Abstract
Background:
Pancreatic ductal adenocarcinoma (PDAC) is a highly lethal malignancy with currently limited early detection options. Extracellular vesicle (EV)-derived microRNAs (miRNAs) have gained interest as non-invasive diagnostic biomarkers due to their stability in circulation and tumour-specific profiles. However, the methodological robustness of existing literature remains unclear.
Objectives:
To systematically evaluate the diagnostic accuracy and methodological quality of studies investigating EV-derived miRNAs for PDAC detection, with a particular focus on adherence to established EV characterisation guidelines.
Design:
Systematic review registered with PROSPERO (CRD42024501503) and conducted in accordance with PRISMA 2020 reporting standards.
Methods:
We searched PubMed, EMBASE, Medline and Cochrane for original human studies published up to February 1, 2025, evaluating EV-derived miRNAs in biofluids from PDAC patients. Eligible studies reported diagnostic accuracy metrics (sensitivity, specificity, Area Under the Curve (AUC)). Methodological quality was assessed using the QUADAS-2 tool, and EV validation was scored against the Minimal Information for Studies of Extracellular Vesicles 2018 (MISEV) 2018 checklist (updated 2023).
Results:
Fifty-six studies were included. Plasma and serum were the most commonly used biofluids. The most frequently evaluated individual miRNAs were miR-21 (13 studies), miR-10b (9 studies), and miR-451a (7 studies). Although several studies reported high diagnostic performance (AUCs up to 0.99), MISEV adherence was limited: only 23.1% of miR-21 studies demonstrated strong EV validation, and >70% of all studies lacked EV quantification or protein marker analysis. Multi-miRNA panels achieved higher AUCs (often > 0.85) but typically scored poorly on EV characterisation. Only 2 of 56 studies included external validation, and 54 studies lacked blinding, contributing to substantial risk of bias.
Conclusion:
EV-derived miRNAs are promising PDAC biomarkers, but progress is hindered by inconsistent methods, poor EV validation, and minimal external verification. Translation to clinical use requires robust EV characterisation, standardised workflows, and prospective multi-cohort studies.
Plain Language Summary
Pancreatic cancer is often diagnosed at a late stage and is one of the hardest cancers to treat. Researchers are exploring ways to detect it earlier, especially through simple blood tests. One promising method involves studying tiny particles in the blood called extracellular vesicles (EVs). These particles can carry small molecules known as microRNAs (miRNAs), which may signal the presence of cancer. In this study, we reviewed 56 research papers that tested EV-derived miRNAs as potential early detection tools for pancreatic cancer. Many of these studies reported promising results, including high accuracy in distinguishing cancer patients from non-cancer patients. However, most did not follow key international quality guidelines called the Minimal Information for Studies of Extracellular Vesicles (MISEV) 2018 standards. These standards help ensure that EVs are real, uncontaminated, and properly measured. We also found that many studies did not use “blinding” which is a process where researchers analysing the samples do not know which ones are from cancer patients. This helps prevent unconscious bias, where expectations might unintentionally affect how results are interpreted. Additionally, very few studies tested their findings in new groups of patients, which is important for confirming that the results are reliable. Our review shows that while EV-miRNAs could be useful for detecting pancreatic cancer earlier, the overall quality of current research is limited. Future studies need to follow international standards more closely, use blinding, and include independent testing before these methods can be confidently used in clinical practice.
