Abstract
MicroRNAs (miRNAs) are endogenous 22-nucleotide RNAs that can play a fundamental regulatory role in the gene expression of various organisms. Current research suggests that miRNAs can assume pivotal roles in carcinogenesis. In this article, through bioinformatics mining and computational analysis, we determine a single miRNA commonly involved in the development of breast, cervical, endometrial, ovarian, and vulvar cancer, whereas we underline the existence of 7 more miRNAs common in all examined malignancies with the exception of vulvar cancer. Furthermore, we identify their target genes and encoded biological functions. We also analyze common biological processes on which all of the identified miRNAs act and we suggest a potential mechanism of action. In addition, we analyze exclusive miRNAs among the examined malignancies and bioinformatically explore their functionality. Collectively, our data can be employed in in vitro assays as a stepping stone in the identification of a universal machinery that is derailed in female malignancies, whereas exclusive miRNAs may be employed as putative targets for future chemotherapeutic agents or cancer-specific biomarkers.
Introduction
RNA is the template basis of transcription and translation in the cell. MicroRNAs (miRNAs) are endogenous 22-nucleotide RNAs that can play fundamental regulatory roles in both animals and plants. miRNAs target messenger RNAs (mRNAs) for degradation or translational repression. Although miRNAs have escaped scientific attention in the past, currently they comprise a large regulatory machinery and thus it is logical that they influence the expression of multiple protein coding genes. 1 Intriguingly, almost half of the annotated human miRNAs lie within chromosomal areas that are considered fragile and thus prone to instability. Furthermore, these chromosomal regions have been linked with various types of cancer in humans. 2 Further evidence suggests that miRNAs can assume the roles of tumor suppressors or oncogenes. For this reason, they are currently referred to as onco-miRNAs. Recently, expression profiling of miRNAs has proven to be the most accurate method of sub-classifying human cancers, superior to protein coding gene profiling. Data regarding the functionality of miRNAs are rapidly expanding: miRNAs such as let-7, miR-15, and miR-16 are putative candidates for cancer therapy. 2 The ongoing challenges include identifying targets regulated by the selected miRNAs, as they bind to their targets via incomplete complementarity.
Female cancers are associated with uncontrolled growth and spread of cancerous cells originating in the female reproductive organs, comprising the cervix, ovaries, uterus, fallopian tubes, vagina, and breasts. It is estimated that there will be 376 190 new cases diagnosed and 73 040 deaths due to female cancers in the United States during the year 2018. 3 Breast cancer is the most common cancer in women in the United States, accounting for an estimated 30% of all new cancer cases in women in 2018. Endometrial cancer is the fourth most common malignancy in American women, accounting for an estimated 7% of new cancer cases in 2018. 3 In the United Kingdom, cervical malignancy accounted for 8% of cancer cases in women aged 25 to 49 years from 2013 to 2015. 4 Cervical cancer is associated with high-risk human papillomavirus (HPV) infection; however, the infection alone may not be sufficient to induce malignancy. 5 Although ovarian cancer is not among the 10 leading sites of new cancer cases in women, it is has a very poor prognosis; it is the fourth most common cancer that women will die from in 2018, expected to be responsible for 5% of all cancer-related deaths in women. This is due to the fact that 60% of cases in the United States are diagnosed at late-stage disease, with a 5-year survival rate of 29%. 3 Finally, vulvar cancer, while being a rare malignancy, is currently on the rise regarding incidence potentially due to its association with HPV infection.6,7
An abundance of scientific research has been published of late regarding the role of miRNAs in gynecological malignancies. For instance, miRNAs such as miR-145 have been identified as central players in cervical carcinogenesis, 8 whereas it has been demonstrated that miR-125b, miR-145, miR-21, and miR-155 have pivotal roles in breast malignancy. 9 miR-200 and let-7 have been identified as key modules in ovarian cancer, 10 and miR-185, miR-210, miR-423, let-7c, miR-205, miR-429 have been shown to be upregulated and associated with oncogenesis, invasion, and metastasis in endometrial carcinoma.11,12 To our knowledge, there is not any available research that underpins the commonalities in the major 5 female malignancies connecting them through an miRNA basis.
