Abstract
Traumatic brain injury (TBI) is not a single disease state but describes an array of conditions associated with insult or injury to the brain. While some individuals with TBI recover within a few days or months, others present with persistent symptoms that can cause disability, neuropsychological trauma, and even death. Understanding, diagnosing, and treating TBI is extremely complex for many reasons, including the variable biomechanics of head impact, differences in severity and location of injury, and individual patient characteristics. Because of these confounding factors, the development of reliable diagnostics and targeted treatments for brain injury remains elusive. We argue that the development of effective diagnostic and therapeutic strategies for TBI requires a deep understanding of human neurophysiology at the molecular level and that the framework of multiomics may provide some effective solutions for the diagnosis and treatment of this challenging condition. To this end, we present here a comprehensive review of TBI biomarker candidates from across the multiomic disciplines and compare them with known signatures associated with other neuropsychological conditions, including Alzheimer’s disease and Parkinson’s disease. We believe that this integrated view will facilitate a deeper understanding of the pathophysiology of TBI and its potential links to other neurological diseases.
Introduction
Traumatic brain injury (TBI) is a broad descriptor for many injuries involving physical harm to the brain. TBI can result from any blunt or penetrating force to the head. 1 Millions of people suffer a TBI each year. 2 TBI is the leading cause of death for persons under age 45, 1 and according to the Centers for Disease Control and Prevention, there were over 60 000 TBI-related deaths in the United States alone in 2019. 3 During the recent wars in Iraq and Afghanistan, the incidence rate of TBI among American warfighters was nearly 20%, with the majority of those cases being concussions (mild TBIs).4,5
TBIs present with a high degree of heterogeneity due to the large variation in TBI-causing insults and injuries, potential genetic predispositions, and other demographic factors, including homelessness and racial/ethnic minority status. 6 Immediate symptoms of TBI, sometimes referred to as post-concussion symptoms (PCSs), commonly include headache, inflammation, confusion, dizziness, and lightheadedness.7-11 TBIs can produce symptoms that appear and persist over longer and unpredictable timescales as well, such as lasting cognitive impairment, loss or disruption of motor function, and behavioral changes.12-14
TBI is likely related to (if not a causative agent of) several other neurological diseases and psychiatric disorders, including Alzheimer’s disease (AD),15-20 Parkinson’s disease (PD),21-24 post-traumatic stress disorder (PTSD),25-27 epilepsy,28,29 chronic traumatic encephalopathy (CTE),30,31 and dementia.32,33 Furthermore, TBIs can have deep and lasting consequences on interpersonal relationships, meaning that TBIs affect many more people than those who experience a TBI themselves. 34 Given its prevalence and severity, TBI is an important public health concern and warrants further study.
It is difficult to estimate the global TBI incidence due to a general scarcity of information, especially from low- and middle-income countries.35,36 TBIs tend to be under-reported (referred to as the “silent epidemic”) in official statistics because many individuals may experience a TBI but not seek medical treatment. 37 Obtaining a substantive count of TBI cases is dependent on accurate diagnostic methods. However, current diagnostics for neurological diseases as a whole are severely limited and usually indirect. 38
Definitive diagnostics for neurological diseases are rare. For example, AD is only definitively diagnosed post-mortem, where an autopsy can reveal the presence or absence of the amyloid-β plaques and tau tangles characteristic of the disease.39,40 Other neurological diseases like PD often rely on differential diagnosis, or the elimination of other possible conditions that can manifest with similar symptoms. 41 As an initial screen for TBI, physicians employ cognitive tests, such as Glasgow Coma Scale (GCS) assessment, the Mini Mental State Examination (MMSE), the Montreal Cognitive Assessment (MoCA), and the Rivermead PCSs Questionnaire (RPQ).11,42-45 Evaluation according to the GCS allows for a standardized assessment of impaired consciousness, as quantified by an individual’s motor, verbal, and eye-opening responses.42,43 GCS scores can be used to grade TBIs as mild, moderate, or severe depending on the observed degree of impaired consciousness, as judged by the test administrator. 46 The U.S. Department of Defense utilizes the duration of post-traumatic amnesia and loss of consciousness with a patient’s GCS score to determine TBI severity. 47 The MMSE and MoCA were designed to be sensitive to even mild cognitive impairment, testing mental faculties such as speech, vision, situational awareness, and recall.44,45 In contrast, the RPQ was designed to measure the severity of various PCSs in affected individuals, including headaches, dizziness, nausea, and poor concentration. 11 Compound tests like the Sport Concussion Assessment Tool 3 combine multiple cognitive tests into a single composite score with the goal of providing a multifaceted assessment within a single test. 48
These cognitive tests are commonly utilized in modern medical practice and research because they are quick, standardized, inexpensive, noninvasive, and easy to administer. Imaging modalities such as computed tomography (CT), positron emission tomography (PET), single-photon emission computed tomography (SPECT), and magnetic resonance imaging can detect some TBIs, such as those that result in skull fractures or intracranial hematomas. However, not all TBIs result in sequelae that can be imaged with these techniques, and exposing patients to ionizing radiation (as with CT, PET, and SPECT) raises other health concerns. 49 The limitations of these diagnostics can be especially poignant for individuals who present with delayed onset of symptoms or PTSD. 50 Furthermore, these clinical diagnostics provide little molecular-level insight into the biochemical changes within an affected individual—changes that are ultimately responsible for the observed effects of a TBI. 51
The myriad physical and biochemical changes that follow a TBI can be classified into primary and secondary phases by the amount of time that lapses before they manifest (Figure 1). The primary injury phase describes changes that occur at the time of the insult. Contusion occurs by a mechanism known as coup-contrecoup; namely, bruising can occur at or near the site of the insult (coup) and on the opposite side of the brain following recoil (contrecoup). 52 These impacts can cause hemorrhaging, tissue damage, and tissue deformation. Cells of the central nervous system (CNS) may be stretched and torn by shear stresses, thereby leaking highly CNS-specific compounds into the cerebrospinal fluid (CSF). 53 Additionally, the blood-brain barrier (BBB) can be disrupted during the primary injury phase, leading to mixing of CSF and blood. This crossover may further lead to the generation of CNS protein-targeting autoantibodies that, upon subsequent injury, may enter the CNS and damage nerve cells.54-56

Representation of the pathophysiological manifestations associated with primary (left) and secondary (right) phases of TBI. The primary phase is characterized by immediate disruption of the BBB, axonal shearing, and compaction that causes necrosis. The secondary phase is characterized by symptoms that can take hours or days to manifest, including edema and cardiac complications.