Keywords
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is the most common malignancy of the pancreas, and one of the deadliest cancers. 1 In the UK around 10 500 patients are diagnosed with the disease every year. 2 Due to steadily increasing incidence and by association mortality, PDAC poses an unmet clinical challenge causing a significant burden on both patients and healthcare providers.3,4 This foreboding outlook is underpinned by nonspecific symptoms, late-stage diagnosis and limited therapeutic options. Around two-thirds of patients have metastatic disease at diagnosis, which is associated with an exceedingly poor 3-year survival of <5%. 5 Research focussing on aetiology and early detection thus offers best hope for changing the course of this disease. Currently, there are no clinically validated biomarkers for early detection of PDAC - Carbohydrate antigen 19-9 (CA19-9) is used in the management of patients with PDAC but offers an inadequate sensitivity (SN), specificity (SP) and positive predictive value (PPV) to warrant its use as an early biomarker. 6
Extracellular vesicles (EVs) are lipid bilayer membrane-enclosed vesicles that are secreted from cells into the extracellular space and can be isolated from plasma, bile, urine and other biofluids. 7 The interest in EVs is rapidly increasing due to improving isolation techniques and their overexpression in cancer. 8 PDAC-derived EVs have already been shown to initiate pre-metastatic niche formation in the liver, a key step in the development of metastatic disease. 9 The EV load can include various bioactive molecules such as lipids, proteins and microRNAs (miRNAs). 10
MicroRNAs (miRNAs) are small (≈22 nucleotide) non-coding RNAs that regulate gene expression, usually through translational repression or by reducing messenger RNA stability. 11 They can act as tumour suppressors or oncogenes and miRNA profiling can be used to distinguish different cancer subtypes. 12 Furthermore, miRNAs have also been shown to function as ligands which directly bind Toll-like receptors and activate pro-metastatic signalling pathways. 13 MiRNAs are particularly interesting biomarkers due to their presence and stability in the circulation. This is especially the case when they are contained within EVs, which can further shield them from enzymatic degradation. 14
This review focuses on EV miRNAs as biomarkers in PDAC, their isolation methodology, downstream target genes and diagnostic accuracy. An additional aim was to gain oversight as to whether current literature is meeting the standards required for consistency and reproducibility as defined in the 2018 guidelines by the International Society for Extracellular Vesicles. 15
Methods
Search Strategy
This systematic review is registered in PROSPERO (CRD42024501503) and adhered to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) 2020 guidelines. 16 It was not feasible to conduct a meta-analysis due to substantial heterogeneity in study methodologies, biofluid sources and reporting of diagnostic performance metrics. The authors conducted comprehensive and systematic search on 4 separate electronic databases: PubMed, Medline, EMBASE, and Cochrane, for papers published between 1946 and February 1, 2025, when the searches were performed. The search terms for each database are provided in Supplemental Information.
Manual retrieval of the references from included articles was also performed to identify additional relevant studies.
The inclusion and exclusion criteria were pre-specified to determine the eligibility of studies for the review. Studies were included if they: (1) investigated PDAC; (2) utilised non-invasive sample types such as plasma, serum, urine, saliva or stool; (3) were original research articles; and (4) involved human participants. Studies were excluded if they: (1) did not focus on PDAC; (2) did not specify the pancreatic cancer subtype; (3) relied on invasive sampling methods (eg, tissue biopsy); (4) did not report SN, SP and AUC; (5) were abstracts, conference proceedings, NHS reports or review articles; (6) were non-English publications; or (7) involved non-human data.
All retrieved articles were imported into Rayyan software for systematic review (https://www.rayyan.ai/ accessed on 1st February 2025). Duplicates were removed, and relevant articles were screened based on title and abstract.
Data Extraction and Quality Assessment
Data extraction was as follows: year of publication, first author, country of study, total number of samples, PDAC stage (if mentioned), number and health status of controls (if stated), biofluid type, method of EV extraction, miRNA concentration measuring method, miRNA biomarker investigated, diagnostic accuracy parameters, proposed biomarker role (diagnostic or prognostic). Discrepancies between the reviewers were resolved through consensus.
The adherence of papers’ methodologies to the Minimal Information for Studies of Extracellular Vesicles (MISEV) 2018 criteria along with updated sections found in 2023 criteria 15 were tested in order to assess methodological rigour. The criteria used to determine numerical scores is provided in Box 1.
ISEV 2018 Modified EV Characterisation Checklist Score Including Updated Sections Aligning with 2023 Version.
