Abstract
B12D family proteins are transmembrane proteins that contain the B12D domain involved in membrane trafficking. Plants comprise several members of the B12D family, but these members’ numbers and specific functions are not determined. This study aims to identify and characterize the members of B12D protein family in plants. Phytozome database was retrieved for B12D proteins from 14 species. The total 66 B12D proteins were analyzed in silico for gene structure, motifs, gene expression, duplication events, and phylogenetics. In general, B12D proteins are between 86 and 98 aa in length, have 2 or 3 exons, and comprise a single transmembrane helix. Motif prediction and multiple sequence alignment show strong conservation among B12D proteins of 11 flowering plants species. Despite that, the phylogenetic tree revealed a distinct cluster of 16 B12D proteins that have high conservation across flowering plants. Motif prediction revealed 41 aa motif conserved in 58 of the analyzed B12D proteins similar to the bZIP motif, confirming that in the predicted biological process and molecular function, B12D proteins are DNA-binding proteins. Cis-regulatory elements screening in putative B12D promoters found various responsive elements for light, abscisic acid, methyl jasmonate, cytokinin, drought, and heat. Despite that, there is specific elements for cold stress, cell cycle, circadian, auxin, salicylic acid, and gibberellic acid in the promoter of a few B12D genes indicating for functional diversification for B12D family members. The digital expression shows that B12D genes of Glycine max have similar expression patterns consistent with their clustering in the phylogenetic tree. However, the expression of B12D genes of Hordeum vulgure appears inconsistent with their clustering in the tree. Despite the strong conservation of the B12D proteins of Viridiplantae, gene association analysis, promoter analysis, and digital expression indicate different roles for the members of the B12D family during plant developmental stages.
Introduction
The coordination of plant growth and reproduction processes is complicated and involves many regulatory proteins. Previous studies in plant transcripts during seed germination have identified various proteins expressed in aleurone and embryo during differentiation called B12D proteins. The first member of B12Ds was identified by screening the transcriptome of barley aleurone and embryo. 1 Thereafter, other members of B12D proteins have been identified and found to be expressed in all plant tissues at different developmental stages.2,3 Moreover, a member of B12D proteins is involved in leaf senescence in plants. 4
B12D family proteins are small transmembrane proteins containing the B12D domain. Six B12D proteins have been identified in the rice genome. 5 On the other hand, at least 8 or 9 B12D proteins were suggested to be expressed in various barley tissues. 2 However, the number of B12D proteins appears to be different across plant species. One of the B12D proteins is previously known to be the subunit NDUFA4 of the mammalian electron transport chain. NDUFA4, also called NADH-ubiquinone reductase complex I subunit MLRQ, 6 is conserved in insects, fungi, and higher metazoans and is involved in ATP synthesis. 7 Subcellular localization reveals that B12D protein from Arabidopsis #AT3G48140 is localized in the mitochondrion, 8 plasma membrane, 9 and peroxisome. 10 However, B12D proteins appear to have a single transmembrane helix and are embedded in the inner mitochondrial membrane.5,6
B12D proteins are expressed in various plant tissues, such as starchy endosperm, pericarp, immature and mature embryos, aleurone, seedling shoots, flowers during heading and early ripening spikelets.2,5,11 B12D genes appear to be preferentially regulated by plant hormones like gibberellic acid, abscisic acid, and ethylene.1,2,12 Additionally, B12D genes are regulated in response to different abiotic and biotic stress.5,12 Rice B12D gene #Os07g41340 is induced by flooding, salt, heat, and cold stress during germination.5,13 Likewise, Os07g41340 is involved in the rice defense response to biotic stress induced by blast disease caused by Magnaporthe oryzae. 14 Moreover, rice B12D gene #Os07g17330 is regulated in response to submergence in 5 rice genotypes. 15
Identifying the B12D gene family in Viridiplantae is essential to characterize this family member and understand their role during plant development. The present study aims to characterize the members of the B12D gene family in Viridiplantae by identification and in silico characterization of gene structure, functional motifs, phylogenetics, screening of cis-regulatory elements in the putative promoters, and digital expression analysis. The result of this study is expected to help uncover the role of B12D gene family members during plant differentiation and maturation.
