Abstract
Background:
Pharmacogenomics (PGx), forming the basis of precision medicine, has revolutionized traditional medical practice. Currently, drug responses such as drug efficacy, drug dosage, and drug adverse reactions can be anticipated based on the genetic makeup of the patients. The pharmacogenomic data of Pakistani populations are limited. This study investigates the frequencies of pharmacogenetic variants and their clinical relevance among ethnic groups in Pakistan.
Methods:
The Pharmacogenomics Knowledge Base (PharmGKB) database was used to extract pharmacogenetic variants that are involved in medical conditions with high (1A + 1B) to moderate (2A + 2B) clinical evidence. Subsequently, the allele frequencies of these variants were searched among multiethnic groups of Pakistan (Balochi, Brahui, Burusho, Hazara, Kalash, Pashtun, Punjabi, and Sindhi) using the 1000 Genomes Project (1KGP) and
Results:
Our search retrieved (n = 29) pharmacogenetic genes and their (n = 44) variants with high to moderate evidence of clinical association. These pharmacogenetic variants correspond to drug-metabolizing enzymes (n = 22), drug-metabolizing transporters (n = 8), and PGx gene regulators, etc. (n = 14). We found 5 pharmacogenetic variants present at >50% among 8 ethnic groups of Pakistan. These pharmacogenetic variants include CYP2B6 (rs2279345, C; 70%-86%), CYP3A5 (rs776746, C; 64%-88%), FLT3 (rs1933437, T; 54%-74%), CETP (rs1532624, A; 50%-70%), and DPP6 (rs6977820, C; 61%-86%) genes that are involved in drug response for acquired immune deficiency syndrome, transplantation, cancer, heart disease, and mental health therapy, respectively.
Conclusions:
This study highlights the frequency of important clinical pharmacogenetic variants (1A, 1B, 2A, and 2B) among multi-ethnic Pakistani populations. The high prevalence (>50%) of single nucleotide pharmacogenetic variants may contribute to the drug response/diseases outcome. These PGx data could be used as pharmacogenetic markers in the selection of appropriate therapeutic regimens for specific ethnic groups of Pakistan.
Keywords
Introduction
The traditional medical practice has been revolutionized by precision medicine. Pharmacogenomics (PGx) plays a central role in the selection of targeted therapies that underpin precision medicine. The premise is that a patient’s response to available treatment options for a particular disease depends upon genomic variations. 1 Pharmacogenetics and PGx are generally used interchangeably in the literature. However, pharmacogenetics focuses on variations in a single gene, while PGx includes variants at the genomic level. The DNA variations in genes encoding drug-metabolizing enzymes and drug transporters explain the differences in pharmacokinetics and pharmacodynamics in response to therapeutic regimens between individuals. The frequencies of such pharmacogenetic variants differ across the continents of the world. 2 Many efforts have been focused on incorporating PGx information for the therapeutic selection of several drugs. It might be available in low-income countries, in the near future. Pharmacogenomic consortiums play a critical role in the generation of such information. These include the Consortium for the Implementation of Clinical Pharmacogenetics Implementation Consortium (CPIC), the Canadian Pharmacogenomics Network for Drug Safety (CPNDS), and other pharmacogenomics initiatives.3-5 To date, 133 clinical guidelines are available for the selection of the right dose of drug. 6 However, in terms of developing countries, genetic data are missing or scarce. To benefit from the pharmacogenomic revolution in these countries, it is essential to explore the available PGx data of their populations and apply it in clinical phenotypes appropriately.
The present study focuses on the collection of PGx data with high to moderate clinical evidence(s) for therapeutic purposes in the Pakistani population. Pakistan is the sixth most populous country in the world with an estimated population of over 207.74 million. 7 The population is diverse and divided into at least 18 major ethnic groups. 8
In the present study, the Pharmacogenomics Knowledge Base (PharmGKB) database was explored for pharmacogenetic variants, which are correlated with clinical phenotypes in Pakistan. The frequencies of the variants were investigated in 8 ethnic groups (Balochi, Brahui, Burusho, Hazara, Kalash, Pashtun/Pathan, Punjabi, and Sindhi) included in the ALlele FREquency Database (ALFRED) and the 1000 Genomes Project (1KGP).9,10 The purpose is to identify and quantify population-specific pharmacogenetic variants for their translation into clinical practices.
Methods
The general methodology of our study is shown in Figure 1.