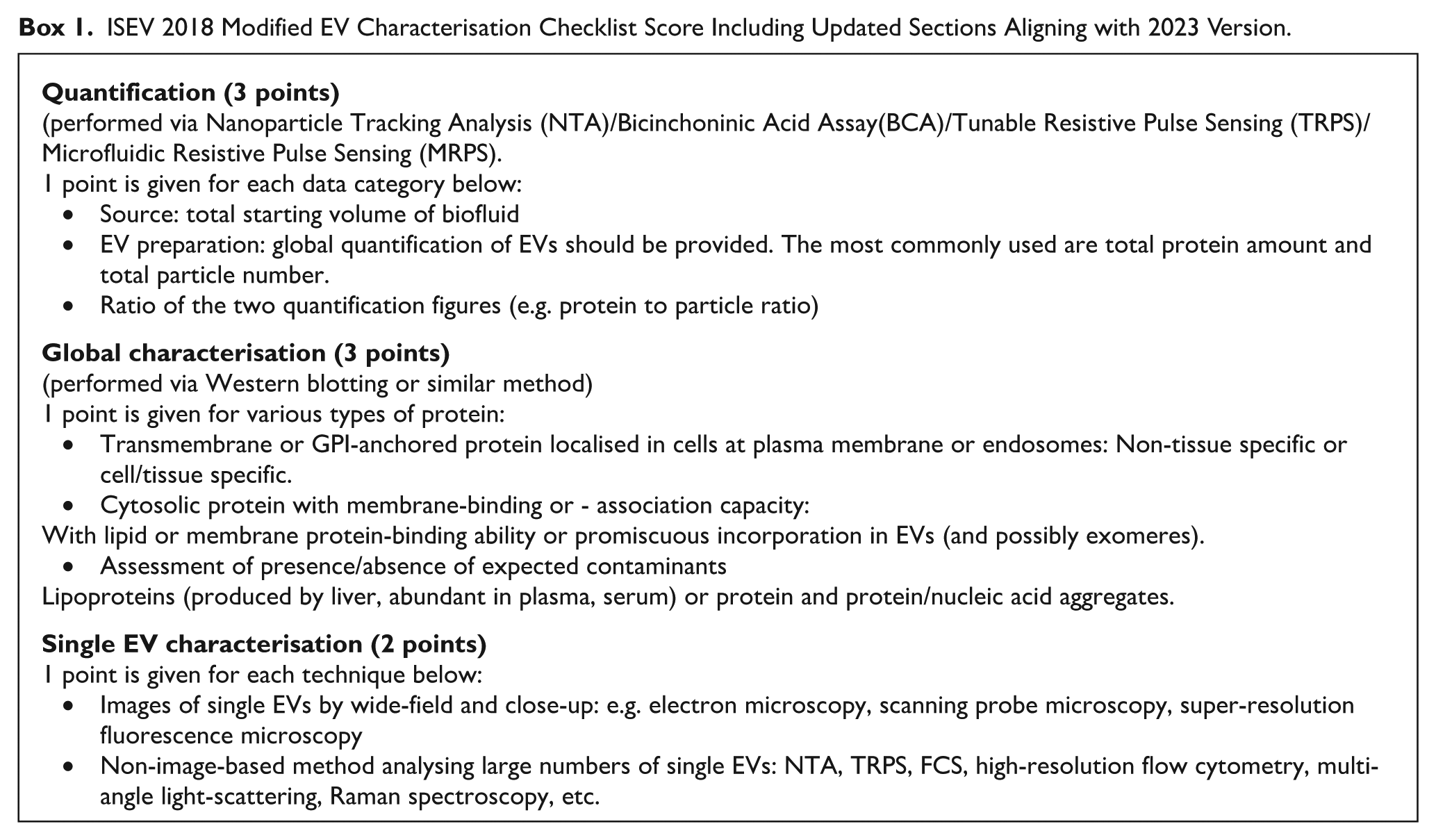

The analysis of the findings from the included manuscripts was conducted, focussing on grouping studies assessing the most commonly reported miRNAs and examining similarities and differences in adherence to MISEV criteria. Throughout the analysis, scores were categorised as low (0-2), moderate (3-4) or strong adherence (5-8). These thresholds reflect the extent to which studies incorporated essential EV characterisation steps, with scores of 5 or above indicating adherence to at least 2 of the 3 major validation criteria (quantification, global characterisation and single EV characterisation). This allowed for thorough evaluation of the robustness of current data exploring diagnostic potential of various miRNA biomarkers in PDAC.