Experimental Procedures
Database sequence retrieval
The Phytozome database (https://phytozome-next.jgi.doe.gov/ 16 ) was retrieved for B12D proteins across Viridiplantae. Fourteen species were selected to identify the B12D protein family in their genomes. The selection of species was based on their classification which the selected species present the main divisions of Viridiplantae. The 14 species are including 3 gymnosperms as follows: Botryococcus braunii v2.1 (chlorophyte), Marchantia polymorpha v3.1 (embryophyte), Selaginella moellendorffii v1.0 (tracheophyte), and 11 angiosperms include 1 Amborellales, Amborella trichopoda v1.0, 5 dicots (Aquilegia coerulea v3.1, Helianthus annuus r1.2, Glycine max Wm82.a2.v1, Populus trichocarpa v3.0, Arabidopsis thaliana TAIR10), and 5 monocots (Musa acuminata v1, Hordeum vulgare r1, Oryza sativa v7.0, Setaria italica v2.2, Zea mays RefGen_V4). For each protein found in the selected 14 species, the CDS (coding sequence), genomic, and amino acid sequences were downloaded from the Phytozome database. To ensure that all retrieved proteins are members of the B12D family, we identified the B12D domain (Pfam #PF06522) using Pfam (http://pfam.sanger.ac.uk) and my hits scan tool (http://myhits.isb-sib.ch/cgi-bin/motif_scan). Chromosome location for each gene was detected using the CDS sequence for each protein as a query in the NCBI tBLASTn against the Whole Genome Shotgun Contigs database. The genome size for each species was retrieved from EnsemblePlants database (https://plants.ensembl.org/index.html 17 ) to investigate the relationship between genome size and the number of B12D members in each plant genome.
In silico characterization and phylogeny
Functional protein association to predict the function of the B12D family proteins was predicted by STRING 11.0 (string-db.org) for the 6 B12D proteins from O. sativa. Transmembrane helix prediction was performed using MEMSAT-SVM. 18 The isoelectric point (PI) and molecular weight (MW) of B12D proteins were calculated by GeneScript (https://www.genscript.com/tools/). Motif prediction in protein sequences conducted by the MEME web server (http://meme-suite.org/tools/meme 19 ). Sequence alignment of conserved motifs was carried out using UniProt UGENE software. 20 Biological processes, molecular function, and cellular components were predicted by FFPred server 21 for A. thaliana and O. sativa B12D proteins. Exon/intron structures were generated by the Gene Structure Display Server (http://gsds.cbi.pku.edu.cn/ 22 ) using the corresponding CDS and genomic sequences for each B12D protein. The phylogenetic tree for the 66 B12D proteins was constructed based on B12D domain sequences by excluding the N and C-terminals sequences. The tree is built using minimum evolution method 23 —with interior branch tests of 1000 replicates using MEGA X software. 24
Cis-regulatory analysis
In the current study, B12D proteins from 5 species, 3 monocots (H. vulgare, S. italica, and O. sativa) and 3 dicots (G. max, P. trichocarpa, and A. thaliana), were selected for regulatory cis-elements screening. The sequences of the putative promoters of the selected B12D genes were obtained using the genomic sequence for each protein available in the Phytozome database as a query in the NCBI tBLASTn against the Whole Genome Shotgun Contigs database. PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/ 25 ) was used to screen cis-elements in the putative promoter sequences 1500 bp upstream of the start codon.
Digital expression of B12D genes
The digital expression analysis for B12D genes from G. max and H. vulgure was conducted using GENEVESTIGATOR v3 software (https://genevestigator.com/ 26 ). The heatmap tree for gene expression in various plant tissues was obtained based on log2 using the “anatomy” category and plant organs/tissues conditions. The heatmap tree for gene expression in various developmental stages was obtained based on log2 using the “development” category and plant growth conditions.
