Abstract
The CCAAT/enhancer binding protein (C/EBP) transcription factors (TFs) regulate many important biological processes, such as energy metabolism, inflammation, cell proliferation etc. A genome-wide gene identification revealed the presence of a total of 99 C/EBP genes in pig and 19 eukaryote genomes. Phylogenetic analysis showed that all C/EBP TFs were classified into 6 subgroups named C/EBPα, C/EBPβ, C/EBPδ, C/EBPε, C/EBPγ, and C/EBPζ. Gene expression analysis showed that the C/EBPα, C/EBPβ, C/EBPδ, C/EBPγ, and C/EBPζ genes were expressed ubiquitously with inconsistent expression patterns in various pig tissues. Moreover, a pig C/EBP regulatory network was constructed, including C/EBP genes, TFs and miRNAs. A total of 27 feed-forward loop (FFL) motifs were detected in the pig C/EBP regulatory network. Based on the RNA-seq data, gene expression patterns related to FFL sub-network were analyzed in 27 adult pig tissues. Certain FFL motifs may be tissue specific. Functional enrichment analysis indicated that C/EBP and its target genes are involved in many important biological pathways. These results provide valuable information that clarifies the evolutionary relationships of the C/EBP family and contributes to the understanding of the biological function of C/EBP genes.
Keywords
Introduction
The C/EBP family includes important regulators that activate or inhibit gene transcription in many tissues and is a subfamily of the basic leucine zipper (bZIP) transcription factor superfamily. 1 The C/EBP family consists of 6 genes: C/EBP alpha (C/EBPα), C/EBP beta (C/EBPβ), C/EBP delta (C/EBPδ), C/EBP epsilon (C/EBPε), C/EBP gamma (C/EBPγ), and C/EBP zeta (C/EBPζ). 2
The first C/EBP protein was identified in rat liver nuclei. 3 Five additional C/EBP TFs had been identified in humans and rats. 2 Advances in the sequencing techniques resulted in the identification of the C/EBP genes in various species and investigation of the evolutionary relationship among the C/EBP genes. Qiu et al 4 identified the C/EBPα, C/EBPβ, and C/EBPδ genes genome-wide in 20 vertebrate genomes, including Atlantic salmon (Salmo salar), Japanese frog (Nanorana pleskei), southern painted turtle (Chrysemys picta bellii), duck (Anas platyrhynchos), and human (Homo sapiens), and analyzed the evolutionary relationships among the C/EBPα, C/EBPβ, and C/EBPδ genes. Thus, understanding of the evolution and function of the C/EBP genes requires genome-wide comparative studies.
Transcription factors play an important regulatory role at the transcriptional level by activating or repressing target genes to regulate gene expression. 5 The C/EBP TFs in the vertebrates regulate many important biological processes, such as cell proliferation, differentiation, signal transduction, inflammation, and energy metabolism.1,4 C/EBPα, C/EBPβ, and C/EBPδ are involved in the regulation of adipocyte differentiation and mammary development.6,7 In the pulmonary epithelium, C/EBPα regulates proliferation and differentiation dependent gene expression, whereas C/EBPβ and C/EBPδ regulate the expression of differentiation markers and are involved in the responses to injury and hormones. 8
Pig and human are highly similar in anatomy, genetics, immunology and physiology, which makes the pig become a valuable animal model to study human complex diseases. 9 It has been reported that C/EBPα, C/EBPβ, and C/EBPδ genes regulate cell proliferation and differentiation in pigs’ liver, fat and muscle tissue, respectively.10-17 In addition, C/EBPβ and C/EBPδ genes regulate the proliferation and differentiation of pig granulosa cells too. 18 The C/EBPζ gene regulates endoplasmic reticulum stress-mediated apoptosis which has been found in many tissues of pigs.19-24 But the expression pattern of C/EBP gene family in pigs’ tissues also has not been systematically analyzed.
In order to further study the function of the pig C/EBP family members, we identified C/EBP genes in pig and others 19 eukaryotic organisms, such as fruit fly, mouse, human and so on, and analyzed physicochemical properties of the proteins, conservative motifs, gene structure, and phylogenetic relationships. We also analyzed the expression patterns of the porcine C/EBP genes, and constructed a transcriptional regulatory network containing the C/EBP genes to investigate the functions of the porcine C/EBP genes. Our study found the C/EBP gene family was divided into 6 main branches, and C/EBPs has high sequence conservation and functional polymorphism. Except C/EBPε gene, the C/EBP genes were expressed ubiquitously with inconsistent expression patterns in various pig tissues. This study provides a comprehensive analysis of the pig C/EBP gene family and important information for subsequent investigation of the functions in pig.