Risk of bias was also conducted following the Quality Assessment of Diagnostic Accuracy Studies (QUADAS-2) tool for evaluating diagnostic studies. 17 The domains of patient selection, index testing and reference standard reporting were assessed, however the flow and timing domain was excluded due to lack of relevance to these studies.
Results
A PRISMA flow diagram demonstrating the search strategy and study selection criteria that were applied according to 18 is shown in Figure 1.
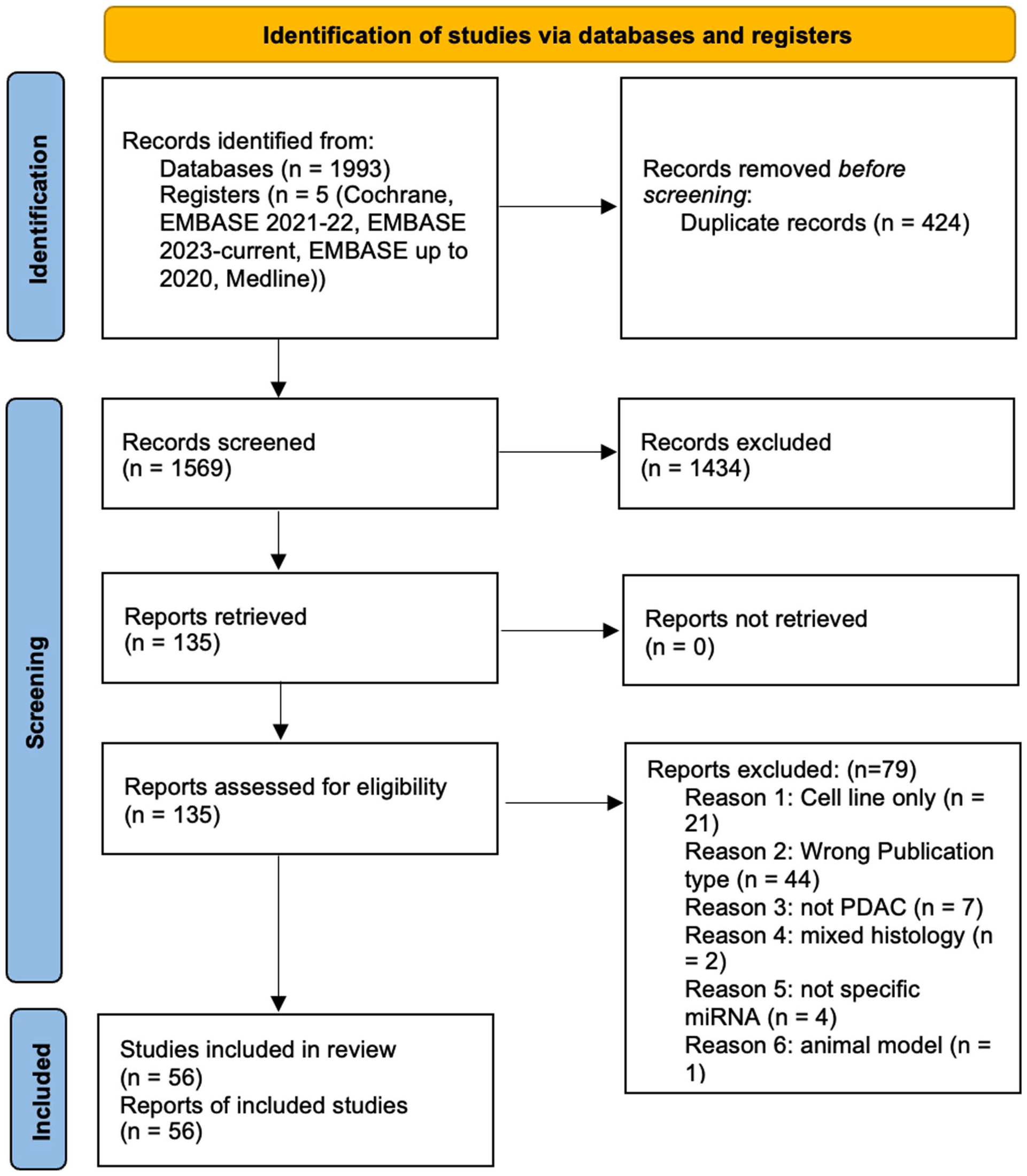
PRISMA flow diagram of the search strategy and study selection.