Investigating B12D genes duplication events
Gene duplication events were analyzed for paralogous clustered together in the phylogenetic tree and located in the same chromosome. The distance between the analyzed genes was not more than 50 kb. 27 The non-synonymous (Ka) to synonymous (Ks) substitutions ratio was calculated to investigate selection pressure for gene duplicates using the DnaSP v5.0 software (http://www.ub.edu/dnasp/ 28 ). Alignment for the coding region of CDS sequences for each gene pair was used to calculate Ka/Ks ratio. A Ka/Ks ratio value >1 is considered a positive selection, and a value <1 is considered a negative selection. 29 The divergence time for each gene duplicate was estimated by dividing the Ks value by the substitution rate, which is equal to 6.1 × 10−9 per 10−6 million years ago. 30
Results
Characteristics of B12D proteins in Viridiplantae
The Phytozome database retrieval for B12D proteins in 14 species of Viridiplantae revealed a total of 66 B12D proteins in the selected species, as shown in Supplemental Table S1. Domain analysis showed that all retrieved proteins contained the B12D domain. The length of the retrieved B12D proteins ranges from 66 to 197 aa, of which 77% are between 86 and 98 aa. The isoelectric point values range from 5.71 to 11.37, of which 90% of analyzed proteins are between 9 and 10.8. The molecular weight ranges from 7.8 to 22.6 kDa, of which 80% of protein are between 9.4 and 11.2 kDa. The number of B12D proteins in each species was from 2 to 9 proteins as follows: 2 B12D proteins in each of B. braunii and A. thaliana, 3 B12D proteins in each of S. moellendorffii and A. trichopoda, 4 B12D proteins in each of M. polymorpha, P. trichocarpa, and S. italica, 5 B12D proteins in M. acuminata, 6 B12D proteins in each of A. coerulea, H. annuus, G. max, O. sativa, and Z. mays, and 9 B12D proteins in H. vulgare. The number of B12D proteins in lower Viridiplantae ranges from 2 to 4 B12D proteins in each species, while in higher Viridiplantae ranges from 4 to 6 B12D proteins in each species, with 2 exceptions: in A. thaliana, 2 B12D proteins and in H. vulgare, 9 B12D proteins. To investigate the relationship between the number of B12D members and the genome size in each species, the genome size data of the selected 14 species was retrieved and presented in Supplemental Table S1. In general, the number of B12D members increases from lower to higher Viridiplantae according to the increasing in genome size. However, the duplication events increase the number of B12D members in M. polymorpha, A. coerulea, and O. sativa despite the limited genome size in these species. Moreover, there are some exceptions as shown in H. annuus and G. max which both have an equal number of B12D members (6 proteins) despite the variation in the genome size (3500 and 1115 Mb, respectively).
The number of exons for each of the retrieved B12D proteins was between 1 and 4, of which 95% have 2 or 3 exons (Figure 1). Despite this similarity in exon numbers and lengths, the length of some introns appears significantly longer in 4 B12D proteins from 4 different species (HanXRQChr09g265771, A.trichopoda_scaffold00055.169, Potri.012G74900, and LOC_Os03g40440). Domain architecture appears conserved among most of the retrieved proteins, in which the B12D domain spans nearly from the first 10 aa at the N-terminus to the last 5 aa at the C-terminus of the proteins that range from 66 to 99 aa. However, longer proteins show a different domain architecture with the B12D domains located near the N-terminus (Supplemental Table S1). Transmembrane helix prediction showed that all retrieved B12D proteins comprise a single transmembrane helix except B12D protein #Potri.010G55300 of P. trichocarpa, which has 3 transmembrane helices (Supplemental Figure S1).

Exon-intron structure for 66 B12D genes from 14 Viridiplantae species visualized by Gene Structure Display Server. The CDS and genomic sequences for each gene were retrieved from the Phytozome database.
Functional association and motifs
Functional protein association by STRING for the 6 B12D proteins from O. sativa showed the data only for 3 B12D proteins: LOC_Os07g41340, LOC_Os07g41350, and LOC_Os06g13680 (Supplemental Figure S2). The other 3 B12D proteins of O. sativa (LOC_Os03g40440, LOC_Os07g17310, and LOC_Os07g17330) have no data about their functional interaction in the STRING database. The 2 proteins, LOC_Os07g41340 LOC_Os07g41350, appear to interact and be associated with the same proteins, including 4 bidirectional sugar transporters (SWEET2A, SWEET3A, SWEET3B, and SWEET4), 2 deoxyribonucleoside kinases, 2 hypoxia-induced proteins, and a spiral 1-like protein. The O. sativa protein, LOC_Os06g13680, interacted with 9 bidirectional sugar transporters (SWEET1A, SWEET2A, SWEET3A, SWEET6A, SWEET1B, SWEET2B, SWEET3B, SWEET6B, and SWEET4) beside a berberine bridge enzyme, which is involved in the biosynthesis of peptidoglycan.