The secondary injury phase describes a variety of changes that occur in the hours and days following a TBI. If the injury results in thrombosis, cerebral ischemia may develop. 57 Cerebral ischemia is hypothesized to increase the concentration of electron transport free radicals in the mitochondria, promoting pathologic reactions with lipids in the inner and outer mitochondrial membranes. 58 Glutamate released from ruptured nerve cells into the surrounding tissues can cause excitotoxicity, thereby leading to a variety of harmful effects, including the release of mitochondrial reactive oxygen species and neuron death.59,60 Cerebral edema, resulting from both cytotoxic and vasogenic edema mechanisms, is observed in secondary TBI because of increased solute uptake following neuronal rupture and BBB disruption.61,62 Upregulated pro- and anti-inflammatory cytokines mediate an inflammatory cascade in response to stress. 63 Moreover, TBI can lead to severe systemic complications, further establishing the importance of understanding TBI and its associated pathophysiology.64,65
Although the primary injury phase of TBI is untreatable, secondary injury occurs on a timescale that allows for mitigation or prevention of secondary symptoms and other TBI-related complications. As examples, hypothermia has been explored as a neuroprotectant for TBI, 66 and α-lipoic acid has been demonstrated to inhibit oxidative stress-related apoptosis through mitochondrial signaling.67-69 The specific biochemical changes that occur in the primary and secondary injury phases of TBI present the opportunity for diagnostic development. For instance, measuring elevated concentrations of highly neuron-specific compounds in serum or highly blood-specific compounds in CSF can be used to deduce a disruption of the BBB. 56 These molecular-level changes form the basis of biomarker-based diagnostics.
Biomarkers, defined as “characteristic features that are objectively measured and evaluated as indicators of biological processes,” have the potential to complement, or eventually supplant, current diagnostics for TBI and other neurological diseases. 70 Biomarker candidates can be found in every branch of multiomics, a broad term encompassing metabolomics, proteomics, transcriptomics, epigenomics, and genomics (Figure 2). Biomarkers may be exogenous, as in the case of toxins secreted by invading pathogens, or endogenous with differential expression, as in the case of TBI. 71 Ideally, biomarkers enable direct quantitation of the body’s biochemical response to an injury or disease. Because standardized biomarker-based diagnostics do not depend on human judgment nearly as much as a cognitive interview, they reduce bias in diagnosis, allowing for better sensitivity and specificity.72,73 Beyond diagnostics, biomarkers hold the potential to predict disease severity and treatment response.74,75

Schematic representation of multiomics, encompassing metabolomics, proteomics, transcriptomics, epigenomics, and genomics, as a unique tiered set of tools to understand TBI. The multiomic disciplines are organized in order of temporal response to a biochemical change from metabolomics (where the most immediate changes are observed) to genomics. Each branch of multiomics investigates different aspects of the central dogma of biology, providing an organized framework to examine the broad biochemical changes that occur in the pathophysiology of neurodegenerative diseases. A holistic view of the multiomic changes that occur in TBI can provide insight the potential links between TBI and other neurological diseases.
Despite their significant potential, biomarkers are not clinical standards for the diagnosis of neurological diseases for several reasons. The field of multiomic biomarker discovery, diagnostic assay development, and data integration for neurological diseases is still relatively young, and most reports of candidate biomarkers require further validation. 56 Validation of biomarker-based diagnostics is intrinsically limited by the associated challenges of current diagnostics described previously. Once a candidate biomarker has been identified, it must be further screened in increasingly large and diverse samples to determine the specific conditions and populations within which the biomarker can be used effectively. Biomarker profiles generally exhibit a temporal dependence, due in part to the biofluid stability of the markers of interest, and sample collection sites can vary for different biomarkers of interest. 56 Additionally, these temporal profiles vary between the different parts of the multiome; although a metabolomic profile can change on the order of minutes to hours, the remainder of the multiome exhibits increasingly delayed responses to external stimuli (the proteome and transcriptome respond slower than the metabolome, the epigenome responds slower than the proteome and transcriptome, and the genome is the most resilient), further complicating cross-comparability and generalizability. 76 Moreover, establishing thresholds based on a patient’s individual history, concerns of over-fitting with biomarker panels, and other statistical concerns further complicate the development and validation of a biomarker assay, to say nothing of the difficulty of crossing the “valley of death” from laboratory development to becoming a clinical standard.77-79
For these reasons and more, although many candidate biomarkers have been identified for TBI, none have been broadly implemented across the medical field for TBI diagnosis. TBI candidate biomarkers have already been reviewed with a focus on their potential clinical applications.56,79-89 However, an equally valuable and much more fundamental application of multiomic biomarker research is gaining insight into the underlying pathophysiology of TBI.