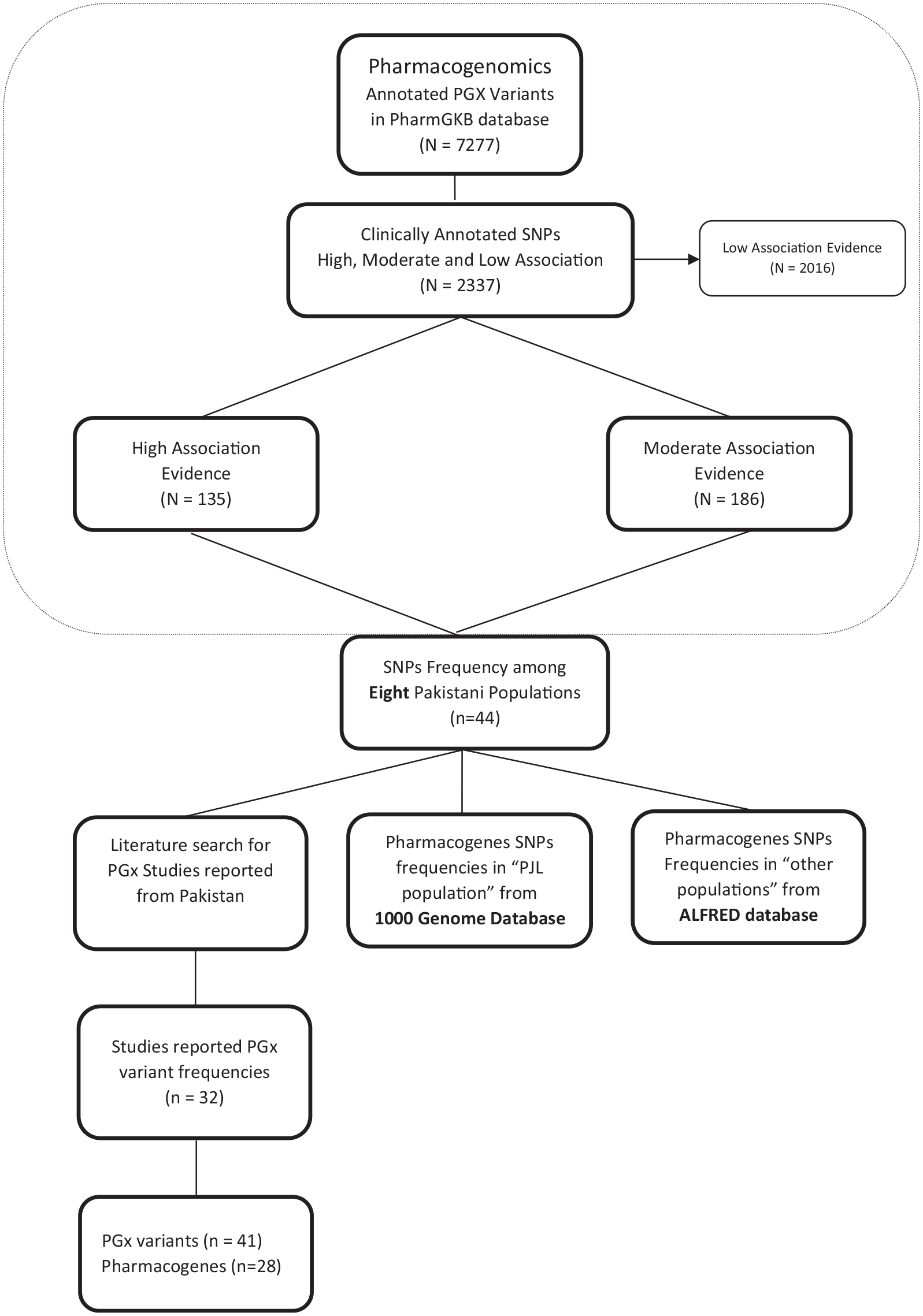
The overall design of the pharmacogenomics study.
Selection of clinical pharmacogenetic variants
A retrospective data analysis was performed for the pharmacogenetic variants associated with clinical outcomes using PharmGKB. The database lists pharmacogenetic variants with clinically annotated datasets. On the basis of clinical relevance, the contribution of variants is classified as high, moderate, or low. The present study was restricted to variants with a high to moderate level of clinical association.
Among the high association variants, level 1A represents the variant-drug combinations, which are currently used in clinical settings as recommended by CPIC and other registered professional societies and endorsed by the PGx guidelines. High association level 1B variants show significant variant-drug combinations with evidence available from more than a single cohort. The moderate association level 2A and 2B variants are the functionally important variants and the variants for which evidence for association is available from certain studies, respectively.6,11
Pharmacogenetic variants (1A + 1B and 2A + 2B) are known for their involvement in the drug-response of certain clinical phenotypes such as heart disease, cancer, etc. The prevalence of these clinically important PGx variants was investigated among ethnic groups of Pakistan.
Data extraction of allele frequencies on the basis of ethnicity for PGx-variants
The ALFRED and 1KGP data were accessed to determine the allele frequencies of selected pharmacogenetic variants (1A + 1B and 2A + 2B). ALFRED database provides the allele frequencies of pharmacogenetic variants in different ethnic groups throughout the world. In the case of Pakistan, data for 7 ethnic groups are available at ALFRED database: Balochi (n = 50), Brahui (n = 50), Burusho (n = 50), Hazara (n = 48), Kalash (n = 50), Pashtun (n = 46), and Sindhi (n = 50). The 1KGP has information on allele and genotype frequencies for a single ethnic group from Pakistan, Punjabis in Lahore (PJL; n = 158).
The allele frequencies of selected pharmacogenetic variants were manually extracted from ALFRED database. For 1KGP database, the Ensembl Rest API Package for R was used for data extraction. 12
The pharmacogenetic variants data of Pakistani population/ethnic groups were also searched in the PubMed database, last accessed on August 19, 2019. The keywords used for searching include genotype, allele, frequency*, Pakistan*. We only selected studies of the human species without language limitations. We only included published articles that contain the allelic and genotypic frequency of pharmacogenomics variants in the Pakistani population.
Principal component analysis
Based on the genotypic frequency of the pharmacogenetic variants, the position of the Pakistani population was investigated among the world’s populations. Therefore, the principal component analysis (PCA) was constructed using the 1KGP browser. Pakistani population was compared with the Asia (South and East Asians), Africa and Europe (Caucasians), and American (Hispanics) populations (list of populations (n = 26); Supplemental Table a). We used PGx genotypic data of the population for the PCA plot, available in 1KGP. Pakistani population; Punjabi, Lahore (PJL) data is only available in 1KGP were used for analysis.