Motif prediction revealed 4 motifs conserved in most of the investigated B12D proteins (Figure 2). The first motif appeared as the basic leucine zipper (bZIP) motif with 41 aa length and was conserved in 58 of the analyzed B12D proteins. The second motif is 21 aa length and located in the N-terminal. The third motif is 15 aa in length and located in the C-terminal. The second and third motifs contain 2 proline residues and appear to be the PxxxP motif required for protein-protein interactions. The fourth motif (LRKFVR) is 6 aa in length and found in 65 of the analyzed B12D proteins close to the C-terminal (Figure 3). Some B12D proteins contain only 1 or 2 of the 4 motifs as shown in the 2 proteins of B. braunii; the proteins Mapoly0021s0146 and Mapoly0021s0147 of M. polymorpha; the proteins HORVU7Hr1G93830 and HORVU1Hr1G46100 of H. vulgare; and the protein Aqcoe3G264200 of A. coerulea.
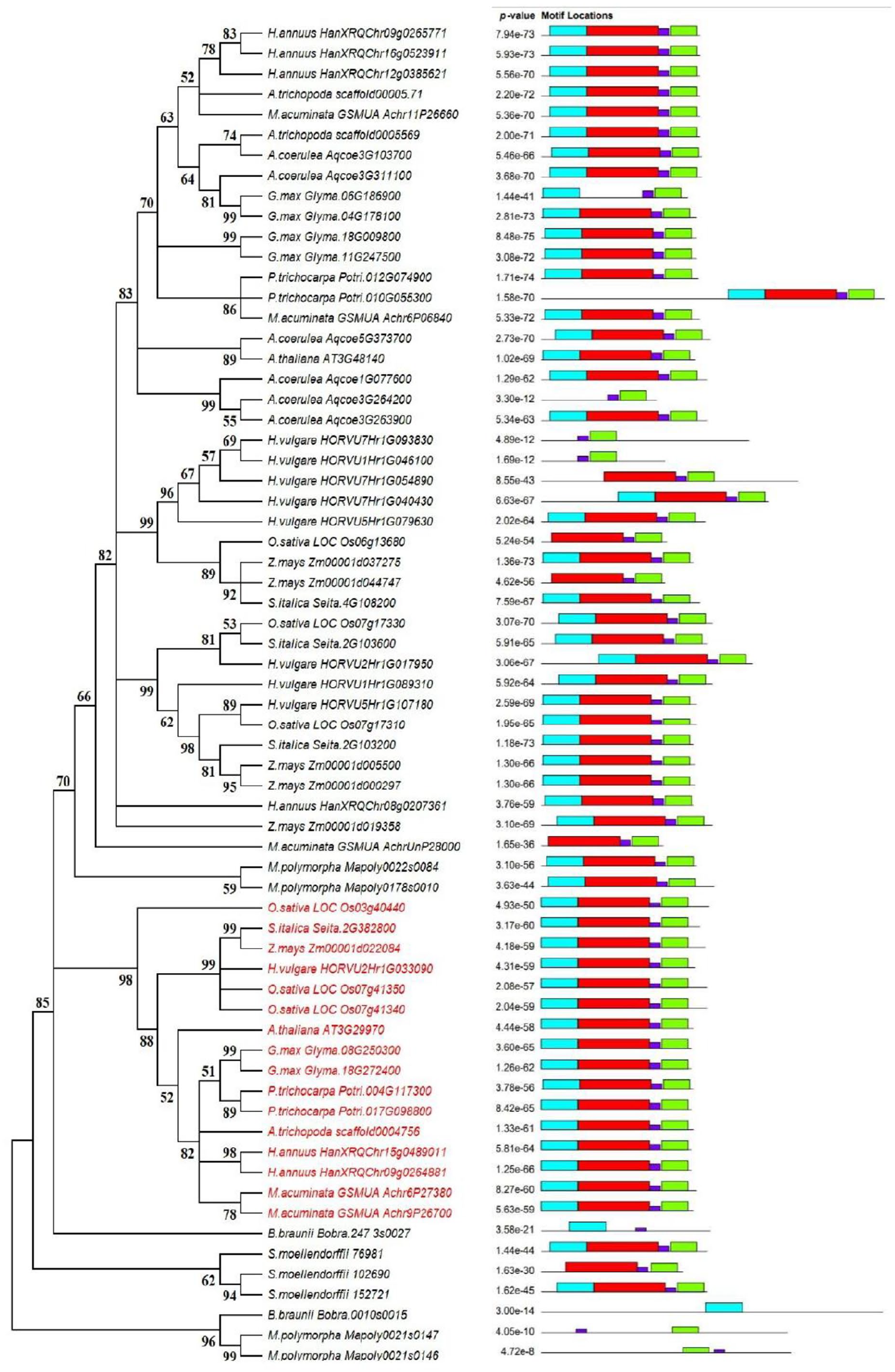
(A) Phylogenetic tree of 66 B12D proteins from 14 Viridiplantae species constructed by minimum evolution method with interior branch tests of 1000 replicates using MEGA X software. The tree was built based on B12D domain sequences and (B) conserved motifs in the 66 B12D proteins predicted by MEME server.