90
With a more complete understanding of the biochemical changes experienced by TBI-affected individuals, diagnostic and therapeutic targets will emerge naturally, as will the potential links between TBI and other neurological diseases (Figure 2).
Metabolomic Biomarkers
Metabolomics involves the study of small molecules involved in metabolism, including lipids (lipidomics being a branch of metabolomics), amino acids, sugars, and more. The metabolome is downstream of the rest of the multiome, meaning that changes are amplified in the metabolome to the largest extent.91,92 The metabolome is the component of the multiome that responds most rapidly to the external environment/stimuli, making it an advantageous target for rapid diagnostics. 93 Common analytical tools in metabolomic biomarker discovery include gas chromatography (GC) and liquid chromatography (LC) for metabolite separation and nuclear magnetic resonance (NMR) spectroscopy and mass spectrometry (MS) for metabolite identification. 94 Hyphenated and tandem analytical methods such as GC-MS, LC-NMR-MS, and MS/MS are popular choices for their throughput and sensitivity.95,96 Many multivariate statistical techniques, such as principal component analysis (PCA), cluster analysis, and logistic regression, are commonly employed in metabolomic data analysis.96,97 These statistical methods serve 2 main purposes: (1) data validation for outliers or misidentified metabolites and (2) identifying trends between sample types, providing meaning to what otherwise would be a list of chemicals.
Metabolomic biomarkers are currently being widely explored for TBI.89,98-107 Because no single metabolite has yet been identified with sufficient diagnostic power, most reports focus on metabolite panels that explain a significant amount of the variance observed between TBI patients/models (typically controlled cortical impact, fluid percussion, and weight-drop models) and controls (typically healthy individuals or “sham” craniotomy models). 108 A selection of metabolites reported with altered levels in TBI is shown in Table 1.
A selection of metabolomic biomarker candidates for TBI.
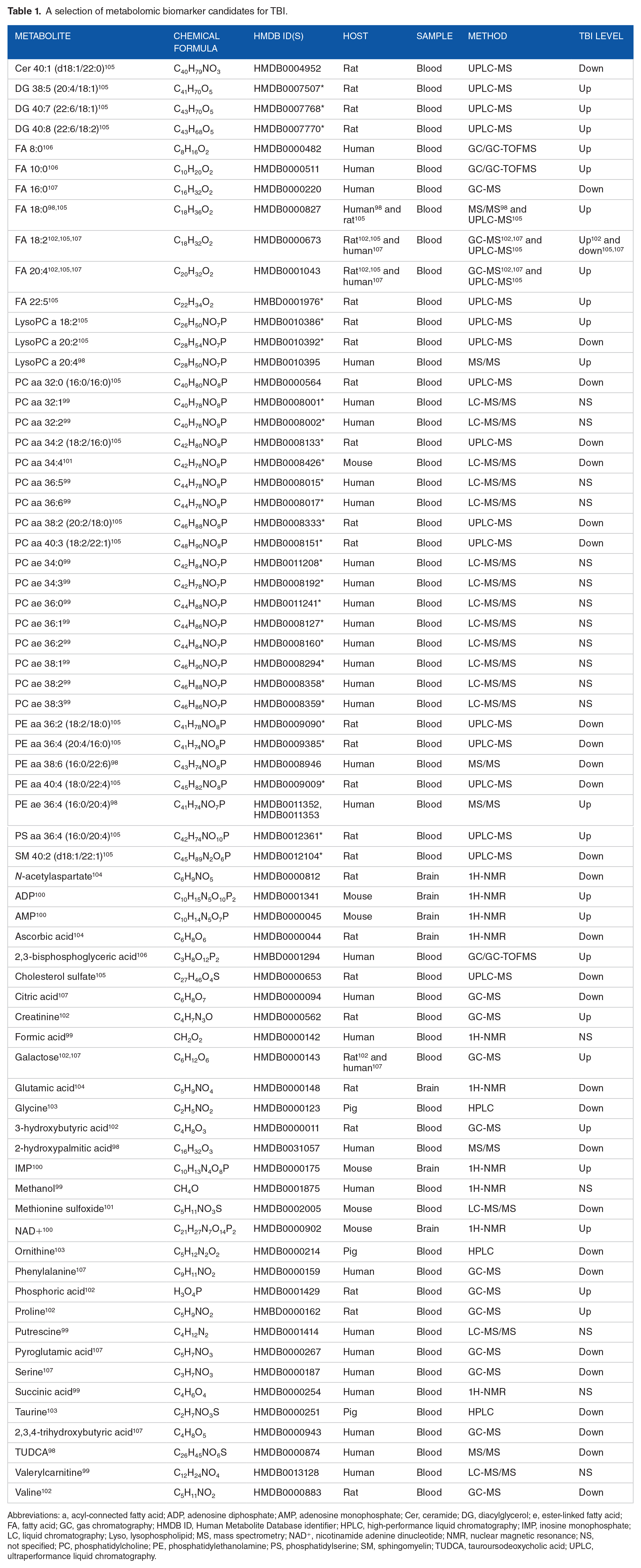
Abbreviations: a, acyl-connected fatty acid; ADP, adenosine diphosphate; AMP, adenosine monophosphate; Cer, ceramide; DG, diacylglycerol; e, ester-linked fatty acid; FA, fatty acid; GC, gas chromatography; HMDB ID, Human Metabolite Database identifier; HPLC, high-performance liquid chromatography; IMP, inosine monophosphate; LC, liquid chromatography; Lyso, lysophospholipid; MS, mass spectrometry; NAD+, nicotinamide adenine dinucleotide; NMR, nuclear magnetic resonance; NS, not specified; PC, phosphatidylcholine; PE, phosphatidylethanolamine; PS, phosphatidylserine; SM, sphingomyelin; TUDCA, tauroursodeoxycholic acid; UPLC, ultraperformance liquid chromatography.