The general steps of the principal component analysis are provided (Supplemental Figure a). Briefly, genotype data from all available world populations were downloaded in the Variant Call Format (VCF) and the Pedigree (PED) file formats from the 1KGP Server. 13 Genotype data were sorted according to the clinical association of pharmacogenetic variants (evidence of high, medium, and low clinical association). We only selected pharmacogenetic variants with high to moderate clinical association using Genome Analysis Toolkits (GATK) Select Variants commands.
The VCF files were converted into Binary Counterpart File (BCF) format using BCF Tools version 1.7. The BCF files were converted to PLINK format (bim, bed, and fam) using Plink 1.9. Finally, the PCA was generated by eigenvalues and eigenvector matrix files using Plink 1.9 (—pca), and the PCA plot was visualized using R-studio.
Results
Clinical pharmacogenetic variants
In the search for pharmacogenetic variants with a known high to moderate effect on an individual’s response to drugs, a total of 321 variants were found. In the present study, we found 44 variants of PGx present in all available ethnic groups of the Pakistani population. These PGx variants belong to drug-metabolizing enzymes (n = 22), drug-metabolizing transporters (n = 8), and others (n = 14; includes transcriptional factors, binding proteins, regulatory variants, intron variants, etc.). The frequency data show the frequency of PGx variants/non-wild alleles among ethnic groups of Pakistan (Tables 1-3).
Frequency data of pharmacogenomic variants* among ethnic groups of Pakistani.
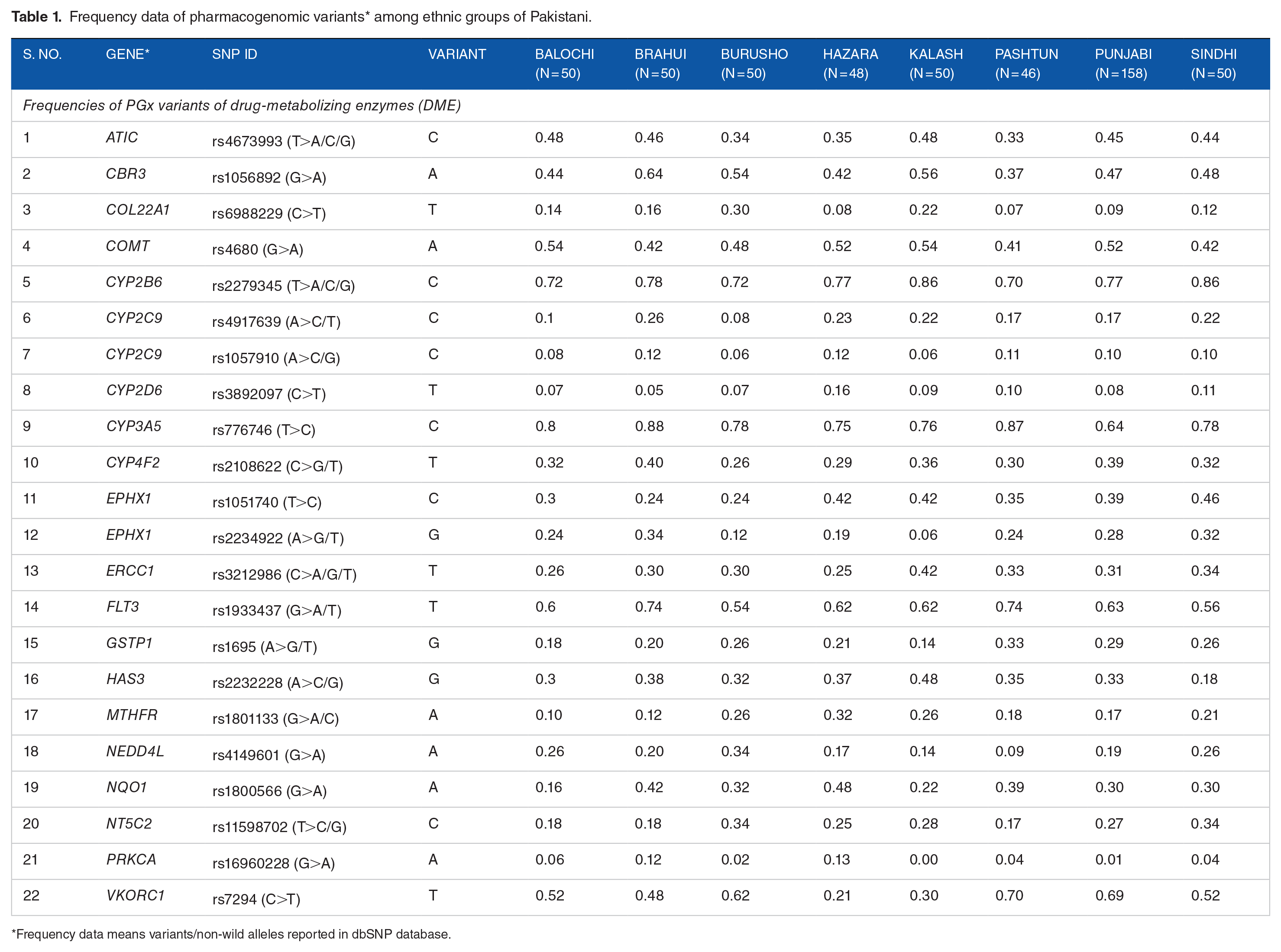
Frequency data means variants/non-wild alleles reported in dbSNP database.
Frequencies of PGx variants of drug metabolizing transporters (DMT).

Frequencies of PGx variants (at gene regulating regions) of drug-metabolizing enzymes/transporters.