In this article, we analyze the miRNA commonalities and differences among female cancers, namely, breast, cervical, endometrial, ovarian, and vulvar. Vaginal cancer was excluded due to the complete lack of experimental data regarding miRNA expression. We created a network association across all 5 cancers and differentially regulated miRNAs. We identified 1 commonly deregulated miRNA in breast, cervical, endometrial, and ovarian cancers, namely, miR-21, whereas miR-145, miR-155, miR-200b, miR-205, miR-34a, miR-92a, and miR-101 were found common in all the examined malignancies with the exception of vulvar cancer. Bioinformatics was employed to identify the putative protein coding targets of the selected miRNAs. Established tumor suppressors such as TP53 and fundamental components of the mTOR cascade such as RICTOR were among the genetic targets we identified. We proceeded by analyzing the biological processes these genes were involved in, which were Ras, ErbB, transforming growth factor (TGF)-β, and Wnt signaling among others, whereas significantly overrepresented were biological processes of embryogenesis and development. Finally, we provide a putative network of interplay among these pathways while also exploring the unique miRNAs attributed to each of the examined malignancies.
Materials and Methods
We identified all miRNAs associated with our targeted malignancies from miRCancer database 13 with the exception of vulvar cancer gene targets. These, as they were not included in the miRCancer database, were manually extracted from research manuscripts.14-18 These research papers were identified through a PubMed advanced search with the following keywords: “vulvar cancer” OR “vulvar carcinogenesis” AND “miRNA” conducted on November 24, 2018. This search retrieved 13 articles out of which 5 publications were deemed appropriate as they provided miRNA targets identified in human cells. An analogous search was conducted for vaginal-cancer-related miRNAs which did not retrieve any exploitable results. We created a visual network and Venn diagrams using Cytoscape V3.5.1. 19 and the online platform for Venn diagram generation of Bioinformatics and Evolutionary Genomics 20 to identify potential commonalities. Tables of the miRCancer database retrieved miRNAs as well as manually extracted targets can be found in Supplemental Table 1. File also includes miRNA orientation where available. We sourced gene targets of the selected miRNAs from miRWalk version 2.0. 21 We employed the miRBase option as the target database, whereas we selected the miRNA (has-miR) input identifier; Validated Target module was used to increase the in vivo significance of our search. In total, 3681 unique genes were identified through this search. The biological function of the genetic targets was analyzed with ClueGo version 2.3.3 plugin for Cytoscape. 22 Statistical analyses were performed using the Bonferroni step-down approach and biological function clusters were selected and visualized in a pie chart only if they met the P-value .001 criterion. Further settings include the following: (1) GO Term Fusion option was selected; (2) statistical options were Enrichment/Depletion (2-sided hypergeometric test) with Bonferroni step-down approach; (3) leading group term was based on the calculated Kappa score; (4) only pathways with P-values less than .001 were taken into account. Graph shown in the figure was constructed using GraphPad Prism version 7. We further analyzed the lists of miRNAs found to be unique in each malignancy as shown in Figure 1A. We identified their genetic targets through miRWalk version 2.0 as explained before (Supplemental Table 6). We performed a GO_Biological analysis using ClueGo version 2.3.3 with the same variables as previously stated (Figure 3). Furthermore, gene lists were also analyzed through KEGG mapper online tool (https://www.genome.jp/kegg/mapper.html) and the 3 most enriched pathways are shown in Supplemental Figure 1. Furthermore, miRNAs found to be specific in each cancer category examined were analyzed regarding GO_Biological Process enrichment as previously discussed. STRING analysis of protein association as shown in Figure 4 was performed through STRING: Functional Protein Association Networks (https://string-db.org) online platform. Addition of connective nodes was not performed. Data were retrieved through text mining, experiments, and databases. Edge boldness represented the confidence of association when medium confidence setting was employed. Disconnected nodes were hidden from the network. To examine whether the common miRNAs were also present in other malignancies, we employed the miRCancer online database (http://mircancer.ecu.edu).