The key methodological characteristics of the incorporated studies, including biofluid type, EV isolation and miRNA quantification methods, control group composition and PDAC staging are summarised in Table 1. It is evident that plasma and serum were the most commonly analysed biofluids, with a smaller number of studies utilising pancreatic juice or urine. A range of EV isolation methods was employed, including ultracentrifugation, polymer-based precipitation kits and size-exclusion chromatography. miRNA extraction was performed using various commercial RNA isolation kits, and RT-qPCR was the most frequently used platform for miRNA quantification. Fewer studies employed microarray or next-generation sequencing approaches. Reporting of input biofluid volumes, elution conditions, and RNA quality control measures varied considerably. Control groups consisted of healthy individuals, patients with benign pancreatic conditions, or mixed non-cancer cohorts.
Study Characteristics of Selected 56 Manuscripts.
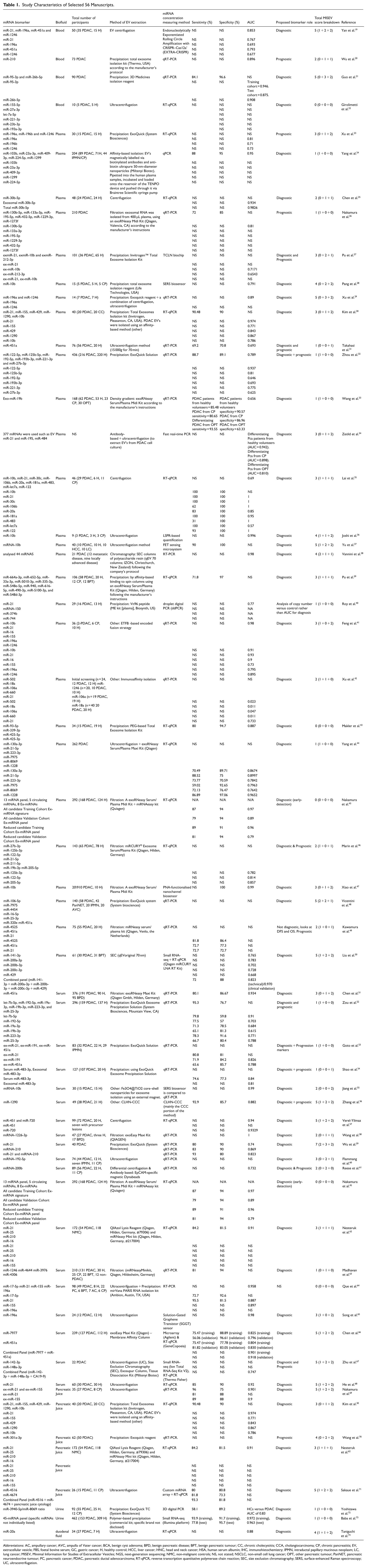
Abbreviations: AC, ampullary cancer; AVC, ampulla of Vater cancer; BCA, benign cyst adenoma; BPD, benign pancreatic disease; BPT, benign pancreatic tumour; CC, chronic cholecystitis; CCA, cholangiocarcinoma; CP, chronic pancreatitis; EV, extracellular vesicle; FBS, foetal bovine serum; GC, gastric cancer; H, healthy control; HCC, liver cancer; HNC, head and neck cancer; HSA, human serum albumin; IHC, immunohistochemistry; IPMN, intraductal papillary mucinous neoplasm; LC, lung cancer; MISEV, Minimal Information for Studies of Extracellular Vesicles; NGS, next-generation sequencing; NMC, non-malignant controls; NS, not stated; NSCLC, non-small cell lung cancer; OPT, other pancreatic tumour; PanNET, pancreatic neuroendocrine tumour; PC, pancreatic cancer; PDAC, pancreatic ductal adenocarcinoma; RT-qPCR, reverse transcription quantitative polymerase chain reaction; SEC, size exclusion chromatography; SERS, surface-enhanced Raman spectroscopy; UC, ultracentrifugation.