Sequence logo for the conserved motifs in the 66 B12D proteins from 14 Viridiplantae species predicted by MEME server.
Gene ontology analysis for A. thaliana and O. sativa proteins revealed a similar biological process and molecular function for the analyzed B12D proteins, which indicates the involvement of B12D proteins in transmembrane transport, and the establishment of localization in cells, and regulation of DNA-templated transcription. The molecular functions for the analyzed B12D proteins include catalytic activity, ion transmembrane transporter activity, nucleic acid binding, and cytokine activity. The cellular components for these proteins include the mitochondrial inner membrane, an integral component of the plasma membrane, endoplasmic reticulum, and extracellular vesicular exosome (Supplemental Tables S4-S9).
Multiple sequence alignment and phylogenetics
Multiple sequence alignment of 66 B12D proteins shows that M. polymorpha B12D proteins Mapoly0021s0146 and Mapoly0021s0147 are less conserved among the aligned 66 proteins, followed by the 2 B12D proteins from B. braunii and the 3 B12D proteins from S. moellendorffii, which are all 3 species from the lower Viridiplantae. The B12D proteins from the other 11 flowering plant species appear to be more conserved, although the alignment shows 2 distinct groups. The first group includes 16 B12D proteins: 3 proteins from O. sativa, 2 proteins of each of M. acuminata, H. annuus, G. max, and P. trichocarpa, and a single B12D protein each of A. trichopoda, A. thaliana, H. vulgare, S. italica, and Z. mays. This group does not include any proteins of A. coerulea. The second group includes the other 41 B12D proteins from these 10 species besides the B12D proteins of A. coerulea (Supplemental Figure S4).
The phylogenetic tree shows the clustering of B12D proteins of the 11 flowering plant species. In contrast, the B12D proteins from the lower Viridiplantae branch from the main clusters include the B12D proteins from higher plants with 85% support except for the 2 M. polymorpha B12D proteins (Mapoly0178s0010 and Mapoly0022s0084) which clustered with flowering plants cluster (Figure 2). The tree shows 2 main clusters: the first cluster branches from a node supported by 98% and includes the 16 B12D proteins that aligned together in the first group of the multiple sequence alignment (shown in red in the tree). These 16 proteins separate into 2 subclusters with 88% support: 1 for monocot proteins and the other for dicot proteins except the 2 proteins (Achr6P27380 and Achr9P26700) of the monocot: M. acuminata and the scaffold00047.156 protein of the Amborellales; A. trichopoda which clustered with dicots. However, B12D protein #AT3G29970 from A. thaliana separated from the dicots subcluster with 52% support. These 16 proteins include 2 B12D proteins of each of M. acuminata, H. annuus, P. trichocarpa, and G. max; single protein of each of A. trichopoda, A. thaliana, H. vulgare, Z. mays, and S. italica; and 3 B12D proteins of O. sativa. However, B12D protein #Os03g40440 from O. sativa forms a separate branch from this cluster with 98% support value. Notably, these cluster of 16 B12D proteins does not include the B12D protein of A. coerulea which is the only exception of 11 flowering plants that is not represented in this cluster.