Commonalities and variations among female malignancies: (A) Venn diagram of miRNAs in female malignancies (breast, ovarian, endometrial, cervical, and vulvar). The central miRNA in all malignancies is miR-21, whereas a further 7 miRNAs (miR-145, miR-200b, miR-205, miR-34, miR-92, miR-101, and miR-155) have been found to be common in all malignancies with the exception of vulvar cancer. The miRNAs in each subset of the Venn diagram can be found in the supplementary table (Sheet 1). (B) Cytoscape network depicting common miRNAs in female malignancies. Red edges indicate the downregulation of the target miRNA, whereas green edges indicate upregulation. miRNAs with unclear pattern of regulation are connected to the relative malignancy by a gray edge. Publications employed to mine these associations are available in the supplemental table (Sheet 2).
Results
In this article, we analyzed in vivo data by employing a bioinformatics approach. We initially gathered miRNA associations with common gynecological cancers, namely, ovarian, cervical, breast, vulvar, and endometrial cancer. These malignancies were selected on the basis of the availability of database information or the literature examining miRNA expression in human tissue experiments. In total, we analyzed 426 associations. Figure 1A presents the Venn diagram commonalities and differences among the examined female malignancies. Interestingly, miR-21 appears to be shared among all malignancies, whereas a cluster or miRNAs, namely, miR-145, miR-200b, miR-155, miR-205, miR-34, miR-92, and miR-101, appear shared between all malignancies with the exception of vulvar cancer (Figure 1A). Such a difference might underline the significance of the different functionalities and structures of epithelial tissues. For example, vulvar epithelium is characterized as squamous in contrast to breast, ovarian, cervical, and endometrial epithelia which are primarily cuboidal or columnar. 23 Furthermore, the location of these epithelia dictates fundamental differences in their functionalities as reviewed in Wira et al. 24 In Figure 1B, we illustrate the central miRNA along with the 7 more core miRNAs of our analysis and present their differential regulation (red—downregulation, green—upregulation, gray—undetermined or contradictory evidence) regarding the linked cancer node (Figure 1B and Supplemental Table 2). Furthermore, we sought to understand the genetic impact of the action of these miRNAs. For this analysis, we employed the miRWalk database, which returned 3680 associations. miR-21 has been found to regulate genes such as BRCA1, whereas miR-34 regulates the TP53 activity (Supplemental Table 4). We further examined the biological processes that target miRNAs were involved in, using ClueGo plugin GO_Biological Process analysis. We set our criteria to a P-value < .001 to mine only statistically significant associations and decrease the amount of analysis noise. Intriguingly, our search returned biological processes such as G1/S transition of mitotic cell cycle, Ras protein signal transduction, ErbB signaling pathway, and Wnt signaling pathway, among others (Figure 2). The genes that mapped to each individual GO_Biological Process were recorded along with the P-value significance of their association with the mapped processes (Supplemental Table 3).

Biological function pie chart of the target genes of the 8 central miRNAs. Network was constructed employing ClueGo version 2.3.2 plugin of Cytoscape V3.5.1. Individual genes attributed to each of the represented GO_Biological Processes can be found in the supplementary material (Sheet 3). All the represented biological processes are significant with a P-value > .001.