From the papers included in this review, the diagnostic potential of 3 miRNAs appeared to be investigated most commonly than: miR-21, miR-10b and miR-451a, with 13, 9 and 7 papers, respectively. The SN, SP and AUC for these 3 miRNAs from each of their respective papers are tabulated in Table 2. Scores representing each study’s adherence to a modified checklist reflecting the set criteria (Box 1) as well as a breakdown of these scores are also shown.
Diagnostic Performance and EV Characterisation Scores of Studies Evaluating miR-21, miR-10b, and miR-451a as Biomarkers for Pancreatic Cancer.

Abbreviation: NS, not stated.
Across the 13 studies investigating
For
Studies assessing
Additional miRNAs, and in particular several miRNA panels, have demonstrated greater diagnostic potential than 3 biomarkers described above, as reflected in their higher AUC values. This is illustrated in Table 3.
Diagnostic Performance and EV Characterisation Scores of Representative Studies Reporting Multi-miRNA Panels.

Abbreviation: NS, not stated.
Biomarker panels in general reported higher AUC values than individual miRNAs, with Nesteruk et al.,
62
Yang et al.,
24
Madhavan et al.,
63
Makler et al.
43
and Baba et al.
73
all reporting AUCs exceeding 0.85. However, these panels consistently exhibited poor adherence to MISEV guidelines, with most scoring only 1 out of 8, primarily due to inadequate global and single-EV characterisation. Therefore, despite demonstrating promising diagnostic potential, the systematic omission of critical EV characterisation steps raises concerns about the reliability of reported diagnostic performance. Among individual miRNAs,
The risk of bias assessment using the QUADAS-2 tool 17 revealed that most studies exhibited methodological concerns. A large proportion of studies were judged to have an unclear or high risk of bias in the domains of patient selection and index test, primarily due to incomplete reporting of inclusion criteria, sample handling and absence of blinding during sample analysis. 54/56 included papers lacked blinding and 37/56 studies did not incorporate independent validation cohorts, increasing the likelihood of overfitting and inflating diagnostic accuracy estimates. The reference standard domain was inconsistently addressed, with several studies failing to specify how PDAC diagnosis was confirmed. Flow and timing were generally adequately reported, although some studies lacked details regarding the timing of biomarker assessment relative to diagnosis. The breakdown of risk of bias assessments across all studies is summarised in Supplemental Figure 1. A new quality assessment checklist summarising recommended methodological standards for future EV-miRNA biomarker studies is provided in Supplemental Table 2.
Discussion
This systematic review demonstrates that despite growing interest in EV-derived miRNAs as non-invasive biomarkers for PDAC, substantial methodological variability limits the interpretability and clinical utility of published findings. A prominent limitation lies in the inconsistent adherence to the MISEV guidelines, which recommend comprehensive characterisation of EV preparations. Majority of included studies failed to report global EV quantification using established metrics such as total protein or particle count, omitted key transmembrane (CD63, CD81, CD9) and cytosolic (TSG101, Alix) protein markers, and neglected single-EV characterisation altogether (Box 1). Such omissions raise the risk of non-EV contaminants like protein aggregates or lipoproteins confounding downstream analyses.75 -77 This issue was particularly evident in studies investigating emerging miRNA candidates or miRNA panels, where reported diagnostic performance often exceeded AUC values of 0.90 despite minimal EV validation, which can result in misleading findings from contamination or technical errors.