The second cluster is separated into multiple subclusters from a node supported with 82%. However, B12D protein from M. acuminata separates from this subcluster from a node supported with 66%. The first and second subclusters each branches from a node supported by 99%. These 2 subclusters include 18 B12D proteins from monocots which each includes at least 1 B12D protein from the 4 species: H. vulgare, Z. mays, S. italica, and O. sativa. The third subcluster branches from a node supported with 83% and includes 13 B12D proteins from the 5 dicot species beside A. trichopoda and M. acuminata.
Cis-elements in the putative promoter of B12D genes
The screening of cis-elements in the promoters for selected B12D genes reveals that the 1500 bp putative promoter for B12D genes includes binding sites for various stress-responsive factors and plant hormones. These include the regulatory elements for gibberellic acid (GARE-motif, CARE, P-box, and TATC-box), the regulatory elements for abscisic acid (ABRE, ABRE2, ABRE4a, and ABRE4), the regulatory elements for auxin (AuxRR-core and TGA-element), the regulatory elements for methyl jasmonate (CGTCA-motif, JERE, and TGACG-motif), the regulatory element for ethylene (ERE), the regulatory element for cytokinin (as-1), the regulatory element for salicylic acid (TCA-element) the regulatory elements for wound (WRE3, WUN-motif, and W box); the regulatory elements for drought (DRE1, DRE core, MYB, MBS, and MYC), the regulatory element for anaerobic induction (ARE), the regulatory element for low-temperature (LTR), the regulatory elements for light (chs-CMA1a, box S, TCCC-motif, TCA-motif, Sp1, MRE, LS7, L-box, GT1-motif, GATA-motif, G-box, I-box, Box II, Box 4, ATC-motif, ATCT-motif, AT1-motif, AE-box, ACE, A-box, 3-AF3 binding site), the regulatory element for anoxic (GC-motif), the regulatory element for heat (STRE), the regulatory element for heat shock (CCAAT-box), the RY-element for seed-specific regulation (the CAT-box, which is specific for meristem, cell cycle, re2f-1, and circadian). Other regulatory elements such as CAAT-box, TATA-box, AP-1, AAGAA-motif, AC-I, ACTCATCCT sequence, unnamed_1, unnamed_2, unnamed_4, unnamed_6, unnamed_16, CTAG-motif, and F-box that are not assigned to any function in the PlantCARE database were found in some screened promotors (Supplemental Table S2). Some regulatory elements such as ABRE, ARE, AT~TATA-box, CAAT-box, CGTCA-motif, G-Box, MYB, MYC, STRE, TATA-box, as-1, and unnamed_4 are found in most analyzed promoters. Moreover, the elements related to light response and drought appear to be extensively present in all analyzed promoters (Figure 4). However, other elements such as re2f-1, circadian, chs-CMA1a, Unnamed_16, O2-site, regulatory elements for low temperature, auxin, salicylic acid, and gibberellic acid were found only in a few promoters (Supplemental Table S2).

Cis-elements in the putative promoter of 31 B12D genes from 6 Viridiplantae species; H. vulgare, S. italica, O. sativa, G. max, P. trichocarpa, and A. thaliana; predicted by PlantCARE database.
Digital expression of B12D genes
Digital expression was analyzed for 6 B12D genes from G. max and 9 B12D genes from H. vulgure. Two genes of G. max., Glyma.06G186900 and Glyma.04G178100, were highly expressed in all plant tissues at all developmental stages. One gene (Glyma.11G247500) was expressed in moderate levels in all tissues except nodule and seed and at a moderate to high level at different developmental stages. One gene (Glyma.18G009800) was expressed at a low level in cotyledon, flower, root, shoot, and leaf and at a high level in the early and middle developmental stages. Two genes (Glyma.18G272400 and Glyma.08G250300) were expressed only on nodules and roots and only during the early developmental stages (Figure 5).

Digital expression analysis of 6 B12D genes of G. max conducted by Genevestigator v3 software. Gene accession numbers are illustrated in the diagrams: (A) expression patterns in various plant tissues and (B) expression patterns in the developmental stages; cotyledon, trifoliate, flowering, pod fill, seeding, maturation.