Metabolites with lipid tails are notated as (number of carbons):(number of carbon-carbon double bonds). For metabolites with multiple assigned lipid tails, the tails are additionally noted in parentheses and separated by a slash.
The authors furnished plausible HMDB IDs for otherwise unidentified metabolites. These assignments are tentative and only for illustrative purposes.
Among the 69 unique metabolites listed in Table 1, only 4—galactose (Human Metabolite Database identifier: HMDB0000143), stearic acid (HMDB0000827), linoleic acid (HMDB0000673), and arachidonic acid (HMDB0001043)—were used in predictive panels in more than one study, possibly due to limited libraries and differences in sample types, sample processing methods, and data acquisition methods.98,102,105,107 Some metabolites measured with altered post-TBI levels remain unidentified, and relatively small sample sizes with large metabolite panels raise concerns of over-fitting. Despite these challenging limitations, there is good agreement in the general trends observed across these reports, such as a general decrease in serum phosphatidylcholines, lending validity to the methods used to gather these data. 105
The metabolomic biomarkers listed in Table 1 can provide insights into the biochemical changes that occur in TBI. For example, the decrease of tauroursodeoxycholic acid (TUDCA), a molecule associated with prevention of apoptosis and protection of mitochondria, helps to explain the cell death and mitochondrial damage observed in TBI at the molecular level. 98 Additionally, methionine sulfoxide, a marker for oxidative stress, was observed to increase over time following a TBI, rationalizing some of the secondary injury symptoms discussed previously.101,109 Viewed collectively, the candidate biomarkers listed in Table 1 indicate that amino acid metabolism (pathways 00250, 00260, 00270, 00280, 00330, 00360, and 00430 in the Kyoto Encyclopedia of Genes and Genomes, KEGG), lipid metabolism (KEGG: 00061, 00062, 00071, 00100, 00561, 00564, 00565, 00590, 00591, 00600, and 01040), carbohydrate metabolism (KEGG: 00010, 00052, and 00053), and the citric acid cycle (KEGG: 00020) may be disrupted, suggesting that broad dysregulation of metabolism occurs in TBI.110-112 Although broad dysregulation does not provide specific insight into linkages between TBI and other neurological diseases, preliminary metabolomics profiling suggests a linkage between TBI and PD through glutamate-mediated excitotoxicity. 23 Ultimately, the metabolome is a rich source of information on physiological alterations. As the field of TBI metabolomics develops, future profiling efforts will continue to advance our understanding of TBI and its relation to other diseases.
Proteomic Biomarkers
Proteomics involves the study of the proteins that are translated or modified by an organism. Analyzing the proteome ideally allows for the identification and quantitation of every protein expressed in a cell, giving deep insights into pathway regulation and therapeutic targets. 113 Proteomic approaches have largely dominated multiomics research over the past few decades, with MS being the most commonly employed analytical technique. 114 MS/MS proteomic approaches can be broadly classified as top-down or bottom-up. Bottom-up proteomics involves early fragmentation or digestion into unique peptides, relying on mass fingerprinting or dissociation information to identify the fragments of the initial protein. 115 Conversely, top-down proteomics involves “soft” MS of an intact protein followed by “hard” MS to measure fragments of the protein, allowing for detection of post-translational modifications and site-specific mutations at the cost of primary sequence information. 116 Among MS approaches, electrospray ionization (ESI) and matrix-assisted laser desorption/ionization (MALDI) are popular as “soft” ionization methods with minimal fragmentation. 117 In addition to chromatographic techniques [high-performance liquid chromatography, ultraperformance liquid chromatography, and hydrophobic interaction liquid chromatography], electrophoretic methods [gel electrophoresis, two-dimensional gel electrophoresis, and capillary electrophoresis (CE)] are broadly employed for proteomic separations.118,119 Similar to metabolomic analyses, hyphenated analytical methods like LC-MS and CE-MS are commonly employed in proteomic analyses for their throughput and sensitivity. 120 In contrast to the predictive panels commonly found in metabolomic data analysis, the field of proteomics generally focuses at the single-protein level to identify and quantify altered protein expression levels. 121
In TBI, many candidate protein biomarkers have been independently studied and extensively reviewed, and the mechanisms that give these markers meaning are generally well understood.56,80,81,86 Therefore, we focus this section of our review on examples of leading proteomic biomarker candidates whose functions and identities provide insights into the biochemical changes that occur following a TBI. A selection of leading protein biomarker candidates for TBI that have been identified with elevated levels in humans is shown in Table 2.
A selection of leading proteomic biomarker candidates for TBI with elevated levels in humans.

Abbreviations: ApoE, apolipoprotein E; APP, amyloid precursor protein; CKBB, brain-type creatine kinase; CSF, cerebrospinal fluid; GFAP, glial fibrillary acidic protein; IL, interleukin; MBP, myelin basic protein; MMP, matrix metalloproteinase; NF, neurofilament; NSE, neuron-specific enolase; S100B, S100 calcium-binding protein β; SBDP, spectrin breakdown product; UCHL1, ubiquitin carboxyl-terminal hydrolase L1.