The 44 PGx variants are involved in the drug response of 12 clinical phenotypes (Table 4). These include drugs used for heart disease and associated factors (n = 13), cancers (n = 9), atherosclerosis (n = 4), mental health therapy (n = 5), drug addiction therapy (n = 4), antiretroviral therapy (n = 2), and pain management therapy (n = 2). Other variants (n = 5) were associated with diabetes type 2, asthma, hepatitis C, rheumatoid arthritis, and organ transplantation. The allele frequencies of these PGx variants involved in the pharmacokinetics of drugs among Pakistani populations are given (Supplemental Table b).
The effect of pharmacogenomic variants in drugs metabolism for clinical phenotypes.

Importantly, the frequency of 5 pharmacogenetic variants was found to be greater than >50% among the 8 ethnic groups of Pakistan. These include CYP2B6 (rs2279345, C; 70%-86%), CYP3A5 (rs776746, C; 64%-88%), FLT3 (rs1933437, T; 54%-74%), CETP (rs1532624, A; 50%-70%) and an intronic variant of DPP6 (rs6977820, C; 70%-86%) genes, which are involved in drug responses in acquired immune deficiency syndrome, organ transplantation, cancer, heart disease, and psychiatric therapy, respectively.
Published pharmacogenetic reports from Pakistani populations
We found a total of 753 articles using keywords. By selecting studies that contain frequencies of pharmacogenetic variants. Finally, 32 articles were retrieved after removing duplicate studies. These studies were classified as pharmacogenomics (n = 7) and genotype-phenotype association studies (n = 25). These published studies reported PGx variants (n = 41) of 28 genes in the Pakistani population. Most of the pharmacogenetic data (total studies, n = 21) are hospital patient data. However, very limited research institutes (total studies, n = 11) reported PGx data.
The total number of pharmacogenetic studies that reported allele frequencies from Pakistan are given in Table 5. The CYP genes are mostly studied. It includes CYP2D6 (5 variants) and 2 variants of each CYP2C9, CYP1A1, CYP2C6, CYPOR, SCNIA, VKORC1, IL-28B, MTHFR, and XRCC. Interestingly, we found only 6 PGx variants associated with disease/drug response, which are clinically classified as high-moderate evidence. It includes CYP2C9 (rs1057910, C; 14%), CYP2D6 (rs3892097, T; 4.5%), MTHFR (rs1801133, T; 15.7%), GTSP1 (rs1695, G; 25.5%), OPRM1 (rs1799971, G; 14.5%), and IFNL3 (rs8099917, G; 22%). The majority of PGx studies were conducted on cancer (n = 6). It includes breast cancers, bladder cancer, esophageal cancer, and renal cancers. Other PGx studies were include metabolic syndrome, diabetes, heart disease, and hepatitis.
Frequencies of pharmacogenomics variants reported in published literature from Pakistani.
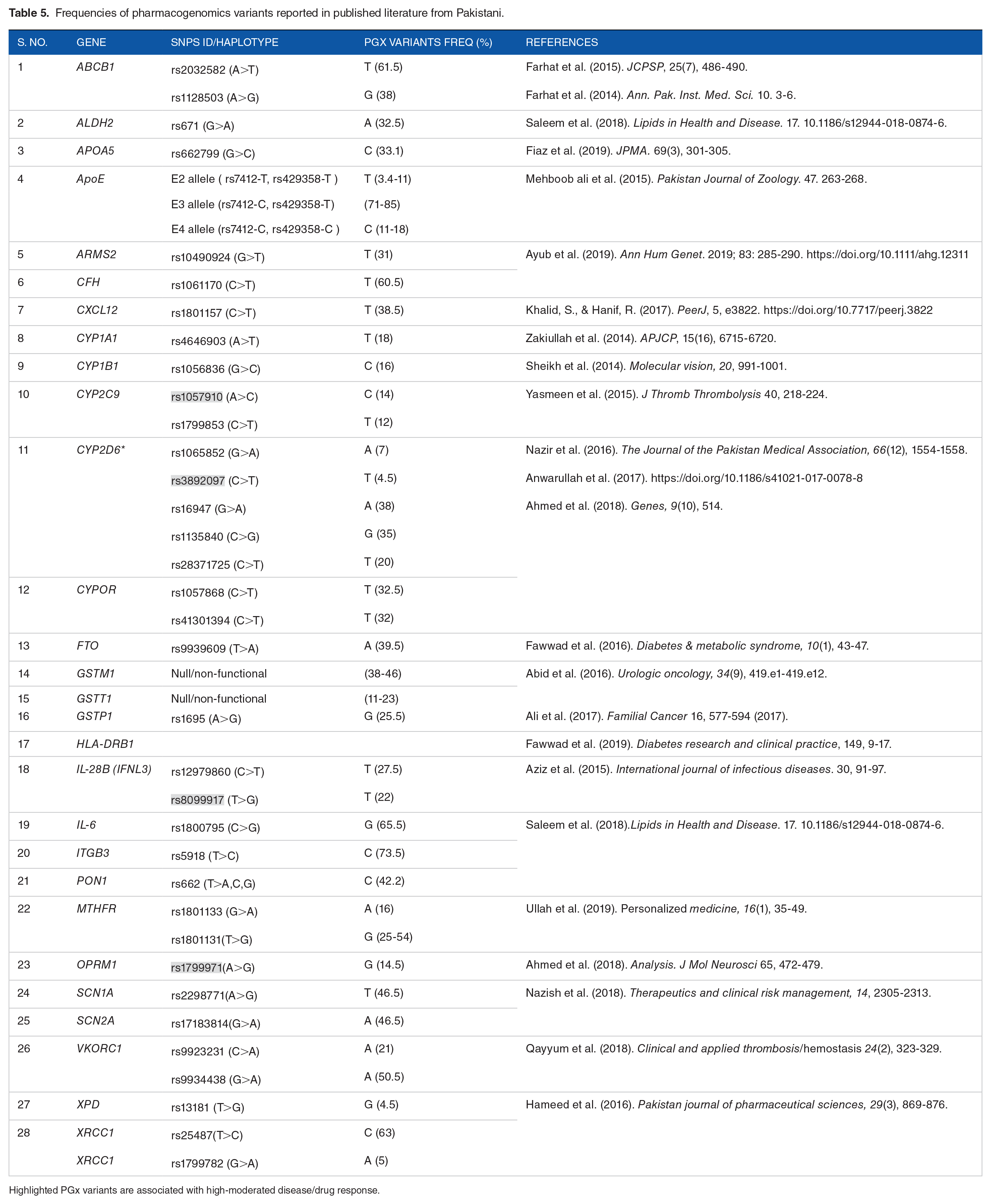
Highlighted PGx variants are associated with high-moderated disease/drug response.