Different expression patterns are seen with H. vulgure genes, in which the gene HORVU7Hr1G40430 is highly expressed in all plant tissues at all developmental stages. In contrast, HORVU1Hr1G89310 and HORVU2Hr1G17950 are expressed from moderate to high at all developmental stages in all plant tissues except shoot apex and spike. Three genes, HORVU7Hr1G93830, HORVU2Hr1G33090, and HORVU7Hr1G54890, show low expression at early developmental stages and high expression at late stages, but different expression patterns at the different tissues. Two other genes, HORVU1Hr1G46100 and HORVU5Hr1G79630, show low expression at early developmental stages and moderate expression at late stages and are expressed highly at lodicule, rachis, and palea while showing no expression to low expression in other tissues (Figure 6).

Digital expression analysis of 9 B12D genes in H. vulgure conducted by Genevestigator v3 software. Gene accession numbers are illustrated in the diagrams: (A) expression patterns in various plant tissues and (B) expression patterns in in the developmental stages; germination, tillering, booting, heading, flowering, spikelet, and ripening.
Gene duplication analysis
B12D gene duplicates were analyzed for selection pressure, and only 4 gene pairs were found on the same chromosomal region (Supplemental Table S1). Ka/Ks values for the analyzed gene duplicates from O. sativa and M. polymorpha were all less than 1, indicating that these 3 duplicates were evolved by negative selection, limiting the function of the genes after duplication. For A. coerulea duplicate, Ka and Ks values are 0, so the Ka/Ks ratio cannot be calculated. The divergence time for the O. sativa gene duplicates is 49.4 and 52.9 million years ago, and for the M. polymorpha gene duplicate is 169.5 million years ago (Supplemental Table S3).
Discussion
The B12D family has transmembrane proteins containing the B12D domain ranging in length from 80 to 98 aa found in plants, animals, and fungi.5,7 Our results show that B12D proteins appear conserved among the Viridiplantae in protein length, amino acid sequence, domain architecture, exon number, transmembrane helices number, and motifs. However, the number of B12D members appears different in the selected 14 species. This variation in gene copy numbers among species occurs during speciation which contributes to the increasing in genome size. In plants, gene copy number divergence between species is found to be caused by evolutionary adaptation to environmental stress. 31 Similarly, the number of B12D proteins in vertebrate species is different which there are at least 2 B12D members in vertebrates. 32 However, there are 3 copies in human of NUDFA4, the MLRQ subunit of mitochondrial NADH-ubiquinone reductase complex I, a member of B12D family. 33
Digital expression analysis shows different expression patterns for the paralogous genes from the same species in higher plants, as shown in G. max and H. vulgare. The phylogenetic tree and expression patterns of G. max B12D genes show that each 2 paralogous are clustered together in the tree and show similar expression patterns. However, expression patterns for H. vulgare B12D genes do not reflect their clustering in the phylogenetic tree. Each 3 paralogous H. vulgare B12D genes show similar expression patterns at the various developmental stages but show different expression patterns in the various tissues. These expression patterns of B12D family members indicate for the diversification in the function of B12D genes which are differentially expressed in the various developmental stages and the various organs of plant. This functional diversification in the highly conserved genes might occur as solution to diversifying a gene regulated by multiple enhancers with a high transcription activity. 34 The sequence conservation among B12D family member that has different roles is caused by the strong selection that results in divergence in the function faster than in the protein sequences as seen in histones and ribosomal RNA gene families.35,36
B12D proteins appear to be DNA binding proteins containing the bZIP motif, which is known to comprise 2 regions: a basic region that is rich in arginine (R), asparagine (N), and lysine (K), and a leucine region that is rich in leucine (L) and isoleucine (I). 37 Similarly, characterization of the wheat bZIP transcription factor family revealed that the conserved bZIP motif has 41 aa in length. It has a similar structure to the bZIP motif conserved across B12D proteins in the present study. 38 The involvement of B12D proteins in the regulation of transcription is confirmed by the predicted biological process of B12D proteins. From these findings, B12D proteins appear to be membrane-bound transcription factors, and their target genes might be nuclear or mitochondrial as some B12D proteins are localized in the mitochondrial inner membrane.5,8,33 Interestingly, some transcription factors are embedded in mitochondrial membranes in a dormant state and start to regulate nuclear genes in response to external or internal signals by translocation to the nucleus during plant growth and reproduction under environmental stress.39,40 However, our knowledge about the target genes of B12D proteins is still very limited.