Materials and Methods
Identification of C/EBP TFs
The General Feature Format Version 3 (GFF3) profiles and the genome and protein sequence files of 20 eukaryotic organisms were downloaded from the ENSEMBL database (http://asia.ensembl.org/index.html). 24 The eukaryotic organisms included nematode (Caenorhabditis elegans), fruit fly (Drosophila melanogaster), sea squirt (Ciona intestinalis), zebrafish (Danio rerio), tropical xenopus (Xenopus tropicalis), Chinese soft-shell turtle (Pelodiscus sinensis), chicken (Gallus gallus), platypus (Ornithorhynchus anatinus), cat (Felis catus), dog (Canis familiaris), horse (Equus caballus), pig (Sus scrofa), cow (Bos taurus), goat (Capra hircus), sheep (Ovis aries), mouse (Mus musculus), rat (Rattus norvegicus), macaque (Macaca mulatta), chimpanzee (Pan troglodytes), and human (Homo sapiens). The HMMER (v3.0) software 25 was used to build the Hidden Markov Model (HMM) of the C/EBP family. The hmmsearch tool search for all putative TFs of the C/EBP family was performed (cutoff value <1E-20). The Conserved Domain Database of NCBI (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) 26 and the SMART tool (http://smart.embl-heidelberg.de/) 27 were used to determine the conserved domains of each C/EBP protein. The longest protein of each C/EBP TF was retained.
Physicochemical parameters, exon-intron structure, and motif analysis
ProtParam (http://web.expasy.org/protparam/) 28 was used to analyze the physicochemical parameters of the C/EBP proteins, including molecular weight (MW), aliphatic index, isoelectric point (pI), and grand average of hydropathicity (GRAVY). The conserved motifs of the C/EBP proteins were identified by the online platform MEME (http://meme-suite.org/). 29 The exon and intron structures of the C/EBP genes were obtained from the ENSEMBL gene annotation information. The diagrams of the gene structures were generated using the Gene Structure Display Server (GSDS v2.0) (http://gsds.gao-lab.org/). 30
Phylogenetic analysis
In this study, we used MEGA (v10.0) 31 to construct Maximum likelihood (ML), Neighbor-Joining (NJ) and Minimum-Evolution (ME) phylogenetic tree of C/EBP TFs based on Jones-Taylor-Thornton model and Games distribution, with 1000 replicate bootstrap tests, and set a cut-off bootstrap value of 65 to define the clades. Representations of the phylogenetic tree were constructed using the iTOL tool (https://itol.embl.de/). 32
Construction of the transcriptional regulatory networks of the pig C/EBP genes
The Position Weight Matrix (PWM) of transcription factors for Sus scrofa was downloaded from the cisbp database (http://cisbp.ccbr.utoronto.ca/). 33 The genomic sequences 2 kb upstream of the translation start site (TSS) and 3ʹ-UTR were extracted by the R package biomaRt 34 for all coding genes. The TFBSTools package of R language 35 was used to predict the target genes of C/EBPs and the transcription factors regulating C/EBP genes with a threshold relScore value of 0.85. The sequences of miRNAs were downloaded from the miRBase database (http://www.mirbase.org/). 36 The miRanda software 37 was used to predict the miRNA targets with a threshold TotScore value of 120. The Cytoscape (v3.7.2) software 38 was used to visualize the gene regulatory network, and the NetworkAnalyzer tool was used to calculate the network topology properties, including the clustering coefficient, network centralization, and network heterogeneity. The R software package basicTrendline (Mei and Yu, 2018) 39 was used to analyze the network node degree distribution.
Expression analysis of the pig C/EBP genes and target genes
The raw RNA sequencing (RNA-seq) data obtained in 27 adult Duroc pig tissue samples were downloaded from the NCBI Sequence Read Archive with the BioProject number PRJNA392949. 40 The tissues included fat, thyroid, lymph, ovary, uterus, breast, spleen, lung, liver, placenta, adrenal gland, brain, gall bladder, gut, heart, kidney, longissimus dorsi, nasopharynx, oesophagus, pancreas, prostate, retina, salivary gland, spinal cord, stomach, testis, and uterus. According to description of Zhao et al, purification of the total RNA from the mixture of equally unrelated pig pool tissues was performed.
After quality control step with FastQc (v0.11.8) (http://www.bioinformatics. babraham.ac.uk/projects/fastqc/), clean data were mapped and genome indexed with Hisat 0.1.6-beta 64-bit 41 to the pig genome (Sus scrofa 11.0). To obtain the expression levels of the genes across 27 tissues, the fragments per kilobase of exon model per million mapped reads (FPKM) values were calculated using Stringtie (v1.0.4, Linux ×86_64). 42 The TBtools software 43 was used to visualize the heatmaps of the gene expression profiles in 27 tissues.