We further inquired whether significant differences existed regarding miRNA enrichment among the examined female malignancies. We analyzed the miRNAs that uniquely mapped to each individual cancer Venn diagram circle (Figure 1A), mined their genetic targets as previously (Supplemental Table 6), and identified the GO_Biological Processes they were involved as stated previously (Figure 3). Although the greatest percentage encoded for processes such as RNA processing or biosynthetic process as expected, tumor-specific characteristics such as vasculature development in ovarian cancer emerged. 25 Although extensive in vitro and in vivo research is needed to confirm the specificity of these miRNAs and judge their putative use as biomarkers of tumor load or tumor response to chemotherapeutic regiments, here we provide an initial in silico filtered and analyzed list of such miRNAs.

Biological function pie chart of the target genes of the cancer-specific miRNAs. Network was constructed employing ClueGo version 2.3.2 plugin of Cytoscape V3.5.1. Target genes employed to mine the biological process associations can be found in Supplemental Table 6. All the represented biological processes are significant with a P-value > .001.
Through this analysis, we identified commonalities as well as differences regarding miRNA expression in major female malignancies and subsequently analyzed their putative cellular function through bioinformatic analysis employing in vivo data. This work allows for a more profound understanding of the roles of miRNAs in gynecological cancers and forms the basis for more focused, in vitro and in vivo research.
Discussion
In this article, bioinformatics was applied to analyze 8 common miRNAs involved in female malignancies, namely, miR-145, miR-200b, miR-155, miR-205, miR-21, miR-34, miR-92, and miR-101.
miR-21 was found to be central in all the examined female malignancies and is one of the first mammalian miRNAs identified that plays a crucial role in an abundance of biological functions and diseases, including embryonic development, inflammation, and cancer. Not surprisingly, miR21 is one of the most frequently upregulated miRNAs in solid tumors. The miRNA-21 gene is located on chromosome 17 (17q23.2), within an intronic region of the coding gene VMP1 (Vacuole membrane protein 1) in the direction of transcription; however, it is transcribed independently through a special promoter. 26 Intriguingly, the same genetic locus is an HPV16 integration site and consequently miR-21 is upregulated in HPV-positive patients. This implies a possible interaction between HPV, one of the most important factors leading to the development of cervical cancer, and onco-miRNAs such as miR-21, meaning that if the expression of these elements is altered, it could lead to carcinogenesis. 27 This onco-miR can be activated via the nuclear factor (NF)-κB and mitogen-activated protein kinase (MAPK) (ERK1/2) pathway, and regulates the expression of programmed cell death protein 4 (PDCD4), RECK (a membrane-anchored inhibitor of metalloproteinases), and tropomyosin 1 (TPM1), a protein with the potential to suppress cell growth and invasiveness of breast carcinoma. Other targets include Sprouty2 (SPRY2), tumor suppressor phosphatase and tensin homolog (PTEN), and CCL20 (which is related to HPV oncoproteins and cervical cancer). 28
In view of the other core miRNAs, miR-34 is a fundamental regulator of tumor suppression, controlling multiple protein targets involved in the cell cycle and apoptosis. Furthermore, it antagonizes cancer cells’ biological processes that lead to metastasis and chemo-resistance. 29 miR-34 has been associated with the development of hematological malignancies via activation of the ERK-Akt pathway. 30 In addition, miR-34 has further been associated with a plethora of other malignancies such as prostate cancer, 31 pancreatic cancer, 32 medulloblastoma, 33 and glioblastoma. 34 Currently, few miR-34 regulators have been delineated, including CD95, MYC, platelet-derived growth factor (PDGF), and protease-activated receptor 2 (PAR2). However, data are more extensive regarding miR-34 targets, comprising NOTCH, C-MYC, CD24, CCL22, and BCL-2, among others.
