Note that this is possible according to our assumption of scoring all inheritance vectors leading to similar IBD-sharing structures equally. In this case, at each locus, the


Two-locus explicitly defined score functions are concept-wise straightforward generalizations of one-locus ones. Moreover, the NCP-optimal score function (10) of Ängquist et al. (2007), for unconditional and conditional two-locus analysis respectively, may be generalized to

Note that the interpretation of these scores as being proportional to probability-based differences with respect to the null and (assumed) alternative hypotheses still hold true.
In some cases where the true disease model λ is fully or partially unknown the usage of the NCP-optimal score function (10) based on an estimate (or assumption)
Algorithm
Begin with choosing

with inheritance distributions under corresponding alternatives

Now, a simple generalization to the previous score in (10) is given by

where
A further generalization arises if adopting a
Subjective or empirically objective perspective; for concepts see e.g. Winkler (1972) and Gelman et al. (2004).
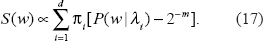
One may note that (16) is the special case of (17) where π = (1/
For illustrational purposes we include a small-scale simulation analyses in this section. We perform power calculations for various settings and present them through

Power calculations for Pedigree 1 and score functions
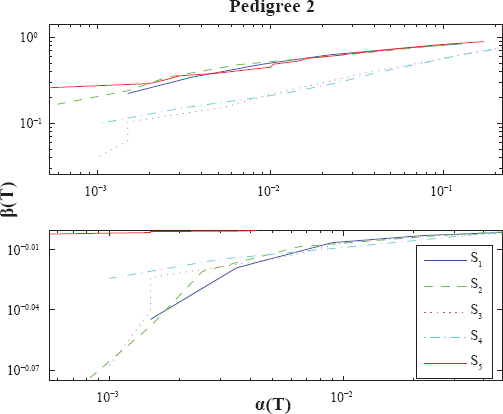
Power calculations for Pedigree 2. See caption of Figure 4.

Power calculations for Pedigree 3. See caption of Figure 4.
Consider a pedigree consisting of two parents of unknown phenotypes and
Further, for each case we use a genome consisting of a single chromosome of length
Finally, we used two genetic models, λ1 and λ2, where both correspond to disease allele frequency
respectively. Here
Results and discussion
It is quite hard to draw very certain conclusions from such a small study, once more note that this section is in some sense a side-track, but a few general observations of some interest may be stated: (i)
Footnotes
Acknowledgements
I send my best regards to Professor Ola Hössjer for prior co-authorship, discussions and ideas that strongly affected my appreciation and views of the concepts constituting this article. Thank you!
I am grateful also towards two anonymous reviewers for several insightful comments and suggestions.