The tissue specificity index (τ) of the C/EBP genes and target genes for C/EBP was also investigated. The tissue specificity index is defined as
The dN and dS analysis of the C/EBP genes and target genes
The data on the nonsynonymous (dN) and synonymous (dS) substitution rates between the pig and human sequences were downloaded from the Ensembl database. 24 The dN/dS and dN+dS values of each gene were calculated. In addition, the dN/dS and dN+dS mean values of the target genes of each C/EBP gene were compared using Kolmogorov-Smirnov (KS) test.
Functional enrichment analysis
The Database for Annotation, Visualization, and Integrated Discovery (DAVID) (https://david.ncifcrf.gov/) 45 was used to annotate the functions of C/EBPs and their target genes with all protein-encoding genes on pig genome as the background genes set, which includes gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis.
Results
Identification of C/EBP genes in the eukaryotes
A total of 99 C/EBP genes were identified in pig and 19 eukaryote genomes. The number of C/EBP genes in each species was slightly different as shown in Table 1. In total, the C/EBP family included C/EBPα, C/EBPβ, C/EBPδ, C/EBPε, C/EBPγ, and C/EBPζ. In particular, the C/EBPα and C/EBPγ genes of all species are located on the same chromosome within a distance of less than 100 kb, except for nematode, fruit fly, Xenopus tropicalis (see Table S1).
Genomic distribution of C/EBP genes on eukaryote genomes.

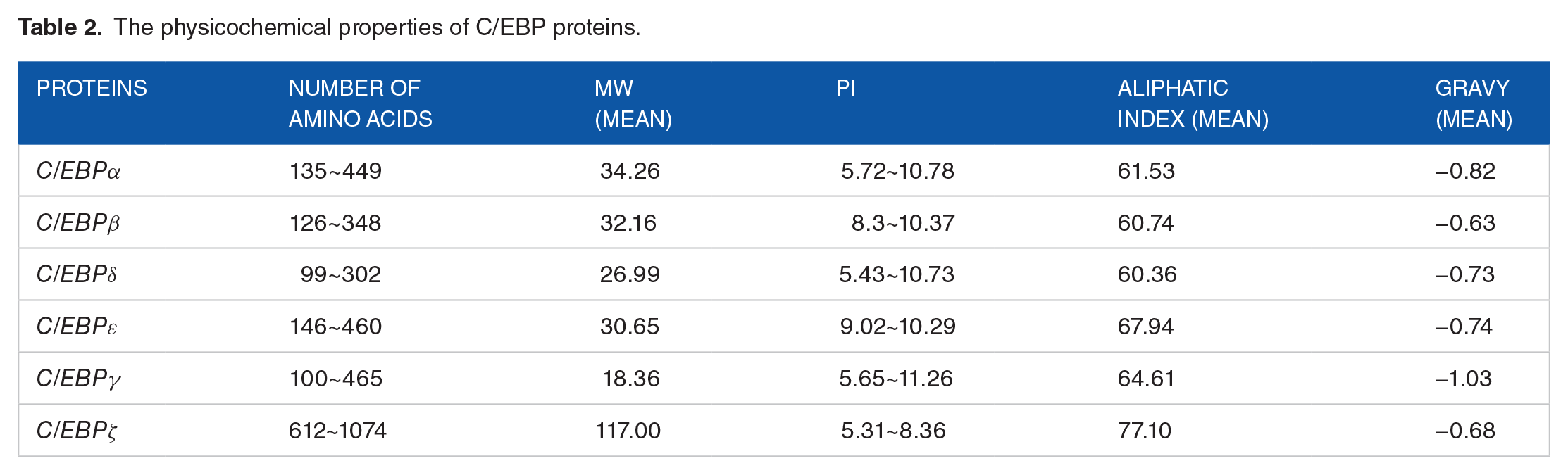
Various physicochemical properties of each C/EBP TF were calculated and are shown in Table 2 and Table S1. The C/EBPζ protein contains the highest number of amino acids, and the C/EBPγ protein consists of the least number of amino acids. The aliphatic index, a measure of thermostability, ranged from 60.36 to 77.10. There are significant differences (P < 2e−16) in the aliphatic index among C/EBP TFs. The GRAVY values of C/EBP proteins are negative indicating hydrophilic properties. The GRAVY values vary from −1.7940 to −0.2800. The pI values of C/EBPβ and C/EBPε proteins in each species are higher than 7, and the pI values of C/EBPζ are less than 7. The pI values of C/EBPγ proteins in each species are higher than 7, except for nematode. The C/EBPα protein of nematode, sea squirt, zebrafish is acidic, and that of other species are alkaline. The C/EBPδ protein is acidic in 1/3 species and alkaline in the remaining 2/3 species.