Our study identified miR-101 as an miRNA involved in female malignancies. miR-101 acts by directly inhibiting TSHZ3 (a smooth muscle cell differentiation transcription factor), HTRA3 (HtrA Serine Peptidase 3 involved in ovarian serous adenocarcinoma and ovarian cystadenocarcinoma development), and FN1 (fibronectin involved in cell migration during embryogenesis and metastasis). In recent publications, it was demonstrated that miR-101 is inhibited in multiple solid malignancies, including osteosarcoma, glioma, papillary thyroid carcinoma, hepatocellular carcinoma, and gallbladder carcinoma. These findings suggest that miR-101 functions as a tumor suppressor.35-39 Current studies have revealed that miR-101 interacts with SOX2 transcription factor in tumor progression suppression. 40
miR-92 is another miRNA that was identified through our bioinformatic analysis as frequent among female malignancies. miR-92 has been investigated in the context of multiple malignancies such as colon cancer, hepatocellular carcinoma, and leukemia.41-44 miR-92 has been demonstrated to hold anti-apoptotic activity in the context of colon cancer. 41 The genetic targets of miR-92 have been involved in cell cycle regulation and signaling, rendering this miRNA important regarding cellular proliferation and mammalian cell development. 45 Lack of miR-92 has been shown to lead to impaired B-cell development, lung hypoplasia, and ventricular septal defects, implicating miR-92 in autoimmunity as well as cancer development.45,46 Recent research has demonstrated that the plasma level of miR-92 is an effective novel biomarker for the detection of leukemia. 47
miR-145 is a member of the miR-143/145 cluster, located on chromosome 5 (5q32-33), encoded within the genes that code for the smooth-muscle-specific contractile proteins. It carries its own promoter and seems to be transcriptionally controlled by p53. Oncogenic signals can reduce the transcription of the miR cluster 143/145, which in turn can increase the tumorigenic events, resulting in cancer. In contrast, anti-oncogenic signals can upregulate this cluster, which can eventually reduce tumorigenesis. 48 Downregulation of miR-145 has been observed in all types of gynecological cancers, with the exception of vulvar, suggesting that the loss of miR-145 is a common early neoplastic event. A number of genes that are important in various cancers have been identified and validated as target genes of miR-145, such as insulin receptor substrate 1 (IRS1), homeobox A9 (HOXA9), sox9 and adducin 3, disintegrin and metalloproteinase domain-containing protein 17 (ADAM17), octamer-binding transcription factor 4 (OCT4), Krüppel-like factor 4 (KLF4), C-MYC14, and MUCIN1. 49 miR-145 also inhibits the activation of the mTOR signaling pathway, commonly deregulated in human cancers, by targeting AKT3. 50
miR-155 was also present in our findings and is processed from the final exon of a non-coding region known as the B-cell Integration Cluster (BIC) on chromosome 21 (21q21.3), which was thought to be a proto-oncogene associated with lymphoma. An abundance of in silico targets are predicted as well as confirmed, including regulatory proteins for hematopoiesis (SHIP-1, AICDA, SPI1, etc) and inflammation (BACH1, FADD, IKBKE, SPI1, SOCS, etc) and known tumor suppressors (CEBPβ, IL17RB, PCCD4, TCF12, ZNF652, LKB1, etc). This multifactorial, p53-regulated onco-miR is found to be consistently upregulated in a variety of cancers such as breast, 51 HPV-positive and HPV-negative cervical cancer, 52 and endometrial adenocarcinoma, where it seems to play a key part in the paracrine role of tumorigenesis by repressing the angiotensin II type 1 receptor (AGTR1), influencing the local renin-angiotensin system (RAS). 53 The multitude of validated miR-155 targets generates interest for possible oncogenic pathways, including its ability to regulate hematopoiesis and lymphocyte homeostasis and tolerance. In addition, there is the possibility of a cross-talk factor relating inflammation and carcinogenesis, with miR-155 being the link between the two. 54