Clustering of Pakistani population-based on PGx variants by principal component analysis
The genotypic frequencies of the pharmacogenetic variants (n = 44) of Pakistan (PJL) from 1KGP were compared with the populations of Africans, Asians (South and East Asians), Europe (Caucasians), and Americas (Hispanics). The PCA results of pharmacogenetic variants show that the Pakistani population lies among South Asian, Hispanic, and Caucasian. Interestingly, both Asian populations (East and South Asian) show differences in terms of genotype frequencies of pharmacogenetic variants. African populations are very distinct from the rest of the world population, as shown in the PCA plot (Supplemental Figure b).
Discussion
This study links clinically important genes with high to moderate associations (1A, 1B, 2A, and 2B) listed in pharmacogenetics databases with their frequencies reported among ethnic groups of Pakistan. We reported 44 PGx variants which correspond to drug-metabolizing enzymes, drug transporters, and others such as gene regulators, etc. Importantly, the frequency of 5 pharmacogenetic variants was found to be greater than >50% among the 8 ethnic groups of Pakistan. These include CYP2B6 (rs2279345, C; 70%-86%), CYP3A5 (rs776746, C; 64%-88%), FLT3 (rs1933437, T; 54%-74%), CETP (rs1532624, A; 50%-70%), and an intronic variant of DPP6 (rs6977820, C; 61%-86%), which are involved in drug responses in acquired immune deficiency syndrome, organ transplantation, cancer, heart disease, and psychiatric therapy, respectively.14-18 These highly prevalent single nucleotide variants (SNVs) contribute to drug metabolisms that can affect the ultimate outcome of the disease.
In addition to 5 SNVs, certain variants have a high-frequency in specific ethnic groups. For example, Kalash population has high-frequency of HAS3 (rs2232228, G; 48%), ERCC1 (rs3212986, T; 42%), and SEMA3C (rs7779029, C; 34%). The Hazara population also showed the highest variant frequency in NQO1 (rs1800566, A; 48%), MTHFR (rs1801133, A; 32%), and CYP2D6 (rs3892097, T; 16%). Similarly, Pashtun population has VKORC1 (rs7294, T; 70%) and GSTP1 (rs1695, G; 33%). The Sindhi and Burusho populations showed a high-frequency of variants for NT5C2 (rs11598702, C; 34%). The Punjabi population showed the highest frequency in VKORC1 (rs7294, T; 69%). Interestingly, the majority of the high-frequency variants were observed in the Brahui population at CBR3 (rs1056892, A; 64%), KIF6 (rs20455, G; 46%), GATM (rs1719247, T; 40%), CYP4F2 (rs2108622, T; 40%), CYP2C9 (rs4917639, C; 26%), CYP2C9 (rs1057910, C; 12%), and SLC28A3 (rs885004, A; 14%). Overall, the highly frequent pharmacogenetic variants were found mostly in Brahui, Kalash, and Hazara populations. On the other hand, Burusho population displays the lowest frequency of variants among all ethnic groups CYP4F2 (rs2108622, T; 26%), CYP2C9 (rs4917639, C; 8%), CYP2C9 (rs1057910, C; 6%), and GP1BA (rs6065, T; 4%). Thus, the presence of either high-frequency variants or retained ancestral alleles may have been due to natural selection based on their geographical location, environmental, and other factors. This PGx knowledge may be used to anticipate which ethnic groups are likely to respond to specific therapeutic drugs.
Recently, a significant increase in alleles of pharmacogenetic variants was reported in healthy Pakistani populations of CYP450 1A2, 2B6, 2C19, 3A5, ALDH3A1, GSTM1, ABCB1, and ABCC2. 19 This study shows a significant difference in the prevalence of variants of drug-metabolizing enzymes and drug transporters compared to other ethnic groups. However, the study had a very low sample size from the main ethnic groups such as Punjabi (n = 8; 5.2%), Pashtun (n = 5; 3.2%), Sindhi (n = 10; 6.5%), and Balochi (not available).