The physicochemical properties of C/EBP proteins.
Phylogenetic relationship analysis
To analyze the phylogenetic relationships between 99 C/EBP genes in 20 species, an unrooted maximum likelihood phylogenetic tree was constructed (Figure 1). All C/EBP TFs were classified into 6 groups and was named C/EBPα, C/EBPβ, C/EBPδ, C/EBPε, C/EBPγ, and C/EBPζ. C/EBP genes were detected in most vertebrates indicating that the C/EBP family members originated in the early stage of vertebrate evolution. In each clade, C/EBP genes of the same order tend to cluster together indicating higher similarity to each other than to other orders of C/EBP genes. C/EBP orthologs of the order primates includes human, macaque, and chimpanzee; the order artiodactyl includes pig, cattle, and goat; the order carnivore includes cat and dog; and the order rodents (rat and mouse) genes are clustered together. We also constructed the un-rooted neighbor-joining tree and minimum-evolution tree shown in Figures S1 and S2. The topological structure of NJ and ME tree were similar with ML tree.

The unrooted maximum likelihood phylogenetic tree of the C/EBP proteins from 20 eukaryotes, with 1000 bootstrap replicates. The numbers on the lines indicate bootstrap support greater than 65%. All C/EBP TFs were classified into 6 groups and was named C/EBPα, C/EBPβ, C/EBPδ, C/EBPε, C/EBPγ, and C/EBPζ.
To investigate the structural features of the C/EBP members, the gene structure and conserved motifs were evaluated by the phylogenetic analysis, as shown in Figure 2. The number of exons in C/EBPζ genes, which contained 15 or 16 exons, was higher than that in the other C/EBP genes. Most of the C/EBPα, C/EBPβ, C/EBPδ, C/EBPε, and C/EBPγ genes contained one or two exons. A total of 10 conserved motifs were identified in C/EBP proteins. All C/EBP TFs contain motif 2, motif 3, and motif 5. Motifs 10 is unique motifs in C/EBPζ proteins and may be associated with the clade-specific functions of the C/EBP proteins (see Table S2).

The maximum likelihood phylogenetic tree, exon-intron structure, and conserved motifs of C/EBP genes. (A) The phylogenetic tree was constructed based on the C/EBP proteins from 20 eukaryotes. (B) Exon-intron structures of C/EBP genes. Yellow boxes indicate translated regions, blue boxes indicate 5′-UTR and 3′-UTR regions, and black lines indicate introns. (C) The conserved motifs of C/EBP proteins. The motifs, numbered 1–10, are displayed in different colored boxes.
Expression analysis of the C/EBP genes
The expression of C/EBP genes was compared in 27 adult Duroc pig tissues. The tissue specificity index (τ) of the C/EBPα, C/EBPβ, C/EBPδ, C/EBPε, C/EBPγ, and C/EBPζ are 0.90, 0.73, 0.68, 0.92, 0.52 and 0.45, respectively. The τ value varies from 0 for housekeeping genes to 1 for tissue-specific genes. So, the C/EBPα and C/EBPε genes may be named as tissue-specific genes, and others C/EBP genes may be the housekeeping genes. The 6 C/EBP genes from pig were classified into 2 groups based on the cluster analysis of the C/EBP gene expression levels in various tissues, as shown in Figure 3. Group I contains only the C/EBPε gene, and other genes are included in group II. The expression levels of the C/EBPε gene were low in all tissues (FPKM < 5), and the gene is specific expressed in the intestine, salivary gland, thyroid, uterus, and lymph. Although C/EBPα, C/EBPβ, C/EBPδ, C/EBPγ, and C/EBPζ are expressed ubiquitously, the expression patterns in various tissues are inconsistent. According to the cluster analysis, group II was classified to 3 sub-groups, including sub-group I (C/EBPβ and C/EBPδ genes), sub-group II (C/EBPα gene), sub-group III (C/EBPγ and C/EBPζ genes). C/EBPα was expressed ubiquitously on 27 pig tissues, and specific expressed in the thyroid, liver, lung, and adipose tissues. C/EBPβ was expressed at the high levels in the thyroid, adrenal gland, lung, adipose, liver, and ovary tissues. C/EBPδ was expressed at the high levels in the thyroid, gall bladder, ovary, and uterus. C/EBPγ and C/EBPζ are widely expressed at similar levels.

Heatmap of C/EBP genes in different tissues of duroc pig. Red represents higher expression levels; blue represents lower expression levels. The color vary from blue for lower expression level to red for higher expression level.
Additionally, the results indicate that the expression patterns in certain tissues, such as brain and spinal cord in the nervous system and ovary and uterus tissues in the female reproductive system, are similar.