The 2 motifs on the N and C terminal contain the conserved PxxxP repeat involved in protein-protein interaction. 41 PxxxP motif is found in the mitochondrial ADP/ATP carrier proteins. It is known to allow close-packing of transmembrane helices alternating closing and opening of the carrier on the 2 sides of the inner mitochondrial membrane. 42
The phylogenetic tree and multiple sequence alignment reveal independent divergence of ancient B12D proteins from the 3 gymnosperm species; B. braunii, M. polymorpha, and S. moellendorffii. The B12D proteins from flowering plants show clear divergence into 2 main clades each includes a member of A. thaliana B12D proteins; AT3G29970 or AT3G48140. Clade AT3G29970 includes 16 B12D proteins while clade AT3G48140 includes 41 B12D proteins. Notably, each of the 2 clades; AT3G29970 and AT3G48140 in the tree shows separate clustering for dicots and monocots B12D proteins except for B12D proteins of the monocot, M. acuminata, which are clustered with dicot species in AT3G48140 clade. Moreover, each of the 2 clades includes at least single B12D protein from the 11 flowering plant species except A. coerulea which all 5 B12D proteins clustered in AT3G48140 clade. Clustering of B12D proteins into 2 clades suggests an independent divergence of these clades from ancient orthologous B12D proteins from lower Viridiplantae. Moreover, the functional association of rice B12D proteins LOC_Os07g41340 and LOC_Os07g41350 that belong to AT3G29970 clade shows different protein associations from the LOC_Os06g13680 protein, confirming that some members of the B12D family have a specific function.
Despite the high conservation of B12D proteins, only 4 pairs of B12D genes from 3 species were found to evolve by duplication. The duplicates from O. sativa and M. polymorpha resulted after experiencing negative selection pressure that silence one of the duplicates. This finding agrees with the study by He et al 5 shows that 1 of B12D duplicate genes (#LOC_Os07g17330) is expressed in various rice tissues but its duplicate (#LOC_Os07g17310) has very low expression, suggesting that some B12D duplicate genes have been silenced during plant evolution. 5
Cis-regulatory elements responsive to abscisic acid, methyl jasmonate, anaerobic induction, cytokinin, light, drought, and heat were found in most analyzed B12D promoters. However, the elements involved in the cell cycle, circadian, O2-site, auxin, regulatory elements for low temperature, and gibberellic acid are found only in the promoters of a small number of B12D genes. This diversity of regulatory elements in promoters of B12D genes indicates that some B12D genes are regulated by specific signals, suggesting that members of the B12D family appear to have different roles during plant growth and stress response. The wide range of stimuli that regulate B12D gene expression indicates rabid evolution in the promoter region in comparison with the change in amino acid sequences. This diversification in the promoter of B12D genes supports the suggestion that the functional divergence of B12D gene family might occur as an adaptation to environmental stress. 31
Conclusion
B12D family proteins are B12D domain-containing proteins include several transmembrane proteins. The number and specific function of each member of B12D proteins are not determined. In this study, we retrieved and characterized B12D proteins from 14 species of Viridiplantae. A high degree of conservation was observed among the analyzed B12D protein from higher plants. Despite this strong conservation, some members of the B12D proteins appear to have a different role during plant developmental stages as revealed from gene association analysis, promoter analysis, and digital expression. Furthermore, comprehensive identification of the B12D family members through functional proteomics, cellular localization, and protein-protein interactions is needed for cognizing the specific function of each member of the B12D family.
Supplemental Material
sj-docx-1-evb-10.1177_11769343221106795 – Supplemental material for In Silico Identification and Characterization of B12D Family Proteins in Viridiplantae
Supplemental material, sj-docx-1-evb-10.1177_11769343221106795 for In Silico Identification and Characterization of B12D Family Proteins in Viridiplantae by Zainab M Almutairi in Evolutionary Bioinformatics
Footnotes
Funding:
The author disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported under Project No. 2021/01/17976, from the Deanship of Scientific Research at Prince Sattam bin Abdulaziz University.
Declaration Of Conflicting Interests:
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