Cardiovascular diseases (CVDs) are the leading cause of death worldwide. The WHO estimated 17.9 million people died from CVDs, 85% of them due to heart attack and strokes. 20 Pakistani populations are at the highest risk of coronary heart disease. According to WHO estimates, the proportional mortality from CVD is 29% in Pakistan. 21 Atherosclerosis is the main cause of heart attacks, stroke, peripheral vascular diseases, etc. Hypertension and dyslipidemia/elevated cholesterol levels are the major contributing factors to atherosclerosis and heart disease. 22 Most variants of PGx (n = 13) are involved in heart disease and associated factors such as anticoagulants, 3-hydroxy-3-methylglutaryl coenzyme A (HMG-CoA) reductase inhibitors, and lipid-lowering drugs. It includes VKORC1 (rs7294, T); CYP4F2 (rs2108622, T); CALU (rs339097, G), CYP2C9 (rs4917639, C/T and rs1057910, C/G) in warfarin pharmacokinetics.23-26 The CETP variants (rs1532624, A) and KIF6 (rs20455, G), are involved in HMG-CoA reductase inhibitors (cholesterol synthesis) and lipid-lowering drugs, respectively.17,27 However, GATM (rs1346268, C and rs1719247, G/T), HAS3 (rs2232228, C/G), CBR3 (rs1056892, A), and SLC28A3 (rs885004, A) with myopathy/cardiomyopathy are reported.28-31
The VKORC1 allele variant (rs7294, T) was found to be the highest in Pashtun and Punjabi (70%) populations and the lowest in Hazara (21%) and Kalash (30%) populations. Allele frequencies for CYP4F2 (rs2108622, T) and CYP2C9 (rs4917639, C) vary from 26% to 40% and 8% to 26% among ethnic groups, respectively. Interestingly, the frequency of CYP2C9 (rs1057910, C) was found to be low (6%-12%). However, CALU (rs339097, G) is not found in any of the Pakistani populations. The frequency of variants of CETP (rs1532624, A) is 50% to 70% in our population. The frequency of KIF6 allele variants (rs20455, G) is 20% to 46% among ethnic groups. The variant frequencies of GATM (rs1346268, C) and (rs1719247, T) are 22% to 54%. The allele frequencies of variant HAS3 (rs2232228, G), CBR3 (rs1056892, A), and SLC28A3 (rs885004, A) are 18% to 48%, 37% to 64% and 4% to 14%, respectively. The GP1BA (rs6065, T) have a decreased risk of aspirin resistance. 32 The frequency of GP1BA (rs6065, T) variants for aspirin metabolism is 4% to 12% in the Pakistani population.
Atherosclerosis
We found 4 variants of pharmacogenetics involved in atherosclerosis; NEDD4L (rs4149601, A), ABCG2 (rs2231142, T), YEATS4 (rs7297610, T), and PRKCA (rs16960228, A). These variants showed an association with drugs prescribed for blood pressure and cholesterol. Importantly, the variants NEDD4L (rs4149601, A) and YEATS4 (rs7297610, T) showed a poor response to hydrochlorothiazide.33,34 The frequency of NEDD4L (rs4149601, A) and YEATS4 (rs7297610, T) is 9% to 34% and 0% to 8%, respectively. On the other hand, PRKCA (rs16960228, A) showed an increased reduction in diastolic blood pressure. 35 The ABCG2 variants (rs2231142, C/T) showed increased plasma rosuvastatin concentrations and better response to treatment. 36 The frequency of PRKCA (rs16960228, A) and ABCG2 (rs2231142, /T) is 0% to 13% in Pakistanis. The alternative allele frequencies of the drugs related to atherosclerosis showed the presence of very low variant frequencies NEDD4L (rs4149601, A) 9% to 34%, PRKCA (rs16960228, A), and ABCG2 (rs2231142, T) 0% to 13% and YEATS4 (rs7297610, T) 0% to 8% among Pakistani populations.
Cancer is the second leading cause of death (~9.6 million) worldwide. In Pakistan, the number of new cancer cases is 0.17 million and the number of deaths is 0.11 million, as reported by The Global Cancer Observatory. 37 Based on incidence and death by cancer type, breast (19%) and lip/oral (11%) cancers are more common, followed by lungs (5.6%), esophagus (4.6%), and leukemia (4%). We found 9 pharmacogenetic variants that are involved in cancer drug metabolism. These are FLT3 (rs1933437, A), FCGR2A (rs1801274, C/G), ERCC1 (rs3212986, T), NQO1 (rs1800566, A), GSTP1 (rs1695, G), NT5C2 (rs11598702, C), MTHFR (rs1801133, A), SEMA3C (rs7779029, C), and CYP2D6 (rs3892097, T).
The FCGR2A (rs1801274, variant C or G) showed a reduced/low drug response of Trastuzumab—anti-HER2. 38 The highest and lowest allele frequency of FCGR2A (rs1801274, C) was reported in Kalash (54%) and Burusho (32%) populations, respectively. The NT5C2 variant (rs11598702, C) has increased clearance levels of gemcitabine drugs. 39 Sindhi and Burusho populations have the highest NT5C2 (rs11598702, C) frequency of 34%, and the lowest frequency (17%) in Pashtun populations. The ERCC1 (rs3212986, A) has been reported to reduce the risk of nephrotoxicity. 40 Other variants of pharmacogenetics FLT3 (rs1933437, A), NQO1 (rs1800566, A), GSTP1 (rs1695, G), MTHFR (rs1801133, A), SEMA3C (rs7779029, C), and CYP2D6 (rs3892097, T) cause adverse drug response such as increased risk of toxicity, leukopenia, thrombocytopenia, neutropenia, and relapses.16,41-44 Allele frequencies ranges are as 54% to 74% for FLT3 (rs1933437, A); 16% to 48% for NQO1 (rs1800566, A); 14% to 33% for GSTP1 (rs1695, G); 10% to 32% for MTHFR (rs1801133, A); 8% to 34% for SEMA3C (rs7779029, C), and 5% to 16% for CYP2D6 (rs3892097, T) among ethnic groups. Interestingly, the Kalash populations showed the highest variant frequency of 54% for FCGR2A (rs1801274, C), 42% for ERCC1 (rs3212986, T), and 34% for SEMA3C (rs7779029, C). The Hazara populations showed 48% NQO1 (rs1800566, A), 32%MTHFR (rs1801133, A), and 16% CYP2D6 (rs3892097 T,) respectively. The Pashtun population also showed the highest variant frequency of 74% for FLT3 (rs1933437, T) and 33% for GSTP1 (rs1695, G).