The majority of the candidate protein biomarkers shown in Table 2 are highly CNS-specific proteins detected in serum at elevated levels following moderate to severe TBI. These proteins are known as peripheral markers of TBI because they indicate (1) damage of CNS cells and (2) disruption of the BBB. 56 Interleukin (IL)-10, an anti-inflammatory cytokine, was included as a representative example of a non-peripheral marker of TBI, as its upregulation in CSF is thought to counter the inflammatory events that occur following TBI. 127 The most extensively studied TBI biomarker is S100 calcium-binding protein β (S100B), with well over 300 reports to date.81,138,139 Notably, serum S100B levels have been used in a clinical setting to rule out mild TBI, albeit in a limited capacity.56,134
A fundamental limitation to the utility of the proteins listed in Table 2 as diagnostic indicators is that none of these are exclusively CNS-specific. For example, outside of the CNS, neuron-specific enolase is expressed in erythrocytes and endocrine cells, glial fibrillary acidic protein is expressed in Leydig cells, and S100B is expressed in adipocytes, chondrocytes, and Langerhans cells.56,140,141 Therefore, rather than qualitative measurement, robust diagnostics must be capable of detecting elevated levels of these proteins in serum following TBI. 142 Circulating protein levels vary from person to person, necessitating a documented individual history to confidently measure increases in biofluid protein levels and further complicating the potential of using proteomic diagnostics in clinical settings. Furthermore, many of these proteins have been observed with elevated levels following injuries that are not TBIs, raising concerns about the specificity of individual markers and prompting the use of biomarker panels. 143
Candidate proteomic biomarkers further shed light on the connections between TBI and other neurological diseases. For example, tau protein aggregation following TBI provides a molecular basis for TBI as a risk factor for AD.135,144 Additionally, cerebral ischemia has been linked to tau acetylation, a modification that can lead to the development of AD-type neuropathology. 145 Moreover, loss of ubiquitin carboxyl-terminal hydrolase L1 in knockout models was recently observed to rescue PD-related defects through suppression of glycolysis. 146 In addition to biochemically linking one of the leading TBI biomarkers to PD, this finding demonstrates that a multiomic understanding of the pathophysiology of TBI is important for investigating how TBI relates to other neurological diseases.
It is worth noting that, though not commonly considered biomarkers of TBI, alterations of expression levels and localization of proteins linked to other neurological diseases have been observed after TBI. For instance, accumulation of α-synuclein following chronic TBI establishes an additional proteomic link to PD. 21 Despite our ability to identify alterations in protein levels that correlate between TBI and other conditions, we still do not fully understand the mechanisms by which these conditions are causally related, if at all. As an example, although amyloid precursor protein (APP) and amyloid-β accumulation have been observed following TBI, the amyloid-β plaques characteristic of AD were not observed by Chen et al.18,147,148 Further complications arise from the consideration that many of these candidate proteomic biomarkers, including amyloid-β and tau, have multiple forms, and the relative ratios of those forms can convey other meaning. 149 These findings suggest that the mechanisms governing the connections between TBI and other neurological diseases may be quite complex, and further research across the multiomic disciplines is necessary to clarify these connections.
Transcriptomic Biomarkers
Transcriptomics is the study of all the RNA transcripts in an individual cell or family of cells, comprising both protein-coding messenger RNA (mRNA) and non-protein-coding RNAs, such as transfer RNA (tRNA), 150 ribosomal RNA (rRNA), 151 microRNA (miRNA),88,152-157 enhancer RNA (eRNA), 158 small interfering RNA (siRNA), 159 P-element induced wimpy testis-interacting RNA (piRNA), 160 small nucleolar RNA (snoRNA), extracellular RNA (exRNA), small Cajal body-specific RNA (scaRNA), and long non-coding RNA (lncRNA). 161 Epigenomics is strictly tied to the transcriptome and is often seen as a subfield of transcriptomics. However, we leave our discussion of epigenomic biomarkers to the following section of this review. Analyzing the transcriptome gives insight into regulation of cellular processes, and the transcriptome is an advantageous diagnostic target due to the relative abundance and stability of RNA in biofluids.82,162 Although early transcriptomic analyses employed hybridization-based microarrays, modern approaches tend to favor techniques like RNA sequencing (RNA-seq), real-time quantitative reverse transcription polymerase chain reaction (RT-qRT-PCR), and labeled-probe hybridization methods like NanoString nCounter for their improved versatility, consistency, and sensitivity.163-167 Modern microarrays are generally performed with a complementary technique—usually RT-qRT-PCR—to confirm the validity of the array data.168,169 As important regulators, miRNAs are key targets in transcriptomic approaches. 170 Aside from direct RNA methods, miRNAs can additionally be studied indirectly through quantitative proteomic analyses, potentially affording deeper information on the regulatory targets of the miRNAs, but these methods are challenging. 171
Similar to metabolite profiling, RNA profiling is currently being widely explored in the realm of TBI. In particular, miRNA profiling holds significant diagnostic potential in TBI, exhibiting clear dependences on TBI severity and time in response to injury.154,172,173 Although miRNA biomarkers in TBI have been extensively reviewed, recent transcriptomic studies have characterized changes in lncRNA, mRNA, and tRNA-derived small RNA (tsRNA) levels associated with brain injury.88,150,154,168,169 A selection of leading transcriptomic candidate biomarkers for TBI is shown in Table 3. 174
A selection of leading transcriptomic biomarker candidates for TBI identified in humans.

Abbreviations: ADAMTS2, a disintegrin and metalloproteinase with thrombospondin motifs 2; ALOX15, arachidonate 15-lipoxygenase; CSF, cerebrospinal fluid; HPR, haptoglobin-related protein; IFIT1, interferon-induced protein with tetratricopeptide repeats 1; miRNA, micro ribonucleic acid; NECAB1, N-terminal EF-hand calcium-binding protein 1; OLAH, oleoyl-(acyl-carrier-protein) hydrolase; OLIG2, oligodendrocyte transcription factor 2; PMR, precursor miRNA; PRSS33, serine protease 33; qRT-PCR, quantitative reverse transcription polymerase chain reaction; RNA-seq, ribonucleic acid sequencing; RT-qRT-PCR: real-time quantitative reverse transcription polymerase chain reaction; SIGLEC8, sialic acid-binding Ig-like lectin 8.