Construction of a transcriptional regulatory network of the C/EBP gene family
The C/EBP family is an important family of transcription factors that regulate the expression of the target genes by binding to the promoter regions to maintain the normal physiological processes in vivo. According to the PWMs of the C/EBPα, C/EBPβ, C/EBPδ, C/EBPε, and C/EBPγ genes from the cisbp database, we predicted 4662, 3164, 8383, 7278, and 1604 target genes regulated by these genes, respectively (see Table 3). A total of 10,270 target genes are regulated by the C/EBP genes for a total of 25,091 regulatory relationships. Binding sites for other transcription factors and miRNAs are present in the regulatory regions of the C/EBP genes.
Regulatory relationships involved in C/EBP family members.

The numbers in brackets is the number of gene families of transcription factors.
In this study, the C/EBP genes are predicted to be regulated by 423 TFs forming 1582 regulatory relationships; mir503 and mir7140 are predicted to regulate C/EBPβ and C/EBPγ, respectively. Additionally, C/EBPβ and C/EBPγ are regulated by C/EBPα, C/EBPδ, and C/EBPε and C/EBPβ and C/EBPε, respectively. Interestingly, C/EBPβ self-regulation is also predicted. Thus, we constructed a pig C/EBP regulatory network (summarized in Figure 4) that includes C/EBP genes, TFs, miRNAs, and target genes. These genes were defined as a node, and the distribution of node degree approximately follows the power-law distribution. Certain network concepts, including the clustering coefficient, network centralization, and network heterogeneity, were calculated to be 0.1890, 0.8160, and 25.1730, respectively. Among them, the clustering coefficient describes the cohesive characteristics of the network. Network centralization and heterogeneity describe the distribution of connectivity (degree) in the network. Generally, if the network is highly heterogeneous, its centralization will be low. Moreover, the higher the heterogeneity of the network, the more uneven the distribution of degree in the network, that is only a few nodes in the network have high connectivity, while most other nodes have low connectivity. So, these results indicated the pig C/EBP regulatory network was a scale-free network.
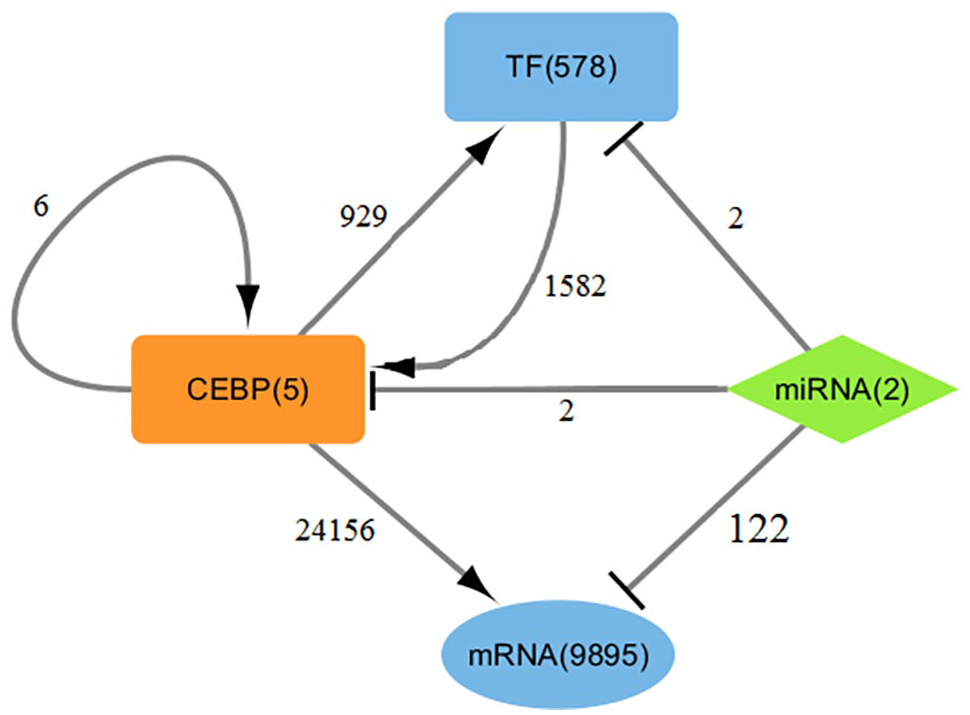
The transcriptional regulatory network with pig C/EBP family genes as the core. The arrows and represent TFs to regulate theirs target genes, and the T-lines represent miRNAs to regulate theirs target genes.