Mental health
A total of 5 clinically significant pharmacogenetic variants were found. These are DPP6 (rs6977820, A/C), GRIK4 (rs1954787, C), EPHX1 (rs1051740, C and rs2234922, G/T), and DRD2 (rs1799978, C) which are associated with drug metabolism, response or recommended dosage to antipsychotics (neuroleptics or major tranquilizers), antidepressants, carbamazepine, and risperidone.18,45-48 The range of alternative allele frequencies for psychotic disorders accounts for 70% to 86% for DPP6 (rs6977820, C), 46% to 67% for GRIK4 (rs1954787, C), 24% to 46% for EPHX1 (rs1051740, C), and 6% to 34% (rs2234922, G/T) and 0% to 12% DRD2 (rs1799978, C) among Pakistani ethnic groups.
Addiction
Treatment of substance abuse
We found 4 variants of pharmacogenetics associated with response to drug addiction therapy. These are OPRD1 (rs678849, T), CHRNA3 (rs578776, A and rs1051730, A), and OPRM1 (rs1799971, G) for buprenorphine, nicotine, and naloxone, respectively.49-51 The range of allele frequencies of OPRD1 (rs678849, T), CHRNA3 (rs578776 A), CHRNA3 (rs1051730 A), and OPRM1 (rs1799971 G) is 44% to 69%, 35% to 74%, 21% to 48%, and 8% to 37%, respectively.
Acquired immuno-deficiency syndrome (AIDs)
The CYP2B6 variant (rs2279345, C) and CCHCR1 (rs746647, G) are 2 pharmacogenetic variants involved in drug response (efavirenz and nevirapine) to HIV treatment. The frequency of the CYP2B6 variant allele (rs2279345, C) is quite high (70%−86%) among all ethnic groups. This may reflect that the Pakistani population (three-fourths) is at risk of low efavirenz levels, particularly (86%) Sindhi and Kalash populations. However, the allele frequency of the CCHCR1 allele (rs746647, G) is 18% to 33%, the risk of adverse risk reaction with nevirapine.
Pain management
Pharmacovariants in COMT (rs4680, A) and (rs2952768, C) have shown an association with opioid drug metabolism.52,53 The variant frequency COMT (rs4680, A) and (rs2952768, C) were found to be 41% to 54% and 28% to 56% in all ethnic groups. The highest frequency of variants of (rs4680, A) is found in the Kalash (54%) and Pashtun (41%) populations. The lowest frequency of variants of COMT (rs2952768, C) is found in the Kalash (28%) and Baloch (62%) populations.
Diabetes
A pharmacogenetic variant of TCF7L2 (rs7903146, G/T) has shown association with glucose metabolism. Patients with ancestral allele C have been reported to be better able to maintain hemoglobin A1c (HbA1c) and fasting plasma glucose levels with sulfonamides compared to patients with the variant allele (T). 54 The frequency of allele variants (rs7903146, T) is 17% to 50% among the Pakistani population. The lowest frequency of variant alleles is reported in the Hazara population (17%) and the highest (50%) in Brahui population.
Asthma
Salbutamol/albuterol is the common drug used for asthma (bronchodilator). It reduced the response of the bronchodilator (FEV1) in patients with an ancestral C allele compared to patients with collagen (COL22A1) gene variant alleles (rs6988229, T). 55 In Pakistan, the frequency of COL22A1 gene variant alleles (rs6988229, T) ranges from 7% to 30%, with the lowest (7%) in Pashtun population and the highest (30%) in Burusho population.
Rheumatoid arthritis
Methotrexate is a chemotherapy agent used to suppress the immune system. Studies have shown an association of the ATIC (5-aminoimidazole-4-carboxamide ribonucleotide formyl transferase/IMP cyclohydrolase) polymorphism (rs4673993, C) with rheumatic drug therapy. 56 Patients with rheumatoid arthritis with variant C treated with methotrexate showed a better response compared to patients with the ancestral T allele. The frequency of variants of ATIC is 33% to 48% in all ethnic groups of Pakistan. The highest frequency (48%) is found in Balochi and Kalash populations and the lowest (33%) in Pashtun population.
Hepatitis C
Peg-interferon alfa and ribavirin are the important drugs used in the treatment of liver disease (HBV and HCV). Interestingly, HCV genotypes and interferon-lambda SNPs contributed to the virological response to the drug. Single interferon-lambda SNPs (INFL3 rs8099917, G) have shown an association with the sustained virologic response in genotype 1, 3, and 4 infected patients. 57 Patients with the variant allele (G) may have a decreased response (lower sustained viral response) to peg-interferon alfa and ribavirin therapy with HCV genotype 1 compared to patients with the ancestral (T) allele. 58 The frequency of INFL3 (G) variants is 10% to 28% among ethnic groups. Baloch showed the highest frequency of variants (28%) and the lowest were found in Hazara (10%) compared to other ethnic populations.