NM and NR codes are derived from the Reference Sequence database. 174
Similar to the metabolomic candidate biomarkers in Table 1, few of the transcriptomic biomarkers listed in Table 3 are reported as among the most significantly different RNAs in multiple reports. However, many miRNAs were measured with the same regulation (up or down) across multiple studies, indicating generally good agreement. 88 Interestingly, both miR-92a and miR-451, in addition to being candidate biomarkers for TBI, have been reported as cancer biomarker candidates, raising concerns about specificity.182,183 Notably, the mRNA NM_000584 codes for IL-8, and 2 tsRNAs (tRF-Ser-FCT-078 and tRF-Thr-AGT-003) were recently reported with post-TBI alterations in mice; taken collectively, these 3 candidate markers (involved in neuroinflammatory responses) indicate that the inflammation observed in the secondary injury phase of TBI may have origins at the transcriptomic level.150,168
Transcriptomic analysis of TBI reveals broad, genome-wide alterations in the expression of 100s of RNAs. 184 Although the difficulties in quantifying the transcriptome currently hinder transcriptomic methods, RNA profiling (particularly using targeted methods such as RNA-seq) holds significant potential as a means to understand TBI and related neurological diseases.164,185-187 For example, down-regulation of synapse signaling-associated gene signatures has been identified in patients with CTE, AD, and both, and many overlapping hippocampal gene signatures have previously been noted between TBI and other neurological diseases.184,188 However, it is challenging to confidently identify gene associations and expression signatures, and results in the literature are sometimes in direct conflict.189,190 As transcriptomic technology matures, it will become an important part of how we understand the pathogenesis of TBI.
Epigenomic Biomarkers
Epigenomics involves the study of modifications that affect gene expression but are independent of the genes themselves. Although all cells in an organism contain the same genetic information, the degree of expression of the genes varies depending on cell type. Some of these cell-specific expression profiles arise from epigenetic regulation of chromatin packaging, such as histone modifications, DNA methylation (DNAm), and modification of regulatory RNAs in the nucleus and mitochondria.87,191 DNAm frequency can be sequence-dependent, with CpG (5′-CG-3′) sites being common sites of methylation. 192 Frequently associated with epigenomics, epitranscriptomics involves regulatory modifications of RNA transcripts, such as RNA methylation (RNAm). 193 Although the basic functions of individual epigenetic modifications are generally understood (eg, DNAm represses transcription, histone acetylation activates transcription), the interactions of multiple regulators acting on a single gene are complex, being influenced by genetic and demographic factors.191,194-196
Epigenomic research generally involves investigating the connections between differential epigenetic modifications and differential gene expression between control and treatment groups. 197 These differences are quantified by genome-wide epigenetic analysis and transcriptomic quantification, as assessed by bisulfite sequencing or array techniques. 184 To reveal associations and determine predictive power, bioinformatics-assisted network and subnetwork association analyses, statistical methods such as PCA, and artificial intelligence approaches can be employed.184,198,199 Epigenomic research in TBI is relatively young, but epigenomic profiling holds significant potential as a means of assessing alterations in gene expression and gaining insight into the specific mechanisms by which gene expression is altered following TBI. 200 A selection of candidate epigenomic biomarkers for TBI, comprising differentially methylated genes with correlated alterations in expression following TBI, is shown in Table 4.
A selection of genes located near DNA methylation sites serving as potential biomarker candidates for TBI.
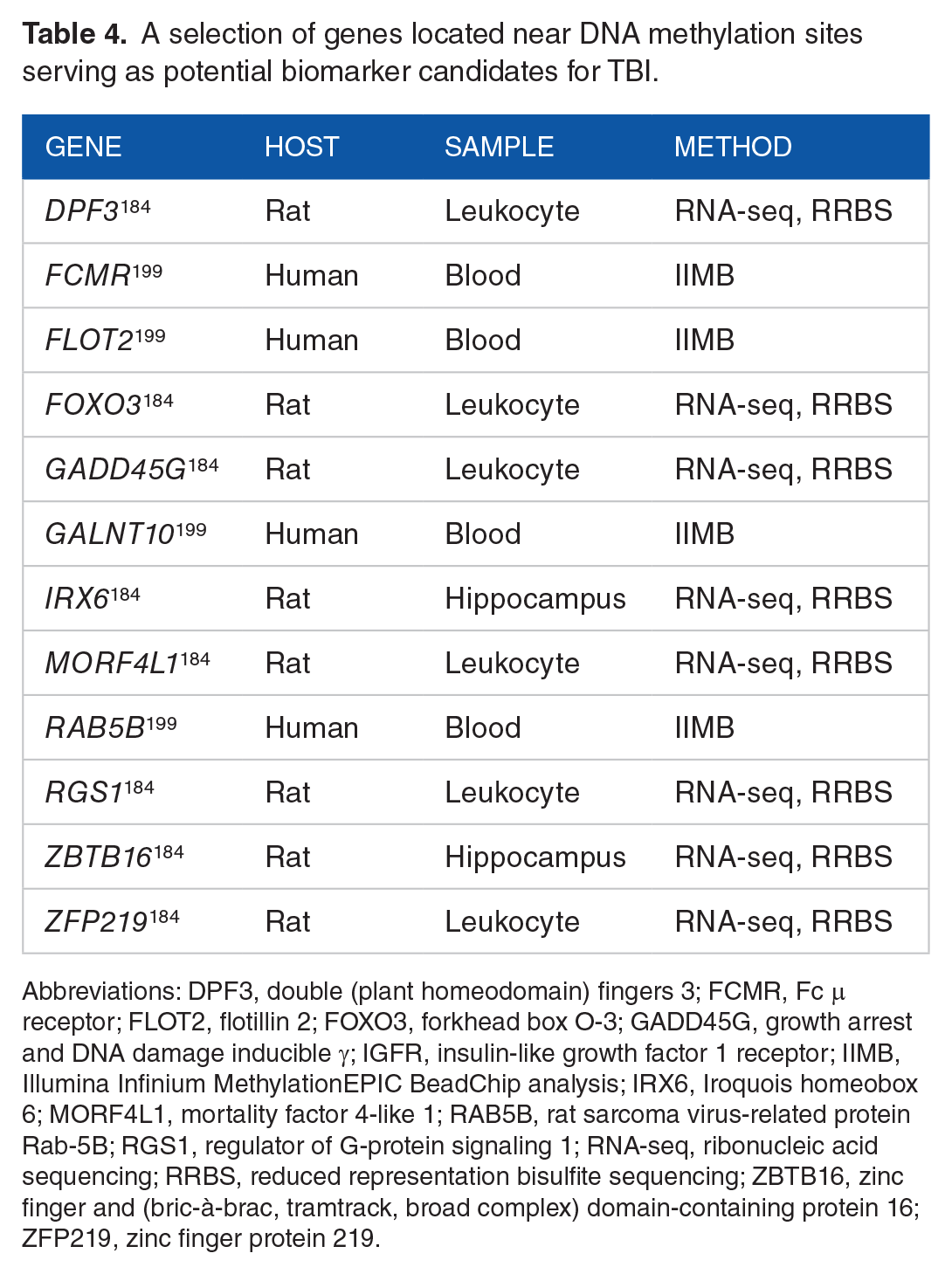