Nonetheless, a subset of studies, particularly those evaluating miR-21and miR-451a did incorporate more rigorous methodological practices, including Western blotting for multiple EV markers and imaging via transmission electron microscopy. These studies tended to produce more reproducible results, highlighting the value of methodological robustness. For example, the most consistent findings for miR-21 came from studies scoring at least 5 out of 8 on the MISEV criteria, incorporating both global quantification and protein-based EV validation. However, the fact that miR-21 studies still exhibited wide variation in reported AUC values, ranging from <0.70 to >0.90 even among high-adherence groups, underscores the impact of other confounding factors, such as biofluid selection, normalisation strategies, and population heterogeneity. Most studies included patients with PDAC across all stages; with a small subset with restricted inclusion of early-stage (I–IIA/B). This variability may have influenced reported diagnostic performance and limited the ability to directly compare findings across studies. Similar heterogeneity was equally apparent in the diagnostic evaluation of miR-10b and miR-451a. For miR-10b, studies predominantly demonstrated moderate methodological adherence, often including some EV validation steps (eg, Western blotting or electron microscopy) without integrating quantification metrics or assessing contamination. These findings suggest that even promising biomarkers may yield misleading results if methodological standards are not uniformly applied.
Our findings collectively underscore an urgent need for standardisation across pre-analytical workflows, EV isolation methods, and miRNA quantification protocols. Harmonisation of these processes would enable more reliable cross-study comparisons and minimise the risk of misleading results arising from technical variability. To support this, journals and funding bodies should mandate compliance with minimum EV characterisation criteria, as outlined in the MISEV guidelines, to ensure methodological transparency and reproducibility.
Beyond EV validation, analytical variability introduced further uncertainty. RNA isolation methods, qPCR platforms and miRNA normalisation strategies were different and were rarely standardised or justified. Additionally, many studies failed to specify the total input volume of biofluid, a detail essential for contextualising miRNA concentration. Together, these factors introduce pre-analytical and analytical bias, which can distort comparisons across studies and compromise biomarker reliability.
Arguably the most critical study design limitation identified in this review was the widespread absence of external validation cohorts. Nearly 70% of included studies relied exclusively on discovery-phase data, with no attempt to assess the miRNAs in independent study populations. This introduces a significant risk of overfitting, whereby predictive performance appears inflated due to tailoring of results to the initial cohort. This limitation was particularly pronounced among studies reporting high AUC values, especially in those investigating miRNA panels, where model complexity increased but methodological transparency decreased. Thus, without proper separation of discovery and validation phases, these findings cannot be generalised.
The QUADAS-2 risk of bias assessment further substantiated these concerns. Nearly all studies lacked blinding during sample processing or analysis, introducing observer bias. Moreover, reference standards for confirming PDAC were often poorly defined or inconsistently applied, with some studies failing to specify whether diagnoses were histologically confirmed. Although the flow and timing of patient inclusion were generally well reported, the cumulative impact of these issues, especially when coupled with the absence of validation cohorts, undermines confidence in reported diagnostic accuracies.
The above-mentioned challenges reflect broader translational barriers that extend beyond EV-miRNA studies. As Peters et al. observed, fewer than 2% of candidate biomarkers in oncology ultimately reach clinical implementation. 78 This failure is frequently attributed to the same issues identified here: inadequate methodological rigour, lack of reproducibility and insufficient multi-cohort validation. The Biomarker Toolkit 78 has been proposed as a structured framework for addressing these gaps, offering a stepwise approach to evaluate biomarker readiness, from analytical validity to clinical utility. Incorporating such tools alongside standardised reporting frameworks like MISEV and QUADAS-2 could significantly improve both the design and interpretability of future studies.