miR-200b, clustered with other members of the miR-200 family in chromosome 1 (1p36), has been reported to be downregulated in various human malignancies. It modulates a wide range of cellular functions such as proliferation, motility, apoptosis, stemness, and epithelial-to-mesenchymal transition (EMT) by targeting, in the latter, 2 of its master regulators: ZEB1 and ZEB2. 55 In endometriotic cells, in which miR-200b is downregulated, the repression of these 2 transcription factor targets seems to control the expression of the anti-metastatic adhesion molecule E-cadherin. This suggests a possible pathway for increased invasive behavior. 56 Other targets include KLF4, FoxG1, a transcriptional repressor with oncogenic potential in cervical cancer, 57 and FUT4, resulting in abnormal protein fucosylation commonly associated with cancer malignancy. 58 miR-205, as a member of the same miR family, shares some of its targets and functions with miR-200b, but is located in the second intron of the LOC642587 locus on chromosome 1. 59 Ectopic expression of miR-205 prompts tumorigenesis. For instance, it is found to be downregulated in breast tumors where it directly targets the human epidermal growth factor receptor (HER)-3 tyrosine protein kinase receptor and interferes with the proliferative pathway mediated by the HER receptor family. 60 On the other hand, studies show that miR-205 is up-modulated in endometrial and ovarian carcinomas. Among its studied targets are transcription factor 21 (TCF21), which inhibits matrix metalloproteinases MMP-2 and MMP-10, 61 VEGF-A (a positive regulator of tumor growth), and tumor suppressors PTEN and SHIP2. 62 Both miR-200b and miR-205 modulate EMT by targeting 2 of its master regulators, ZEB1 and ZEB2, 55 and target NOTCH pathway components, such as JAG1.
In this work, we focused on the core similarities but also the differences regarding miRNA deregulation among the major female malignancies. An initial question arose regarding the existence of a common regulatory pathway underpinning these commonalities. To mine such a pathway, we employed the genes that mapped to the 4 key pathways that emerged as commonly enriched in our GO_Biological Process analysis, namely, ErbB, Ras, Wnt, and TGF-β pathways (Figure 2). Our generated STRING analysis indicated that such an interconnectivity existed, potentially underlining an interplay between the major pathways leading to tumorigenesis under the influence of the centrally identified miRNAs (Figure 4). Further in vitro and in vivo research is obviously required to fully delineate the nature of interplay among the depicted protein players and underpin potentially novel interactions among the major pathways deregulated in cancer.

Putative association of interconnectivity among target miRNA-regulated pathways. Here we employed genes that mapped to ErbB, Wnt, Ras, and TGF-β signaling GO_Biological Process (Supplemental Table 3) categories to see whether connectivity in a protein level is possible through STRING database analysis. We used data originating from text mining, experiments, and databases. Thickness of edges indicates the strength of confidence in the represented associations (see figure key), whereas protein nodes were colored according to the pathway they belonged to (see figure key).
Another question that arose from our work was whether the common miRNAs identified as central in female malignancies were also deregulated in other cancers. To address this, we used our core miRNAs as probes and mined the mirCancer database (Figure 5 and Supplemental Table 5). We identified malignancies such as hepatocellular, colon, and lung cancers carrying a similar miRNA profile out of the 54 malignancies retrieved. Intriguingly, these malignancies carry a high metastatic potential, similar to that of breast, cervical, and ovarian cancers. 63 This finding may indicate that the signaling pathway depicted previously (Figure 4) may play a role in the metastatic potential of these tumors. Obviously, this association is not the only one that brings these malignancies together. A classic example is the CA125 biomarker that is clinically employed to diagnose and stage ovarian cancer that has been found to be also significantly increased in other malignancies such as lung, liver, and colon cancer.64-66 This might indicate a more profound connectivity between these malignancies in a fundamental molecular level.