FFLs related to the C/EBP genes
According to the sequence, we predicted that mir503 and mir7140 regulate C/EBPβ and C/EBPγ, respectively. The miR503 and C/EBPβ genes coregulate 14 target genes forming 14 miR503-C/EBPβ-target gene FFL motifs, and miR7140 and C/EBPγ coregulate 11 target genes, including 11 miR7140-C/EBPγ-target gene FFL motifs. Additionally, miR503-ELF3-C/EBPβ and miR7140-ARID5B-C/EBPγ motifs were identified. The miR503-ELF3-C/EBPβ motif is involved in the miR503 and ELF3 coregulation of the C/EBPβ gene, and the miR503 gene targets the ELF3 and C/EBPβ genes. The miR7140-ARID5B-C/EBPγ motif is included in 3 regulatory relationships: miR7140→ARID5B, ARID5B→C/EBPγ, and miR7140→C/EBPγ. So, a total of 27 FFL motifs were identified in the pig C/EBP regulatory network. The combinations of all FFL motifs were used to construct the FFL sub-network (see Figure 5). There are C/EBPβ-binding sites in 5ʹ-untranslated region (5ʹ-UTR) of C/EBPγ. Thus, C/EBPβ and C/EBPγ co-regulated 10 target genes in FFL sub-network.

Subnetwork composed of 27 FFLs. The arrows and represent TFs to regulate theirs target genes, and the T-lines represent miRNAs to regulate theirs target genes. There are 2 miRNA-TF-C/EBP FFL motifs and 25 miRNA-C/EBP-target gene FFL motifs, including miR7140-C/EBPβ-ARID5B. In addition, there are 12 TF-TF-target motifs in this network.
Based on the data of RNA-seq, gene expression patterns related to this FFL sub-network were analyzed in 27 adult tissues of Duroc pig. We explored the tissue-specific target genes for C/EBP in FFL sub-network. The results indicate that target genes regulated by C/EBPβ and C/EBPγ have variable expression patterns in various tissues (see Table 4, Figures S3–S29). For example, the ATP synthase F1 subunit alpha (ATP5F1A) gene is a target gene of C/EBPβ and C/EBPγ, and form a FFL motif which is expressed ubiquitously. The glutamate decarboxylase-like protein 1 (GADL1) gene is muscle-specific express gene (τ= 0.99). We suggest that the C/EBPβ-C/EBPγ-GADL1 motif may play an important role in the muscle tissue. Some FFL motifs may be tissue-specific. Based on the target genes expression pattern, we estimated that the number of FFL motifs in each tissue may be significantly different; however, miRNA expression patterns were not evaluated in the present study.
Statistical table of distribution of FFLs in various tissues.

The dN and dS analysis of the C/EBP genes and target genes
The data on the nonsynonymous (dN) and synonymous (dS) substitution rates between the human and pig sequences were downloaded from the Ensembl database. The dN/dS values of the C/EBP genes ranged from 0.02 to 0.21 indicating that pig C/EBP genes underwent purifying selection. The dN+dS value of the C/EBPδ gene was 1.52, which was higher than that of 5 other C/EBP genes (0.31–0.67) indicating that the C/EBPδ gene evolved rapidly and had an increased mutation rate.
The dN+dS mean values of the target genes of C/EBPα, C/EBPβ, C/EBPδ, C/EBPε, and C/EBPγ are 0.58, 0.48, 0.56, 0.53, and 0.51, respectively. The dN/dS mean values of the target genes are 0.166, 0.175, 0.17, 0.177, and 0.166, respectively. The dN/dS and dN+dS mean values of the target genes of each C/EBP gene were compared using Kolmogorov-Smirnov (KS) test. The results indicate that the dN/dS distributions of the target genes of C/EBPα are similar to that of C/EBPβ, C/EBPε and C/EBPγ (P < .05), respectively. The very low dN/dS values suggest strong negative selection on all C/EBP genes, which may remain due to genetic drift or persistence. The dN+dS value distribution of the C/EBPα target genes is similar to that of the C/EBPδ target genes and is significantly higher than that of other C/EBP target genes (P < .05). The results indicate that the target genes of C/EBPα appear to be evolving rapidly.
Functional enrichment analysis of the C/EBP genes and target genes
We used the DAVID software to analyze the functions of the pig C/EBP genes. The results indicate that the functions are associated with many biological processes, including macrophage differentiation (GO: 0030225), inner ear development (GO: 0048839), positive regulation of osteoblast differentiation (GO: 0045669), transcriptional misregulation in cancer pathways (ssc05202), and tuberculosis pathways (ssc05152) (Table 5).
The results of functional enrichment analysis of porcine C/EBP family members.