Conclusion
In summary, EV-derived miRNA biomarkers for PDAC offer significant diagnostic potential, particularly given their non-invasive nature, stability and biological relevance. However, current evidence base is undermined by methodological inconsistency, poor adherence to EV characterisation standards, and a pervasive lack of validation cohorts. These shortcomings have led to inflated diagnostic performance metrics and limited confidence in study findings. To advance the field, future research must prioritise methodological rigour through robust EV validation, use of independent multi-cohort designs and adoption of biomarker readiness frameworks. Only by addressing these limitations can EV miRNA biomarkers be credibly positioned for clinical translation for PDAC detection.
Supplemental Material
sj-docx-1-bmi-10.1177_11772719251381960 – Supplemental material for Extracellular Vesicle-Derived miRNAs as Diagnostic Biomarkers for Pancreatic Ductal Adenocarcinoma: A Systematic Review of Methodological Rigour and Clinical Applicability
Supplemental material, sj-docx-1-bmi-10.1177_11772719251381960 for Extracellular Vesicle-Derived miRNAs as Diagnostic Biomarkers for Pancreatic Ductal Adenocarcinoma: A Systematic Review of Methodological Rigour and Clinical Applicability by Ryhan Divyang Patel, Bhavik Patel and Tatjana Crnogorac-Jurcevic in Biomarker Insights
Footnotes
Acknowledgements
Rebecca Jones, Liaison Librarian, Imperial College London.
Abbreviations
AC: ampullary cancer; AUC: area under the (ROC) curve; BCA: bicinchoninic acid (assay); BCA (benign): benign cyst adenoma; BPD: benign pancreatic disease; BPT: benign pancreatic tumour; CA19-9: carbohydrate antigen 19-9; CCA: cholangiocarcinoma; cfDNA: circulating cell-free DNA; CP: chronic pancreatitis; CRISPR-Cas12a: clustered regularly interspaced short palindromic repeats–CRISPR associated protein 12a; EV: extracellular vesicle; FBS: foetal bovine serum; FCS: fluorescence correlation spectroscopy; GC: gastric cancer; H: healthy control; HCC: hepatocellular carcinoma (liver cancer); HNC: head and neck cancer; HAS: human serum albumin; IHC: immunohistochemistry; IPMN: intraductal papillary mucinous neoplasm; LC: lung cancer; miRNA: microRNA; MISEV: Minimal Information for Studies of Extracellular Vesicles; mRNA: messenger RNA; MRPS: microfluidic resistive pulse sensing; NGS: next-generation sequencing; NMC: non-malignant controls; NTA: nanoparticle tracking analysis; OPT: other pancreatic tumour; PanNET: pancreatic neuroendocrine tumour; PC: pancreatic cancer; PDAC: pancreatic ductal adenocarcinoma; PPV: positive predictive value; PROSPERO: International Prospective Register of Systematic Reviews; PRISMA: Preferred Reporting Items for Systematic Reviews and Meta-Analyses; QUADAS-2: Quality Assessment of Diagnostic Accuracy Studies-2; ROC: receiver operating characteristic; RT-qPCR: reverse transcription quantitative polymerase chain reaction; SEC: size-exclusion chromatography; SERS: surface-enhanced Raman spectroscopy; SN: sensitivity; SP: specificity; TEM: transmission electron microscopy; TRPS: tunable resistive pulse sensing; UC: ultracentrifugation.
Ethical Considerations
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Author Contributions
RDP – Conceptualisation, searching, methodology, writeup – original draft, writeup – revisions; BP – Conceptualisation, searching, writeup – original draft; TCJ – Writeup – revisions, supervision.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
AI Statement
The authors confirm that no artificial intelligence (AI) tools were used for language refinement, content generation, or modification of scientific data in this manuscript.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