Abbreviations: DPF3, double (plant homeodomain) fingers 3; FCMR, Fc μ receptor; FLOT2, flotillin 2; FOXO3, forkhead box O-3; GADD45G, growth arrest and DNA damage inducible γ; IGFR, insulin-like growth factor 1 receptor; IIMB, Illumina Infinium MethylationEPIC BeadChip analysis; IRX6, Iroquois homeobox 6; MORF4L1, mortality factor 4-like 1; RAB5B, rat sarcoma virus-related protein Rab-5B; RGS1, regulator of G-protein signaling 1; RNA-seq, ribonucleic acid sequencing; RRBS, reduced representation bisulfite sequencing; ZBTB16, zinc finger and (bric-à-brac, tramtrack, broad complex) domain-containing protein 16; ZFP219, zinc finger protein 219.
Currently, less common epigenomic modifications such as ubiquitination and sumoylation remain relatively underexplored, and TBI epigenomic studies are rare.
201
However, the epigenome is a valuable target for the deep insights into gene regulation that can be gained from its study. For example, of the genes listed in Table 4,
Due to the scarcity of epigenomic research in TBI, few specific systems-level conclusions can be drawn. However, alterations in N6-methyladenosine-related RNAm have been linked to both TBI and AD, providing a link between the two at the epitranscriptomic level.193,207,208 The importance of methylation additionally establishes enzymes involved in methylation and demethylation, such as fat mass and obesity-associated protein, as targets of therapeutic relevance following TBI.193,209,210 Ultimately, the epigenome is a rich source of information, and future epigenomic efforts will shape our understanding of TBI and potential links to other neurodegenerative diseases.
Genomic Biomarkers
Genomics involves the study of the set of genes contained in an organism’s DNA. The genome can provide information about an individual’s risk of developing certain diseases and inform individualized treatment options.211,212 For monogenic diseases caused by variation in a single gene, genomic biomarker identification can be straightforward. However, most diseases (eg, AD, PD) are polygenic, are influenced by environmental factors (largely thought to affect the epigenome), present heterogeneous phenotypes, and do not follow classical Mendelian genetic patterns.213,214 All of these factors complicate the identification of biomarkers indicative of genetic susceptibility to specific diseases.212,215 Genomic analyses can be broadly classified as genome-wide association studies (GWAS) or candidate gene studies. In GWAS, the entire genome is considered without any previously formed hypotheses about potentially associated genes, making GWAS inherently less biased and more comprehensive. 216 In candidate gene studies, a single gene of interest is interrogated for single nucleotide polymorphisms (SNPs) associated with a particular disease.215,216 Because the genome is contained within all cells of the body, almost any sample type can be used to extract DNA for biomarker identification. The advancement of high-throughput assays, including SNP arrays that detect specific alleles hybridized with fluorescently labeled oligonucleotide probes, has allowed for the large-scale collection of genomic data that can be used to identify potential biomarkers. After a smaller number of genes have been identified as potentially relevant to a specific condition using microarrays, quantitative polymerase chain reaction (PCR) assays can be used for higher specificity, sensitivity, and clinical relevance. 217 Genetic biomarkers are most commonly identified from genome association studies by multiple hypothesis testing, where each individual genetic marker is analyzed for significant association with a certain phenotype.218,219
Genetic factors are known to contribute to the likelihood of an individual presenting with neurological diseases such as AD and PD.220-222 For TBI, genomic biomarkers may help to understand how an individual’s body will respond to and recover from injury, potentially leading to improved risk assessment, individualized monitoring, and tailored treatment plans. As discussed previously, genetic variation likely plays a role in the high degree of heterogeneity observed in TBI through influence on an individual’s pre-injury status, neural repair efficiency, and secondary injury symptoms.216,223 A selection of candidate genomic biomarkers associated with impact, recovery status, or both following TBI is presented in Table 5.