The functional enrichment analysis of the target genes regulated by the C/EBP genes showed that the target genes of C/EBPα, C/EBPβ, C/EBPε, and C/EBPγ are associated with nucleoplasm (GO: 0005654) and extracellular exosome (GO: 0070062). The target genes of C/EBPδ and C/EBPγ are involved in the transforming growth factor beta (TGFβ) receptor signaling pathway. The target genes of C/EBPδ are involved in the platelet-derived growth factor receptor signaling (GO: 0048008) (Table 6).
The results of functional enrichment analysis of porcine C/EBP target genes.

Discussion
The characteristics and phylogenetic relationship analysis of the C/EBP gene family
Improved genome sequencing and annotation enabled identification of all C/EBP TFs in the eukaryotic organism genomes. This study identified all C/EBP TFs in pig and others eukaryotic organisms and found that the C/EBP genes are ubiquitous in the majority of the vertebrates. There are differences in biochemical parameters among C/EBP TFs. These biochemical parameters affect protein solubility, sub-cellular localization and interaction. 46 The physical and chemical properties of different C/EBPs also quite indicated the differences in the functionalities of the C/EBPs. Some biochemical parameters have very difference in different species. For example, the C/EBPα of nematode, sea squirt zebrafish is acidic while in other species is basic. The C /EBPδ is acidic in 1/3 of the species and alkaline in the remaining 2/3. These results indicate that some C/EBP genes have low functional conservation among different eukaryotes.47,48
In lower eukaryotes, such as nematode, fruit fly, there are C/EBPα and C/EBPγ genes. But in vertebrate species, more C/EBP genes distributed on genome. It indicated the C/EBP gene family may have been expanded in early stage of vertebrate species evolution. The molecular phylogenetic tree showed that the C/EBP family members originated in the early stage of vertebrate evolution. Similarly, Qiu et al 4 analyzed the evolution of the C/EBPα, C/EBPβ, and C/EBPδ genes in 20 vertebrate genomes and demonstrated that the C/EBP TFs originated early in vertebrate evolution.
According to the phylogenetic tree, all C/EBP TFs were classified into 6 subgroups. However, Qiu et al analyzed the phylogenetic trees of C/EBPα, C/EBPβ, and C/EBPδ homologs in 2 fish, 2 amphibian, 2 reptile, 11 avian, and 3 mammalian species. The results indicated that the C/EBPα and C/EBPβ genes are clustered into 1 group. This difference may be due to different species and members of the C/EBP family.
The expression pattern and function of the C/EBP gene family of pig
Based on the data of RNA-seq, the C/EBPα, C/EBPβ, C/EBPδ, C/EBPγ, and C/EBPζ genes are expressed in all 27 tissues of adult Duroc pig, and C/EBPε is expressed only in 5 tissues. Uhlén et al 49 studied the expression patterns of the C/EBP genes in 32 human tissues and demonstrated that the C/EBPβ, C/EBPδ, C/EBPγ, and C/EBPζ genes are expressed ubiquitously. Summers et al 50 created a pig gene expression atlas based on the meta-analysis and demonstrated that C/EBPε is expressed only in a few tissues, and other C/EBP genes are expressed in the majority of tissues. Shyamsunder et al 51 and Studd et al 52 found the C/EBPε gene is a major regulator of myeloid and lymphoid cell differentiation, respectively. Our results found the C/EBPε gene shown the exceptionally low expression levels in 27 pig tissues, and it also named as thyroid and lymph tissue-specific genes. So, it indicated that C/EBPε gene may only perform functions in specific tissues related to immune system, such as myeloid and lymphoid cell.The differences in the exon-intron, motif structures and gene expression patterns among various clades provided some indications that the C/EBP genes may have a variety of physiological functions. The difference number of exons and motifs might be the basis of C/EBP genes functional diversity. Functional enrichment analysis demonstrated that pig C/EBP genes perform many important biological functions, involved in the inner ear development (GO: 0048839), positive regulation of osteoblast differentiation (GO: 0045669), and the tuberculosis pathway (ssc05152). The C/EBPζ gene contained the highest number of exons and motifs, which indicated that it may take some specific roles. Previous studies showed that C/EBPζ is involved in cochlear cell apoptosis in rats and may co-regulate the inner ear development with GATA binding protein 2 (GATA2).53,54
We used cluster analysis to explore similar expression patterns in multiple tissues among C/EBP genes. There were similar expression between C/EBPβ and C/EBPδ genes, belonged to the same sub-group. It has been reported that both C/EBPβ and C/EBPδ were involved in the chondrocyte differentiation, heart development and early regulation of fat formation.55-57The sub-group III includes C/EBPγ and C/EBPζ genes, which it has been reported that they are important regulators of cellular stress response networks to maintain the basic life activities of cells. 58
Regulation of gene expression is one of the most important and complex issues in biology. Many studies suggested that tissue specificity of gene expression or gene regulatory network took biologically important roles. In this study, we found the C/EBPα may be named as fat tissue-specific gene. Chen et al 59 demonstrated that C/EBPα gene is the lipogenic marker of the porcine adipocytes that promotes adipocyte differentiation. In this study, we also calculated the tissue specific index of the C/EBP family target genes. Our results indicated that C/EBPβ genes may be the housekeeping gene, which are expressed at the high levels in many tissues, as previous studies11,13,18 had shown. But some of its target genes might name as tissue-specific genes, such as ELF3, GADL1, SRGAP3. Combined with the FFL sub-network, we deem some FFL motifs may be tissue-specific. Sonawane et al 60 reported that transcription factors primarily participate in tissue-specific regulatory processes via alterations in their targeting patterns.