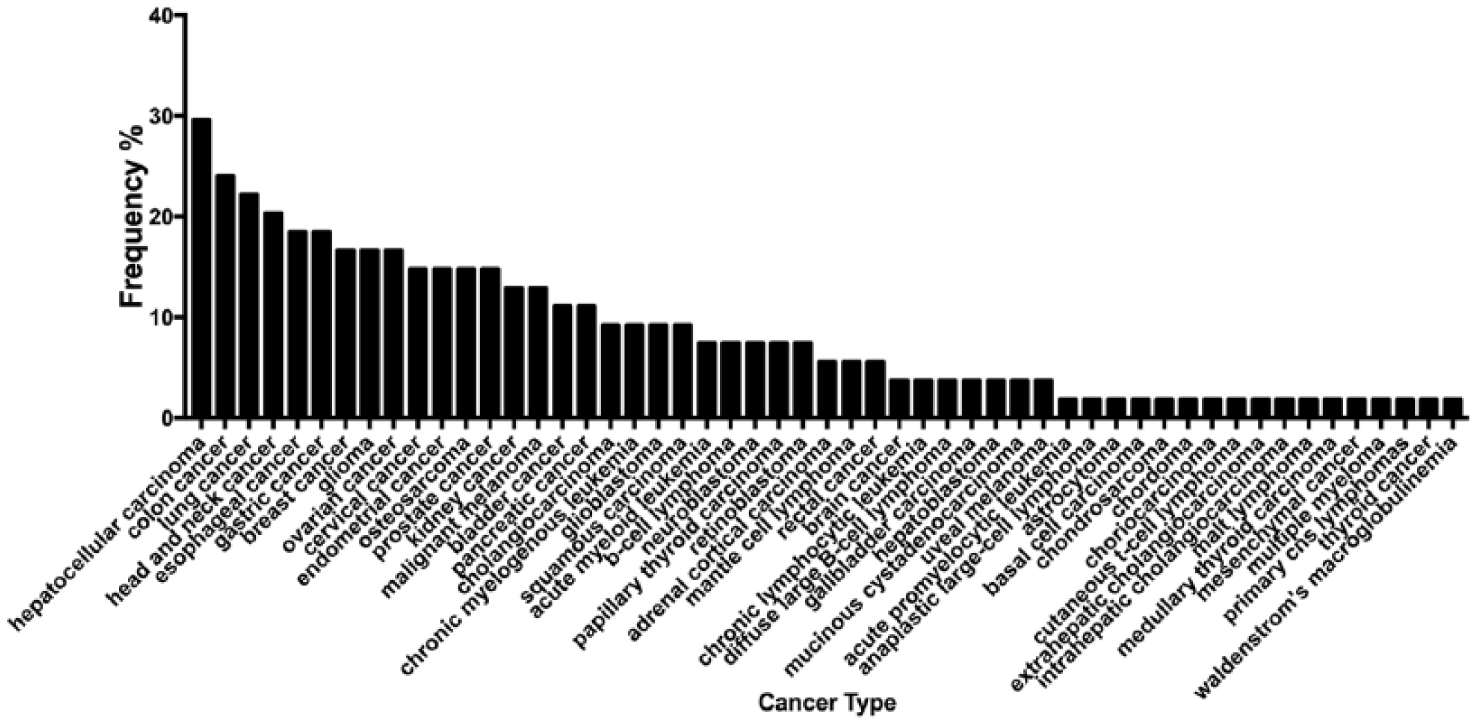
Selected miRNA association with other malignancies and their most central genetic targets. Selected miRNAs were inserted in the miRCancer database to retrieve all the malignancies that they have been associated with. This chart presents the most frequent cancers that have appeared in our search. A total of 54 types of cancer were retrieved and frequencies were calculated on the basis of how many times has each type been associated with any of the selected miRNAs. Graph was generated using GraphPad Prism version 7; the dataset used to create it can be found in Supplemental Table 5.