Organ transplantation
The pharmacogenetic variant CYP3A5 (rs776746) allele has shown evidence of a strong association with immunosuppressive drug metabolism (tacrolimus). Transplant patients with the variant allele CYP3A5 (rs776746, C) have shown a reduction in tacrolimus metabolism, resulting in increased plasma drug levels. 59 Therefore, patients are at high risk for drug-related toxicity and need a low dose of tacrolimus. The overall frequency of CYP3A5 (rs776746, C) was found to be much higher (64%-88%) in all ethnic groups, compared to the values reported in other South Asian populations. The Punjabis population has the lowest prevalence (64%) and Brahui has the highest prevalence (88%) of this variant. This indicates that two-thirds of Pakistan’s population would exhibit poor metabolizers of immunosuppressant drugs.
The genotypic data of 44 PGx variants are reported for Punjabi at 1KGP. The Punjabis constitute about half (45%) of the total Pakistani population. The Punjabi identity is primarily linguistic, and all those who speak this Indo-Aryan language as their first language are classified as Punjabis. 59 The Punjabis are further divided into many castes, tribes, and clans, residing in the most populous province of Pakistan, Punjab. Due to the large admixture found in this area, it often makes it difficult to classify indigenous Punjabi castes/tribes. Therefore, using the Punjabi population from Lahore in the principal component analysis as a reference for the Pakistani population may not truly represent the total pool of PGx in our country. The availability of other population pharmacogenetic data would better locate the position of Pakistani ethnic groups among the world population.
The principal component analysis plot shows that the Pakistani population is located among the South Asian population groups. Interestingly, the frequencies of pharmacogenetic variants in South Asia were distant from those in East Asia. The population cluster pattern also showed that African populations are distinct from the rest of the world’s populations, indicating high ancestral alleles. Human migration is believed to have started from Africa to Europe and Asia. Thus, low variant frequencies are present in Africans followed by Europe and Asia. Recently, it has been reported that due to the presence of the variant allele CYP3A5 (rs776746, C) in Asians, the tacrolimus concentration/dose (Co/D) ratio is significantly lower in the CYP3A5 expresser group compared to the non-expresser in Asian and European populations at any posttransplant period. 60
There are certain limitations in our study. First, we only selected PGx variants with high to moderate clinical evidence and their frequency among Pakistani populations (Punjabi, Sindhi, Pashtun, Balochi, Hazara, Kalash, Burusho, and Brahui). Low clinical evidence PGx variants were excluded from the study; primarily due to limited studies. Interestingly, it should be noted that most of the PGx-based studies are conducted on non-Asian populations. Further studies involving Asian populations may expand our knowledge about clinically significant variants.
To date, the PGx data for the Pakistani population are very limited. This may be due to the lack of trained manpower and financial constraints. 61 Additionally, there is no national database of pharmacogenetics available in Pakistan. Therefore, this study highlights clinically important pharmacogenetic variants and their frequencies in Pakistani populations and may provide targeted therapeutic drugs based on the genetic makeup of the patients. Ethnic specific PGx variants knowledge may help in selecting the right therapeutic drugs and its dosage for specific clinical phenotypes. This will impact on translational of PGx knowledge into clinical practice.
Supplemental Material
sj-docx-3-evb-10.1177_11769343221095834 – Supplemental material for The Prevalence of Pharmacogenomics Variants and Their Clinical Relevance Among the Pakistani Population
Supplemental material, sj-docx-3-evb-10.1177_11769343221095834 for The Prevalence of Pharmacogenomics Variants and Their Clinical Relevance Among the Pakistani Population by Abdul Rafay Khan, Sayed Hajan Shah, Sadia Ajaz, Sadaf Firasat, Aiysha Abid and Ali Raza in Evolutionary Bioinformatics
Supplemental Material
sj-docx-4-evb-10.1177_11769343221095834 – Supplemental material for The Prevalence of Pharmacogenomics Variants and Their Clinical Relevance Among the Pakistani Population
Supplemental material, sj-docx-4-evb-10.1177_11769343221095834 for The Prevalence of Pharmacogenomics Variants and Their Clinical Relevance Among the Pakistani Population by Abdul Rafay Khan, Sayed Hajan Shah, Sadia Ajaz, Sadaf Firasat, Aiysha Abid and Ali Raza in Evolutionary Bioinformatics
Supplemental Material
sj-pdf-2-evb-10.1177_11769343221095834 – Supplemental material for The Prevalence of Pharmacogenomics Variants and Their Clinical Relevance Among the Pakistani Population
Supplemental material, sj-pdf-2-evb-10.1177_11769343221095834 for The Prevalence of Pharmacogenomics Variants and Their Clinical Relevance Among the Pakistani Population by Abdul Rafay Khan, Sayed Hajan Shah, Sadia Ajaz, Sadaf Firasat, Aiysha Abid and Ali Raza in Evolutionary Bioinformatics
Supplemental Material
sj-xlsx-1-evb-10.1177_11769343221095834 – Supplemental material for The Prevalence of Pharmacogenomics Variants and Their Clinical Relevance Among the Pakistani Population
Supplemental material, sj-xlsx-1-evb-10.1177_11769343221095834 for The Prevalence of Pharmacogenomics Variants and Their Clinical Relevance Among the Pakistani Population by Abdul Rafay Khan, Sayed Hajan Shah, Sadia Ajaz, Sadaf Firasat, Aiysha Abid and Ali Raza in Evolutionary Bioinformatics
Footnotes
Funding:
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests:
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
AR and AK designed the study, AK, AR, SA, SHS, and SF search clinical data and analyze ethnic data. AK wrote the manuscript. SA, AA, and AR made substantial edits to the manuscript. All authors reviewed and approved the manuscript.
Disclaimer
It may please be noted that neither the manuscript nor any part of this manuscript is under consideration for publication elsewhere.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