The transcriptional regulatory network
Complex regulatory mechanisms of gene expression in eukaryotes control the development, physiology, and pathology. 5 In this study, we constructed a C/EBP gene regulatory network to provide some information to explain the regulatory mechanisms of the pig C/EBP genes. The results indicate that the C/EBP gene regulatory network is a typical scale-free network. A key advantage of the scale-free topology is the increased tolerance to random failures, which helps the animals adjust to a stressful environment and maintains C/EBPs to perform its biological functions. 61 Furthermore, human TF and miRNA regulatory network were also reported to correspond to approximately scale-free topology.62,63
In the regulatory gene network, transcription factors play an important role at the gene transcription level due to the regulation of certain target genes. Each TF contains different conservation motifs to target various genes. In this study, the target genes of each porcine C/EBP TFs were different. In the Database of Human Transcription Factor Targets, 64 the target genes of the C/EBP TFs are also different. Comparison with the target genes of the human C/EBP TFs indicated that porcine C/EBPδ regulates more target genes, but the number of the target genes of other porcine C/EBP TFs is less than that in humans. Selection of different target genes during evolution to meet the needs of normal homeostatic development and growth or to adapt to various environmental stresses may differ in different species.
The biological network is composed of motifs, and the FFL is an important motif. In this study, 27 FFLs were detected in the C/EBP genes of the transcriptional regulatory sub-network. The FFL-containing C/EBP genes play important roles in multiple human tissues or cells. Sun et al 65 reported that miR223->C/EBPβ->LMO2 FFL in human myeloid cells can regulate cell proliferation and maintain normal differentiation and development. Ponomarev et al 66 have reported that MiR124-C/EBPα-PU.1 FFL is related to the transition between an activated phenotype and a quiescent state of macrophages. Shi et al 67 reported that 28 FFLs of C/EBPα, C/EBPβ, and C/EBPδ may be associated with occurrence of human hypertrophic cardiomyopathy. Although these FFLs has not been reported in pig tissues or cells, we deem that further experiment, such as ChIP-seq, would help identifying this FFL sub-networks, and demonstrating the regulation function of the C/EBP gene family in pig.
Conclusion
In the present study, we identified all C/EBP TFs of pig and others eukaryotic organisms and demonstrated the conservation and evolutionary relationship of the C/EBP family genes. The porcine C/EBP genes are expressed generally, except C/EBPε gene, and participate in many important biological functions. The C/EBP gene regulatory network was constructed to provide information on the regulatory mechanisms of the pig C/EBP genes. Some FFL motifs may be tissues-specific. Thus, our study provides potential functional insight into the roles of the C/EBP genes in pig.
Supplemental Material
sj-docx-1-evb-10.1177_11769343211041382 – Supplemental material for Genome-Wide Phylogenetic Analysis, Expression Pattern, and Transcriptional Regulatory Network of the Pig C/EBP Gene Family
Supplemental material, sj-docx-1-evb-10.1177_11769343211041382 for Genome-Wide Phylogenetic Analysis, Expression Pattern, and Transcriptional Regulatory Network of the Pig C/EBP Gene Family by Chaoxin Zhang, Tao Wang, Tongyan Cui, Shengwei Liu, Bing Zhang, Xue Li, Jian Tang, Peng Wang, Yuanyuan Guo and Zhipeng Wang in Evolutionary Bioinformatics
Footnotes
Funding:
The authors would like to thank financially supported by Natural Science Foundation of China (No. 32070571), the Academic Backbone Project of Northeast Agricultural University (No.15XG14), NEAU Research Founding for Excellent Young Teachers (2010RCB29).
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author contribution statement
ZPW and CXZ conceived the study, and ZPW, CXZ, TW and SWL participated in its design. TYC, BZ, XL, JT and PW were involved in the acquisition of data, CXZ, TW and TYC performed all data analysis. ZPW and CXZ drafted the manuscript, and SWL, YYG, BZ, XL, JT, PW and TYC contributed to the writing and editing. All authors read and approved the final manuscript.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