A selection of genomic biomarker candidates associated with differential outcomes in TBI.

Abbreviations: ABC, adenosine triphosphate-binding cassette; ACE, angiotensin-converting enzyme; AD, Alzheimer’s disease; ANKK1, ankyrin repeat- and kinase domain-containing 1; APOE, apolipoprotein E; AQP4, aquaporin 4; BDNF, brain-derived neurotrophic factor; CNS, central nervous system; COMT, catechol-O-methyltransferase; CSF, cerebrospinal fluid; GABA, γ-aminobutyric acid; GCS, Glasgow coma scale; GOS, Glasgow outcome scale; MME, membrane metalloendopeptidase; SNP, single nucleotide polymorphism; TP53, tumor protein p53.
Gene names are italicized, but protein names are not.
Genetic studies tend to be limited in statistical power and population generalizability due to small samples consisting mostly of Caucasian individuals assigned male at birth.223,255 Nonetheless, the genome remains an invaluable source of information for predicting outcomes on an individual basis and for deep insights into the biochemical mechanisms underlying TBI and related neurological diseases. For example, the ε4 allele of
The genome is upstream of the rest of the multiome, meaning that the genome holds direct influence on the epigenome, the transcriptome, the proteome, and the metabolome. SNPs in coding regions play a dominant role in determining functional activity, and SNPs in non-coding regions can affect regulation of gene expression.262,263 Although TBI does not induce change in the genome itself, understanding genome-wide associations at the molecular level is essential to characterizing the mechanisms underpinning the complex pathophysiology of TBI and may shed light on coincidence of downstream neurological pathologies.
Limitations
Our review has several limitations that are worth noting. We focus exclusively on the identities of candidate biomarkers in TBI. Although considerations of injury severity, time course, biomarker validity, and more are valuable in understanding TBI overall, those considerations remain outside the scope of this review, in part due to the general scarcity of knowledge on those topics for biomarkers of TBI (for further information, see the Limitations section of the review by Gan et al). 86 Additionally, our review does not substantively discuss clinical implications in an effort to maintain our focus on molecular-level, mechanistic understanding. Therefore, discussions regarding clinically relevant biomarker parameters, such as sensitivity, specificity, statistical power, and population generalizability, are largely absent here. 77
The biomarkers discussed here are limited to those within the scope of the multiomic disciplines. It is worth noting that there are biomarkers outside of the multiome—such as cell-free plasma DNA levels and electroencephalography power and coherence—that have been reported with high predictive accuracy for TBI.264,265 Despite its incompleteness, multiomics provides a useful framework for understanding the system-wide biochemical changes that occur during TBI at the molecular level.
Conclusion
TBI is a spectrum of diseases, and the biomarker profile associated with it can change markedly based on the severity of the insult, the health history of the individual under consideration, and the timeline between injury and diagnosis. Thus, to understand, identify, and treat TBI, a multiomic assessment of the condition is likely required. Recent TBI biomarker research has led to the identification of many new candidate biomarkers across the multiomic disciplines. In this review, we have examined the leading biomarker candidates from each major component of the multiome. From the identities of these biomarkers, we have explored (1) the fundamental insights that can be gained from the identities of these biomarkers into the pathophysiology of TBI and (2) the potential mechanistic connections between TBI and other neurological diseases. From a strong mechanistic understanding, molecular diagnostic and therapeutic targets will emerge naturally, theoretically allowing for more targeted and effective clinical tests and treatments for TBI.
It is important to note that each part of the multiome has a temporal dependence for turnover, and the major question of which significant biochemical alterations causally link TBI to other neurodegenerative diseases—if any—remains open. Current biomarker candidates do not provide enough information to fully understand the complex biochemical changes that occur following a TBI, nor do they allow for confident identification of the potential mechanistic connections between TBI and other neurological diseases. Furthermore, the TBI-specificity and population generalizability of current targets are generally insufficient for broad clinical use. During the course of assembling this review, we identified a gap in longitudinal studies of TBI progression—particularly from a molecular framework. Ultimately, “big science” large-scale collaborative efforts across the multiomic disciplines will continue to be necessary to achieve a holistic understanding of TBI.
Footnotes
Acknowledgements
The authors gratefully acknowledge Mr. Davis Bobbitt for helpful discussions in the planning stages of the review. Figures were created with ![]() . This work was performed at the Los Alamos National Laboratory, which is operated by Triad National Security, LLC, for the National Nuclear Security Administration of the U.S. Department of Energy (Contract No. 89233218CNA000001). Many thanks to Samitaur Medical Technologies Inc., Mr. and Mrs. Frederick and Laurie Samitaur Smith for their support of TBI research.
. This work was performed at the Los Alamos National Laboratory, which is operated by Triad National Security, LLC, for the National Nuclear Security Administration of the U.S. Department of Energy (Contract No. 89233218CNA000001). Many thanks to Samitaur Medical Technologies Inc., Mr. and Mrs. Frederick and Laurie Samitaur Smith for their support of TBI research.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a Congressionally Directed Medical Research Program of the U.S. Department of Defense (R-00674-19-0: Biomarkers for Detection and Treatment of Traumatic Brain Injury; PI HM). The views expressed in this article are those of the authors and do not reflect the official policy or position of the U.S. Department of Defense nor the U.S. Government.
Declaration of Conflicting Interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
PAK, HM, and LML conceived the review. HM secured funding. PAK, SCM, and KDL wrote the review. All authors revised, edited, and have approved of the final version of the review.
Ethics Statement
No institutional review board approval was required for this work because no human subjects research was performed as part of this review.
Informed Consent Statement
No informed consent was required for this work because no human subjects research was performed as part of this review.