Furthermore, we sought to identify whether miRNAs that were found specific to individual female cancers in our work have also been shown to possess similar tumor specificity in other research. For example, miR-222 found to be specific for endometrial cancer in our bioinformatic analysis has also been shown to carry prognostic value as shown in endometrial cancer patient tissue sample analysis. 67 Another in vitro work has underlined its importance in endometrial cancer proliferation and invasion. 68 A large number of studies have reported the usefulness of miRNAs as diagnostic, prognostic, or predictive biomarkers for breast cancer. In our study, the largest retrieved list, consisting of 102 involved miRNAs, pertained only to this type of female malignancy. Among these miRNAs, many have been already recognized as promising biomarkers, such as let-7. 69 Its involvement in the regulation of breast cancer tumor-initiating cells (T-ICs) 70 makes it a good prognostic and diagnostic biomarker candidate. Another example is miR-125 that targets directly HER2, 71 whereas miR-122 showed both diagnostic and predictive values for the response to chemotherapy. 69 On the other hand, several miRNAs on the same list, such as miR-597, miR-3188, and miR-1229, have been correlated with breast cancer72-74 but insufficiently studied and not included in the previously described molecular fingerprints of breast cancer. As for the ovarian cancer, 32 specific miRNAs were sourced. The upregulation of miR-223 has been recognized as a useful diagnostic biomarker and also indicative of short overall survival, 75 whereas miR-532, miR-199, and miR-223 showed diagnostic potential for differential staging of ovarian carcinoma. 76 However, miR-411 and miR-551a need to be further investigated. The former has only been correlated with drug resistance,77,78 whereas its possible involvement in oncogenesis remains unclear. Regarding the latter, there is evidence of tumor-suppressive function 79 although it has not been included in miRNA signatures of the ovarian cancer. The output data about cervical cancer contained only 24 miRNAs. Many among them have been previously described in miRNA panels serving as early and later detection biomarkers of cervical intraepithelial neoplasia and cervical cancer. Namely, miR-484, miR-181, and miR-135 are commonly deregulated in low-grade neoplasia (CIN1), whereas miR-376 and miR-99 were found to be deregulated from the CIN2 stage and until invasive carcinoma.80,81 Moreover, miR-337, miR-634, miR-374c, and other retrieved miRNAs seem to have an impact on cell proliferation and invasion in cervical cancer cells.82-84 Therefore, they should be considered in future efforts of defining such panels.
Due to the nature of our research, which was solely in silico, no in vitro validation was conducted and therefore our results can be purely used as a guide for further investigation. Another limitation of our approach is that it does not include any form of normalization of the used data as they were retrieved through a database mining process. In general, normalization remains highly controversial when analyzing the expression of circulating miRNAs as there is no consensus yet on the suggested miRs and small nucleolar RNAs (snoRNAs) which could be used as endogenous controls. 85
In summary, here we provide further putative targets of fundamental research as well as guidelines as to which miRNAs might be found to carry tumor specificity regarding others. The emergence of miRNAs as gene expression modulators renders them ideal novel diagnostic and prognostic indicators as well as potential therapeutic targets. In addition, to their tissue specificity, miRNAs’ unique characteristics such as stability, ease of detection, and association with clinicopathological prognostic parameters render them exemplary targets in cancer diagnostics and therapeutics. miR-specific antagonists, miRNA replacement therapy, miRNA manipulation to increase responsiveness to therapeutic interventions, or miRNAs as prognostic, diagnostic, or treatment-responsive markers have started to emerge in applied clinical research.86,87 Here we have provided an extensive list of putative cancer-specific as well as universal cancer targets that may be employed in a clinical setting after extensive fundamental research. Future aspects would include in vitro analysis via RNA sequencing of malignancy samples to validate our in silico targets and provide novel proof of the universal miRNA basis in female malignancies.
Supplemental Material
Supplemental_Material – Supplemental material for MicroRNAs in Female Malignancies
Supplemental material, Supplemental_Material for MicroRNAs in Female Malignancies by Themis Liolios, Stavroula Lila Kastora and Giorgia Colombo in Cancer Informatics
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
TL wrote and drafted the manuscript. SLK conceptualized the manuscript, helped in methodology of the data, validated, formal analysed and investigated the manuscript. Also, contributed to the writing and original draft preparation, visualization, supervision of the manuscript. GC helped in data curated, also contributed to the writing, review and editing of the manuscript.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
